Changes in Heterotrophic Picoplankton Community Structure after Induction of a Phytoplankton Bloom under Different Light Regimes
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and Cell Counts
2.2. Molecular Analysis
2.3. Data Processing and Analysis
3. Results
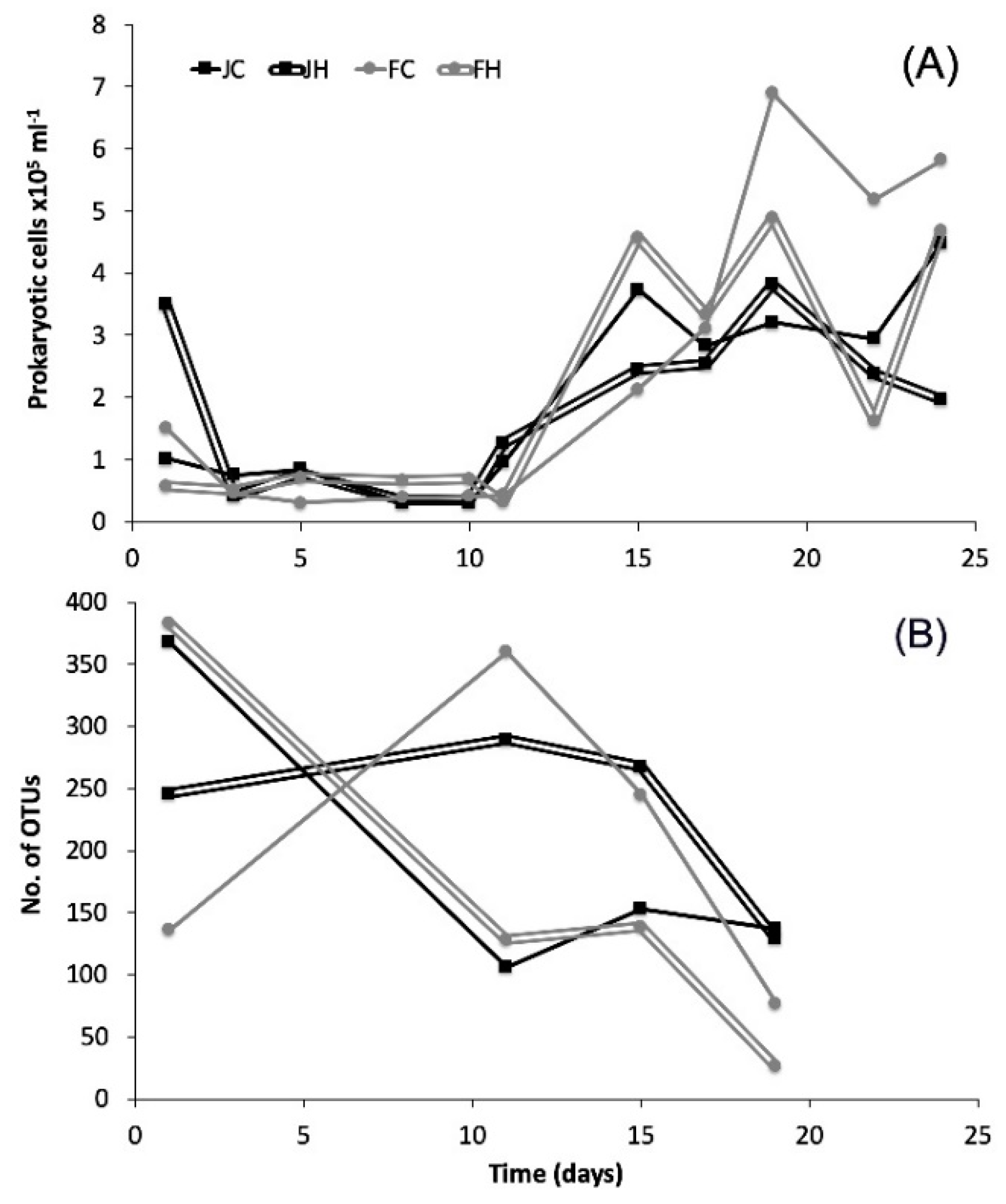
3.1. Heterotrophic Prokaryotescell Abundance
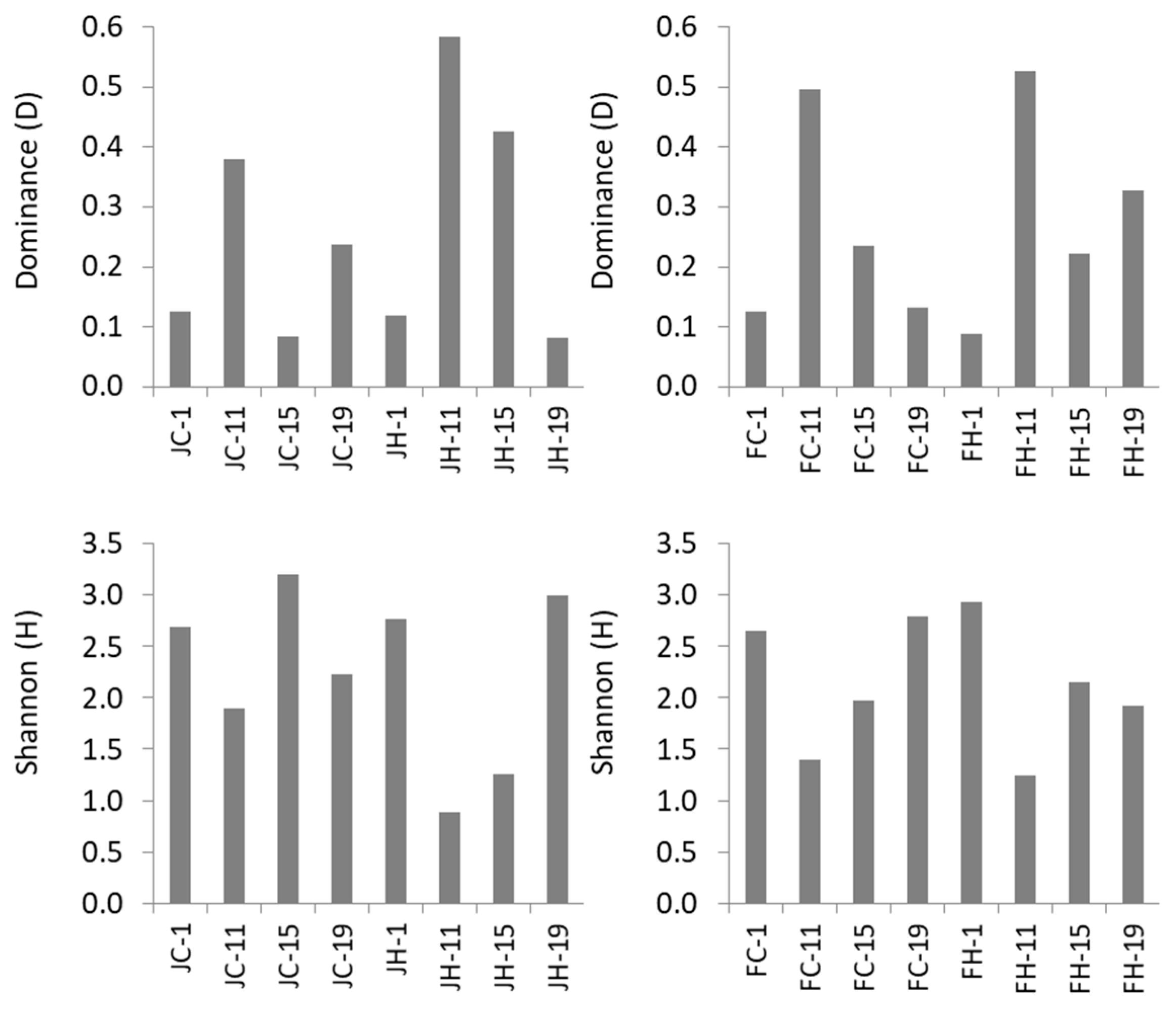
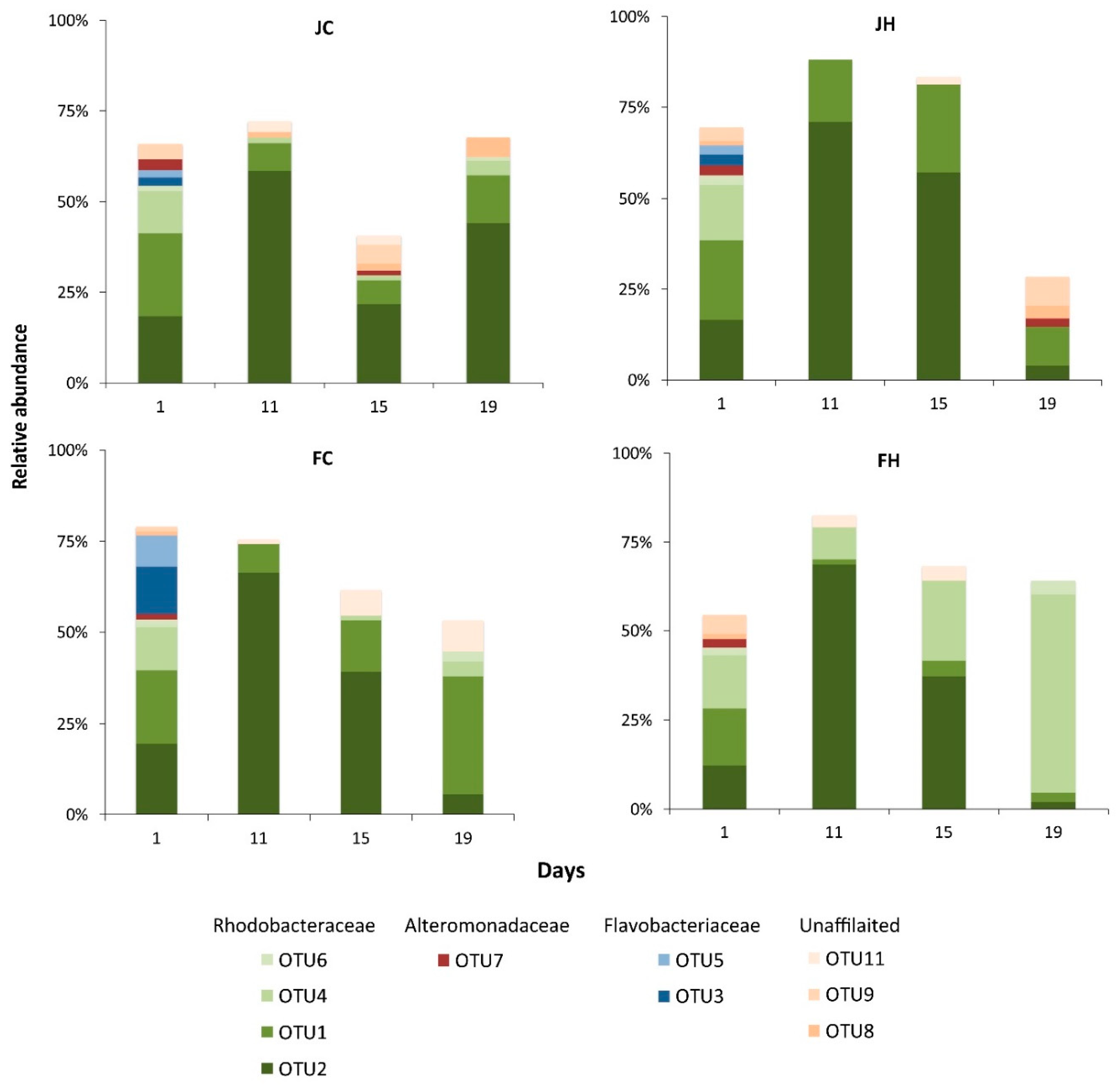
3.2. Diversity and Dynamics of Bacteria and Archaea
4. Discussion
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Tada, Y.; Nakaya, R.; Goto, S.; Yamashita, Y.; Suzuki, K. Distinct bacterial community and diversity shifts after phytoplankton-derived dissolved organic matter addition in a coastal environment. J. Exp. Mar. Biol. Ecol. 2017, 495, 119–128. [Google Scholar] [CrossRef]
- Fenchel, T. Biogeography for Bacteria. Science 2003, 301, 925–926. [Google Scholar] [CrossRef] [PubMed]
- Hanson, C.A.; Fuhrman, J.A.; Horner-Devine, M.C.; Martiny, J.B.H. Beyond biogeographic patterns: Processes shaping the microbial landscape. Nat. Rev. Microbiol. 2012, 10, 497–506. [Google Scholar] [CrossRef] [PubMed]
- Thorton, D.C.O. Dissolved organic matter (DOM) release by phytoplankton in the contemporary and future ocean. Eur. J. Phycol. 2014, 49, 20–46. [Google Scholar] [CrossRef]
- Paver, S.F.; Hayek, K.R.; Crabb, D.B.; Davis-Richardson, A.G.; Rosario-Passapera, R.; Gano, K.A.; Fagen, J.R.; Brown, C.T.; Giongo, A.; Triplett, E.W.; et al. Interactions between specific phytoplankton and bacteria affect lake bacterial community succession. Environ. Microbiol. 2013, 15, 2489–2504. [Google Scholar] [CrossRef]
- Bagatini, I.L.; Eiler, A.; Bertilsson, S.; Klaveness, D.; Tessarolli, L.P.; Vieira, A.A.H. Host-Specificity and Dynamics in Bacterial Communities Associated with Bloom-Forming Freshwater Phytoplankton. PLoS ONE 2014, 9, e85950. [Google Scholar] [CrossRef]
- Pinhassi, J.; Winding, A.; Binnerup, S.; Zweifel, U.; Riemann, B.; Hagström, A. Spatial variability in bacterioplankton community composition at the Skagerrak-Kattegat Front. Mar. Ecol. Prog. Ser. 2003, 255, 1–13. [Google Scholar] [CrossRef]
