First Insights into the Microbiology of Three Antarctic Briny Systems of the Northern Victoria Land
Abstract
1. Introduction
2. Materials and Methods
2.1. Brine Sample Collection
2.2. Available Dataset
2.3. Predictive Functional Profiling
2.4. Statistical Analyses
3. Results
3.1. Main Features of Analyzed Antarctic Brines: A Comparison
3.1.1. Chemical Data
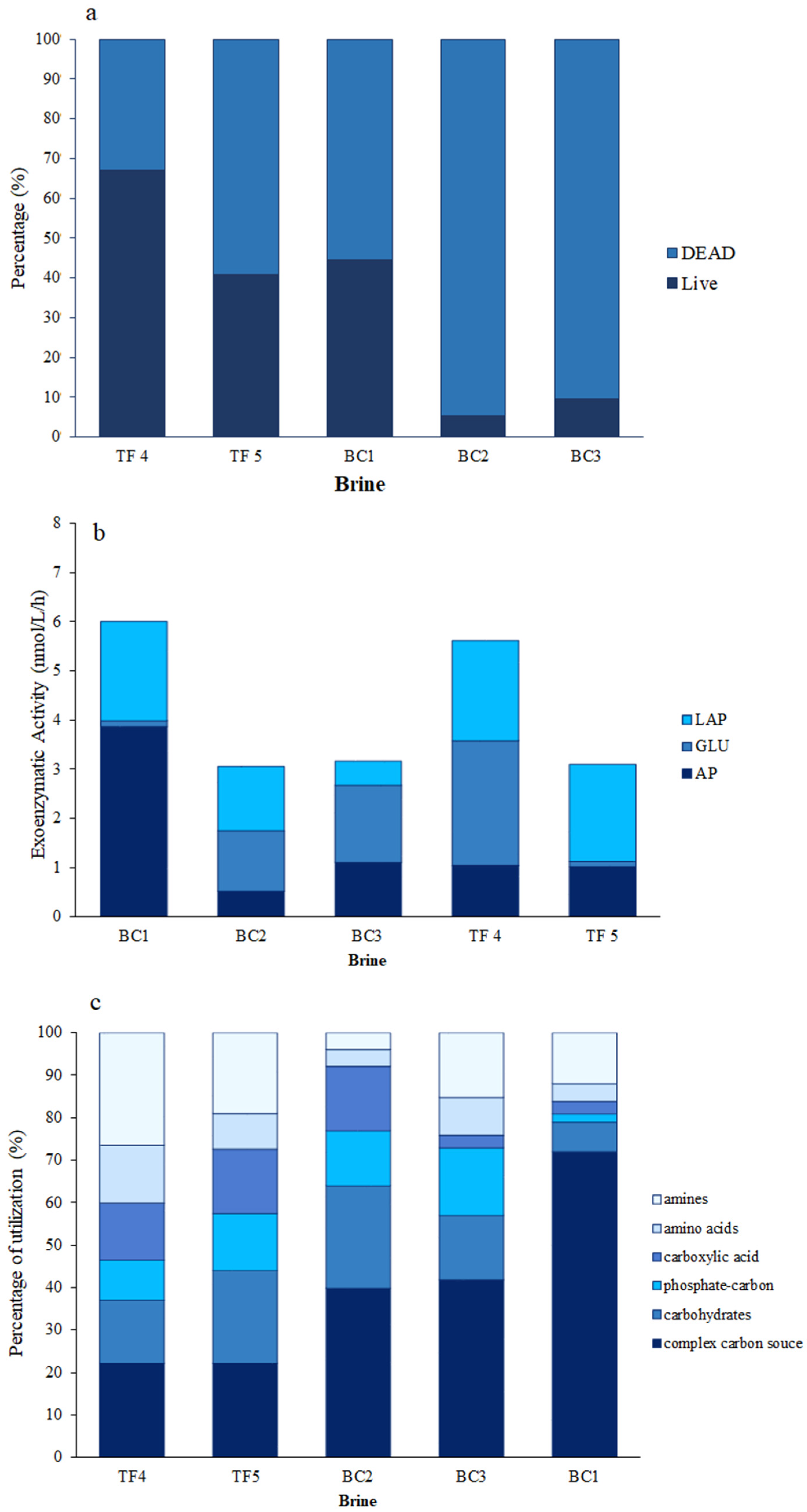
3.1.2. Microbial Abundance
3.1.3. Microbial Activities
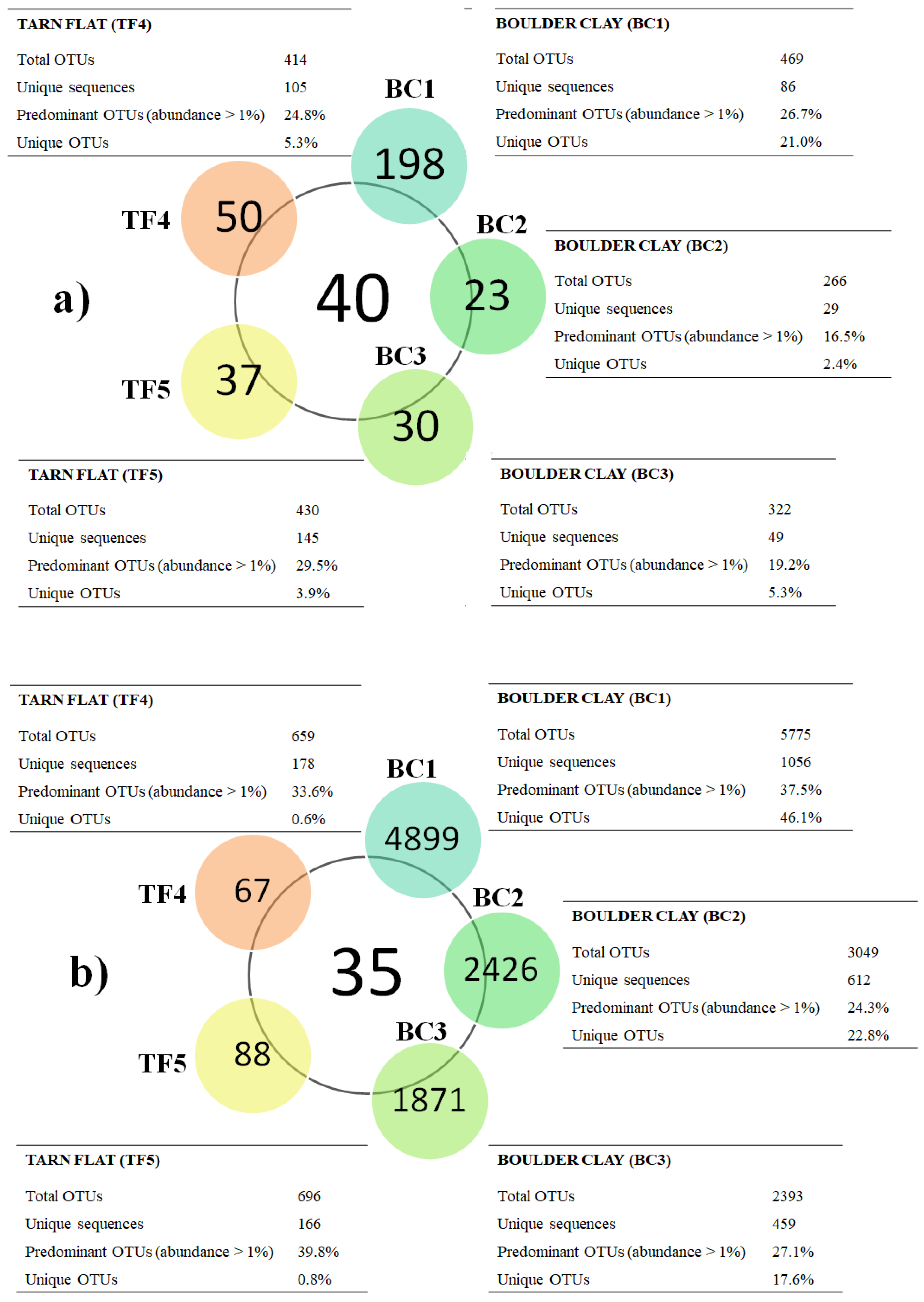
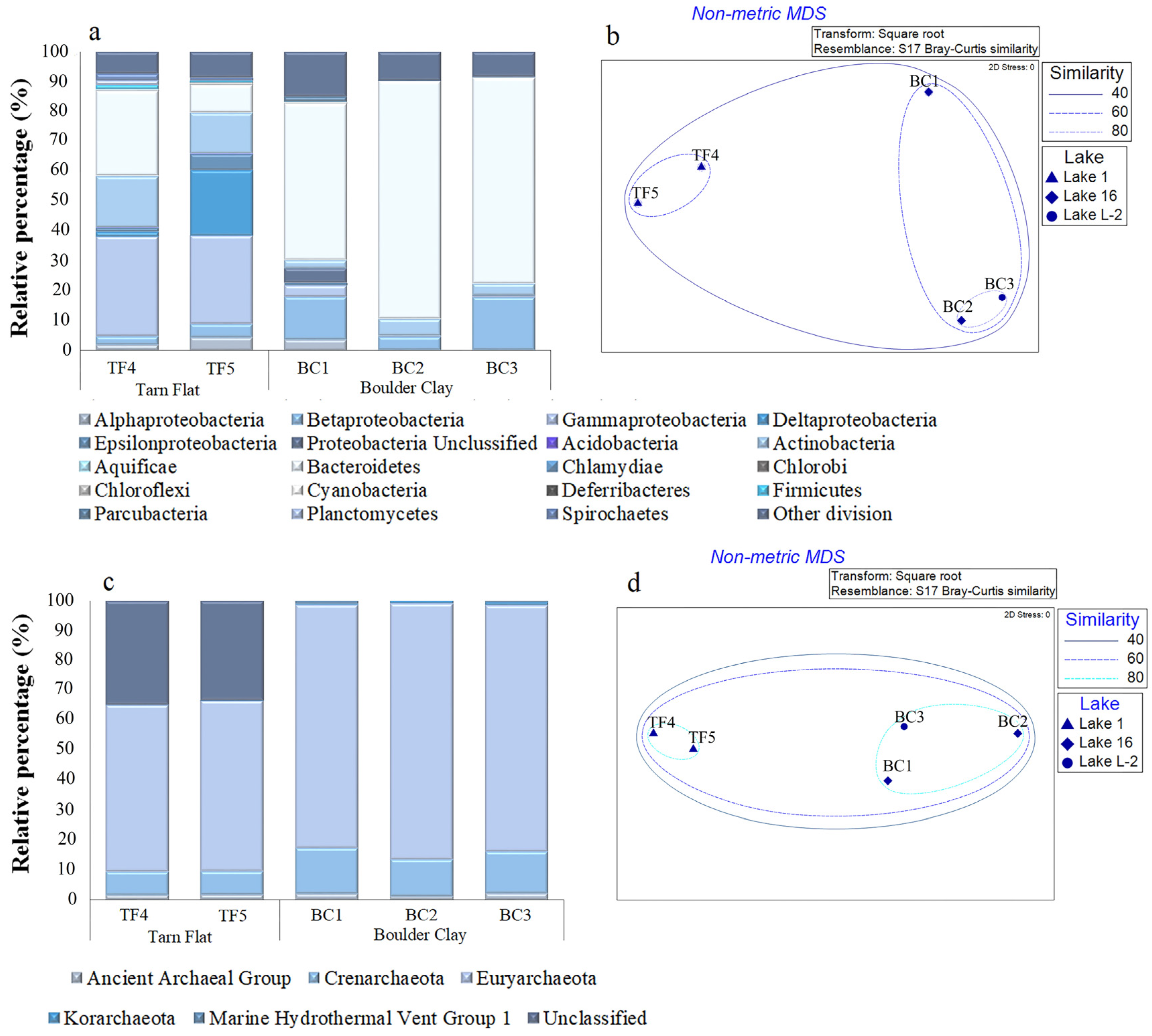
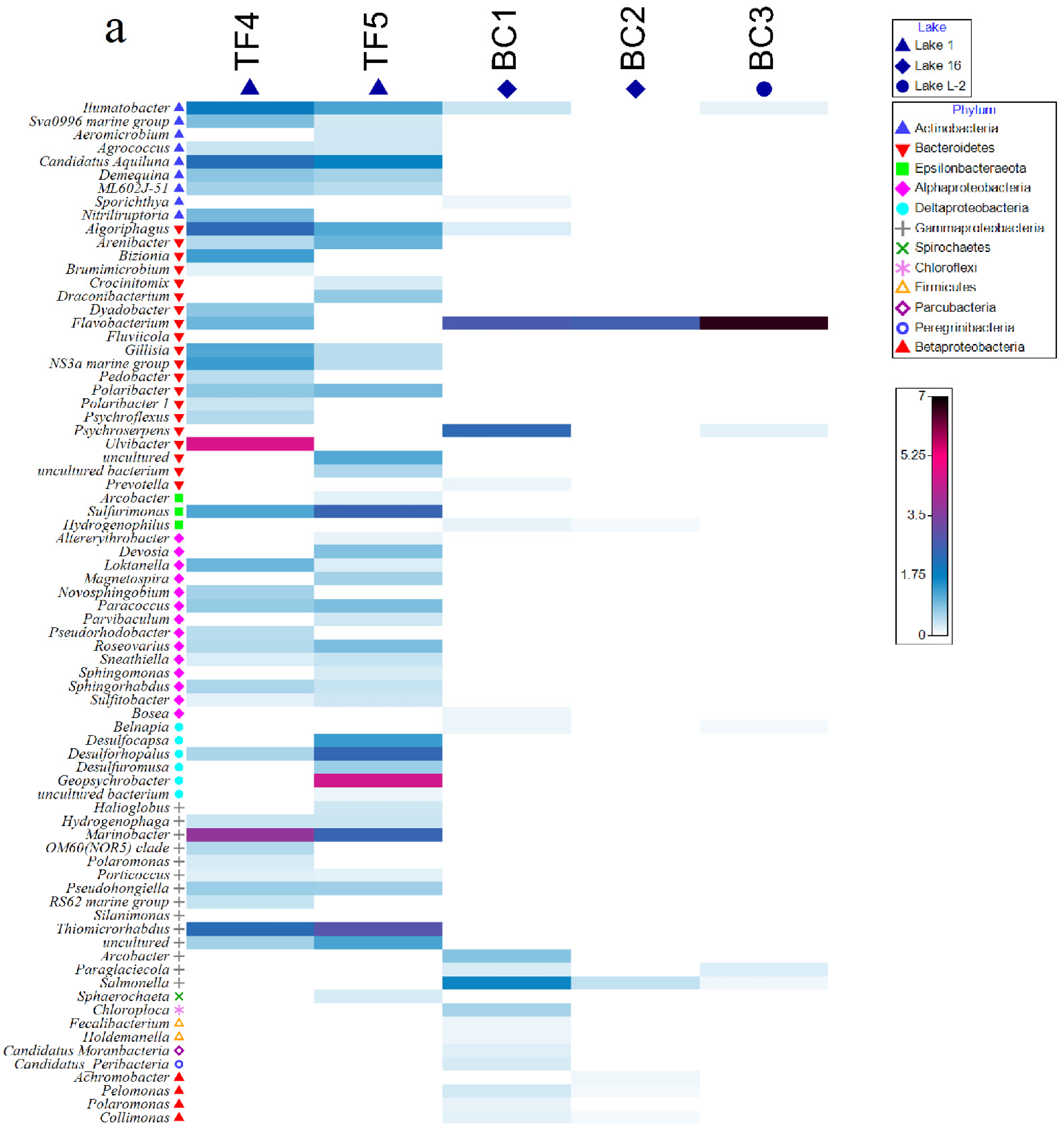
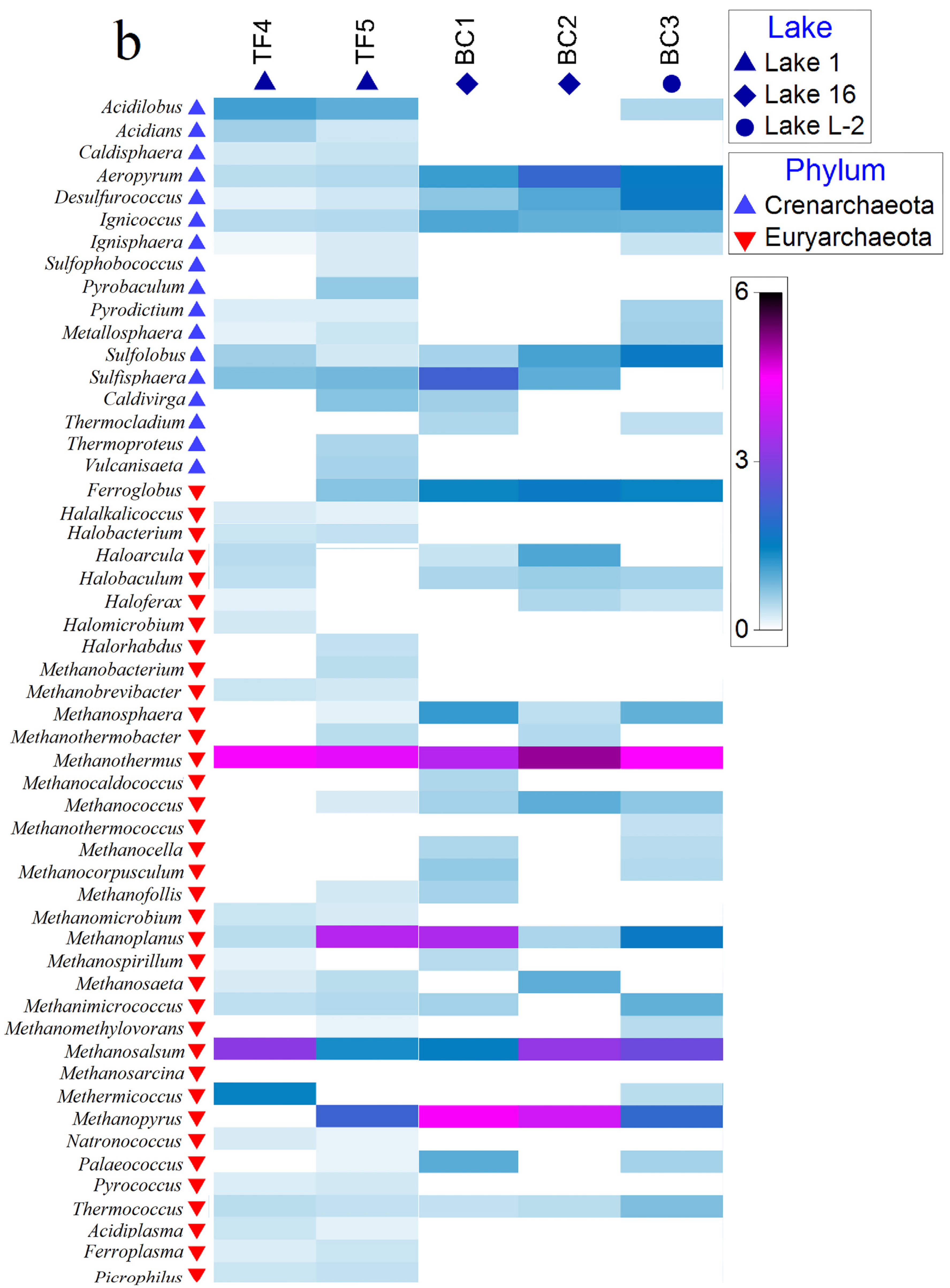
3.1.4. Prokaryotic Community Composition
3.1.5. Overall Comparison among Brine Samples with Respect to Previous Data
3.2. Predicted Functional Genes
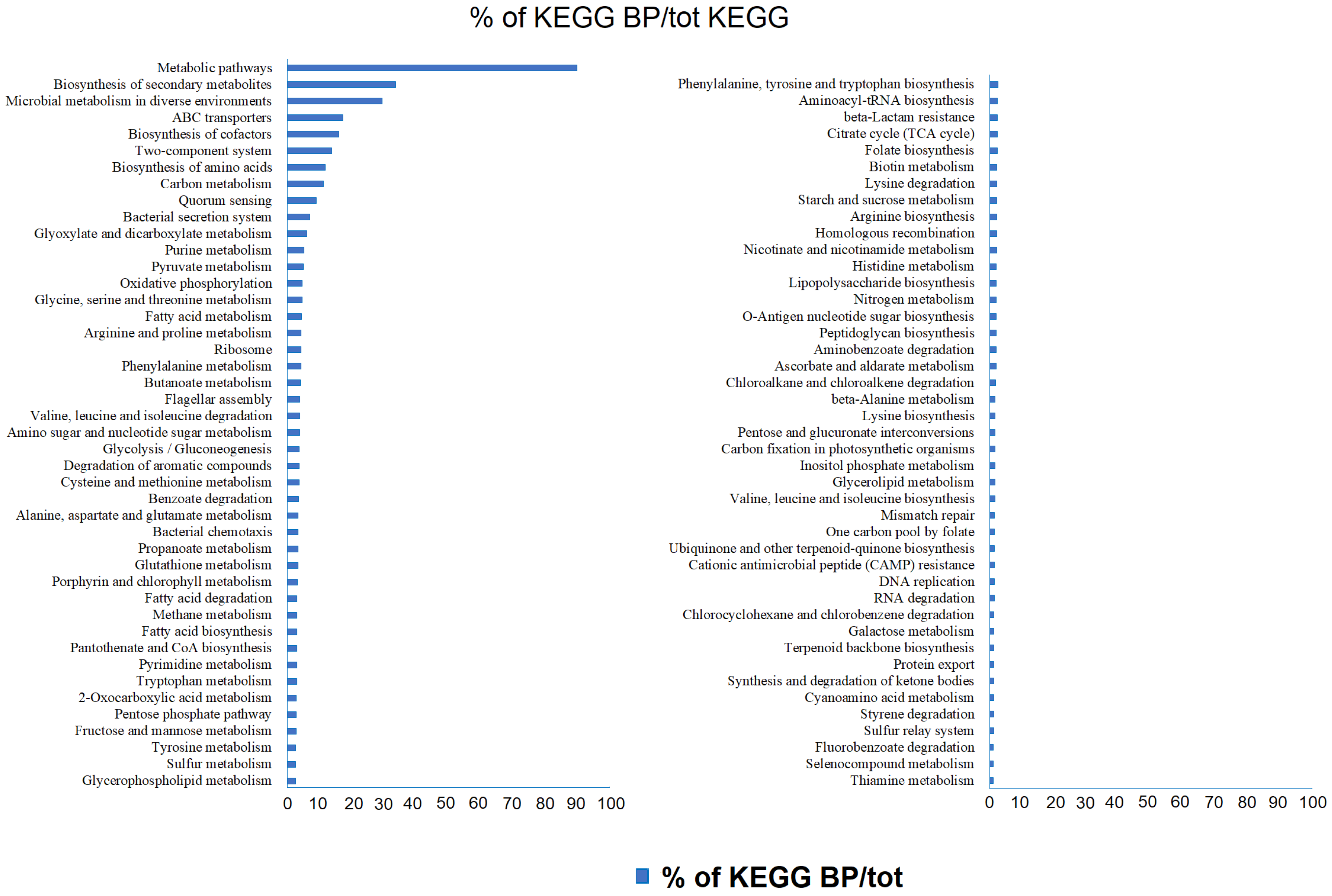
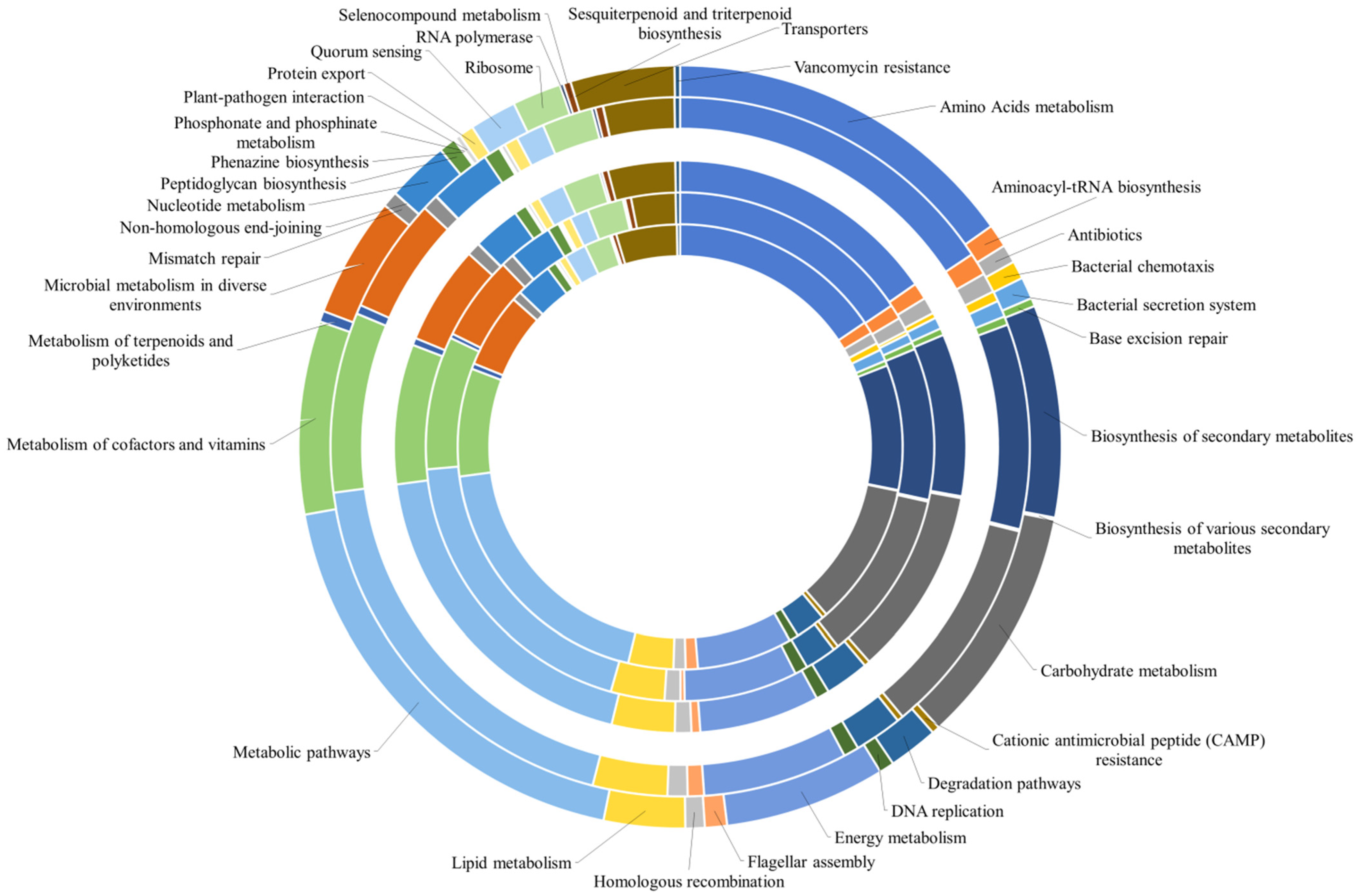
3.2.1. General Aspects
3.2.2. Metabolism of Amino Acids
3.2.3. Metabolism of Carbohydrates
3.2.4. Metabolism of Cofactors and Vitamins
3.2.5. Energy Metabolism
3.2.6. Transporters
3.2.7. Degradation Pathways
3.2.8. Biosynthesis of Secondary Metabolites
3.2.9. Microbial Metabolism in Diverse Environments
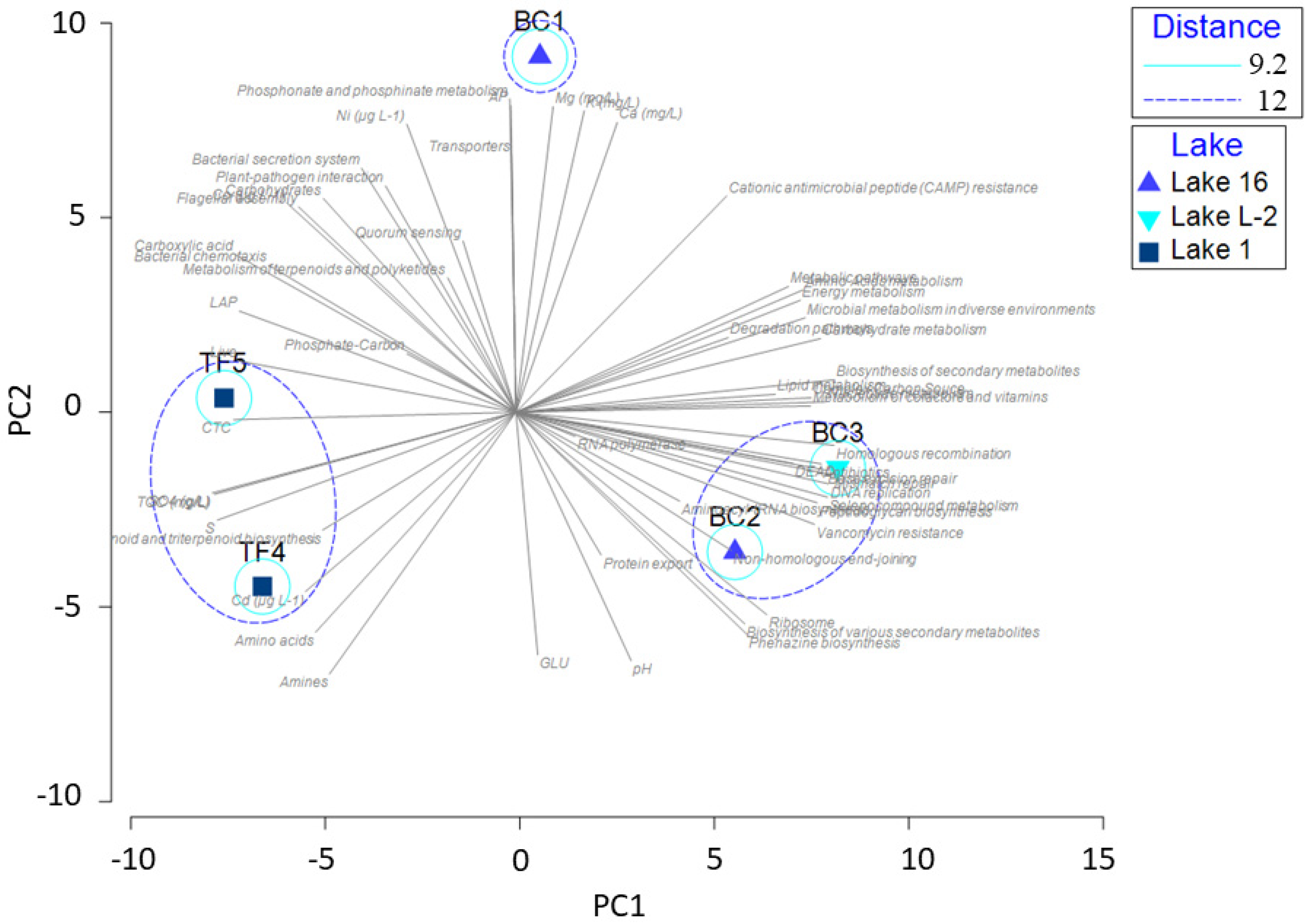
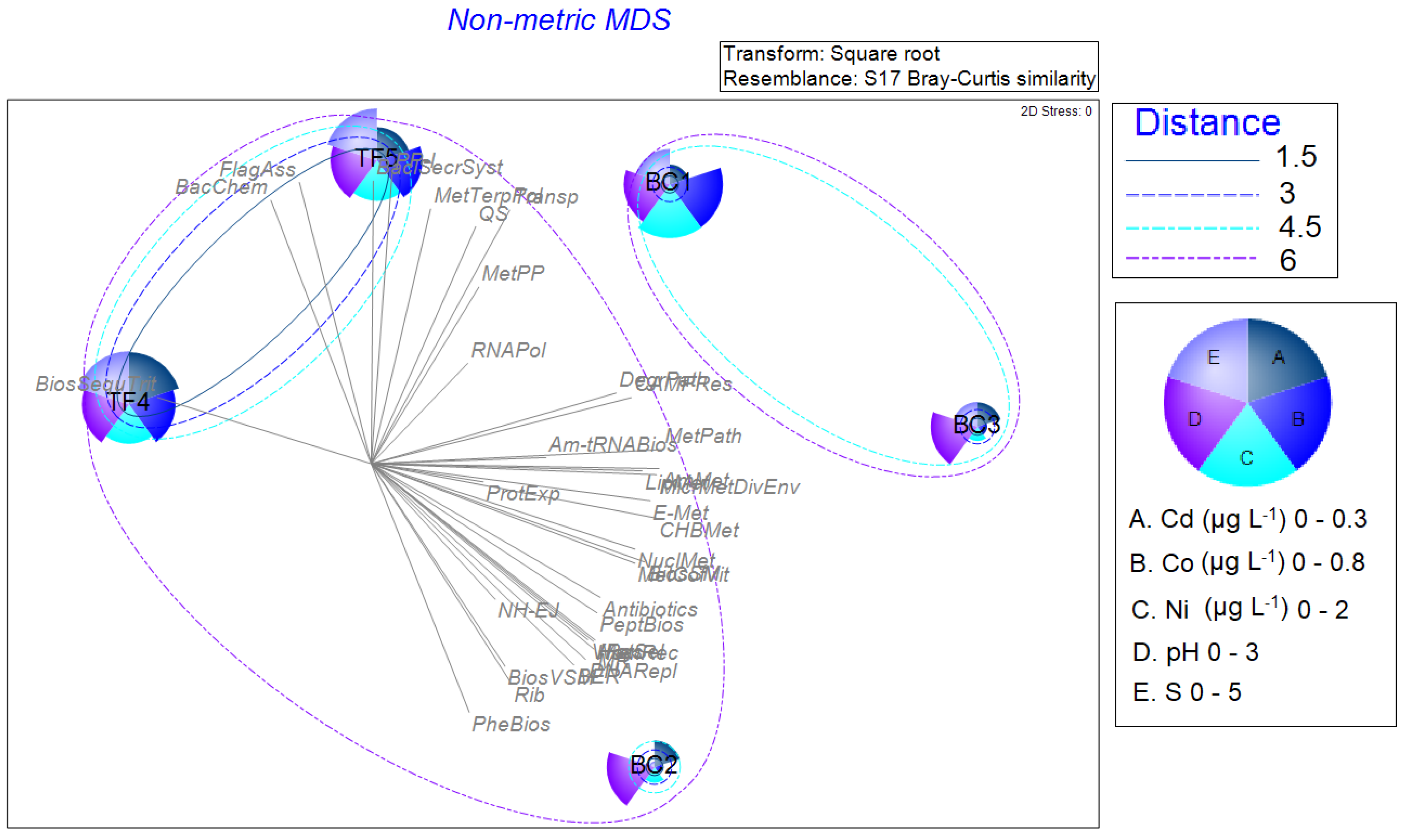
3.3. Statistical Analyses on the Entire Dataset
4. Discussion
4.1. Microbial Abundance, Biomass, and Morphometric Traits
4.2. Microbial Activities
4.3. Prokaryotic Community Compositions
4.4. Prediction of the Metabolic Profiles
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- French, H.M.; Guglielmin, M. Frozen ground phenomena in the vicinity of Terra Nova Bay, Northern Victoria Land, Antarctica: A preliminary report. Geogr. Ann. 2000, 82, 513–526. [Google Scholar] [CrossRef]
- Guglielmin, M.; Lewkowicz, A.G.; French, H.M.; Strini, A. Lake-ice blisters, Terra Nova Bay area, Northern Victoria Land, Antarctica. Geogr. Ann. Ser. A Phys. Geogr. 1999, 91, 99–111. [Google Scholar] [CrossRef]
