DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha)
Abstract
1. Introduction
2. Materials and Methods
2.1. Psyllid Collection
2.2. Identification of Insects Based on Morphology and Host Plant Association
2.3. Molecular Analysis
2.4. COI Barcode Data Analysis
3. Results and Discussion
3.1. Species Delimitation
3.1.1. Field Collection, Host Plant Association and Initial Morphological Examination
3.1.2. COI Genetic Analysis
3.1.3. Morphological Reexamination of the Cryptic Taxa Detected
3.1.4. A Species Delimitation Method Integrating Host Plant, COI and Morphology
3.2. An Updated Biodiversity of the New Zealand Psylloidea Reveals New Insights into Psyllid Biology and Ecology
3.2.1. The Genus Psylla Showed Examples of Putative Host Plant Specificity and a Syntopic Distribution
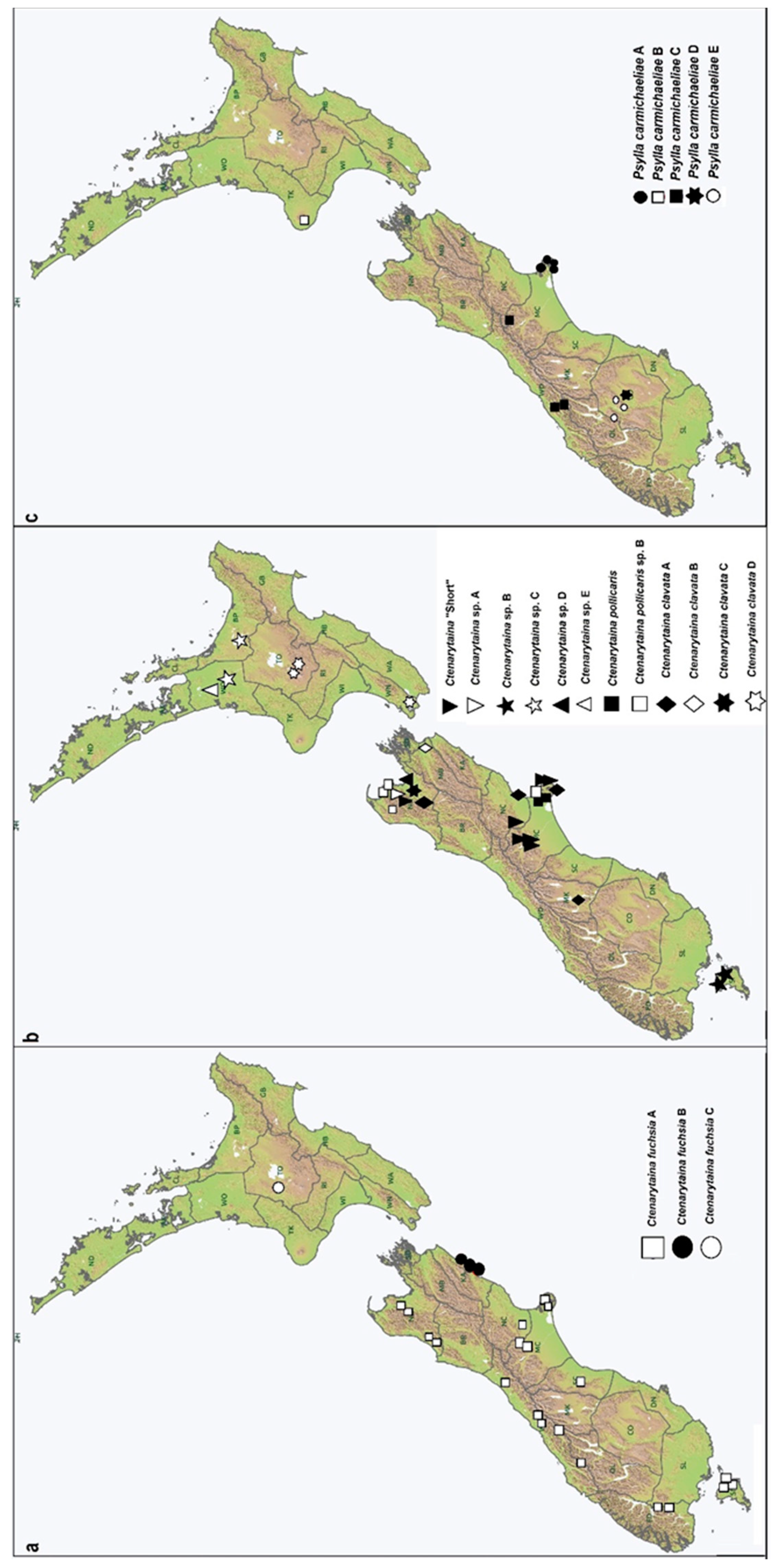
3.2.2. The Ctenarytaina fuchsiae Complex and Its Distribution Show a Strict Geographical Isolation
3.2.3. Are Psyllid Cryptic Taxa Associated with Host Plant Cryptic Species?
3.2.4. Other Instances of Multiple Taxa on the Same Host Plant
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Burckhardt, D.; Ouvrard, D. A revised classification of the jumping plant–lice (Hemiptera: Psylloidea). Zootaxa 2012, 3509, 1–34. [Google Scholar] [CrossRef]
- Ouvrard, D. Psyl’list—The World Psylloidea Database. Available online: Http://www.Hemiptera-databases.Com/psyllist (accessed on 17 April 2018).
- Hollis, D. Australian Psylloidea: Jumping Plant Lice and Lerp Insects; Australian Biological Resources Study: Canberra, Australia, 2004; pp. 1–216. [Google Scholar]
- Linné, C. Systema Naturae per Regna tri Naturae, Secundum Classes, Ordines, Genera, Species, cum Characteribus, Differentiis, Synonymis, Locis, 10th ed.; Impensis Direct: Stockholm, Sweden, 1758; pp. 1–824. [Google Scholar]
- EPPO/CABI. Quarantine Pests for Europe, 2nd ed.; Smith, I.M., McNamara, D.G., Scott, P.R., Holderness, M., Eds.; CABI International: Wallingford, UK, 1997; p. 1425. [Google Scholar]
- Aubert, B. Trioza erytreae Del Guercio and Diaphorina citri Kuwayama (Homoptera: Psylloidea), the two vectors of citrus greening disease: Biological aspects and possible control strategies. Fruits 1987, 42, 149–162. [Google Scholar]
- Capoor, S.P.; Rao, D.G.; Viswanath, S.M. Diaphorina citri Kuwayama, a vector of the greening disease of citrus in India. Indian J. Agric. Sci. 1967, 37, 572–576. [Google Scholar]