- Tada, Y.; Taniguchi, A.; Sato-Takabe, Y.; Hamasaki, K. Growth and succession patterns of major phylogenetic groups of marine bacteria during a mesocosm diatom bloom. J. Oceanogr. 2012, 68, 509–519. [Google Scholar] [CrossRef]
- Zhou, J.; Richlen, M.L.; Sehein, T.R.; Kulis, D.M.; Anderson, D.M.; Cai, Z. Microbial Community Structure and Associations During a Marine Dinoflagellate Bloom. Front. Microbiol. 2018, 9, 1201. [Google Scholar] [CrossRef]
- Meziti, A.; Kormas, K.A.; Moustaka-Gouni, M.; Karayanni, H. Spatially uniform but temporally variable bacterioplankton in a semi-enclosed coastal area. Syst. Appl. Microbiol. 2015, 38, 358–367. [Google Scholar] [CrossRef]
- Hoppe, H.G.; Giesenhagen, H.C.; Köppe, R.; Hansen, H.P.; Göcke, K. Impact of change in climate and policy from 1988 to 2007 on environmental and microbial variables at the time series station Boknis Eck, Baltic Sea. Biogeosciences 2013, 10, 4529–4546. [Google Scholar] [CrossRef]
- Tsiola, A.; Tsagaraki, T.M.; Giannakourou, A.; Nikolioudakis, N.; Yücel, N.; Herut, B.; Pitta, P. Bacterial Growth and Mortality after Deposition of Saharan Dustand Mixed Aerosols in the Eastern Mediterranean Sea: A Mesocosm Experiment. Front. Mar. Sci. 2017, 3, 281. [Google Scholar] [CrossRef]
- Stefanidou, N.; Genitsaris, S.; Lopez-Bautista, J.; Sommer, U.; Moustaka-Gouni, M. Effects of heat shock and salinity changes on coastal Mediterranean phytoplankton in a mesocosm experiment. Mar. Biol. 2018, 165, 154. [Google Scholar] [CrossRef]
- Shade, A.; Read, J.S.; Youngblut, N.D.; Fierer, N.; Knight, R.; Kratz, T.K.; Lottig, N.R.; Roden, E.E.; Stanley, E.H.; Stombaugh, J.; et al. Lake microbial communities are resilient after a whole-ecosystem disturbance. ISME J. 2012, 6, 2153–2167. [Google Scholar] [CrossRef] [PubMed]
- Allison, S.D.; Martiny, J.B.H. Resistance, resilience, and redundancy in microbial communities. Proc. Natl. Acad. Sci. USA 2008, 105, 11512–11519. [Google Scholar] [CrossRef] [PubMed]