- Borruso, L.; Sannino, C.; Selbmann, L.; Battistel, D.; Zucconi, L.; Azzaro, M.; Turchetti, B.; Buzzini, P.; Guglielmin, M. A thin ice layer segregates two distinct fungal communities in Antarctic brines from Tarn Flat (Northern Victoria Land). Sci. Rep. 2018, 8, 6582. [Google Scholar] [CrossRef]
- Papale, M.; Lo Giudice, A.; Conte, A.; Rizzo, C.; Rappazzo, C.; Maimone, G.; Caruso, G.; La Ferla, R.; Azzaro, M.; Gugliandolo, C.; et al. Microbial assemblages in pressurized Antarctic brine pockets (Tarn Flat, Northern Victoria Land): A hotspot of biodiversity and activity. Microorganisms 2019, 7, 333. [Google Scholar] [CrossRef]
- Azzaro, M.; Maimone, G.; La Ferla, R.; Cosenza, A.; Rappazzo, A.C.; Caruso, G.; Paranhos, R.; Cabral, A.S.; Forte, E.; Guglielmin, M. The prokaryotic community in an extreme Antarctic environment: The brines of Boulder Clay lakes (Northern Victoria Land). Hydrobiologia 2021, 848, 1837–1857. [Google Scholar] [CrossRef]
- Preston, L.; Dartnell, L. Planetary habitability: Lessons learned from terrestrial analogues. Int. J. Astrobiol. 2014, 13, 81–98. [Google Scholar] [CrossRef]
- Porcino, N.; Cosenza, A.; Azzaro, M. A review on the geochemistry of lakes in Victoria Land (Antarctica). Chemosphere 2020, 251, 126229. [Google Scholar] [CrossRef] [PubMed]
- Murray, A.E.; Kenig, F.; Fritsen, C.H.; McKay, C.P.; Cawley, K.M.; Edwards, R.; Kuhn, E.; McKnight, D.M.; Ostrom, N.E.; Peng, V.; et al. Microbial life at −13 °C in the brine of an ice-sealed Antarctic lake. Proc. Natl. Acad. Sci. USA 2012, 109, 20626–20631. [Google Scholar] [CrossRef] [PubMed]
- Dugan, H.A.; Doran, P.T.; Wagner, B.; Kenig, F.; Fritsen, C.H.; Arcone, S.A.; Kuhn, E.; Ostrom, N.E.; Warnock, J.P.; Murray, A.E. Stratigraphy of Lake Vida, Antarctica: Hydrologic implications of 27 m of ice. Cryosphere 2015, 9, 439–450. [Google Scholar] [CrossRef]
- Forte, E.; Dalle Fratte, M.; Azzaro, M.; Guglielmin, M. Pressurized brines in continental Antarctica as a possible analogue of Mars. Sci. Rep. 2016, 6, 33–58. [Google Scholar] [CrossRef] [PubMed]
