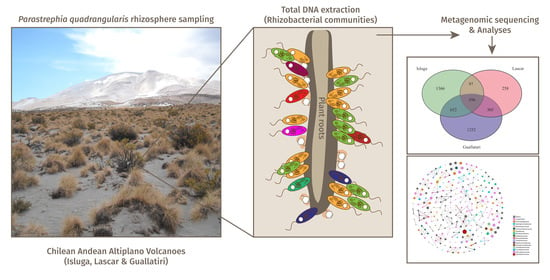
Composition and Potential Functions of Rhizobacterial Communities in a Pioneer Plant from Andean Altiplano
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sampling Sites
2.2. DNA Extraction, Library Preparation and 16S rRNA Amplicon Sequencing
2.3. Bioinformatic Processing of the Sequences and Statistical Analysis of the Data
3. Results
3.1. Chemical Properties of Rhizosphere Soils
3.2. Alpha Diversity and Taxonomic Assignments of Rhizobacterial Community
3.3. Shared OTUs and Predicted Functions of Rhizobacterial Community Members
3.4. Co-Occurrence Networks and Putative Keystone Taxa across Rhizobacterial Community Members
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Connell, J.H.; Slatyer, R.O. Mechanisms of succession in natural communities and their role in community stability and organization. Am. Nat. 1977, 111, 1119–1144. [Google Scholar] [CrossRef]
- Navarro-Noya, Y.E.; Jan-Roblero, J.; del González-Chávez, M.; Hernández-Gama, R.; Hernández-Rodríguez, C. Bacterial communities associated with the rhizosphere of pioneer plants (Bahia xylopoda and Viguiera linearis) growing on heavy metals-contaminated soils. Antonie Leeuwenhoek. 2010, 4, 335–349. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Schmid, M.; van Tuinen, D.; Berg, G. Plant-driven selection of microbes. Plant. Soil. 2009, 321, 235–257. [Google Scholar] [CrossRef]
- Ciccazzo, S.; Esposito, A.; Borruso, L.; Brusetti, L. Microbial communities and primary succession in high altitude mountain environments. Ann. Microbiol. 2016, 66, 43–60. [Google Scholar] [CrossRef]
- Sun, X.; Zhou, Y.; Tan, Y.; Wu, Z.; Lu, P.; Zhang, G.; Yu, F. Restoration with pioneer plants changes soil properties and remodels the diversity and structure of bacterial communities in rhizosphere and bulk soil of copper mine tailings in Jiangxi Province, China. Environ. Sci. Pollut. Res. 2018, 25, 22106–22119. [Google Scholar] [CrossRef]
- Liu, F.; Hewezi, T.; Lebeis, S.L.; Pantalone, V.; Grewal, P.S.; Staton, M.E. Soil indigenous microbiome and plant genotypes cooperatively modify soybean rhizosphere microbiome assembly. BMC Microbiol. 2019, 19, 201. [Google Scholar] [CrossRef]
- Ciccazzo, S.; Esposito, A.; Rolli, E.; Zerbe, S.; Daffonchio, D.; Brusetti, L. Safe-sites effects on rhizosphere bacterial communities in a high-altitude alpine environment. BioMed. Res. Int 2014, 2014, 480170. [Google Scholar] [CrossRef]
- Ciccazzo, S.; Esposito, A.; Rolli, E.; Zerbe, S.; Daffonchio, D.; Brusetti, L. Different pioneer plant species select specific rhizosphere bacterial communities in a high mountain environment. Springer Plus 2014, 3, 391. [Google Scholar] [CrossRef] [Green Version]
- Stern, C.R. Active Andean volcanism: Its geologic and tectonic setting. Rev. geol. Chile. 2004, 31, 161–206. [Google Scholar] [CrossRef]
- Tapia, J.; Murray, J.; Ormachea, M.; Tirado, N.; Nordstrom, D.K. Origin, distribution, and geochemistry of arsenic in the Altiplano-Puna plateau of Argentina, Bolivia, Chile, and Perú. Sci. Total Environ. 2019, 678, 309–325. [Google Scholar] [CrossRef]
- Garreaud, R.; Vuille, M.; Clement, A.C. The climate of the Altiplano: Observed current conditions and mechanisms of past changes. Palaeogeogr. Palaeoclimatol. Palaeoecol. 2003, 194, 3–22. [Google Scholar] [CrossRef] [Green Version]
- Rundel, P.W.; Palma, B. Preserving the unique Puna ecosystems of the Andean Altiplano. Mt. Res. Dev. 2000, 3, 262–271. [Google Scholar] [CrossRef] [Green Version]
- Tapia, J.; González, R.; Townley, B.; Oliveros, V.; Álvarez, F.; Aguilar, G.; Menzies, A.; Calderón, M. Geology and geochemistry of the Atacama Desert. Antonie Leeuwenhoek 2018, 111, 1273–1291. [Google Scholar] [CrossRef]
- Sernageomin. Chile: Territorio volcánico. In Servicio Nacional Geología y Minería; Sernageomin: Santiago, Chile, 2018; p. 135. Available online: https://www.sernageomin.cl/pdf/LIBROdevolcanes_SERNAGEOMIN.pdf (accessed on 10 July 2021).
