Meiofauna Life on Loggerhead Sea Turtles-Diversely Structured Abundance and Biodiversity Hotspots That Challenge the Meiofauna Paradox
Abstract
1. Introduction
2. Material and Methods
2.1. Study Area and Epibiont Sampling
2.2. Meiofauna and Nematode Processing
2.3. Data Analyses
3. Results
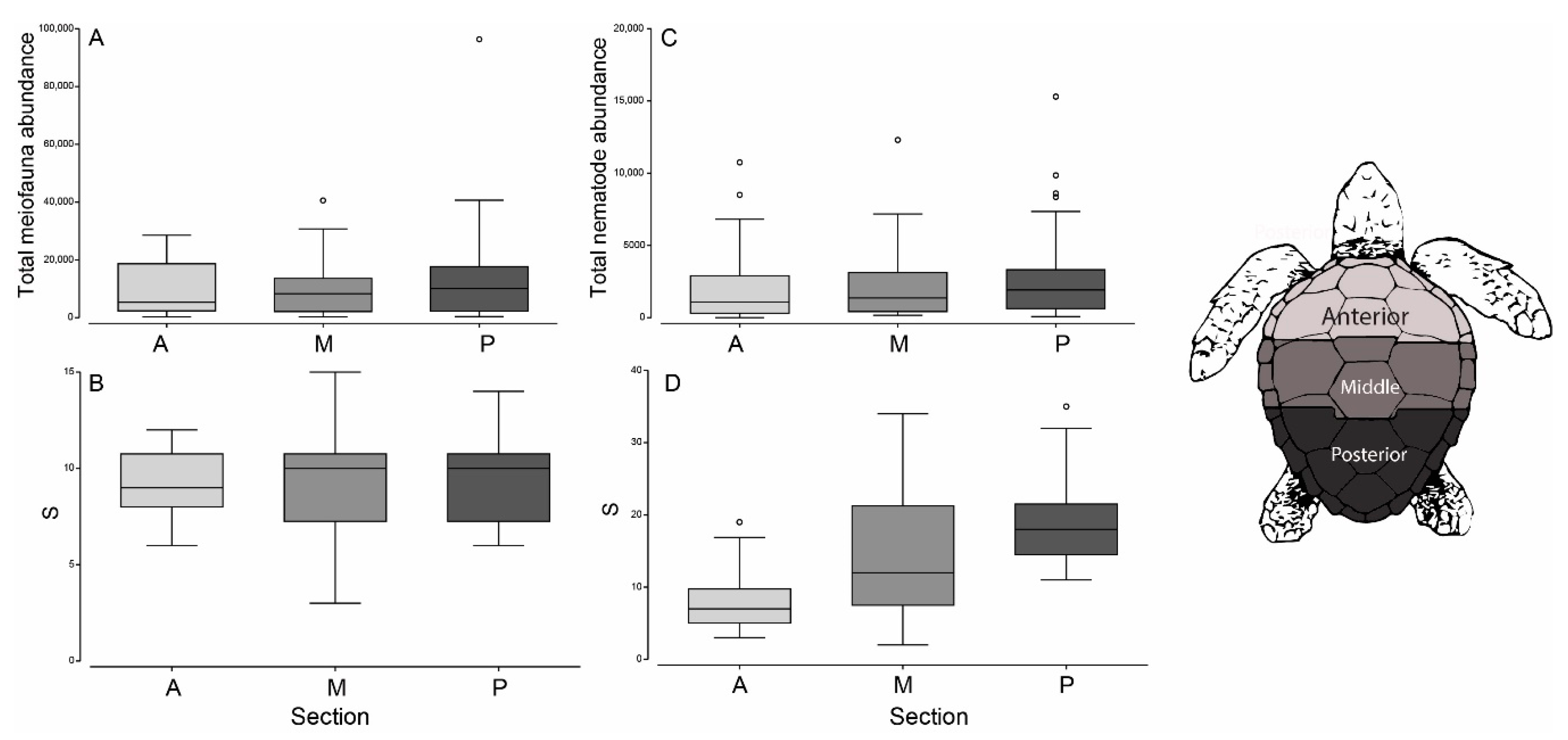
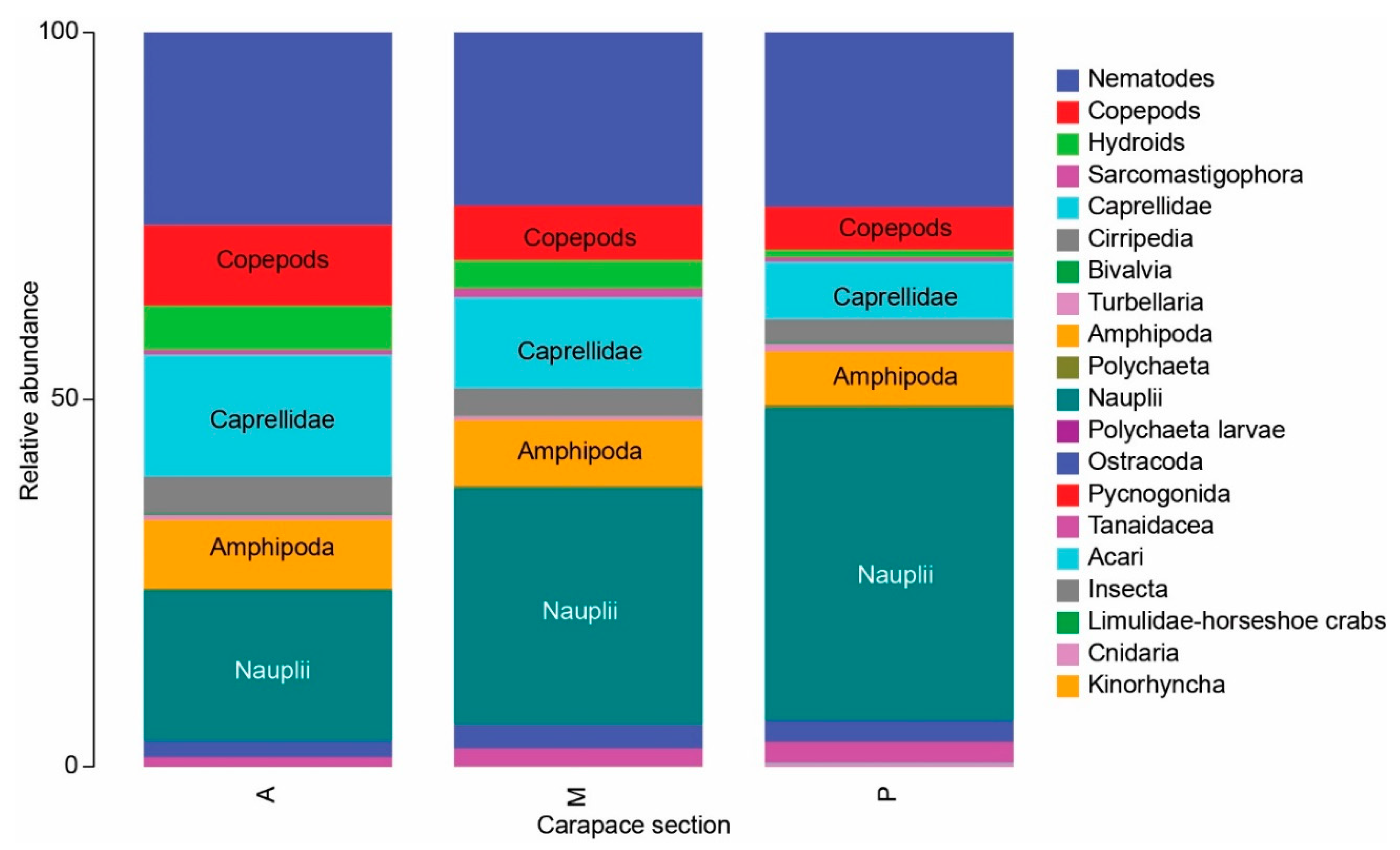
3.1. Meiofauna Communities
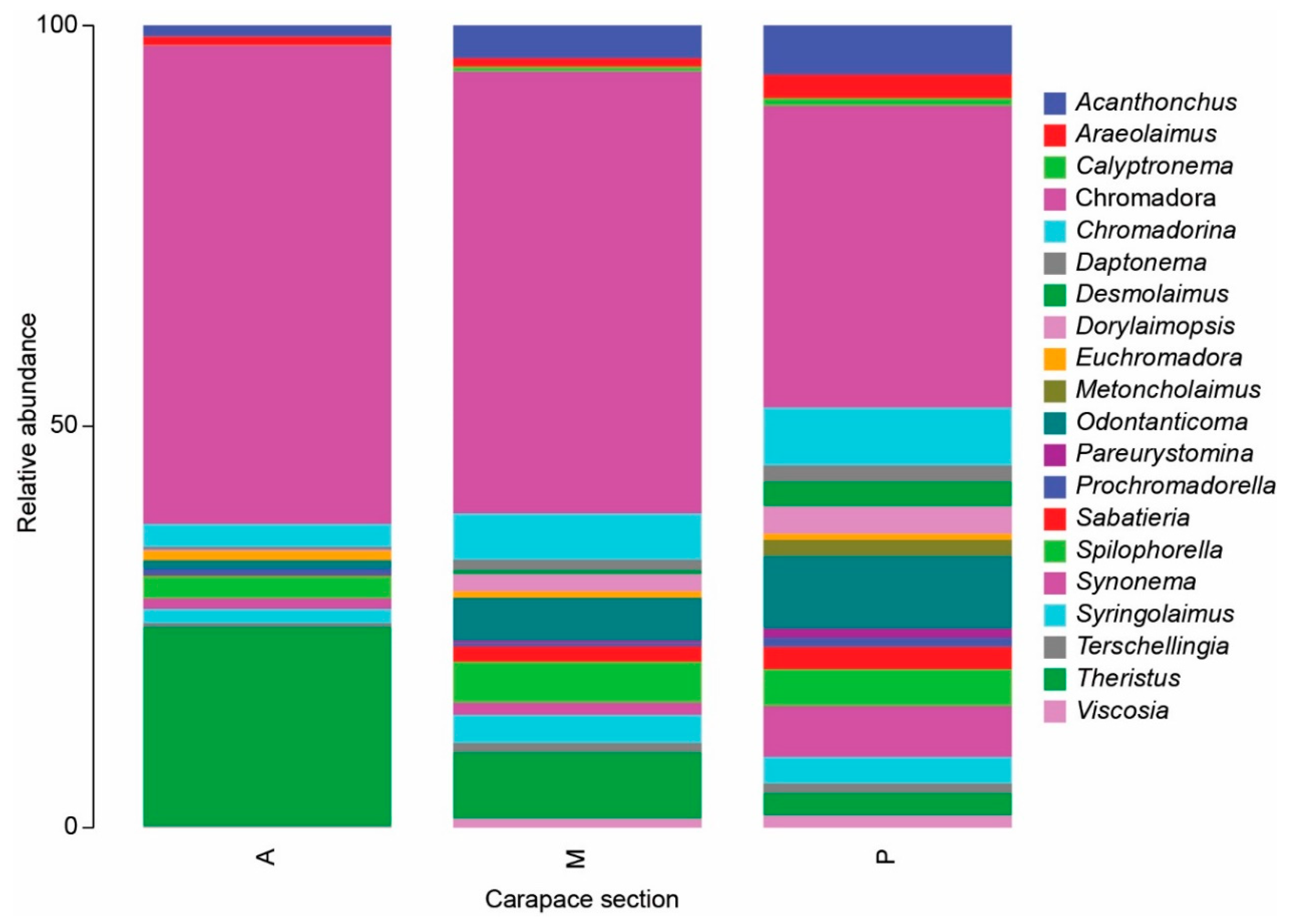
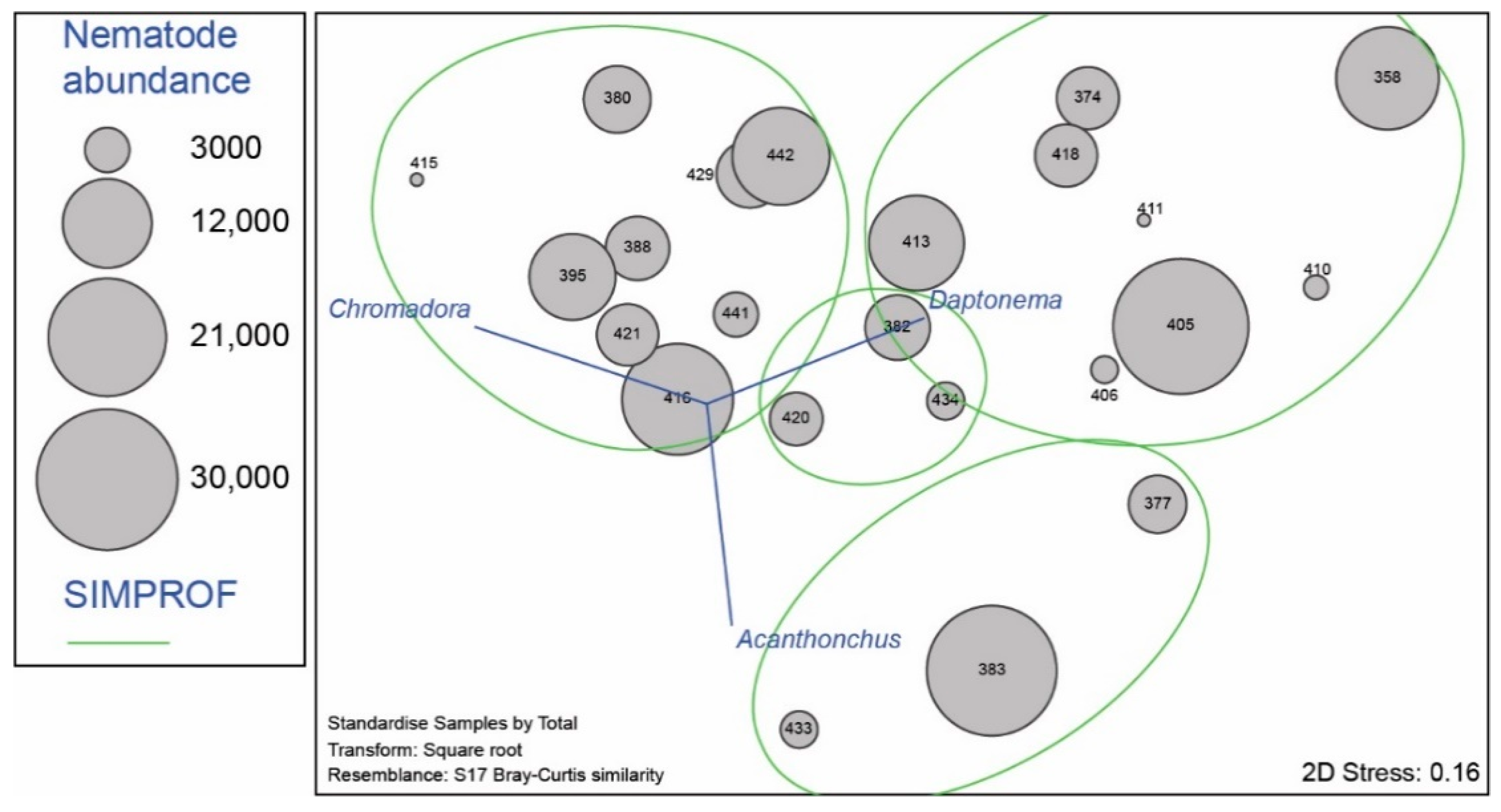
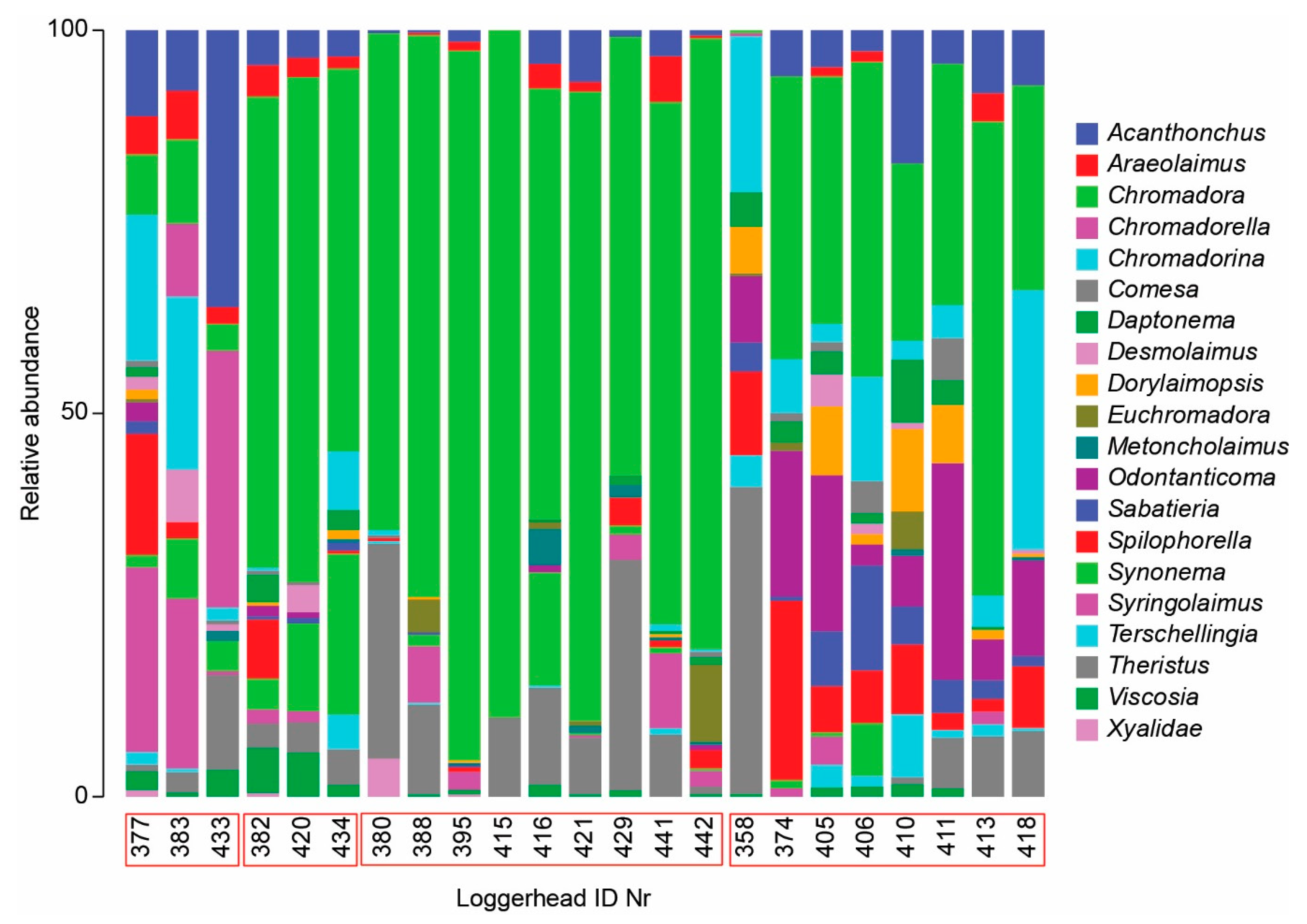
3.2. Nematode Communities
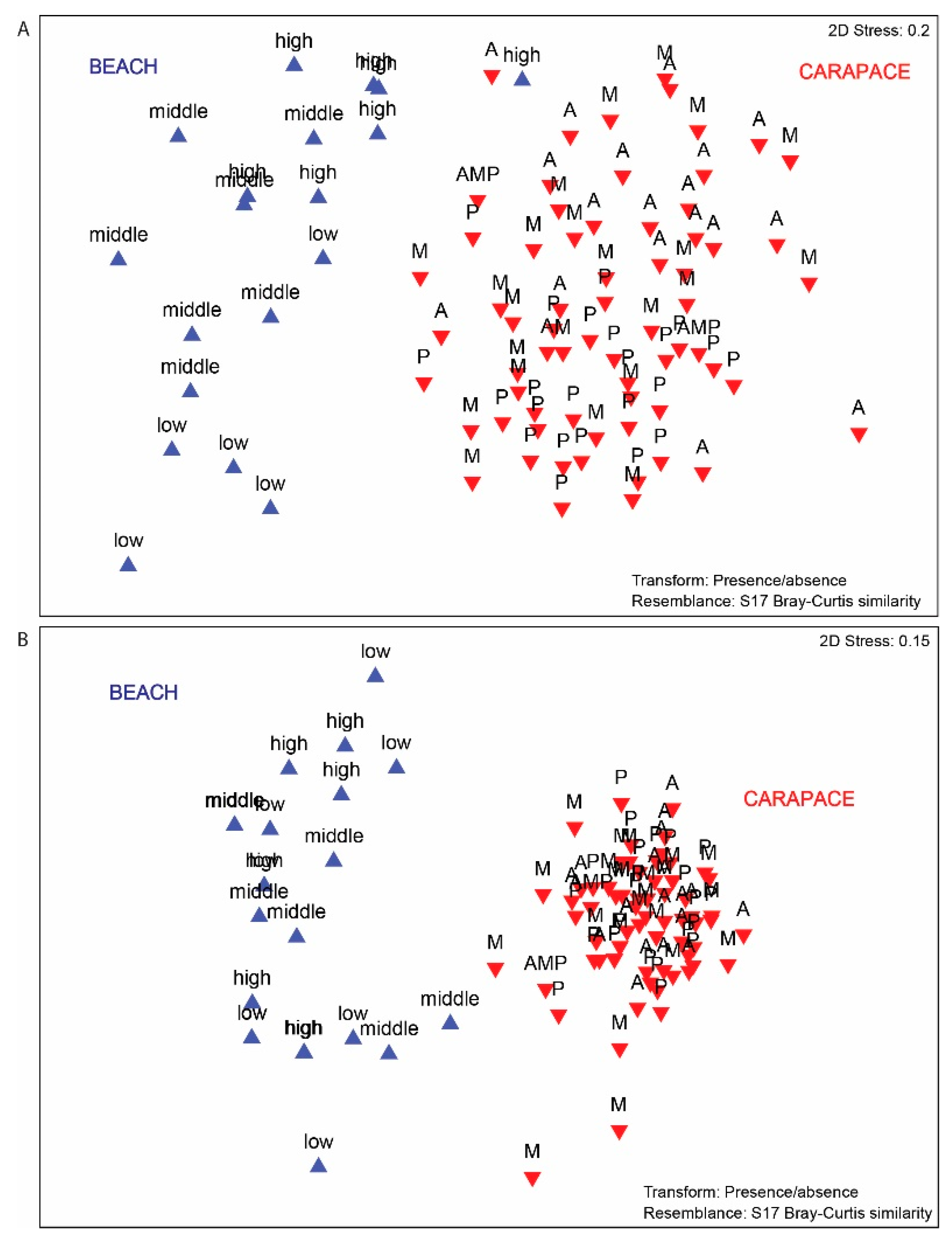
3.3. Beach vs. Carapace Samples
4. Discussion
4.1. Diversity and Structure of Meiofauna on Loggerhead Carapaces
4.2. Meiofauna Paradox
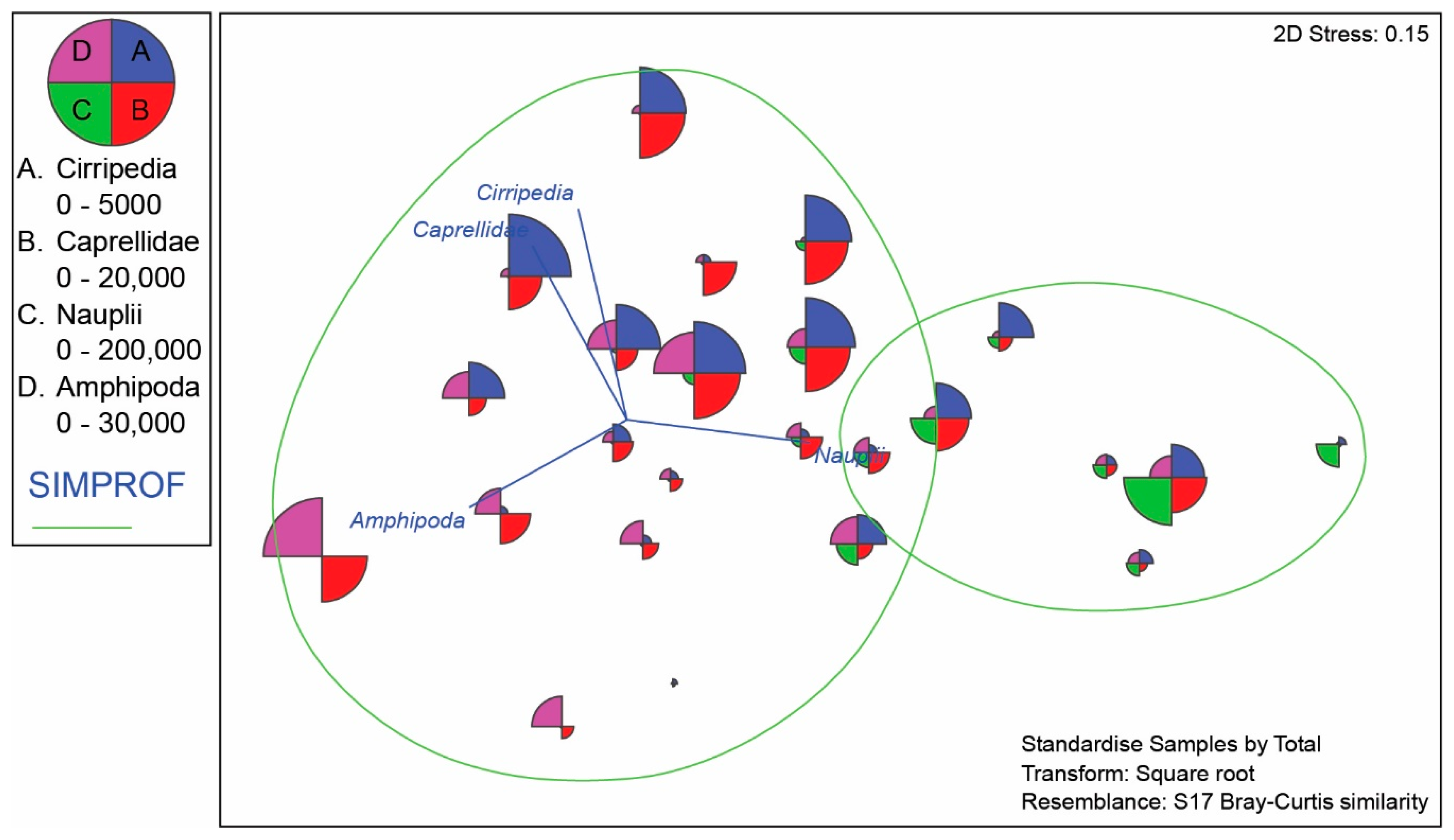
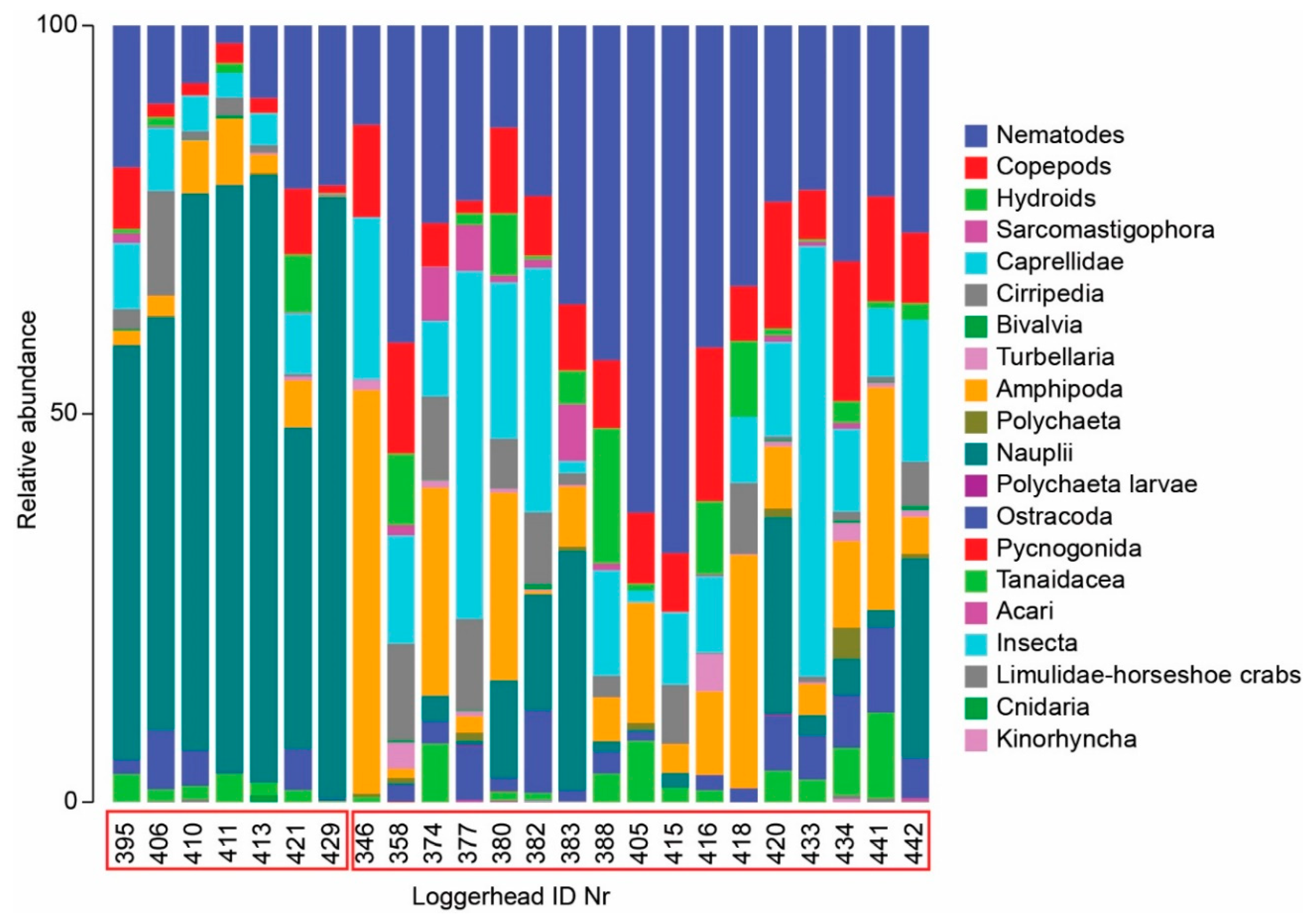
4.3. Meiofauna and Nematode Epibiont Community Structure across Turtle Individuals
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Frick, M.G.; Pfaller, J.B. Sea Turtle Epibiosis. In The Biology of Sea Turtles, Volume III; CRC Press: Boca Raton, FL, USA, 2013; pp. 399–426. [Google Scholar]
- Robinson, N.J.; Figgener, C.; Gatto, C.; Lazo-Wasem, E.A.; Paladino, F.V.; Tomillo, P.S.; Zardus, J.D.; Pinou, T. Assessing potential limitations when characterising the epibiota of marine megafauna: Effect of gender, sampling location, and inter-annual variation on the epibiont communities of olive ridley sea turtles. J. Exp. Mar. Biol. Ecol. 2017, 497, 71–77. [Google Scholar] [CrossRef]
- Correa, G.V.V.; Ingels, J.; Valdes, Y.V.; Fonseca-Genevois, V.G.; Farrapeira, C.M.R.; Santos, G.A.P. Diversity and composition of macro- and meiofaunal carapace epibionts of the hawksbill sea turtle (Eretmochelys imbricata Linnaeus, 1822) in Atlantic waters. Mar. Biodivers. 2014, 44, 391–401. [Google Scholar] [CrossRef]
- dos Santos, G.A.P.; Corrêa, G.V.V.; Valdes, Y.; Apolônio Silva de Oliveira, D.; Fonsêca-Genevois, V.G.; Silva, A.C.; Pontes, L.P.; Dolan, E.; Ingels, J. Eretmochelys imbricata shells present a dynamic substrate for a facilitative epibiont relationship between macrofauna richness and nematode diversity, structure and function. J. Exp. Mar. Biol. Ecol. 2018, 502, 153–163. [Google Scholar] [CrossRef]
- Dodd, C.K., Jr. Synopsis of the Biological Data on the Loggerhead Sea Turtle Caretta caretta (Linnaeus 1758); Florida Cooperative Fish and Wildlife Research Unit: Gainesville, FL, USA, 1988. [Google Scholar]
- Dodd, C.K.; Byles, R. Post-nesting movements and behavior of loggerhead sea turtles (Caretta caretta) departing from east-central Florida nesting beaches. Chelonian Conserv. Biol. 2003, 4, 530–536. [Google Scholar]
