Genetic, Bio-Agronomic, and Nutritional Characterization of Kale (Brassica Oleracea L. var. Acephala) Diversity in Apulia, Southern Italy
Abstract
1. Introduction
2. Materials and Methods
2.1. Germplasm Collection
2.2. Phenotypic Evaluation
2.3. DNA Extraction and SSR Marker analysis
2.4. Bioactive Compounds and Antioxidant Activity
3. Results
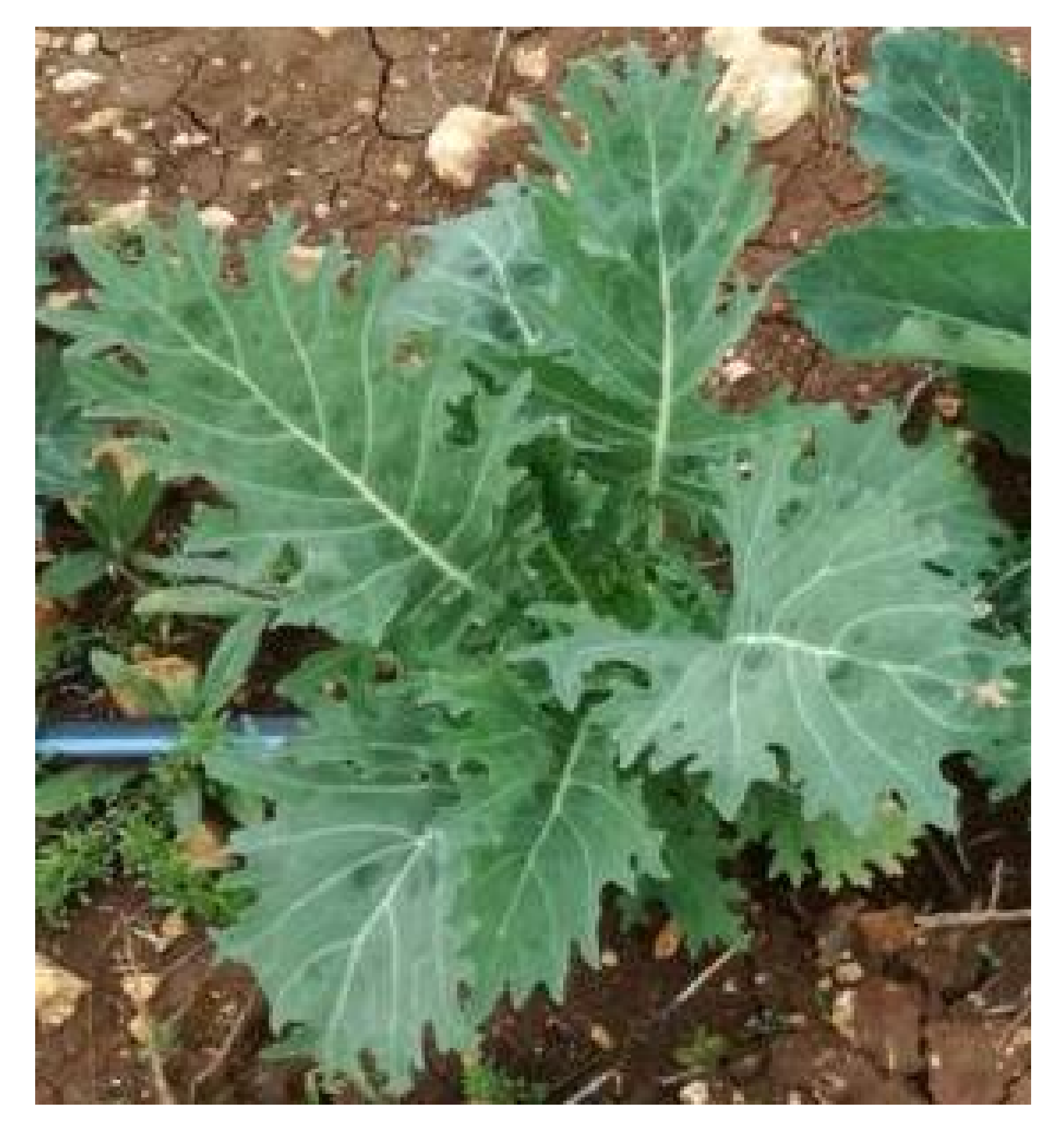
3.1. Establishment of a Kale Germplasm Collection
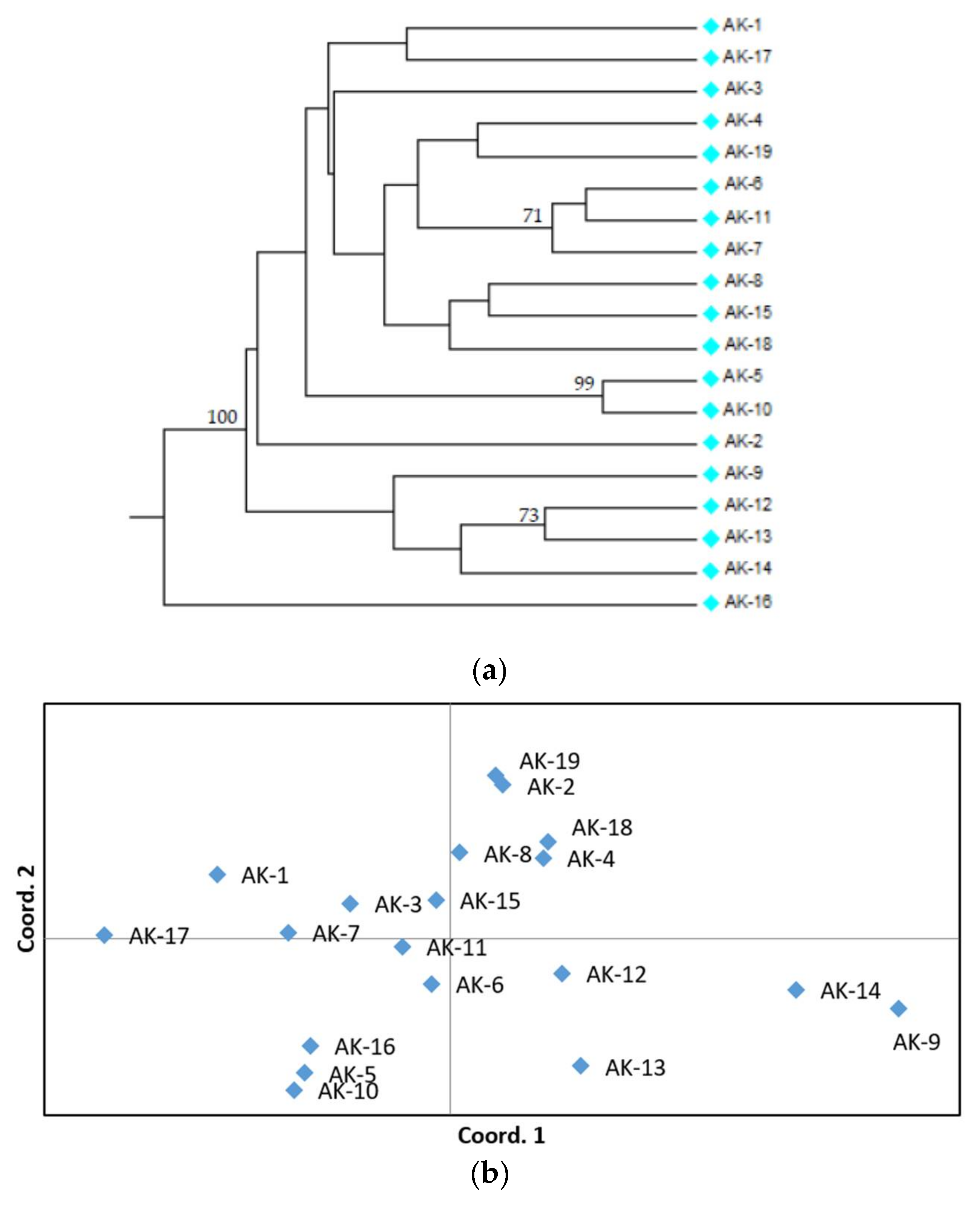
3.2. SSR Polymorphism and Genetic Relationships among Accessions
3.3. Phenotypic Variation
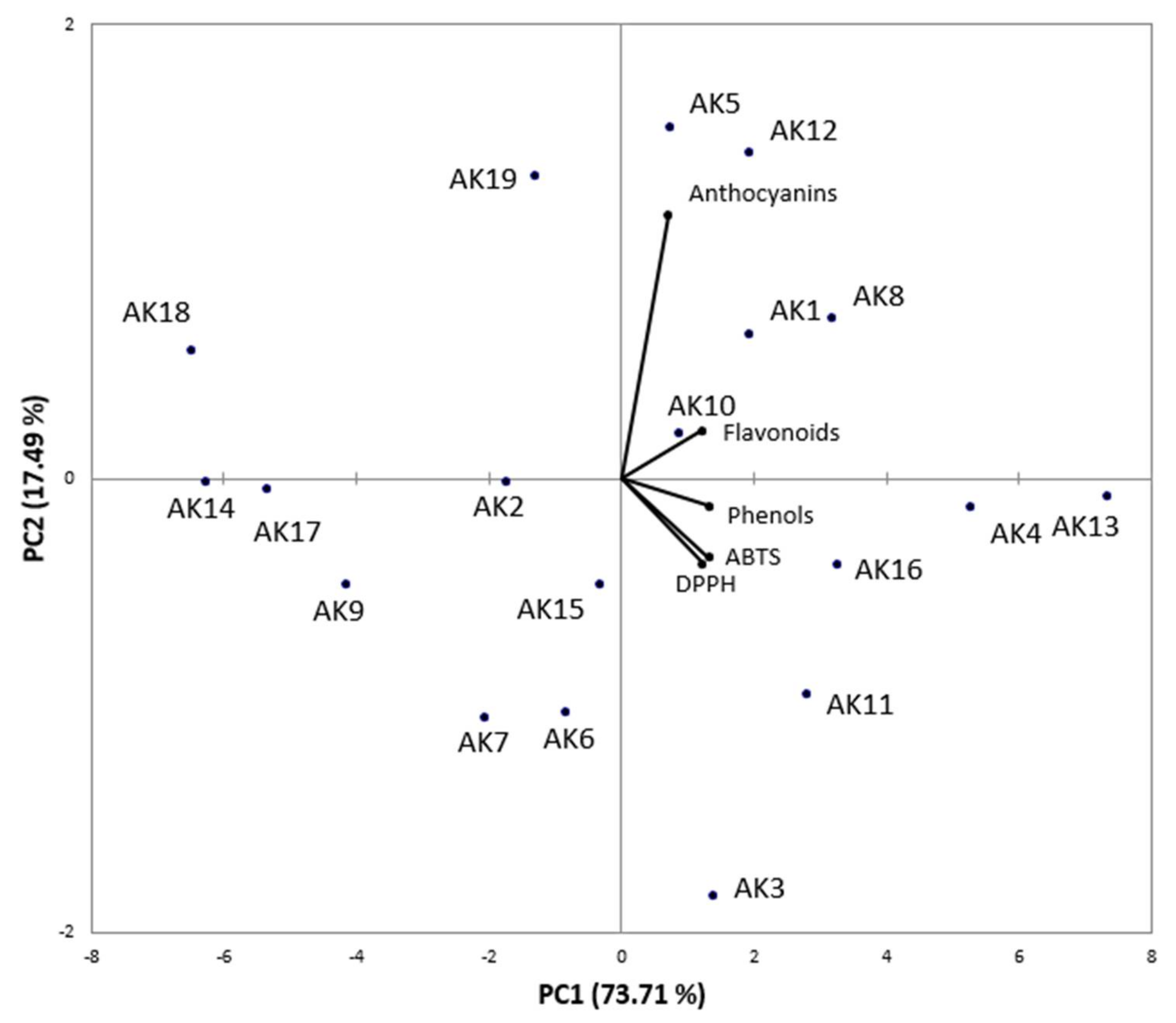
3.4. Bioactive Compounds and Correlation with Antioxidant Activity
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Balkaya, A.; Yanmaz, R. Promising kale (Brassica oleracea var. acephala) populations from Black Sea region in Turkey. N. Z. J. Crop Hortic. Sci. 2005, 33, 1–7. [Google Scholar] [CrossRef]
- Dixon, G.R. Vegetable Brassicas and Related Crucifers; Crop Production Science in Horticulture Series, 14; CABI: Wallingford, UK, 2007; Volume 60, pp. 205–211. ISBN 9780851993959. [Google Scholar]
- Cartea, M.E.; Picoaga, A.; Soengas, P.; Ordás, A. Morphological characterization of kale populations from northwestern Spain. Euphytica 2003, 29, 25–32. [Google Scholar] [CrossRef]
- Giorgetti, L.; Giorgi, G.; Cherubini, E.; Gervasi, P.G.; Della Croce, C.M.; Longo, V.; Bellani, L. Screening and identification of major phytochemical compounds in seeds, sprouts and leaves of Tuscan black kale Brassica oleracea (L.) ssp acephala (DC) var. sabellica L. Nat. Prod. Res. 2017, 23, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Christensen, S.; von Bothmer, R.; Poulsen, G.; Maggioni, L.; Phillip, M.; Andersen, B.A. AFLP analysis of genetic diversity in leafy kale (Brassica oleracea L. convar. acephala (DC.) Alef.) landraces, cultivars and wild populations in Europe. Genet. Res. Crop Evol. 2011, 58, 657–666. [Google Scholar] [CrossRef]