- Downing, A.L.; Leibold, M.A. Species richness facilitates ecosystem resilience in aquatic food webs. Freshw. Biol. 2010, 55, 2123–2137. [Google Scholar] [CrossRef]
- Ruiz-González, C.; Simó, R.; Sommaruga, R.; Gasol, J.M. Away from darkness: A review on the effects of solar radiation on heterotrophic bacterioplankton activity. Front. Microbiol. 2013, 4, 131. [Google Scholar] [CrossRef]
- Yamamichi, M.; Kazama, T.; Tokita, K.; Katano, I.; Doi, H.; Yoshida, T.; Hairston, N.G., Jr.; Urabe, J. A shady phytoplankton paradox: When phytoplankton increases under low light. Proc. R. Soc. B 2018, 285, 20181067. [Google Scholar] [CrossRef]
- Sommer, U.; Lengfellner, K.; Lewandowska, A. Experimental induction of a coastal spring bloom early in the year by intermittent high-light episodes. Mar. Ecol. Prog. Ser. 2012, 446, 61–71. [Google Scholar] [CrossRef]
- Paver, S.F.; Kent, A.D. Direct and Context-dependent effects of light, temperature, and phytoplankton shape bacterial community composition. Ecosphere 2017, 8, e01948. [Google Scholar] [CrossRef]
- Alonso-Sáez, L.; Gasol, J.M.; Lefort, T.; Hofer, J.; Sommaruga, R. Effect of Natural Sunlight on Bacterial Activity and Differential Sensitivity of Natural Bacterioplankton Groups in Northwestern Mediterranean Coastal Waters. Appl. Environ. Microbiol. 2006, 72, 5806–5813. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-González, C.; Lefort, T.; Massana, R.; Simó, R.; Gasol, J.M. Diel changes in bulk and single-cell bacterial heterotrophic activity in winter surface waters of the northwestern Mediterranean Sea. Limnol. Oceanogr. 2012, 57, 29–42. [Google Scholar] [CrossRef]
- Sánchez, O.; Koblížek, M.; Gasol, J.M.; Ferrera, I. Effects of grazing, phosphorus and light on the growth rates of major bacterioplankton taxa in the coastal NW Mediterranean. Environ. Microbiol. Rep. 2017, 9, 300–309. [Google Scholar] [CrossRef] [PubMed]
- Schwalbach, M.S.; Brown, M.; Fuhrman, J.A. Impact of light on marine bacterioplankton community structure. Aquat. Microb. Ecol. 2005, 39, 235–245. [Google Scholar] [CrossRef]