- Frick, M.G.; Williams, K.L.; Bolten, A.B.; Bjorndal, K.A.; Martins, H.R. Foraging ecology of oceanic-stage loggerhead turtles Caretta caretta. Endanger. Species Res. 2009, 9, 91–97. [Google Scholar] [CrossRef]
- Carr, A. So Excellent a Fishe: A Natural History of Sea Turtles; University Press of Florida: Ganinesville, FL, USA, 1967. [Google Scholar]
- Robinson, N.J.; Majewska, R.; Lazo-Wasem, E.A.; Nel, R.; Paladino, F.V.; Rojas, L.; Zardus, J.D.; Pinou, T. Epibiotic diatoms are universally present on all sea turtle species. PLoS ONE 2016, 11. [Google Scholar] [CrossRef]
- Schärer, M.T. A survey of the epibiota of Eretmochelys imbricata (Testudines: Cheloniidae) of Mona Island, Puerto Rico. Rev. Biol. Trop. 2003, 51, 87–90. [Google Scholar]
- Schratzberger, M.; Ingels, J. Meiofauna matters: The roles of meiofauna in benthic ecosystems. J. Exp. Mar. Biol. Ecol. 2018, 502, 12–25. [Google Scholar] [CrossRef]
- Baguley, J.; Coull, B.; Chandler, G. Meiobenthos. In Encyclopedia of Ocean Sciences; Cochran, J., Bokuniewicz, H., Yager, P., Eds.; Elsevier: Amsterdam, The Netherlands; Academic Press: Cambridge, MA, USA, 2019; Volume 2. [Google Scholar]
- Appeltans, W.; Ahyong, S.T.; Anderson, G.; Angel, M.V.; Artois, T.; Bailly, N.; Bamber, R.; Barber, A.; Bartsch, I.; Berta, A.; et al. The Magnitude of Global Marine Species Diversity. Curr. Biol. 2012, 22, 2189–2202. [Google Scholar] [CrossRef]
- Lambshead, P.J.D. Recent developments in marine benthic biodiversity research. Oceanis 1993, 19, 5–24. [Google Scholar]
- Lambshead, P.J.D. Marine Nematode Biodiversity. In Nematology: Advances and Perspectives Vol. 1: Nematode Morphology, Physiology and Ecology; Chen, Z.X., Chen, S.Y., Dickson, D.W., Eds.; CABI Publishing: London, UK, 2004; Volume 1, pp. 436–467. [Google Scholar]
- Lambshead, P.J.D.; Boucher, G. Marine nematode deep-sea biodiversity–hyperdiverse or hype? J. Biogeogr. 2003, 30, 475–485. [Google Scholar] [CrossRef]
- Ullberg, J.; Olafsson, E. Free-living marine nematodes actively choose habitat when descending from the water column. Mar. Ecol. Prog. Ser. 2003, 260, 141–149. [Google Scholar] [CrossRef]
- Ullberg, J.; Olafsson, E. Effects of biological disturbance by Monoporeia affinis (Amphipoda) on small-scale migration of marine nematodes in low-energy soft sediments. Mar. Biol. 2003, 143, 867–874. [Google Scholar] [CrossRef]
- Coomans, A. Nematode systematics: Past, present and future. Nematology 2000, 2, 3–7. [Google Scholar] [CrossRef]