- Syfert, M.M.; Serbina, L.; Burckhardt, D.; Knapp, S.; Percy, D.M. Emerging new crop pests: Ecological modelling and analysis of the South American potato psyllid Russelliana solanicola (Hemiptera: Psylloidea) and its wild relatives. PLoS ONE 2017, 12, e0167764. [Google Scholar] [CrossRef] [PubMed]
- Liefting, L.W.; Weir, B.S.; Pennycook, S.R.; Clover, G.R.G. ‘Candidatus Liberibacter solanacearum’, associated with plants in the family solanaceae. Int. J. Syst Evol. Microbiol. 2009, 59, 2274–2276. [Google Scholar] [CrossRef] [PubMed]
- Teulon, D.A.J.; Workman, P.J.; Thomas, K.L.; Krause Nielsen, M.-C. Bactericera cockerelli: Incursion, dispersal and current distribution on vegetable crops in New Zealand. N. Z. Plant. Prot. 2009, 62, 136–144. [Google Scholar]
- Vereijssen, J.; Smith, G.R.; Weintraub, P.G. Bactericera cockerelli (Hemiptera: Triozidae) and Candidatus Liberibacter solanacearum in potatoes in New Zealand: Biology, transmission, and implications for management. J. Integr. Pest. Manag. 2018, 9, 1–21. [Google Scholar] [CrossRef]
- Taylor, G.S.; Fagan-Jeffries, E.P.; Austin, A.D. A new genus and twenty new species of Australian jumping plant-lice (Psylloidea: Triozidae) from Eremophila and Myoporum (Scrophulariaceae: Myoporeae). Zootaxa 2016, 4073, 1–84. [Google Scholar] [CrossRef] [PubMed]
- Ferris, G.F.; Klyver, F.D. Report upon a collection of chermidae (Homoptera) from New Zealand. Trans. N. Z. Inst. 1932, 63, 34–61. [Google Scholar]
- Tuthill, L.D. On the psyllidae of New Zealand (Homoptera). Pac. Sci. 1952, 6, 83–125. [Google Scholar]
- Dale, P.J. A review of the Psylloidea (Insecta: Hemiptera) of the New Zealand Subregion. Unpublished Ph.D. Thesis, University of Auckland, Auckland, New Zealand, 1985. [Google Scholar]
- Macfarlane, R.P.; Andrew, I.G.; Berry, J.A.; Johns, P.M.; Hoare, R.J.B.; Lariviere, M.-C.; Grenslade, P.; Henderson, R.C.; Smithers, C.N.; Palma, R.L.; et al. Phylum Arthropoda, subphylum Hexapoda: Protura, springtails, Diplura, insects. N. Z. Inventory Biodiv. 2010, 2, 233–467. [Google Scholar]
- Martoni, F.; Burckhardt, D.; Armstrong, K. An annotated checklist of the psyllids of New Zealand (Hemiptera: Psylloidea). Zootaxa 2016, 4144, 556–574. [Google Scholar] [CrossRef] [PubMed]
- Munoz, J.; Felicisimo, A.M.; Cabezas, F.; Burgaz, A.R.; Martinez, I. Wind as a long-distance dispersal vehicle in the Southern Hemisphere. Science 2004, 304, 1144–1147. [Google Scholar] [CrossRef] [PubMed]
- Dijkstra, K.D.B. Gone with the wind: Westward dispersal across the Indian Ocean and island speciation in Hemicordulia dragonflies (Odonata: Corduliidae). Zootaxa 2007, 1438, 27–48. [Google Scholar] [CrossRef]