- Rizzo, C.; Conte, A.; Azzaro, M.; Papale, M.; Rappazzo, A.C.; Battistel, D.; Roman, M.; Lo Giudice, A.; Guglielmin, M. Cultivable bacterial communities in brines from perennially ice-covered and pristine Antarctic lakes: Ecological and biotechnological implications. Microorganisms 2020, 8, 819. [Google Scholar] [CrossRef]
- Lo Giudice, A.; Conte, A.; Papale, M.; Rizzo, C.; Azzaro, M.; Guglielmin, M. Prokaryotic diversity and metabolically active communities in brines from two perennially ice-covered Antarctic lakes. Astrobiology 2021, 21, 551–565. [Google Scholar] [CrossRef] [PubMed]
- Oyewusi, H.A.; Wahab, R.A.; Edbeib, M.F.; Mohamad, M.A.N.; Hamid, A.A.A.; Kaya, Y.; Huyop, F. Functional profiling of bacterial communities in Lake Tuz using 16S rRNA gene sequences. Biotechnol. Biotechnol. Equip. 2021, 35, 1–10. [Google Scholar] [CrossRef]
- Sannino, C.; Borruso, L.; Mezzasoma, A.; Battistel, D.; Zucconi, L.; Selbmann, L.; Azzaro, M.; Onofri, S.; Turchetti, B.; Buzzini, P.; et al. Intra- and inter-cores fungal diversity suggests interconnection of different habitats in an Antarctic frozen lake (Boulder Clay, Northern Victoria Land). Environ. Microbiol. 2020, 22, 3463–3477. [Google Scholar] [CrossRef] [PubMed]
- Douglas, G.M.; Maffei, V.J.; Zaneveld, J.; Yurgel, S.N.; Brown, J.N.; Taylor, C.M.; Huttenhower, C.; Langille, M.G.I. PICRUSt2: An improved and extensible approach for metagenome inference. bioRxiv 2019, 672295. [Google Scholar] [CrossRef]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Thurber, R.L.V.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, 353–361. [Google Scholar] [CrossRef]
- Heberle, H.; Meirelles, G.V.; da Silva, F.R.; Telles, G.P.; Minghim, R. InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015, 16, 169. [Google Scholar] [CrossRef]
- La Ferla, R.; Azzaro, M.; Michaud, L.; Caruso, G.; Lo Giudice, A.; Paranhos, R.; Cabral, A.S.; Conte, A.; Cosenza, A.; Maimone, G.; et al. Prokaryotic abundance and activity in permafrost of the Northern Victoria Land and Upper Victoria Valley (Antarctica): A study case. Microb. Ecol. 2017, 74, 402–415. [Google Scholar] [CrossRef]