- Šamec, D.; Urlić, B.; Salopek-Sondi, B. Kale (Brassica oleracea var. acephala) as a superfood: Review of the scientific evidence behind. Crit. Rev. Food Sci. Nutr. 2018, 20, 1–37. [Google Scholar] [CrossRef]
- Greifenberg, A. Cavolo da foglia (Brassica oleracea L. Var. acephala DC.). In Orticoltura, 1st ed.; Bianco, V.V., Pimpini, F., Eds.; Pàtron: Bologna, Italy, 1990; pp. 1–985. ISBN 9788855520997. [Google Scholar]
- Elia, A.; Santamaria, P. Biodiversity in vegetable crops: A heritage to save. The case of the Puglia region. Ital. J. Agron. 2013, 8, 21–34. [Google Scholar] [CrossRef]
- Tautz, D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res. 1989, 17, 6463–6471. [Google Scholar] [CrossRef] [PubMed]
- Ragusa, L.; Branca, F.; Licciardello, F.; Restuccia, C.; Melilli, M.G.; Argento, S.; Raccuia, S.A. Development of modified atmosphere packages on the quality of sicilian kale (Brassica oleracea var. acephala) shoots. Acta Hortic. 2013, 1005, 315–321. [Google Scholar] [CrossRef]
- Sikora, E.; Bodziarczyk, I. Composition and antioxidant activity of kale (Brassica oleracea L. var. acephala) raw and cooked. Acta Sci. Pol. Technol. Aliment. 2012, 11, 239–248. [Google Scholar] [PubMed]
- Velasco, P.; Cartea, M.E.; Gonzalez, C.; Vilar, M.; Ordás, A. Factors affecting the glucosinolate content of kale (Brassica oleracea acephala group). J. Agric. Food Chem. 2007, 55, 955–962. [Google Scholar] [CrossRef] [PubMed]
- Ambrosone, C.B.; McCann, S.E.; Freudenheim, J.L.; Marshall, J.R.; Zhang, Y.; Shields, P.G. Breast cancer risk in premenopausal women is inversely associated with consumption of broccoli, a source of isothiocyanates, but is not modified by GST genotype. J. Nutr. 2004, 134, 1134–1138. [Google Scholar] [CrossRef] [PubMed]