- Pérez, F.L. Steady as a rock: Biogeomorphic influence of nurse rocks and slope processes on kūpaoa (Dubautia menziesii) shrubs in Haleakalā Crater (Maui, Hawai’i). Geomorphology. 2017, 295, 631–644. [Google Scholar] [CrossRef]
- Lambrinos, J.G.; Kleier, C.C.; Rundel, P.W. Plant community variation across a Puna landscape in the Chilean Andes. Rev. Chil. Hist. Nat. 2006, 79, 233–243. [Google Scholar] [CrossRef] [Green Version]
- Menoyo, E.; Lugo, M.N.; Teste, F.P.; Ferrero, M.F. Grass dominance drives rhizospheric bacterial communities in a desertic shrub and grassy steppe Highland. Pedobiologia 2017, 62, 36–40. [Google Scholar] [CrossRef]
- Acuña, J.J.; Jaisi, D.; Campos, M.; Mora, M.L.; Jorquera, M.A. ACCD-producing rhizobacteria from an Andean Altiplano native plant (Parastrephia quadrangularis) and their potential to alleviate salt stress in wheat seedling. Appl. Soil. Ecol. 2019, 136, 184–190. [Google Scholar] [CrossRef]
- Araya, J.P.; González, M.; Cardinale, M.; Schnell, S.; Stoll, A. Microbiome dynamics associated with the Atacama flowering Desert. Front Microbiol. 2020, 10, 3160. [Google Scholar] [CrossRef] [Green Version]
- Inostroza, N.G.; Barra, P.J.; Wick, L.Y.; Mora, M.L.; Jorquera, M.A. Effect of rhizobacterial consortia from undisturbed arid- and agro-ecosystems on wheat growth under different conditions. Lett. Appl. Microbiol. 2017, 64, 158–163. [Google Scholar] [CrossRef] [Green Version]
- Mandakovic, D.; Maldonado, J.; Pulgar, R.; Cabrera, P.; Gaete, A.; Urtuvia, V.; Seeger, M.; Cambiazo, V.; Gonzalez, M. Microbiome analysis and bacterial isolation from Lejía Lake soil in Atacama Desert. Extremophiles 2018, 22, 665–673. [Google Scholar] [CrossRef]
- Maza, F.; Maldonado, J.; Vásquez-Dean, J.; Mandakovic, D.; Gaete, A.; Cambiazo, V.; González, M. Soil bacterial communities from the Chilean Andean highlands: Taxonomic composition and culturability. Front. Bioeng. Biotechnol. 2019, 7, 10. [Google Scholar] [CrossRef]
- Radojevic, M.; Bashkin, V. Practical Environmental Analysis; Royal Society of Chemistry: London, UK, 1999. [Google Scholar]
- Walkley, A.; Black, I.A. An examination of the Degtjareff method for determining soil organic matter and a proposed modification of the chromic acid titration method. Soil Sci. 1934, 37, 29–38. [Google Scholar] [CrossRef]
- Murphy, J.; Riley, J.P. A modified single solution method for the determination of phosphate in natural waters. Anal. Chim. Acta. 1962, 27, 31–36. [Google Scholar] [CrossRef]
- Ogram, A.; Sayler, G.S.; Barkay, T. The extraction and purification of microbial DNA from sediments. J. Microbiol. Methods. 1987, 7, 57–66. [Google Scholar] [CrossRef]
- Wasimuddin; Schlaeppi, K.; Ronchi, F.; Leib, S.L.; Erb, M.; Ramette, A. Evaluation of primer pairs for microbiome profiling from soils to humans within the One Health framework. Mol. Ecol. Resour. 2020, 20, 1558–1571. [Google Scholar] [CrossRef]