- Papale, M.; Conte, A.; Mikkonen, A.; Michaud, L.; La Ferla, R.; Azzaro, M.; Caruso, G.; Paranhos, R.; Cabral, A.S.; Maimone, G.; et al. Prokaryotic assemblages within permafrost active layer at Edmonson Point (Northern Victoria Land, Antarctica). Soil Biol. Biochem. 2018, 123, 165–179. [Google Scholar] [CrossRef]
- Kalcheva, H.; Beshkova, M.; Pehlivanov, L.; Kalcev, R. Bacterioplankton dynamics and the influence of environmental factors on it in the Srebarna Lake. In Proceedings of the The Third International Scientific Conference BALWOIS, Ohrid, Republic of Macedonia, 27–31 May 2008. [Google Scholar]
- Young, K.D. The selective value of bacterial shape. Microbiol. Mol. Biol. Rev. 2006, 70, 660–703. [Google Scholar] [CrossRef]
- Gentile, G.; Maimone, G.; La Ferla, R.; Azzaro, M.; Catalfamo, M.; Genovese, M.; Santisi, S.; Maldani, M.; Macrì, A.; Cappello, S. Phenotypic variations of Oleispira antarctica RB8T in different growth conditions. Curr. Microbiol. 2020, 77, 3414–3421. [Google Scholar] [CrossRef]
- Rezaeinejad, S.; Ivanov, V. Heterogeneity of Escherichia coli population by respiratory activity and membrane potential of cells during growth and long-term starvation. Microbiol. Res. 2011, 166, 129–135. [Google Scholar] [CrossRef]
- Posch, T.; Pernthaler, J.; Alfreider, A.; Psenner, R. Cell-specific respiratory activity of aquatic bacteria studied with the tetrazolium reduction method, cyto-clear slides, and image analysis. Appl. Environ. Microbiol. 1997, 63, 867–873. [Google Scholar] [CrossRef]
- Kenarova, A.; Encheva, M.; Chipeva, V. Physiological diversity of bacterial communities from different soil locations on Livingston Island, South Shetland archipelago, Antarctica. Pol. Biol. 2013, 36, 223–233. [Google Scholar] [CrossRef]
- Loferer-Krössbacher, M.; Klima, J.; Psenner, R. Determination of bacterial cell dry mass by transmission electron microscopy and densitometric image analysis. Appl. Environ. Microbiol. 1998, 64, 688–694. [Google Scholar] [CrossRef]