- Agogué, H.; Joux, F.; Obernosterer, I.; LeBaron, P. Resistance of Marine Bacterioneuston to Solar Radiation. Appl. Environ. Microbiol. 2005, 71, 5282–5289. [Google Scholar] [CrossRef]
- Schlesinger, W.H. Biogeochemistry: Treatise on Geochemistry, 1st ed.; Elsevier: Amsterdam, The Netherlands, 2005; Volume 8. [Google Scholar]
- Parker, M.S.; Armbrust, E.V. Synergistic effects of light, temperature, and nitrogen source on transcription of genes for carbon and nitrogen metabolism in the centric diatom Thalassiosira pseudonana (Bacillariophyceae). J. Phycol. 2005, 41, 1142–1153. [Google Scholar] [CrossRef]
- Blanchet, M.; Fernandez, C.; Joux, F. Photoreactivity of riverine and phytoplanktonic dissolved organic matter and its effects on the dynamics of a bacterial community from the coastal Mediterranean Sea. Prog. Oceanogr. 2018, 163, 82–93. [Google Scholar] [CrossRef]
- Von Scheibner, M.; Dörge, P.; Biermann, A.; Sommer, U.; Hoppe, H.G.; Jürgens, K. Impact of warming on phyto-bacterioplankton coupling and bacterial community composition in experimental mesocosms. Environ. Microbiol. 2014, 16, 718–733. [Google Scholar] [CrossRef]
- Dowd, S.; Callaway, T.R.; Wolcott, R.D.; Sun, Y.; McKeehan, T.; Hagevoort, R.G.; Edrington, T.S. Evaluation of the bacterial diversity in the feces of cattle using 16S rDNA bacterial tag-encoded FLX amplicon pyrosequencing (bTEFAP). BMC Microbiol. 2008, 8, 125. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Quince, C.; Lanzén, A.; Curtis, T.P.; Davenport, R.J.; Hall, N.; Head, I.M.; Read, L.F.; Sloan, W.T. Accurate determination of microbial diversity from 454 pyrosequencing data. Nat. Methods 2009, 6, 639–641. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Reeder, J.; Knight, R. The ‘rare biosphere’: A reality check. Nat. Rev. Microbiol. 2009, 6, 636–637. [Google Scholar] [CrossRef] [PubMed]
- Behnke, A.; Engel, M.; Christen, R.; Nebel, M.; Klein, R.R.; Stoeck, T. Depicting more accurate pictures of protistan community complexity using pyrosequencing of hypervariable SSU rRNA gene regions. Environ. Microbiol. 2010, 13, 340–349. [Google Scholar] [CrossRef] [PubMed]