- Gohl, D.M.; Vangay, P.; Garbe, J.; MacLean, A.; Hauge, A.; Becker, A.; Trevor, J.; Clayton, G.J.B.; Johnson, T.J.; Hunter, R.; et al. Systematic improvement of amplicon marker gene methods for increased accuracy in microbiome studies. Nat. Biotechnol. 2016, 34, 942–949. [Google Scholar] [CrossRef] [Green Version]
- Al-Ghalith, G.A.; Hillmann, B.; Ang, K.; Shields-Cutler, R.; Knights, D. SHI7 is a self-learning pipeline for multipurpose short-read DNA quality control. mSystems 2018, 24, e00202-17. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Zhang, Q.; Staley, C.; Gao, H.; Ishii, S.; Wei, X.; Liu, J.; Cheng, J.; Hao, M.; Sadowsky, M.J. Impact of long-term grazing exclusion on soil microbial community composition and nutrient availability. Biol. Fertil. Soils. 2019, 55, 121–134. [Google Scholar] [CrossRef]
- Qiu, L.; Zhang, Q.; Zhu, H.; Reich, P.B.; Banerjee, S.; van der Heijden, M.G.A.; Sadowsky, M.J.; Ishii, S.; Jia, X.; Shao, M.; et al. Erosion reduces soil microbial diversity, network complexity and multifunctionality. ISME J. 2021, 15, 2474–2489. [Google Scholar] [CrossRef]
- Schloss, P.D. Amplicon sequence variants artificially split bacterial genomes into separate clusters. mSphere 2021, 6, e00191-21. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. Uchime improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [Green Version]
- Al-Ghalith, G.A.; Montassier, E.; Ward, H.N.; Knights, D. NINJA-OPS: Fast accurate marker gene alignment using concatenated ribosomes. PLoS Comput. Biol. 2016, 12, e1004658. [Google Scholar] [CrossRef] [PubMed]
- Louca, S.; Parfrey, L.W.; Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science. 2016, 353, 1272–1277. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Wang, H.Z.; Dsouza, M.; Lou, J.; He, Y.; Dai, Z.M.; Brookes, P.C.; Xu, J.; Gilbert, J.A. Geographic patterns of co-occurrence network topological features for soil microbiota at continental scale in eastern China. ISME J. 2016, 10, 1891–1901. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Acuña, J.J.; Inostroza, N.G.; Durán, P.; Mora, M.L.; Sadowsky, M.J.; Jorquera, M.A. Niche differentiation in the composition, predicted function, and co-occurrence networks in bacterial communities associated with Antarctic vascular plants. Front. Microbiol. 2020, 11, 1036. [Google Scholar] [CrossRef] [PubMed]
- Berry, D.; Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 2014, 5, 219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. ICWSM Conf. 2009, 8, 361–362. [Google Scholar]
- Yoshitake, S.; Fujiyoshi, M.; Watanabe, K.; Masuzawa, T.; Nakatsubo, T.; Koizumi, H. Successional changes in the soil microbial community along a vegetation development sequence in a subalpine volcanic desert on Mount Fuji, Japan. Plant. Soil. 2013, 364, 261–272. [Google Scholar] [CrossRef]
- Alsharif, W.; Saad, M.M.; Hirt, H. Desert Microbes for Boosting Sustainable Agriculture in Extreme Environments. Front. Microbiol. 2020, 11, 1666. [Google Scholar] [CrossRef]