- Derycke, S.; Remerie, T.; Backeljau, T.; Vierstraete, A.; Vanfleteren, J.; Vincx, M.; Moens, T. Phylogeography of the Rhabditis (Pellioditis) marina species complex: Evidence for long-distance dispersal, and for range expansions and restricted gene flow in the northeast Atlantic. Mol. Ecol. 2008, 17, 3306–3322. [Google Scholar] [CrossRef] [PubMed]
- Bhadury, P.; Austen, M.; Bilton, D.; Lambshead, P.; Rogers, A.; Smerdon, G. Evaluation of combined morphological and molecular techniques for marine nematode (Terschellingia spp.) identification. Mar. Biol. 2008, 154, 509–518. [Google Scholar] [CrossRef]
- Worsaae, K.; Kerbl, A.; Vang, Á.; Gonzalez, B.C. Broad North Atlantic distribution of a meiobenthic annelid–against all odds. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Giere, O. Meiobenthology: The Microscopic Motile Fauna of Aquatic Sediments, 2nd ed.; Springer: Berlin, Germany, 2009; p. 527. [Google Scholar]
- Cerca, J.; Purschke, G.; Struck, T.H. Marine connectivity dynamics: Clarifying cosmopolitan distributions of marine interstitial invertebrates and the meiofauna paradox. Mar. Biol. 2018, 165, 123. [Google Scholar] [CrossRef]
- Pfaller, J.B.; Bjorndal, K.A.; Reich, K.J.; Williams, K.L.; Frick, M.G. Distribution patterns of epibionts on the carapace of loggerhead turtles, Caretta caretta. Mar. Biodivers. Rec. 2008, 1. [Google Scholar] [CrossRef]
- National Marine Fisheries Service and US Fish and Wildlife Service. Recovery Plan for the Northwest Atlantic Population of the Loggerhead Sea Turtle (Caretta caretta), Second Revision; National Marine Fisheries Service: Silver Spring, MD, USA, 2008. [Google Scholar]
- Hart, K.M.; Lamont, M.M.; Fujisaki, I.; Tucker, A.D.; Carthy, R.R. Common coastal foraging areas for loggerheads in the Gulf of Mexico: Opportunities for marine conservation. Biol. Conserv. 2012, 145, 185–194. [Google Scholar] [CrossRef]
- Hart, K.M.; Lamont, M.M.; Sartain, A.R.; Fujisaki, I. Migration, Foraging, and Residency Patterns for Northern Gulf Loggerheads: Implications of Local Threats and International Movements. PLoS ONE 2014, 9, e103453. [Google Scholar] [CrossRef] [PubMed]
- Shamblin, B.M.; Bolten, A.B.; Abreu-Grobois, F.A.; Bjorndal, K.A.; Cardona, L.; Carreras, C.; Clusa, M.; Monzón-Argüello, C.; Nairn, C.J.; Nielsen, J.T. Geographic patterns of genetic variation in a broadly distributed marine vertebrate: New insights into loggerhead turtle stock structure from expanded mitochondrial DNA sequences. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, M.M.P.B.; Gredzens, C.; Bateman, B.L.; Boettcher, R.; Ceriani, S.A.; Godfrey, M.H.; Helmers, D.; Ingram, D.K.; Kamrowski, R.L.; Pate, M.; et al. Conservation hotspots for marine turtle nesting in the United States based on coastal development. Ecol. Appl. 2016, 26, 2708–2719. [Google Scholar] [CrossRef] [PubMed]