- Kunin, V.; Engelbrektson, A.; Ochman, H.; Hugenholtz, P. Wrinkles in the rare biosphere: Pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ. Microbiol. 2010, 12, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Lemos, L.N.; Fulthorpe, R.R.; Triplett, E.W.; Roesch, L.F. Rethinking microbial diversity analysis in the high throughput sequencing era. J. Microbiol. Methods 2011, 86, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Glöckner, F.O. SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Smoot, M.E.; Ono, K.; Ruscheinski, J.; Wang, P.L.; Ideker, T. Cytoscape 2.8: New features for data integration and network visualization. Bioinformatics 2011, 27, 431–432. [Google Scholar] [CrossRef]
- Prosser, J.I. Replicate or lie. Environ. Microbiol. 2010, 12, 1806–1810. [Google Scholar] [CrossRef]
- Karayanni, H.; Meziti, A.; Spatharis, S.; Genitsaris, S.; Courties, C.; Kormas, K.A. Changes in Microbial (Bacteria and Archaea) Plankton Community Structure after Artificial Dispersal in Grazer-Free Microcosms. Microorganisms 2017, 5, 31. [Google Scholar] [CrossRef] [PubMed]
- Brinkhoff, T.; Giebel, H.A.; Simon, M. Diversity, ecology, and genomics of the Roseobacter clade: A short overview. Arch. Microbiol. 2008, 189, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Lefort, T.; Gasol, J.M. Global-scale distributions of marine surface bacterioplankton groups along gradients of salinity, temperature, and chlorophyll: A meta-analysis of fluorescence in situ hybridization studies. Aquat. Microb. Ecol. 2013, 70, 111–130. [Google Scholar] [CrossRef]
- Fandino, L.; Riemann, L.; Steward, G.; Long, R.; Azam, F. Variations in bacterial community structure during a dinoflagellate bloom analyzed by DGGE and 16S rDNA sequencing. Aquat. Microb. Ecol. 2001, 23, 119–130. [Google Scholar] [CrossRef]