- Navarro, G.; Arrázola, S.; Atahuachi, M.; De la Barra, N.; Mercado, M.; Ferreira, W.; Moraes, M. Libro Rojo De la Flora Amenazada de Bolivia; Ministerio de Medio Ambiente y Agua Viceministerio de Medio Ambiente, Biodiversidad, Cambios Climaticos y de Gestion y Desarrollo Forestal: Cochabamba, Bolivia, 2012. [Google Scholar]
- Jorquera, M.A.; Maruyama, F.; Ogram, A.V.; Navarrete, O.U.; Lagos, L.M.; Inostroza, N.G.; Acuña, J.J.; Rilling, J.I.; de La Luz Mora, M. Rhizobacterial Community Structures Associated with Native Plants Grown in Chilean Extreme Environments. Microb. Ecol. 2016, 72, 633–646. [Google Scholar] [CrossRef]
- Fernández-Gómez, B.; Maldonado, J.; Mandakovic, D.; Gaete, A.; Gutierrez, R.A.; Mass, A.; Cambiazo, V.; Gonzalez, M. Bacterial communities associated to Chilean altiplanic native plants from the Andean grasslands soils. Sci Rep. 2019, 9, 1042. [Google Scholar] [CrossRef]
- Fuentes, A.; Herrera, H.; Charles, T.C.; Arriagada, C. Fungal and Bacterial Microbiome Associated with the Rhizosphere of Native Plants from the Atacama Desert. Microorganisms. 2020, 8, 209. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Acuña, J.J.; Inostroza, N.G.; Mora, M.L.; Radic, S.; Sadowosky, M.J.; Jorquera, M.A. Endophytic bacterial communities associated with roots and leaves of plants growing in Chilean extreme environments. Sci Rep. 2019, 9, 4950. [Google Scholar] [CrossRef] [Green Version]
- Ibeyaima, A.; Rana, J.; Dwivedi, A.K.; Saini, N.; Gupta, S.; Sarethy, I.P. Pseudonocardiaceae sp. TD-015 from the Thar Desert, India: Antimicrobial Activity and Identification of Antimicrobial Compounds. Curr. Bioact. Compd. 2018, 14, 112–118. [Google Scholar] [CrossRef]
- Sghaier, H.; Hezbri, K.; Ghodhbane-Gtari, F.; Pujic, P.; Sen, A.; Daffonchio, D.; Boudabous, A.; Tisa, L.S.; Klenk, H.-P.; Armengaud, J.; et al. Stone-dwelling actinobacteria Blastococcus saxobsidens, Modestobacter marinus and Geodermatophilus obscurus proteogenomes. ISME J. 2016, 10, 21–29. [Google Scholar] [CrossRef]
- Qin, S.; Bian, G.K.; Zhang, Y.J.; Xing, K.; Cao, C.L.; Liu, C.H.; Dai, C.-C.; Li, W.-J.; Jiang, J.H. Modestobacter roseus sp. nov., an endophytic actinomycete isolated from the coastal halophyte Salicornia europaea Linn., and emended description of the genus Modestobacter. Int. J. Syst. Evol. Microbiol. 2013, 63, 2197–2202. [Google Scholar] [CrossRef]
- Golinska, P.; Montero-Calasanz, M.C.; Świecimska, M.; Yaramis, A.; Igual, J.M.; Bull, A.T.; Goodfellow, M. Modestobacter excelsi sp. nov., a novel actinobacterium isolated from a high altitude Atacama Desert soil. Sys. Appl. Microbiol. 2020, 43, 1–9. [Google Scholar] [CrossRef]
- Bull, A.T.; Idris, H.; Sanderson, R.; Asenjo, J.; Andrews, B.; Goodfellow, M. High altitude, hyper-arid soils of the Central-Andes harbor mega-diverse communities of actinobacteria. Extremophiles 2018, 22, 47–57. [Google Scholar] [CrossRef] [Green Version]