- Cartea, M.; Francisco, M.; Soengas, P.; Velasco, P. Phenolic compounds in Brassica vegetables. Molecules 2011, 16, 251–280. [Google Scholar] [CrossRef] [PubMed]
- Heber, D.; Bowerman, S. Applying science to changing dietary patterns. J. Nutr. 2001, 131, 3078–3081. [Google Scholar] [CrossRef]
- Verhoeven, D.T.; Goldbohm, R.A.; Van Poppel, G.; Verhagen, H.; Van den Brandt, P.A. Epidemiological studies on brassica vegetables and cancer risk. Cancer Epidemiol. Biomark. Prev. 1996, 5, 733–748. [Google Scholar]
- Gawlik-Dziki, U. Effect of hydrothermal treatment on the antioxidant properties of broccoli (Brassica oleracea var. botrytis italica) florets. Food Chem. 2008, 109, 393–401. [Google Scholar] [CrossRef] [PubMed]
- De Pascale, S.; Maggio, A.; Pernice, R.; Fogliano, V.; Barbieri, G. Sulphur fertilization may improve the nutritional value of Brassica rapa L. subsp. Sylvestris. Eur. J. Agron. 2017, 26, 418–424. [Google Scholar] [CrossRef]
- Kusznierewicz, B.; Śmiechowska, A.; Bartoszek, A.; Namieśnik, J. The effect of heating and fermenting on antioxidant properties of white cabbage. Food Chem. 2008, 108, 853–861. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. Isolation of plant DNA from fresh tissue. Focus 1990, 12, 13–14. [Google Scholar]
- Suwabe, K.; Iketani, H.; Nunome, T.; Kage, T.; Hirai, M. Isolation and characterization of microsatellites in Brassica rapa L. Theor. Appl. Genet. 2002, 104, 1092–1098. [Google Scholar] [CrossRef] [PubMed]
- Lowe, A.J.; Moule, C.; Trick, M.; Edwards, K.J. Efficient large-scale development of microsatellites for marker and mapping applications in Brassica crop species. Theor. Appl. Genet. 2004, 108, 1103–1112. [Google Scholar] [CrossRef] [PubMed]