- Zubkov, M.V.; Fuchs, B.M.; Archer, S.D.; Kiene, R.P.; Amann, R.; Burkill, P.H. Linking the composition of bacterioplankton to rapid turnover of dissolved dimethylsulphoniopropionate in an algal bloom in the North Sea. Environ. Microbiol. 2001, 3, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Fuhrman, J.A.; Hagström, A. Bacterial and archaeal community structure and its patterns. In Microbial Ecology of the Oceans, 2nd ed.; Kirchman, D.L., Ed.; Wiley-Blackwell: Hoboken, NJ, USA, 2008; pp. 45–90. [Google Scholar]
- Newton, R.J.; Griffin, L.E.; Bowles, K.M.; Meile, C.; Gifford, S.; Givens, C.E.; Howard, E.C.; King, E.; Oakley, C.A.; Reisch, C.R.; et al. Genome characteristics of a generalist marine bacterial lineage. ISME J. 2010, 4, 784–798. [Google Scholar] [CrossRef] [PubMed]
- Brenner, D.J.; Krieg, N.R.; Staley, J.T. The Proteobacteria: Part C: The Alpha-, Beta-, Delta-, and Epsilonproteobacteria. In Bergey’s Manual of Systematic Bacteriology, 2nd ed.; Garriy, G.M., Ed.; Springer: New York, NY, USA, 2005; pp. 161–229. [Google Scholar]
- Allers, E.; Pinhassi, J.; Gasol, J.M.; Šimek, K.; Pernthaler, J.; Gómez-Consarnau, L.; Gómez-Consarnau, L. Response of Alteromonadaceae and Rhodobacteriaceae to glucose and phosphorus manipulation in marine mesocosms. Environ. Microbiol. 2007, 9, 2417–2429. [Google Scholar] [CrossRef]
- Biermann, A.; Engel, A.; Riebesell, U. Changes in organic matter cycling in a plankton community exposed to warming under different light intensities. J. Plankton Res. 2014, 36, 658–671. [Google Scholar] [CrossRef]
- Hansell, D.A.; Carlson, C.A. Biogeochemistry of Marine Dissolved Organic Matter, 1st ed.; Academic Press: San Diego, CA, USA, 2002. [Google Scholar]
- Jiao, N.; Herndl, G.J.; Hansell, D.A.; Benner, R.; Kattner, G.; Wilhelm, S.W.; Kirchman, D.L.; Weinbauer, M.G.; Luo, T.; Chen, F.; et al. Microbial production of recalcitrant dissolved organic matter: Long-term carbon storage in the global ocean. Nat. Rev. Microbiol. 2010, 8, 593–599. [Google Scholar] [CrossRef]
- Sarmento, H.; Gasol, J.M. Use of phytoplankton-derived dissolved organic carbon by different types of bacterioplankton. Environ. Microbiol. 2012, 14, 2348–2360. [Google Scholar] [CrossRef]