- Close, R.C.; Moar, N.T.; Tomlinson, A.I.; Lowe, A.D. Aerial dispersal of biological material from Australia to New Zealand. Int. J. Biometeorol. 1978, 22, 1–19. [Google Scholar] [CrossRef]
- Tomlinson, A.I. Meteorological aspects of trans-Tasman insect dispersal. N. Z. Entomol. 1973, 5, 253–268. [Google Scholar] [CrossRef]
- Buckley, T.R.; Krosch, M.; Leschen, R.A.B. Evolution of New Zealand insects: Summary and prospectus for future research. Aust. Entomol. 2015, 54, 1–27. [Google Scholar] [CrossRef]
- Maskell, W.M. On some species of Psyllidae in New Zealand. Trans. N. Z. Inst. 1890, 22, 157–170. [Google Scholar]
- Šulc, K. Trioza cockerelli n. Sp, novinka ze severní ameriky, mající i hospodářský význam [Trioza cockerelli n. Sp., a novelty from North America, being also of economic importance]. Acta Soc. Entomol. Bohem. 1909, 6, 102–108. [Google Scholar]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. B-Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef] [PubMed]
- Hebert, P.D.N.; Gregory, T.R. The promise of DNA barcoding for taxonomy. Syst. Biol. 2005, 54, 852–859. [Google Scholar] [CrossRef] [PubMed]
- Hall, A.A.G.; Morrow, J.L.; Fromont, C.; Steinbauer, M.J.; Taylor, G.S.; Johnson, S.N.; Cook, J.M.; Riegler, M. Codivergence of the primary bacterial endosymbiont of psyllids versus host switches and replacement of their secondary bacterial endosymbionts. Environ. Microbiol. 2016, 18, 2591–2603. [Google Scholar] [CrossRef] [PubMed]
- Martoni, F.; Bulman, S.R.; Pitman, A.R.; Armstrong, K. Elongation factor-1α accurately reconstructs relationships amongst psyllid families (Hemiptera: Psylloidea), with possible diagnostic implications. J. Econ. Entomol. 2017, 110, 2618–2622. [Google Scholar] [CrossRef] [PubMed]
- Percy, D.M. Making the mosts of your host: The Metrosideros-feeding psyllids (Hemiptera: Psylloidea) of the Hawaiian Islands. ZooKeys 2017, 649, 1–163. [Google Scholar] [CrossRef] [PubMed]
- Percy, D.M.; Crampton-Platt, A.; Sveinsson, S.; Lemmon, A.R.; Moriarty Lemmon, E.; Ouvrard, D.; Burckhardt, D. Resolving the psyllid tree of life: Phylogenomic analyses of the superfamily Psylloidea (Hemiptera). Syst. Entomol. 2018. [Google Scholar] [CrossRef]
- Crosby, T.K.; Dugdale, J.S.; Watt, J.C. Area codes for recording specimen localities in the New Zealand subregion. N. Z. J. Zool. 1998, 25, 175–183. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit i from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–297. [Google Scholar] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Srivathsan, A.; Meier, R. On the inappropriate use of Kimura-2-parameter (K2P) divergences in the DNA–barcoding literature. Cladistics 2012, 28, 190–194. [Google Scholar] [CrossRef]