- El-Esawi, M.A.; Germaine, K.; Bourke, P.; Malone, R. Genetic diversity and population structure of Brassica oleracea germplasm in Ireland using SSR markers. C. R. Biol. 2016, 339, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Smouse, P.E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research: An update. Bioinformatics 2012, 28, 2537–2539. [Google Scholar] [CrossRef] [PubMed]
- Kalinowski, S.T.; Taper, M.L.; Marshall, T.C. Revising how the computer program CERVUS accommodates genotyping error increases success in paternity assignment. Mol. Ecol. 2007, 16, 1099–1106. [Google Scholar] [CrossRef] [PubMed]
- Takezaki, N.; Nei, M.; Tamura, K. POPTREEW: Web version of POPTREE for constructing population trees from allele frequency data and computing some other quantities. Mol. Biol. Evol. 2014, 31, 1622–1624. [Google Scholar] [CrossRef] [PubMed]
- Cosmai, L.; Caponio, F.; Pasqualone, A.; Paradiso, V.M; Summo, C. Evolution of the oxidative stability, bio-active compounds and color characteristics of non-thermally treated vegetable pâtés during frozen storage. J. Sci. Food Agric. 2017, 9, 4904–4911. [Google Scholar] [CrossRef] [PubMed]
- Pasqualone, A.; Bianco, A.M.; Paradiso, V.M.; Summo, C.; Gambacorta, G.; Caponio, F.; Blanco, A. Production and characterization of functional biscuits obtained from purple wheat. Food Chem. 2015, 180, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Downes, K.; Chope, G.A.; Terry, L.A. Postharvest application of ethylene and methylcyclopropene either before or after curing affects onion (Allium cepa L.) bulb. Postharvest Biol. Technol. 2010, 55, 36–44. [Google Scholar] [CrossRef]