- Deming, J.W. Psychrophiles and polar regions. Curr. Opin. Microbiol. 2002, 5, 301–309. [Google Scholar] [CrossRef]
- Wagner, D.; Kobabe, S.; Liebner, S. Bacterial community structure and carbon turnover in permafrost-affected soils of the Lena Delta, North Eastern Siberia. Can. J. Microbiol. 2009, 55, 73–83. [Google Scholar] [CrossRef][Green Version]
- Laybourn-Parry, J.; Pearce, D.A. The biodiversity and ecology of Antarctic lakes: Models for evolution. Philos. Trans. R. Soc. B Biol. Sci. 2007, 362, 2273–2289. [Google Scholar] [CrossRef] [PubMed]
- Huston, A.L.; Methe, B.; Deming, J.W. Purification, characterization, and sequencing of an extracellular cold-active aminopeptidase produced by marine psychrophile Colwellia psychrerythraea strain 34H. Appl. Environ. Microbiol. 2004, 70, 3321–3328. [Google Scholar] [CrossRef] [PubMed]
- Ortega, G.; Lain, A.; Tadeo, X.; Lopez-Mendez, B.; Castano, D.; Millet, O. Halophilic enzyme activation induced by salts. Sci. Rep. 2011, 1, 6. [Google Scholar] [CrossRef]
- Karan, R.; Capes, M.D.; DasSarma, S. Function and biotechnology of extremophilic enzymes. Biochemistry 2011, 76, 686–693. [Google Scholar]
- Lei, F.; Zhao, Q.; Sun-Waterhouse, D.; Zhao, M. Characterization of a salt-tolerant aminopeptidase from marine Bacillus licheniformis SWJS33 that improves hydrolysis and debittering efficiency for soy protein isolate. Food Chem. 2016, 214, 347–353. [Google Scholar] [CrossRef]
- Salwan, R.; Sharma, V.; Chand Kasan, R.; Gulati, A. Bioprospecting psychrotrophic bacteria for serine-type proteases from the cold areas of the Western Himalayas. Curr. Microbiol. 2020, 77, 795–806. [Google Scholar] [CrossRef]
- Qin, Y.; Huang, Z.; Liu, Z. A novel cold-active and salt-tolerant α-amylase from marine bacterium Zunongwangia profunda: Molecular cloning, heterologous expression and biochemical characterization. Extremophiles 2014, 18, 271–281. [Google Scholar] [CrossRef]