- Collins, R.A.; Boykin, L.M.; Cruickshank, R.H.; Armstrong, K. Barcoding’s next top model: An evaluation of nucleotide substitution models for specimen identification. Methods Ecol. Evol. 2012, 3, 457–465. [Google Scholar] [CrossRef]
- Puillandre, N.; Lambert, A.; Brouillet, S.; Achaz, G. Abgd, automatic barcode gap discovery for primary species delimitation. Mol. Ecol. 2012, 21, 1864–1877. [Google Scholar] [CrossRef] [PubMed]
- Pons, J.; Barraclough, T.; Gomez-Zurita, J.; Cardoso, A.; Duran, D.; Hazell, S.; Kamoun, S.; Sumlin, W.; Vogler, A. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst. Biol. 2006, 55, 595–609. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Kapli, P.; Pavlidis, P.; Stamatakis, A. A general species delimitation method with applications to phylogenetic placements. Bioinformatics 2013, 29, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Kapli, P.; Lutteropp, S.; Zhang, J.; Kobert, K.; Pavlidis, P.; Stamatakis, A.; Flouri, T. Multi-rate poisson tree processes for singlelocus species delimitation under maximum likelihood and markov chain monte carlo. Bioinformatics 2017, 33, 1630–1638. [Google Scholar] [PubMed]
- Drummond, A.J.; Suchard, M.A.; Xie, D.; Rambaut, A. Bayesian phylogenetics with Beauti and the Beast 1.7. Mol. Biol. Evol. 2012, 29, 1969–1973. [Google Scholar] [CrossRef] [PubMed]
- Michonneau, F. Cryptic and not-so-cryptic species in the complex “Holothuria (Thymiosycia) imaptiens” (Forsskål, 1775) (Echinodermata: Holothuroidea: Holothuriidae). bioRxiv 2015, 014225. [Google Scholar]
- Rambaut, A. Figtree v1.4.3. 2016. Available online: http://tree.Bio.Ed.Ac.Uk/software/figtree/ (accessed on 4 October 2016).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Ezard, T.; Fujisawa, T.; Barraclough, T.G. Splits: Species’ Limits by Threshold Statistics. R Package Version 1.0-18/r45. Available online: http://r-forge.R-project.Org/projects/splits/ (accessed on 15 January 2009).
- Burckhardt, D.; Ouvrard, D.; Queiroz, D.; Percy, D. Psyllid host-plants (Hemiptera: Psylloidea): Resolving a semantic problem. Fla. Entomol. 2014, 97, 242–246. [Google Scholar] [CrossRef]
- Wonglersak, R.; Cronk, Q.C.; Percy, D. Salix transect of Europe: Structured genetic variation and isolation-by-distance in the nettle psyllid, Trioza urticae (Psylloidea, Hemiptera), from Greece to Arctic Norway. Biodivers. Data J. 2017, 5, e10824. [Google Scholar] [CrossRef] [PubMed]