- Huang, D.; Ou, B.; Prior, R.L. The chemistry behind antioxidant capacity assays. J. Agric. Food Res. 2005, 53, 1841–1856. [Google Scholar] [CrossRef] [PubMed]
- Kasprzak, K.; Oniszczuk, T.; Wójtowicz, A.; Waksmundzka-Hajnos, M.; Olech, M.R.; Polak, R.; Oniszczuk, A. Phenolic acid content and antioxidant properties of extruded corn snacks enriched with kale. J. Anal. Methods Chem. 2018, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Leja, M.; Kamińska, I.; Kołton, A. phenolic compounds as the major antioxidants in red cabbage. Folia Hort. 2010, 22, 19–24. [Google Scholar] [CrossRef]
- Mageney, V.; Neugart, S.; Albach, D.C. A guide to the variability of flavonoids in Brassica oleracea. Molecules 2017, 22, 252. [Google Scholar] [CrossRef] [PubMed]
- Naguib, A.E.-M.M.; El-Baz, F.K.; Salama, Z.A.; Hanaa, H.A.E.B; Ali, H.F.; Gaafar, A.A. Enhancement of phenolics, flavonoids and glucosinolates of Broccoli (Brassica olaracea, var. Italica) as antioxidants in response to organic and bio-organic fertilizers. J. Saudi Soc. Agric. Sci. 2012, 11, 135–142. [Google Scholar] [CrossRef]
- Jaiswal, A.K.; Abu-Ghannam, N.; Gupta, S. A comparative study on the polyphenolic content, antibacterial activity and antioxidant capacity of different solvent extracts of Brassica oleracea vegetables. Int. J. Food Sci. Technol. 2012, 47, 223–231. [Google Scholar] [CrossRef]

| Locus | Na | Ne | Size Range (bp) | Ho | He | I | F | PIC |
|---|---|---|---|---|---|---|---|---|
| BRMS-005 | 2 | 1.630 | 162–166 | 0.183 | 0.355 | 0.514 | 0.481 | 0.373 |
| BRMS-015 | 5 | 2.009 | 253–265 | 0.343 | 0.439 | 0.714 | 0.223 | 0.587 |
| BRMS-016 | 5 | 2.124 | 125–205 | 0.530 | 0.480 | 0.825 | −0.079 | 0.623 |
| BRMS-019 | 2 | 1.372 | 266–268 | 0.211 | 0.218 | 0.332 | 0.039 | 0.298 |
| BRMS-033 | 2 | 1.116 | 244–247 | 0.069 | 0.078 | 0.123 | 0.114 | 0.086 |