- Wang, Q.F.; Miao, J.L.; Hou, Y.H.; Ding, Y.; Wang, G.D.; Li, G.Y. Purification and characterization of an extracellular cold-active serine protease from the psychrophilic bacterium Colwellia sp. NJ341. Biotechnol. Lett. 2005, 27, 1195–1198. [Google Scholar] [CrossRef]
- Tchigvintsev, A.; Tran, H.; Popovic, A.; Kovacic, F.; Brown, G.; Flick, R.; Hajighasemi, M.; Egorova, O.; Somody, J.C.; Tchigvintsev, D.; et al. The environment shapes microbial enzymes: Five cold-active and salt-resistant carboxylesterases from marine metagenomes. Appl. Microbiol. Biotechnol. 2015, 99, 2165–2178. [Google Scholar] [CrossRef] [PubMed]
- Cox, G.F.N.; Weeks, W.F. CRREL Report 82-30, Equations for determining the gas and brine volumes in sea ice samples. J. Glaciol. 1983, 29, 306–316. [Google Scholar] [CrossRef]
- Hoppe, H.G.; Arnosti, C.; Herndl, G.F. Ecological significance of bacterial enzymes in marine environment. In Microbial Enzymes in the Environment Activity, Ecology and Applications; Burns, R.C., Dick, R.P., Eds.; Marcel Dekker Inc.: New York, NY, USA, 2002; pp. 73–107. [Google Scholar]
- Maxwell, S.G. Acquisition, Degradation, and Cycling of Organic Matter within Sea-Ice Brines by Bacteria and Their Viruses. Ph.D. Thesis, University of Washington, Washington, DC, USA, 2020; pp. 1–168. [Google Scholar]
- Stedmon, C.A.; Thomas, D.N.; Papadimitriou, S.; Granskog, M.A.; Dieckmann, G.S. Using fluorescence to characterize dissolved organic matter in Antarctic sea ice brines. J. Geophys. Res. Biogeosci. 2011, 116, G03027. [Google Scholar] [CrossRef]
- Cox, M.; Lehninger, A.L.; Nelson, D.R. Lehninger Principles of Biochemistry; Worth Publishers: New York, NY, USA, 2000; pp. 306–308. [Google Scholar]
- Sun, S.; Jones, R.B.; Fodor, A.A. Inference-based accuracy of metagenome prediction tools varies across sample types and functional categories. Microbiome 2020, 8, 46. [Google Scholar] [CrossRef]
- Chistoserdova, L.; Laukel, M.; Portais, J.C.; Vorholt, J.A.; Lidstrom, M.E. Multiple formate dehydrogenase enzymes in the facultative methylotroph Methylobacterium extorquens AM1 are dispensable for growth on methanol. J. Bacteriol. 2004, 186, 22–28. [Google Scholar] [CrossRef]