- Shen, J.; Wyness, A.J.; Claire, M.W.; Zerkle, A.L. Spatial variability of microbial communities and salt distributions across a latitudinal aridity gradient in the Atacama Desert. Microb. Ecol. 2021, 82, 442–458. [Google Scholar] [CrossRef]
- Idris, H.; Goodfellow, M.; Sanderson, R.; Asenjo, J.A.; Bull, A.T. Actinobacterial Rare Biospheres and Dark Matter Revealed in Habitats of the Chilean Atacama Desert. Sci. Rep. 2017, 7, 8373. [Google Scholar] [CrossRef] [Green Version]
- Wu, D.; Senbayram, M.; Moradi, G.; Mörchen, R.; Knief, C.; Klumpp, E.; Jones, D.L.; Well, R.; Chen, R.; Bol, R. Microbial potential for denitrification in the hyperarid Atacama Desert soils. Soil. Bio. Biochem. 2021, 157, 108248. [Google Scholar] [CrossRef]
- Mapelli, F.; Marasco, R.; Fusi, M.; Scaglia, B.; Tsiamis, G.; Rolli, E.; Fodelianakis, S.; Bourtzis, K.; Ventura, S.; Tambone, F.; et al. The stage of soil development modulates rhizosphere effect along a High Arctic desert chronosequence. ISME J. 2018, 12, 1188–1198. [Google Scholar] [CrossRef]
- Astorga-Eló, M.; Zhang, Q.; Larama, G.; Stoll, A.; Sadowsky, M.J.; Jorquera, M.A. Composition, predicted functions and co-occurrence networks of rhizobacterial communities impacting flowering desert events in the Atacama Desert, Chile. Front. Microbiol. 2020, 11, 571. [Google Scholar] [CrossRef]





| Samples | Guallatiri | Isluga | Lascar |
|---|---|---|---|
| Sampling point | |||
| Location coordinates | 18°29′2.98″ S 69°8′24.36″ W | 19°12′12.20″ S 68°46′17.53″ W | 23°21′12.00″ S 67°48′40.64″ W |
| Altitude (MASL) a | 4386 | 3983 | 4353 |
| Chemical properties | |||
| N (mg kg −1) | 10 | 10 | 12 |
| P (mg kg −1) | 23 | 37 | 22 |
| K (mg kg −1) | 145 | 207 | 217 |
| Organic matter (g kg −1) | 1.48 | 1.62 | 1.82 |
| pHH2O | 5.59 | 4.77 | 5.93 |
| CEC (cmol+ kg −1) ¥ | 2.05 | 2.26 | 3.47 |
| Sum of bases | 1.91 | 1.98 | 3.44 |
| Cu (mg kg −1) | 0.80 | 0.29 | 2.97 |
| Zn (mg kg −1) | 0.30 | 0.35 | 0.70 |
| S (mg kg −1) | 13 | 98 | 44 |
| Richness | |||
| Sobs † | 1325 ± 510 A | 1137 ± 437 A | 830 ± 13 A |
| ACE ‡ | 1478 ± 630 A | 1281 ± 503 A | 912 ± 10 A |
| Chao1 | 1463 ± 643 A | 1249 ± 518 A | 889 ± 13 A |
| Alpha diversity | |||
| Coverage (%) | 98.73 ± 0.98 A | 98.94 ± 0.66 A | 99.35 ± 0 A |
| Shannon | 6.09 ± 0.42 A | 5.86 ± 0.44 A | 5.31 ± 0.02 A |
| Simpson | 0.007 ± 0.005 A | 0.009 ± 0.005 A | 0.014 ± 0.002 A |
| Genus | Closest Relative Taxonomic Affiliation | Guallatiri (%) | Isluga (%) | Lascar (%) |
|---|---|---|---|---|
| Modestobacter | Actinobacteria; Actinobacteria; Actinomycetales; Geodermatophilaceae | 2.6 | 2.7 | 5.5 |
| Segetibacter | Bacteroidetes; Saprospirae; Saprospirales; Chitinophagaceae | 2.5 | 1.6 | 2.7 |
| Actinomycetospora | Actinobacteria; Actinobacteria; Actinomycetales; Pseudonocardiaceae | 2.2 | 1.2 | 2.8 |
| Kaistobacter | Proteobacteria; Alphaproteobacteria; Sphingomonadales; Sphingomonadaceae | 1.53 | 1.71 | 2.28 |
| Nitrososphaera | Crenarchaeota; Thaumarchaeota; Nitrososphaerales; Nitrososphaeraceae | 1.96 | 1.79 | 1.74 |