- Jin, Q.; He, L.J.; Zhang, A.B. A simple 2d non-parametric resampling statistical approach to assess confidence in species identification in DNA barcoding-an alternative to likelihood and bayesian approaches. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Luo, A.R.; Lan, H.Q.; Ling, C.; Zhang, A.B.; Shi, L.; Ho, S.Y.W.; Zhu, C.D. A simulation study of sample size for DNA barcoding. Ecol. Evol. 2015, 5, 5869–5879. [Google Scholar] [CrossRef] [PubMed]
- Ross, H.A.; Murugan, S.; Li, W.L.S. Testing the reliability of genetic methods of species identification via simulation. Syst. Biol. 2008, 57, 216–230. [Google Scholar] [CrossRef] [PubMed]
- Heenan, P.B. Taxonomic revision of Carmichaelia (Fabaceae-Galegeae) in New Zealand 1. N. Z. J. Bot. 1995, 33, 455–475. [Google Scholar] [CrossRef]
- Heenan, P.B. A taxonomic revision of Carmichaelia (Fabaceae-Galegeae) in New Zealand 2. N. Z. J. Bot. 1996, 34, 157–177. [Google Scholar] [CrossRef]
- Stringer, I.A.N.; Hitchmough, R.A.; Lariviere, M.C.; Eyles, A.C.; Teulon, D.A.J.; Dale, P.J.; Henderson, R.C. The conservation status of New Zealand Hemiptera. N. Z. Entomol. 2012, 35, 110–115. [Google Scholar] [CrossRef]
- Yen, A.L.; Finlay, K.J.; Weiss, J.; Vereijssen, J. Understanding the Significance of Natural Pathways into Australia and New Zealand; Plant Biosecurity Cooperative Research Centre: Bruce, Australia, 2014; p. 182. [Google Scholar]
- De Lange, P.J. A revision of the New Zealand Kunzea ericoides (Myrtaceae) complex. Phytokeys 2014, 40, 1–185. [Google Scholar] [CrossRef] [PubMed]
- Stephens, J.M.C.; Molan, P.C.; Clarkson, B.D. A review of Leptospermum scoparium (Myrtaceae) in New Zealand. N. Z. J. Bot. 2005, 43, 431–449. [Google Scholar] [CrossRef]
- Goldberg, J.; Trewick, S.A.; Paterson, A.M. Evolution of New Zealand’s terrestrial fauna: A review of molecular evidence. Philos. Trans. R. Soc. B 2008, 363, 3319–3334. [Google Scholar] [CrossRef] [PubMed]
- Morrow, J.L.; Hall, A.A.G.; Riegler, M. Symbionts in waiting: The dynamics of incipient endosymbiont complementation and replacement in minimal bacterial communities of psyllids. Microbiome 2017, 5, 58. [Google Scholar] [CrossRef] [PubMed]
- Martoni, F. Biodviersity, Evolution and Microbiome of the New Zealand Psylloidea (Hemiptera: Sternorrhyncha). Ph.D. Thesis, Lincoln University, Lincoln, New Zealand, 2017. [Google Scholar]
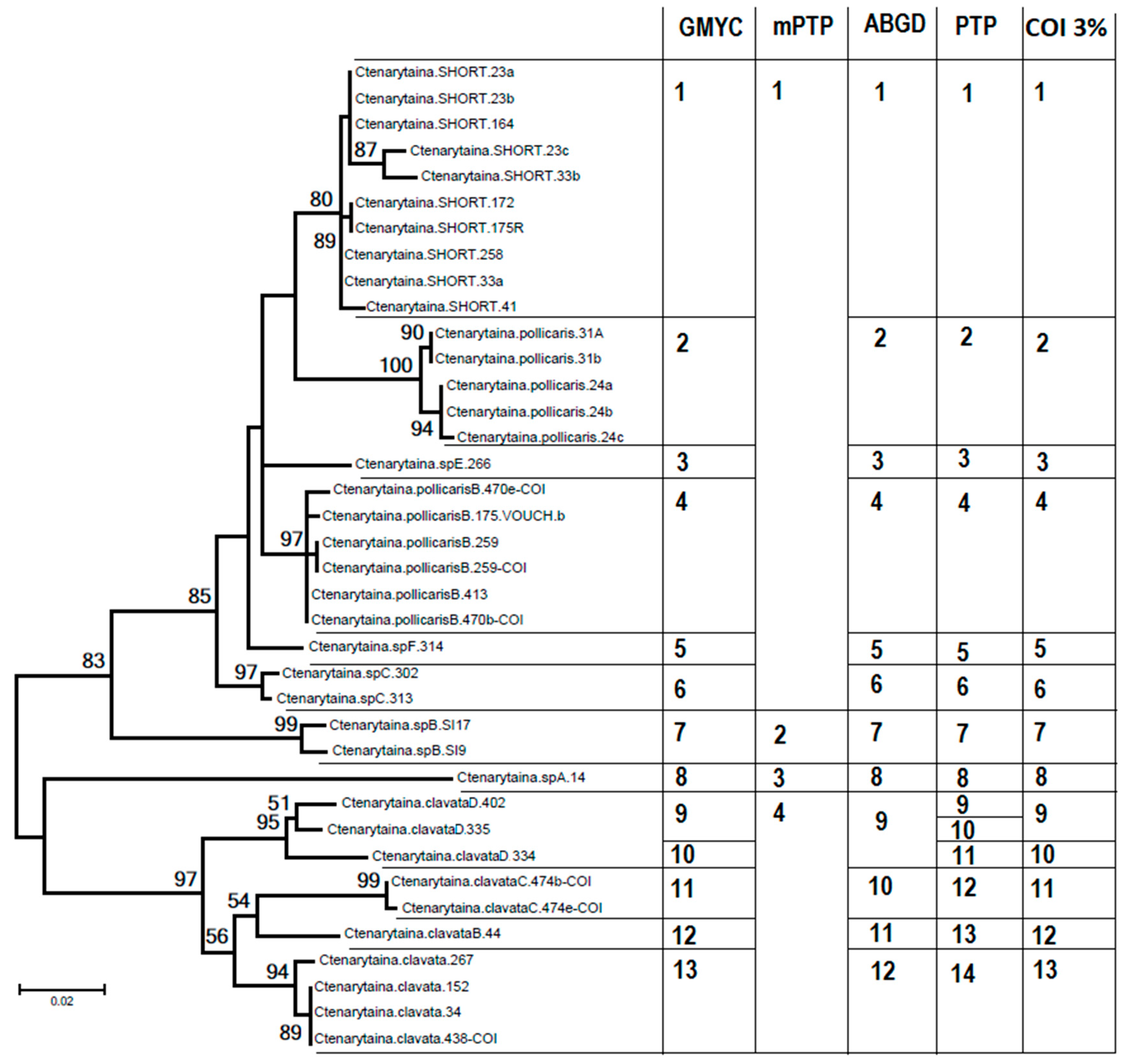