- Bolten, A.B. Techniques for Measuring Sea Turtles. In Research and Management Techniques for the Conservation of Sea Turtles; Eckert, K.L., Bjornal, K.A., Abreu-Grobois, F.A., Donnelly, M., Eds.; IUCN/SSC Marine Turtle Specialist Group Publication No.4: Washington, DC, USA, 1999; pp. 110–114. [Google Scholar]
- Yoder, M.; De Ley, I.T.; King, I.W.; Mundo-Ocampo, M.; Mann, J.; Blaxter, M.; Poiras, L.; De Ley, P. DESS: A versatile solution for preserving morphology and extractable DNA of nematodes. Nematology 2006, 8, 367–376. [Google Scholar] [CrossRef]
- Somerfield, P.J.; Warwick, R.M. Meiofauna Techniques. In Methods for the Study of Marine Benthos; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2013; pp. 253–284. [Google Scholar] [CrossRef]
- Higgins, R.P.; Thiel, H. Introduction to the Study of Meiofauna; Smithsonian Institution Press: London, UK, 1988. [Google Scholar]
- Somerfield, P.; Warwick, R. Meiofauna in Marine Pollution Monitoring Programmes: A Laboratory Manual; MAFF Directorate of Fisheries Research Technical Series: Lowestoft, UK, 1996; p. 71. [Google Scholar]
- Bain, O.; Baldwin, J.G.; Beveridge, I.; Bezerra, T.C.; Braeckman, U.; Coomans, A.; Decraemer, W.; Derycke, S.; Durette-Desset, M.-C.; Fonseca, G. Nematoda; Walter de Gruyter: Berlin, Germany, 2014; Volume 2. [Google Scholar]
- Guilini, K.; Bezerra, T.N.; Eisendle-Flöckner, U.; Deprez, T.; Fonseca, G.; Holovachov, O.; Leduc, D.; Miljutin, D.; Moens, T.; Sharma, J.; et al. NeMys: World Database of Free-Living Marine Nematodes. Available online: http://nemys.ugent.be (accessed on 1 May 2018).
- Platt, H.M.; Warwick, R.M. Free-Living Marine Nematodes. Part II. British Chromadorids: Pictorial Key to World Genera and Notes for the Identification of British Species. Synop. British Fauna 1988, 38, 502. [Google Scholar]
- Hurlbert, S.H. The nonconcept of species diversity: A critique and alternative parameters. Ecology 1971, 52, 577–586. [Google Scholar] [CrossRef]
- Hill, M.O. Diversity and evenness: A unifying notation and its consequences. Ecology 1973, 54, 427–432. [Google Scholar] [CrossRef]
- Heip, C.; Herman, P.; Soetaert, K. Indices of Diversity and Evenness. Oceanis 1998, 24, 61–87. [Google Scholar]
- Clarke, K.R.; Gorley, R.N. PRIMER v7: User Manual/Tutorial; PRIMER-E: Plymouth, UK, 2015; p. 296. [Google Scholar]
- Somerfield, P.J.; Clarke, K.R. Inverse analysis in non-parametric multivariate analyses: Distinguishing groups of associated species which covary coherently across samples. J. Exp. Mar. Biol. Ecol. 2013, 449, 261–273. [Google Scholar] [CrossRef]