| BRMS-036 | 5 | 2.357 | 143–155 | 0.921 | 0.562 | 0.895 | −0.657 | 0.530 |
| BRMS-042-2 | 3 | 1.727 | 230–248 | 0.480 | 0.387 | 0.589 | −0.245 | 0.473 |
| BRMS-050 | 5 | 2.518 | 125–192 | 0.727 | 0.578 | 0.993 | −0.275 | 0.630 |
| OL11-G11 | 4 | 1.954 | 152–228 | 0.404 | 0.461 | 0.740 | 0.115 | 0.584 |
| OL12-A04 | 7 | 2.692 | 147–169 | 0.492 | 0.583 | 1.017 | 0.174 | 0.743 |
| Ra2-A01 | 2 | 1.333 | 106–108 | 0.205 | 0.201 | 0.304 | −0.031 | 0.267 |
| Ra2-E12 | 4 | 2.133 | 160–168 | 0.555 | 0.507 | 0.808 | −0.099 | 0.567 |
| Mean | 4 | 1.914 | 0.427 | 0.404 | 0.654 | −0.039 | 0.480 |
| Source | df | Sum of Squares | Variance | Percentage on Total Variance (%) |
|---|---|---|---|---|
| Among Accessions | 18 | 513.7 | 2.9 | 34 |
| Within Accessions | 132 | 747.0 | 5.7 | 66 |
| Tota | 150 | 1260.7 | 8.5 |
| Source of Variation | df | PH | HT | FT | Y1 | Y2 | SW |
|---|---|---|---|---|---|---|---|
| Accession | 18 | *** | ** | *** | * | ** | n.s. |
| Block | 2 | n.s. | *** | *** | n.s. | n.s. | n.s. |
| df: degree of freedom |
| Accessions | PH (cm) | HT (Days) | FT (Days) | Y1 (g/Plant) | Y2 (g/Plant) |
|---|---|---|---|---|---|
| AK-1 | 34.2 ± 2.0 a | 152 ± 0.9 e | 170 ± 2.0 e | 55.6 ± 16.2 a,b | 38.6 ± 11.9 c |
| AK-2 | 29.6 ± 0.8 a,b,c | 156 ± 0.9 c | 174 ± 0.9 c | 188.8 ± 26.6 a | 58.6 ± 14.3 b,c |
| AK-3 | 30.6 ± 0.5 a,b | 156 ± 1.0 c | 174 ± 1.4 c | 118.0 ± 25.8 a,b | 46.4 ± 23.0 b,c |
| AK-4 | 30.4 ± 2.1 a,b | 152 ± 1.8 e | 170 ± 0.8 e | 123.2 ± 57.2 a,b | 69.2 ± 20.5 a,b,c |
| AK-5 | 32.4 ± 2.2 a,b | 153 ± 0.9 d | 171 ± 1.0 d | 63.4 ± 16.2 a,b | 58.0 ± 12.7 b,c |
| AK-6 | 25.4 ± 1.6 a,b,c,d | 148 ± 1.1 h | 166 ± 1.4 h | 69.2 ± 21.2 a,b | 70.8 ± 19.7 a,b,c |
| AK-7 | 26.0 ± 0.7 a,b,c,d | 150 ± 1.2 g | 168 ± 0.9 g | 93.2 ± 8.4 a,b | 176.4 ± 9.2 a |