| Species Delimitation Method | Taxa | Reference |
|---|---|---|
| GMYC | 94–96 | [40] |
| PTP | 107–107 | [41] |
| mPTP | 81–81 | [42] |
| ABGD | 90 | [39] |
| 3% Threshold | 89 | [12,29,49] |
| Taxon | Location | Host Plant | |
|---|---|---|---|
| Family Psyllidae | |||
| 1 | Acizzia acaciaebaileyanae | MC [1], NSW [1] | Acacia baileyana |
| 2 | Acizzia acaciae | AK [1], WO [1], MC [2], SL [1], WN [1] | Acacia melanoxylon |
| 3 | Acizzia albizziae | MB [1], NN [2], MC [5], WD [2], TK [1], NSW [1] | Acacia sp. |
| 4 | Acizzia conspicua | WI [1] | Acacia sp. |
| 5 | Acizzia dodonaeae | KA [1], NN (4), TO [1], WN [1], DN [1] | Dodonaea viscosa |
| 6 | Acizzia exquisita | GB [1] | Acacia sp. |
| 7 | Acizzia hakae | MC [1], NN [1], SC [1], WN [1] | Acacia sp., Grevillea sp. |
| 8 | Acizzia jucunda | MB [1], NN [1], GB [1] | Acacia sp. |
| 9 | Acizzia solanicola | AK [1] | Solanum melongena |
| 10 | Acizzia sp.* | MC [2], SA [1] | Acacia baileyana |
| 11 | Acizzia uncatoides | MC [4], CO [2], OL [1], BR [1], TK [1], WI [1], NN [6] | Acacia sp. |
| 12 | Acizzia “Waitakere” ˠ | GB [1], WN [1] | Acacia sp. |
| 13 | Arytainilla spartiophila | MC [3], OL [1] | Cytisus scoparius |
| 14 | Baeopelma foersteri | SL [1], SC [1] | Alnus glutinosa |
| 15 | Psylla apicalis A | DN [1], SL [1], FD [1], MC [2], OL [1] | Sophora microphylla |
| 16 | Psylla apicalis B * | DN [2], CO [2], OL [1], WD [1], BR [1], NN [3], MC [1], SL [1] | Sophora microphylla |
| 17 | Psylla carmichaeliae A | MC [4] | Carmichaelia australis |
| 18 | Psylla carmichaeliae B * | TK [1] | Carmichaelia sp. |
| 19 | Psylla carmichaeliae C * | WD [2], NC [1] | Carmichaelia sp. |
| 20 | Psylla carmichaeliae D * | CO [1] | Carmichaelia compacta |
| 21 | Psylla carmichaeliae E * | CO [2], OL [1] | Carmichaelia petri |
| Family Calophyidae | |||
| 22 | Calophya schini | HB [1], MC [2] | Schinus molle |
| Family Homotomidae | |||
| 23 | Mycopsylla fici | AK [2], NSW [2] | Ficus macrophylla |
| Family Liviidae | |||
| 24 | Psyllopsis fraxini | SL [1], SC [1], BP [1] | Fraxinus excelsior |
| 25 | Psyllopsis fraxinicola | MC [2], FD [1], NSW [1] | Fraxinus excelsior |
| Family Aphalaridae | |||
| 26 | Anoeconeossa communis | WO [1] | Eucalyptus sp. |
| 27 | Anomalopsylla insignita | MC [3] | Olearia paniculata |
| 28 | Anomalopsylla “Pollen Island” ˠ | MC [1] | Olearia odorata |
| 29 | Atmetocranium myersi | MC [1] | Weinmannia racemosa |
| 30 | Blastopsylla occidentalis | AK [1], WO [2] | Eucalyptus sp. |
| 31 | Cardiaspina fiscella | WO [1], WI [1] | Eucalyptus sp. |