- Grossart, H.; Simon, M. Interactions of planktonic algae and bacteria: Effects on algal growth and organic matter dynamics. Aquat. Microb. Ecol. 2007, 47, 163–176. [Google Scholar] [CrossRef]
- Lindh, M.V.; Riemann, L.; Baltar, F.; Romero-Oliva, C.; Salomon, P.S.; Granéli, E.; Pinhassi, J. Consequences of increased temperature and acidification on bacterioplankton community composition during a mesocosm spring bloom in the Baltic Sea. Environ. Microbiol. Rep. 2013, 5, 252–262. [Google Scholar] [CrossRef]
- Subramani, R.; Aalbersberg, W. Marine actinomycetes: An ongoing source of novel bioactive metabolites. Microbiol. Res. 2012, 167, 571–580. [Google Scholar] [CrossRef] [PubMed]
- Beardsley, C.; Pernthaler, J.; Wosniok, W.; Amann, R. Are Readily Culturable Bacteria in Coastal North Sea Waters Suppressed by Selective Grazing Mortality? Appl. Environ. Microbiol. 2003, 69, 2624–2630. [Google Scholar] [CrossRef] [PubMed]
- Smayda, T.J. Cryptic planktonic diatom challenges phytoplankton ecologists. Proc. Natl. Acad. Sci. USA 2011, 108, 4269–4270. [Google Scholar] [CrossRef]
- Ye, W.; Liu, X.; Lin, S.; Tan, J.; Pan, J.; Li, D.; Yang, H. The vertical distribution of bacterial and archaeal communities in the water and sediment of Lake Taihu. FEMS Microbiol. Ecol. 2009, 70, 263–276. [Google Scholar] [CrossRef] [PubMed]
- Kormas, K.A.; Vardaka, E.; Moustaka-Gouni, M.; Kontoyanni, V.; Petridou, E.; Gkelis, S.; Neofitou, C. Molecular detection of potentially toxic cyanobacteria and their associated bacteria in lake water column and sediment. World J. Microbiol. Biotechnol. 2010, 26, 1473–1482. [Google Scholar] [CrossRef]
- Cardman, Z.; Arnosti, C.; Durbin, A.; Ziervogel, K.; Cox, C.; Steen, A.D.; Teske, A. Verrucomicrobia Are Candidates for Polysaccharide-Degrading Bacterioplankton in an Arctic Fjord of Svalbard. Appl. Environ. Microbiol. 2014, 80, 3749–3756. [Google Scholar] [CrossRef]