- Anderson, M.J.; Gorley, R.N.; Clarke, K.R. PERMANOVA+ for PRIMER: Guide to Software and Statistical Methods; PRIMER-E Ltd.: Plymouth, UK, 2008; p. 214. [Google Scholar]
- De Oliveira, D.A.; Derycke, S.; Da Rocha, C.M.; Barbosa, D.F.; Decraemer, W.; Dos Santos, G.A. Spatiotemporal variation and sediment retention effects on nematode communities associated with Halimeda opuntia (Linnaeus) Lamouroux (1816) and Sargassum polyceratium Montagne (1837) seaweeds in a tropical phytal ecosystem. Mar. Biol. 2016, 163, 102. [Google Scholar] [CrossRef]
- Gobin, J.F. Free-living marine nematodes of hard bottom substrates in Trinidad and Tobago, West Indies. Bull. Mar. Sci. 2007, 81, 73–84. [Google Scholar]
- Moore, P. The nematode fauna associated with holdfasts of kelp (Laminaria hyperborea) in north-east Britain. J. Mar. Biol. Assoc. U. K. 1971, 51, 589–604. [Google Scholar] [CrossRef]
- Kito, K. Phytal marine nematode assemblage on Sargassum muticum Agardh with reference to the structure and seasonal. Mar. Ecol.-Prog. Ser. 1982, 37, 19. [Google Scholar]
- Da Rocha, C.M.C.; Venekey, V.; Bezerra, T.N.C.; Souza, J.R.B. Phytal Marine Nematode Assemblages and their Relation with the Macrophytes Structural Complexity in a Brazilian Tropical Rocky Beach. Hydrobiologia 2006, 553, 219–230. [Google Scholar] [CrossRef]
- Derycke, S.; Van Vynckt, R.; Vanoverbeke, J.; Vincx, M.; Moens, T. Colonization patterns of Nematoda on decomposing algae in the estuarine environment: Community assembly and genetic structure of the dominant species Pellioditis marina. Limnol. Oceanogr. 2007, 52, 992–1001. [Google Scholar] [CrossRef]
- Gwyther, J.; Fairweather, P.G. Colonisation by epibionts and meiofauna of real and mimic pneumatophores in a cool temperate mangrove habitat. Mar. Ecol.-Prog. Ser. 2002, 229, 137–149. [Google Scholar] [CrossRef]
- Zhinan, Z. Phytal meiofauna of a rocky shore at the Cape d’Aguilar marine reserve, Hong Kong. Mar. Flora Fauna Hong Kong South. China IV 1997, 4, 205. [Google Scholar]
- Atilla, N.; Wetzel, M.A.; Fleeger, J.W. Abundance and colonization potential of artificial hard substrate-associated meiofauna. J. Exp. Mar. Biol. Ecol. 2003, 287, 273–287. [Google Scholar] [CrossRef]
- Raes, M.; De Troch, M.; Ndaro, S.G.M.; Muthumbi, A.; Guilini, K.; Vanreusel, A. The structuring role of microhabitat type in coral degradation zones: A case study with marine nematodes from Kenya and Zanzibar. Coral Reefs 2007, 26, 113–126. [Google Scholar] [CrossRef]
- Raes, M.; Decraemer, W.; Vanreusel, A. Walking with worms: Coral-associated epifaunal nematodes. J. Biogeogr. 2008, 35, 2207–2222. [Google Scholar] [CrossRef]