| 32 | Creiis lituratus | WO [1] | Eucalyptus sp. |
| 33 | Cryptoneossa triangula | WI [1], SA [1] | Eucalyptus sp. |
| 34 | Ctenarytaina clavata A | MC [1], NN [1], NC [1], MK [1] | Kunzea ericoides |
| 35 | Ctenarytaina clavata B * | MB [1] | Kunzea ericoides |
| 36 | Ctenarytaina clavata C * | NN [1] | Leptospermum scoparius |
| 37 | Ctenarytaina clavata. D * | TO [2], WN [1] | Kunzea ericoides |
| 38 | Ctenarytaina eucalypti | MC [2], NC [3], WA [1], SL [3], GB [1], TO [2], SC [1], DN [1], FD [2], SI [2], VIC [1], SA [1] | Eucalyptus globulus |
| 39 | Ctenarytaina fuchsiae A | MC [4], FD [2], SC [1], NC [1], WD [5], NN [5], SI [7] | Fuchsia excorticata |
| 40 | Ctenarytaina fuchsiae B * | KA [3] | Fuchsia excorticata |
| 41 | Ctenarytaina fuchsiae C * | TO [1] | Fuchsia excorticata |
| 42 | Ctenarytaina longicauda | AK [2] | Lophostemon confertus |
| 43 | Ctenarytaina pollicaris | MC [2] | Leptospermum scoparium |
| 44 | Ctenarytaina pollicaris B * | MC [1], NN [3] | Leptospermum scoparium |
| 45 | Ctenarytaina “Short” ˠ | MC [5], NC [1], NN [1] | Leptospermum scoparium |
| 46 | Ctenarytaina sp. A * | NN [1] | Olearia paniculata |
| 47 | Ctenarytaina spatulata | NC [1], MC [2], SC [1], FD [3], SL [1], AK [1], WO [2], WI [2], SI [1], TO [1] | Eucalyptus nicholii |
| 48 | Ctenarytaina sp. B * | SI [2] | Kunzea ericoides |
| 49 | Ctenarytaina sp. C * | BP [1], WO [1] | Kunzea ericoides |
| 50 | Ctenarytaina sp. D * | NN [1] | Kunzea ericoides |
| 51 | Ctenarytaina sp. E * | WO [1] | Kunzea ericoides |
| 52 | Ctenarytaina thysanura | SC [1] | Eucalyptus sp. |
| 53 | Ctenarytaina sp. unknown ˠ | AK [2], WN [1] | Syzygium sp. |
| 54 | Eucalyptolyma maideni | SA [1] | Eucalyptus sp. |
| 55 | Glycaspis granulata | AK [1], WO [1], WI [1] | Eucalyptus sp. |
| Family Triozidae | |||
| 56 | Bactericera cockerelli | AK [1], BR [1], MC [1] | Solanum tuberosum |
| 57 | Casuarinicola australis | ND [1], QLD [1] | Casuarina sp. |
| 58 | Trioza acuta A | MC [3], MB [1] | Ozothamnus leptophyllus |
| 59 | Trioza acuta B * | MC [1] | Ozothamnus leptophyllus |
| 60 | Trioza eugeniae (T. adventicia) | MC [1], HB [1], SA [2] | Syzygium smithii |
| 61 | Trioza bifida | MC [6], NC [1], MB [1], SI [2], DN [1] | Olearia sp., Pseudowintera sp., Hebe sp., Dracophyllum sp. |
| 62 | Trioza “Brenda May” ˠ | SL [1], FD [1] | Olearia ilicifolia |
| 63 | Trioza colorata | MC [1], NC [1] | Halocarpus bidwillii |
| 64 | Trioza compressa | NN [4] | Olearia, Fuchsia |
| 65 | Trioza curta | NN [1] | Metrosideros |