- Stibal, M.; Wadham, J.L.; Lis, G.P.; Telling, J.; Pancost, R.D.; Dubnick, A.; Sharp, M.J.; Lawson, E.C.; Butler, C.E.H.; Hasan, F.; et al. Methanogenic potential of Arctic and Antarctic subglacial environments with contrasting organic carbon sources. Glob. Chang. Biol. 2012, 18, 3332–3345. [Google Scholar] [CrossRef]
- Ortiz, M.; Bosch, J.; Coclet, C.; Johnson, J.; Lebre, P.; Salawu-Rotimi, A.; Vikram, S.; Makhalanyane, T.; Cowan, D. Microbial nitrogen cycling in Antarctic soils. Microorganisms 2020, 8, 1442. [Google Scholar] [CrossRef] [PubMed]
- Santos, A.A.; Venceslau, S.S.; Grein, F.; Leavitt, W.D.; Dahl, C.; Johnston, D.T.; Pereira, I.A. A protein trisulfide couples dissimilatory sulfate reduction to energy conservation. Science 2015, 350, 1541–1545. [Google Scholar] [CrossRef] [PubMed]
- Griesbeck, C.; Hauska, G.; Schütz, M. Biological Sulfide-Oxidation: Sulfide-Quinone Reductase (SQR), the Primary Reaction. In Recent Research Developments in Microbiology; Pandalai, S.G., Ed.; Research Signpost: Trivadrum, India, 2000; Volume 4, pp. 129–203. [Google Scholar]
- Karr, E.A.; Sattley, W.M.; Jung, D.O.; Madigan, M.T.; Achenbach, L.A. Remarkable diversity of phototrophic purple bacteria in a permanently frozen Antarctic lake. Appl. Environ. Microbiol. 2003, 69, 4910–4914. [Google Scholar] [CrossRef]
- Karr, E.A.; Sattley, W.M.; Rice, M.R.; Jung, D.O.; Madigan, M.T.; Achenbach, L.A. Diversity and distribution of sulfate-reducing bacteria in permanently frozen Lake Fryxell, McMurdo Dry Valleys, Antarctica. Appl. Environ. Microbiol. 2005, 71, 6353–6359. [Google Scholar] [CrossRef]
- Christner, B.C.; Priscu, J.C.; Achberger, A.M.; Barbante, C.; Carter, S.P.; Christianson, K.; Michaud, A.B.; Mikucki, J.A.; Mitchell, A.C.; Skidmore, M.L.; et al. WISSARD Science Team. Subglacial Lake Whillans: A microbial ecosysytem beneath the West Antarctic Ice Sheet. Nature 2014, 512, 310–313. [Google Scholar] [CrossRef]