- Fonsêca-Genevois, V.D.; Somerfield, P.J.; Neves, M.H.B.; Coutinho, R.; Moens, T. Colonization and early succession on artificial hard substrata by meiofauna. Mar. Biol. 2006, 148, 1039–1050. [Google Scholar] [CrossRef][Green Version]
- Copley, J.T.P.; Flint, H.C.; Ferrero, T.J.; Van Dover, C.L. Diversity of meiofauna and free-living nematodes in hydrothermal vent mussel beds on the northern and southern East Pacific Rise. J. Mar. Biol. Assoc. U. K. 2007, 87, 1141–1152. [Google Scholar] [CrossRef][Green Version]
- Jensen, P. Ecology of benthic and epiphytic nematodes in brackish waters. Hydrobiologia 1984, 108, 201–217. [Google Scholar] [CrossRef]
- Zekely, J.; Gollner, S.; Van Dover, C.L.; Govenar, B.; Le Bris, N.; Nemeschkal, H.L.; Bright, M. Nematode communities associated with tubeworm and mussel aggregations on the East Pacific Rise. Cah. Biol. Mar. 2006, 47, 477–482. [Google Scholar]
- Trotter, D.; Webster, J. Distribution and abundance of marine nematodes on the kelp Macrocystic integrifolia. Mar. Biol. 1983, 78, 39–43. [Google Scholar] [CrossRef]
- Coull, B.; Creed, E.; Eskin, R.; Montagna, P.; Palmer, M.; Wells, J. Phytal Meiofauna from the Rocky Intertidal at Murrells Inlet, South Carolina/Colman, J. 1940. On the fauna inhabiting intertidal seaweeds. J. Mar. Biol. Assoc. UK, 24: 129-183. Trans. Am. Microsc. Soc. 1983, 102, 380–389. [Google Scholar] [CrossRef]
- Traunspurger, W.; Rothhaupt, K.-O.; Peters, L.; Wetzel, M. Community development of free-living aquatic nematodes in littoral periphyton communities. Nematology 2005, 7, 901–916. [Google Scholar] [CrossRef]
- Meylan, A. Spongivory in hawksbill turtles: A diet of glass. Science 1988, 239, 393–395. [Google Scholar] [CrossRef]
- Fujii, J.A.; McLeish, D.; Brooks, A.J.; Gaskell, J.; Van Houtan, K.S. Limb-use by foraging marine turtles, an evolutionary perspective. PeerJ 2018, 6, e4565. [Google Scholar] [CrossRef]
- Preen, A.R. Infaunal mining: A novel foraging method of loggerhead turtles. J. Herpetol. 1996, 30, 94–96. [Google Scholar] [CrossRef]
- Derycke, S.; Backeljau, T.; Moens, T. Dispersal and gene flow in free-living marine nematodes. Front. Zool. 2013, 10, 1. [Google Scholar] [CrossRef] [PubMed]
- Derycke, S.; De Ley, P.; Tandingan De Ley, I.; Holovachov, O.; Rigaux, A.; Moens, T. Linking DNA sequences to morphology: Cryptic diversity and population genetic structure in the marine nematode Thoracostoma trachygaster (Nematoda, Leptosomatidae). Zool. Scr. 2010, 39, 276–289. [Google Scholar] [CrossRef]