- Teira, E.; Gasol, J.M.; Aranguren-Gassis, M.; Fernández, A.; González, J.; Lekunberri, I.; Aranguren-Gassis, M.; Fernández-Carrera, A.; Álvarez-Salgado, X.A.; Aranguren-Gassis, M.; et al. Linkages between bacterioplankton community composition, heterotrophic carbon cycling and environmental conditions in a highly dynamic coastal ecosystem. Environ. Microbiol. 2008, 10, 906–917. [Google Scholar] [CrossRef]
- Murray, A.; Arnosti, C.; De La Rocha, C.; Grossart, H.; Passow, U. Microbial dynamics in autotrophic and heterotrophic seawater mesocosms. II. Bacterioplankton community structure and hydrolytic enzyme activities. Aquat. Microb. Ecol. 2007, 49, 123–141. [Google Scholar] [CrossRef]
- Nydahl, A.; Panigrahi, S.; Wikner, J. Increased microbial activity in a warmer and wetter climate enhances the risk of coastal hypoxia. FEMS Microbiol. Ecol. 2013, 85, 338–347. [Google Scholar] [CrossRef] [PubMed]
- Panigrahi, S.; Nydahl, A.; Anton, P.; Wikner, J. Strong seasonal effect of moderate experimental warming on plankton respiration in a temperate estuarine plankton community. Estuar. Coast. Shelf Sci. 2013, 135, 269–279. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karayanni, H.; Kormas, K.A.; Moustaka-Gouni, M.; Sommer, U. Changes in Heterotrophic Picoplankton Community Structure after Induction of a Phytoplankton Bloom under Different Light Regimes. Diversity 2019, 11, 195. https://doi.org/10.3390/d11100195
Karayanni H, Kormas KA, Moustaka-Gouni M, Sommer U. Changes in Heterotrophic Picoplankton Community Structure after Induction of a Phytoplankton Bloom under Different Light Regimes. Diversity. 2019; 11(10):195. https://doi.org/10.3390/d11100195
Chicago/Turabian StyleKarayanni, Hera, Konstantinos A. Kormas, Maria Moustaka-Gouni, and Ulrich Sommer. 2019. "Changes in Heterotrophic Picoplankton Community Structure after Induction of a Phytoplankton Bloom under Different Light Regimes" Diversity 11, no. 10: 195. https://doi.org/10.3390/d11100195
APA StyleKarayanni, H., Kormas, K. A., Moustaka-Gouni, M., & Sommer, U. (2019). Changes in Heterotrophic Picoplankton Community Structure after Induction of a Phytoplankton Bloom under Different Light Regimes. Diversity, 11(10), 195. https://doi.org/10.3390/d11100195