- Mikucki, J.; Pearson, A.; Johnston, D.T.; Turchyn, A.V.; Farquhar, J.; Schrag, D.P.; Anbar, A.D.; Priscu, J.C.; Lee, P.A. A contemporary microbially maintained subglacial ferrous ocean. Science 2009, 324, 397–400. [Google Scholar] [CrossRef]
- Sattley, W.M.; Madigan, M.T. Isolation, characterization, and ecology of cold-active, chemolithotrophic, sulfur-oxidizing bacteria from perennially ice-covered Lake Fryxell, Antarctica. Appl. Environ. Microbiol. 2006, 72, 5562–5568. [Google Scholar] [CrossRef]
- Sattley, W.M.; Madigan, M.T. Temperature and nutrient induced responses of Lake Fryxell sulfate-reducing prokaryotes and description of Desulfovibrio lacusfryxellense sp. nov., a pervasive, cold-active, sulfate-reducing bacterium from Lake Fryxell, Antarctica. Extremophiles 2010, 14, 357–366. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Papale, M.; Rizzo, C.; Caruso, G.; La Ferla, R.; Maimone, G.; Lo Giudice, A.; Azzaro, M.; Guglielmin, M. First Insights into the Microbiology of Three Antarctic Briny Systems of the Northern Victoria Land. Diversity 2021, 13, 323. https://doi.org/10.3390/d13070323
Papale M, Rizzo C, Caruso G, La Ferla R, Maimone G, Lo Giudice A, Azzaro M, Guglielmin M. First Insights into the Microbiology of Three Antarctic Briny Systems of the Northern Victoria Land. Diversity. 2021; 13(7):323. https://doi.org/10.3390/d13070323
Chicago/Turabian StylePapale, Maria, Carmen Rizzo, Gabriella Caruso, Rosabruna La Ferla, Giovanna Maimone, Angelina Lo Giudice, Maurizio Azzaro, and Mauro Guglielmin. 2021. "First Insights into the Microbiology of Three Antarctic Briny Systems of the Northern Victoria Land" Diversity 13, no. 7: 323. https://doi.org/10.3390/d13070323
APA StylePapale, M., Rizzo, C., Caruso, G., La Ferla, R., Maimone, G., Lo Giudice, A., Azzaro, M., & Guglielmin, M. (2021). First Insights into the Microbiology of Three Antarctic Briny Systems of the Northern Victoria Land. Diversity, 13(7), 323. https://doi.org/10.3390/d13070323