| 66 | Trioza dacrydii | NN [1] | Halocarpus bidwillii |
| 67 | Trioza decurvata | MB [1], TO [1], NN [1] | Dracophyllum sp. |
| 68 | Trioza discariae | MC [2], NC [1] | Discaria toumatou |
| 69 | Trioza doryphora | MC [4] | Olearia ilicifolia, Coprosma sp. |
| 70 | Trioza emarginata | NC [1] | Coprosma sp. |
| 71 | Trioza falcata A | MC [1], NC [3] | Aristotelia fruticosa |
| 72 | Trioza falcata B * | NN [1] | Aristotellia fruticosa |
| 73 | Trioza fasciata | AK [1], NN [1] | Muehlenbeckia complexa |
| 74 | Trioza “Fortrose” ˠ | NN [1] | Elaeocarpus hookerianus |
| 75 | Trioza gourlayi | NN [1] | Olearia virgata |
| 76 | Trioza hebicola | NN [1] | Hebe sp. |
| 77 | Trioza irregularis | MC [8], NN [2] | Pseudopanax arboreus, Schefflera digitata. |
| 78 | Trioza “Massey” ˠ | MC [1], NN [1] | Olearia sp. |
| 79 | Trioza obscura | MB [1], NN [1] | Hebe sp. |
| 80 | Trioza “Omahuta” ˠ | NN [3] | Metrosiderosumbrellata, Brachygliottis repanda |
| 81 | Trioza panacis | MC [2], WN [1] | Pseudopanax crassifolius |
| 82 | Trioza “Price’s Valley” ˠ | MC [2] | Plagianthus regius |
| 83 | Trioza sp. A * | NN [3] | Pittosporum divaricatum |
| 84 | Trioza sp. B ? | NN [1] | Olearia arborescens |
| 85 | Trioza sp. C * | MC [1] | Pseudopanax edgerleyi |
| 86 | Trioza sp. D ? | MC [1] | Olearia virgata |
| 87 | Trioza subacuta | MC [2] | Olearia avicennifolia |
| 88 | Trioza subvexa | NN [4] | Olearia avicennifolia |
| 89 | Trioza vitreoradiata | SL [3], FD [1], CL [1], MC [3], NN [2], GB [1], TK [2], MB [2], AK [1] | Pittosporum crassifolium, Pittosporum spp. |
| 90 | Triozid sp. ˠ | AK [2], CL [1], NSW [1] | Casuarina sp. |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martoni, F.; Bulman, S.; Pitman, A.; Taylor, G.; Armstrong, K. DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha). Diversity 2018, 10, 50. https://doi.org/10.3390/d10030050
Martoni F, Bulman S, Pitman A, Taylor G, Armstrong K. DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha). Diversity. 2018; 10(3):50. https://doi.org/10.3390/d10030050
Chicago/Turabian StyleMartoni, Francesco, Simon Bulman, Andrew Pitman, Gary Taylor, and Karen Armstrong. 2018. "DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha)" Diversity 10, no. 3: 50. https://doi.org/10.3390/d10030050
APA StyleMartoni, F., Bulman, S., Pitman, A., Taylor, G., & Armstrong, K. (2018). DNA Barcoding Highlights Cryptic Diversity in the New Zealand Psylloidea (Hemiptera: Sternorrhyncha). Diversity, 10(3), 50. https://doi.org/10.3390/d10030050