| Mycobacterium | Actinobacteria; Actinobacteria; Actinomycetales; Mycobacteriaceae | 1.71 | 0.87 | 1.31 |
| Solibacter | Acidobacteria; Solibacteres; Solibacterales; Solibacteraceae | 1.49 | 1.06 | 0.99 |
| DA101 | Verrucomicrobia; Spartobacteria; Chthoniobacterales; Chthoniobacteraceae | 2.52 | 0.32 | 0.45 |
| Methylobacterium | Proteobacteria; Alphaproteobacteria; Rhizobiales; Methylobacteriaceae; | 0.45 | 1.73 | 0.74 |
| Geodermatophilus | Actinobacteria; Actinobacteria; Actinomycetales; Geodermatophilaceae | 0.22 | 0.77 | 1.52 |
| Streptomyces | Actinobacteria; Actinobacteria; Actinomycetales; Streptomycetaceae | 0.62 | 1.20 | 0.49 |
| Amycolatopsis | Actinobacteria; Actinobacteria; Actinomycetales; Streptomycetaceae | 0.95 | 0.87 | 0.35 |
| Aeromicrobium | Actinobacteria; Actinobacteria; Actinomycetales; Nocardioidaceae | 0.63 | 1.15 | 0.11 |
| Bradyrhizobium | Proteobacteria; Alphaproteobacteria; Rhizobiales; Bradyrhizobiaceae | 0.96 | 0.47 | 0.43 |
| Sphingomonas | Proteobacteria; Alphaproteobacteria; Sphingomonadales; Sphingomonadaceae | 0.64 | 0.53 | 0.64 |
| Rhodoplanes | Proteobacteria; Alphaproteobacteria; Rhizobiales; Hyphomicrobiaceae | 0.70 | 0.59 | 0.41 |
| Pseudonocardia | Actinobacteria; Actinobacteria; Actinomycetales; Pseudonocardiaceae | 0.60 | 0.64 | 0.21 |
| Kribbella | Actinobacteria; Actinobacteria; Actinomycetales; Nocardioidaceae | 0.36 | 0.87 | 0.17 |
| Burkholderia | Proteobacteria; Betaproteobacteria; Burkholderiales; Burkholderiaceae | 0.24 | 0.19 | 0.95 |
| Devosia | Proteobacteria; Alphaproteobacteria; Rhizobiales; Hyphomicrobiaceae | 0.24 | 0.53 | 0.28 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Q.; Araya, M.M.; Astorga-Eló, M.; Velasquez, G.; Rilling, J.I.; Campos, M.; Sadowsky, M.J.; Jorquera, M.A.; Acuña, J.J. Composition and Potential Functions of Rhizobacterial Communities in a Pioneer Plant from Andean Altiplano. Diversity 2022, 14, 14. https://doi.org/10.3390/d14010014
Zhang Q, Araya MM, Astorga-Eló M, Velasquez G, Rilling JI, Campos M, Sadowsky MJ, Jorquera MA, Acuña JJ. Composition and Potential Functions of Rhizobacterial Communities in a Pioneer Plant from Andean Altiplano. Diversity. 2022; 14(1):14. https://doi.org/10.3390/d14010014
Chicago/Turabian StyleZhang, Qian, Macarena M. Araya, Marcia Astorga-Eló, Gabriela Velasquez, Joaquin I. Rilling, Marco Campos, Michael J. Sadowsky, Milko A. Jorquera, and Jacquelinne J. Acuña. 2022. "Composition and Potential Functions of Rhizobacterial Communities in a Pioneer Plant from Andean Altiplano" Diversity 14, no. 1: 14. https://doi.org/10.3390/d14010014
APA StyleZhang, Q., Araya, M. M., Astorga-Eló, M., Velasquez, G., Rilling, J. I., Campos, M., Sadowsky, M. J., Jorquera, M. A., & Acuña, J. J. (2022). Composition and Potential Functions of Rhizobacterial Communities in a Pioneer Plant from Andean Altiplano. Diversity, 14(1), 14. https://doi.org/10.3390/d14010014