- Boeckner, M.; Sharma, J.; Proctor, H. Revisiting the meiofauna paradox: Dispersal and colonization of nematodes and other meiofaunal organisms in low-and high-energy environments. Hydrobiologia 2009, 624, 91–106. [Google Scholar] [CrossRef]
- de Oliveira, D.A.S.; Decraemer, W.; Moens, T.; dos Santos, G.A.P.; Derycke, S. Low genetic but high morphological variation over more than 1000 km coastline refutes omnipresence of cryptic diversity in marine nematodes. BMC Evol. Biol. 2017, 17, 71. [Google Scholar]
- Lins, L.; Vanreusel, A.; van Campenhout, J.; Ingels, J. Selective settlement of deep-sea canyon nematodes after resuspension—An experimental approach. J. Exp. Mar. Biol. Ecol. 2013, 441, 110–116. [Google Scholar] [CrossRef]
- Ullberg, J.; Zoologiska, I.; Stockholms, U. Dispersal in Free-Living, Marine, Benthic Nematodes: Passive or Active Processes? Zoologiska Institutionen University: Stockholm, Sweden, 2004. [Google Scholar]
- Plotkin, P.T.; Spotila, J.R. Post-nesting migrations of loggerhead turtles Caretta caretta from Georgia, USA: Conservation implications for a genetically distinct subpopulation. Oryx 2002, 36, 396–399. [Google Scholar] [CrossRef]
- Marcovaldi, M.Â.; Lopez, G.G.; Soares, L.S.; Lima, E.H.; Thomé, J.C.; Almeida, A.P. Satellite-tracking of female loggerhead turtles highlights fidelity behavior in northeastern Brazil. Endanger. Species Res. 2010, 12, 263–272. [Google Scholar] [CrossRef][Green Version]
- Hawkes, L.A.; Witt, M.J.; Broderick, A.C.; Coker, J.W.; Coyne, M.S.; Dodd, M.; Frick, M.G.; Godfrey, M.H.; Griffin, D.B.; Murphy, S.R. Home on the range: Spatial ecology of loggerhead turtles in Atlantic waters of the USA. Divers. Distrib. 2011, 17, 624–640. [Google Scholar] [CrossRef]
- Foley, A.M.; Schroeder, B.A.; Hardy, R.; MacPherson, S.L.; Nicholas, M. Long-term behavior at foraging sites of adult female loggerhead sea turtles (Caretta caretta) from three Florida rookeries. Mar. Biol. 2014, 161, 1251–1262. [Google Scholar] [CrossRef]
- Griffin, D.B.; Murphy, S.R.; Frick, M.G.; Broderick, A.C.; Coker, J.W.; Coyne, M.S.; Dodd, M.G.; Godfrey, M.H.; Godley, B.J.; Hawkes, L.A. Foraging habitats and migration corridors utilized by a recovering subpopulation of adult female loggerhead sea turtles: Implications for conservation. Mar. Biol. 2013, 160, 3071–3086. [Google Scholar] [CrossRef]
- Girard, C.; Tucker, A.D.; Calmettes, B. Post-nesting migrations of loggerhead sea turtles in the Gulf of Mexico: Dispersal in highly dynamic conditions. Mar. Biol. 2009, 156, 1827–1839. [Google Scholar] [CrossRef]
- SWOT. The State of the World’s Sea Turtles. 2020, p. 54. Available online: www.seaturtlestatus.org (accessed on 1 April 2020).
- Wallace, B.P.; DiMatteo, A.D.; Bolten, A.B.; Chaloupka, M.Y.; Hutchinson, B.J.; Abreu-Grobois, F.A.; Mortimer, J.A.; Seminoff, J.A.; Amorocho, D.; Bjorndal, K.A.; et al. Global Conservation Priorities for Marine Turtles. PLoS ONE 2011, 6, e24510. [Google Scholar] [CrossRef] [PubMed]
- Robinson, N.J.; Lazo-Wasem, E.A.; Paladino, F.V.; Zardus, J.D.; Pinou, T. Assortative epibiosis of leatherback, olive ridley and green sea turtles in the Eastern Tropical Pacific. J. Mar. Biol. Assoc. U. K. 2017, 97, 1233–1240. [Google Scholar] [CrossRef]
- Pereira, S.; Lima, E.; Ernesto, L.; Mathews, H.; Ventura, A. Epibionts associated with Chelonia mydas from northern Brazil. Mar. Turt. Newsl. 2006, 111, 17–18. [Google Scholar]
- Casale, P.; D’Addario, M.; Freggi, D.; Argano, R. Barnacles (Cirripedia, Thoracica) and Associated Epibionts from Sea Turtles in the Central Mediterranean. Crustaceana 2012, 85, 533–549. [Google Scholar] [CrossRef]
- Limpus, C. Global overview of the status of marine turtles: A 1995 viewpoint. Biol. Conserv. Sea Turt. 1995, 2, 605–609. [Google Scholar]
- Ross, J.P. Historical decline of loggerhead, ridley, and leatherback sea turtles. In Biology and Conservation of Sea Turtles; Smithsonian Institution Press: Washington, DC, USA, 1982; pp. 189–195. [Google Scholar]
- Hays, G.C. Good news for sea turtles. Trends Ecol. Evol. 2004, 19, 349–351. [Google Scholar] [CrossRef]
- Hirayama, R. Oldest known sea turtle. Nature 1998, 392, 705–708. [Google Scholar] [CrossRef]
- Harvey, V.L.; LeFebvre, M.J.; deFrance, S.D.; Toftgaard, C.; Drosou, K.; Kitchener, A.C.; Buckley, M. Preserved collagen reveals species identity in archaeological marine turtle bones from Caribbean and Florida sites. R. Soc. Open Sci. 2019, 6, 191137. [Google Scholar] [CrossRef]
- Bowen, B.W.; Nelson, W.S.; Avise, J.C. A molecular phylogeny for marine turtles: Trait mapping, rate assessment, and conservation relevance. Proc. Natl. Acad. Sci. USA 1993, 90, 5574–5577. [Google Scholar] [CrossRef]
- Miller, J.D. Reproduction in sea turtles. Biol. Sea Turt. 1997, 1, 51–82. [Google Scholar]
- Pfaller, J.B.; Alfaro-Shigueto, J.; Balazs, G.H.; Ishihara, T.; Kopitsky, K.; Mangel, J.C.; Peckham, S.H.; Bolten, A.B.; Bjorndal, K.A. Hitchhikers reveal cryptic host behavior: New insights from the association between Planes major and sea turtles in the Pacific Ocean. Mar. Biol. 2014, 161, 2167–2178. [Google Scholar] [CrossRef]
- Hosono, T.; Minami, H. Stable isotope analysis of epibiotic caprellids (Amphipoda) on loggerhead turtles provides evidence of turtle’s feeding history. In New Frontiers in Crustacean Biology; Brill: Leiden, The Netherlands, 2011; pp. 299–309. [Google Scholar]
- Reich, K.J.; Bjorndal, K.A.; Frick, M.G.; Witherington, B.E.; Johnson, C.; Bolten, A.B. Polymodal foraging in adult female loggerheads (Caretta caretta). Mar. Biol. 2010, 157, 113–121. [Google Scholar] [CrossRef]
- Detjen, M.; Sterling, E.; Gómez, A. Stable isotopes in barnacles as a tool to understand green sea turtle (Chelonia mydas) regional movement patterns. Biogeosci. Discuss. 2015, 12, 4655–4669. [Google Scholar] [CrossRef]
- Ceriani, S.A.; Weishampel, J.F.; Ehrhart, L.M.; Mansfield, K.L.; Wunder, M.B. Foraging and recruitment hotspot dynamics for the largest Atlantic loggerhead turtle rookery. Sci. Rep. 2017, 7, 1–13. [Google Scholar] [CrossRef]

| Sample Size | Minimum | Maximum | Average | SD | Total |
|---|---|---|---|---|---|
| Anterior | 3 | 19 | 8.1 | 4.5 | 51 |
| Middle | 2 | 34 | 14.0 | 8.4 | 79 |
| Posterior | 11 | 35 | 19.2 | 6.5 | 92 |
| Entire carapace | 2 | 50 | 26.6 | 11.0 | 111 |
| df | SS | MS | Pseudo-F | P(Perm) | Perms | Pairwise Groups | t | P(Perm) | Perms | |
|---|---|---|---|---|---|---|---|---|---|---|
| S-sections | 2 | 1285.3 | 642.66 | 13.933 | 0.001 | 883 | A, M | 2.7304 | 0.009/0.01 (mc) | 124 |
| Residual | 62 | 2859.7 | 46.124 | A, P | 6.3076 | 0.001/0.001 (mc) | 125 | |||
| M, P | 2.3529 | 0.021/0.027 (mc) | 73 | |||||||
| ES(51)-sections | 2 | 408.78 | 204.39 | 11.713 | 0.002 | 999 | A, M | 2.441 | 0.017 | 998 |
| Residual | 62 | 1081.9 | 17.45 | A, P | 5.2855 | 0.001 | 995 | |||
| M, P | 2.355 | 0.024 | 998 | |||||||
| N1-sections | 2 | 295.65 | 147.83 | 7.4929 | 0.001 | 999 | A, M | 2.252 | 0.017 | 996 |
| Residual | 62 | 1223.2 | 19.729 | A, P | 3.7725 | 0.001 | 998 | |||
| M, P | 1.9165 | 0.059 | 996 | |||||||
| N2-sections | 2 | 108.69 | 54.343 | 5.6309 | 0.009 | 999 | A, M | 1.8712 | 0.073 | 998 |
| Residual | 62 | 598.35 | 9.6508 | A, P | 3.0995 | 0.001 | 998 | |||
| M, P | 1.8111 | 0.078 | 997 | |||||||
| Ninf-sections | 2 | 15.959 | 7.9794 | 4.3242 | 0.013 | 999 | A, M | 1.5188 | 0.153 | 998 |
| Residual | 62 | 114.41 | 1.8453 | A, P | 2.7645 | 0.004 | 998 | |||
| M, P | 1.611 | 0.109 | 997 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ingels, J.; Valdes, Y.; Pontes, L.P.; Silva, A.C.; Neres, P.F.; Corrêa, G.V.V.; Silver-Gorges, I.; Fuentes, M.M.P.B.; Gillis, A.; Hooper, L.; et al. Meiofauna Life on Loggerhead Sea Turtles-Diversely Structured Abundance and Biodiversity Hotspots That Challenge the Meiofauna Paradox. Diversity 2020, 12, 203. https://doi.org/10.3390/d12050203
Ingels J, Valdes Y, Pontes LP, Silva AC, Neres PF, Corrêa GVV, Silver-Gorges I, Fuentes MMPB, Gillis A, Hooper L, et al. Meiofauna Life on Loggerhead Sea Turtles-Diversely Structured Abundance and Biodiversity Hotspots That Challenge the Meiofauna Paradox. Diversity. 2020; 12(5):203. https://doi.org/10.3390/d12050203
Chicago/Turabian StyleIngels, Jeroen, Yirina Valdes, Letícia P. Pontes, Alexsandra C. Silva, Patrícia F. Neres, Gustavo V. V. Corrêa, Ian Silver-Gorges, Mariana M.P.B. Fuentes, Anthony Gillis, Lindsay Hooper, and et al. 2020. "Meiofauna Life on Loggerhead Sea Turtles-Diversely Structured Abundance and Biodiversity Hotspots That Challenge the Meiofauna Paradox" Diversity 12, no. 5: 203. https://doi.org/10.3390/d12050203
APA StyleIngels, J., Valdes, Y., Pontes, L. P., Silva, A. C., Neres, P. F., Corrêa, G. V. V., Silver-Gorges, I., Fuentes, M. M. P. B., Gillis, A., Hooper, L., Ware, M., O’Reilly, C., Bergman, Q., Danyuk, J., Sanchez Zarate, S., Acevedo Natale, L. I., & dos Santos, G. A. P. (2020). Meiofauna Life on Loggerhead Sea Turtles-Diversely Structured Abundance and Biodiversity Hotspots That Challenge the Meiofauna Paradox. Diversity, 12(5), 203. https://doi.org/10.3390/d12050203