| AK-8 | 28.0 ± 1.8 a,b,c | 158 ± 1.3 b | 176 ± 0.8 b | 103.2 ± 39.3 a,b | 86.2 ± 17.1 a,b,c |
| AK-9 | 25.4 ± 1.6 a,b,c,d | 120 ± 1.2 i | 138 ± 0.7 i | 35.6 ± 33.0 b | 56.0 ± 21.1 b,c |
| AK-10 | 25.6 ± 0.7 a,b,c,d | 150 ± 1.5 g | 168 ± 1.1 g | 113.4 ± 35.3 a,b | 67.4 ± 18.8 a,b,c |
| AK-11 | 26.8 ± 0.4 a,b,c,d | 148 ± 1.4 h | 166 ± 1.2 h | 75.2 ± 22.5 a,b | 54.8 ± 7.2 b,c |
| AK-12 | 24.4 ± 2.5 b,c,d | 153 ± 0.9 d | 171 ± 0.9 d | 94.8 ± 43.4 a,b | 86.6 ± 16.1 a,b,c |
| AK-13 | 20.6 ± 1.4 c,d | 150 ± 0.9 g | 168 ± 0.9 g | 45.8 ± 26.8 a,b | 62.2 ± 19.9 b,c |
| AK-14 | 18.6 ± 0.8 d | 156 ± 1.1 c | 174 ± 0.9 c | 85.8 ± 39.4 ab | 149.8 ± 24.7 a,b |
| AK-15 | 25.2 ± 1.8 a,b,cd | 158 ± 1.8 b | 176 ± 1.0 b | 34.6 ± 14.8 b | 50.4 ± 26.5 b,c |
| AK-16 | 26.6 ± 0.2 a,b,c,d | 158 ± 1.2 b | 176 ± 1.8 b | 150.0 ± 29.2 a,b | 123.4 ± 15.7 a,b,c |
| AK-17 | 30.6 ± 0.8 a,b | 159 ± 1.2 a | 177 ± 1.2 a | 117.8 ± 15.3 a,b | 88.8 ± 48.8 a,b,c |
| AK-18 | 27.0 ± 2.4 a,b,c,d | 151 ± 0.9 f | 169 ± 1.4 f | 114.8 ± 20.6 a,b | 86.2 ± 26.2 a,b,c |
| AK-19 | 26.0 ± 1.0 a,b,c,d | 151 ± 0.9 f | 169 ± 1.0 f | 61.2 ± 11.5 a,b | 68.6 ± 12.2 a,b,c |
| Mean | 27.0 ± 0.9 | 151.5 ± 1.9 | 169.5 ± 1.8 | 91.8 ± 9.2 | 78.9 ± 8.2 |
| Accessions | Total Phenols (mg Gallic ac./g dw) | Total Flavonoids (mg Quercetin/g dw) | Total Anthocyanins (mg 3-o Glucoside/100 g dw) | ABTS Assay (μmol eq Trolox/g dw) | DPPH Assay (μmol eq Trolox/g dw) |
|---|---|---|---|---|---|
| AK-1 | 3.9 ± 0.2 a,b | 76.5 ± 3.2 a,b | 17.2 ± 0.6 a,b,c,d | 26.5 ± 1.9 a,b,c | 34.6 ± 0.2 a,b,c |
| AK-2 | 3.5 ± 0.1 a,b | 57.1 ± 1.0 c,d,e | 14.9 ± 2.1 a,b,c,d | 24.6 ± 3.0 a,b | 36.3 ± 3.6 a,b,c |
| AK-3 | 3.9 ± 0.2 a,b | 62.8 ± 0.4 b,c,d,e | 9.8 ± 1.4 d | 29.7 ± 2.4 a,b | 45.4 ± 4.1 a,b,c |
| AK-4 | 4.1 ± 0.2 a,b | 81.9 ± 2.2 a | 16.5 ± 3.4 a,b,c,d | 29.9 ± 2.4 a,b,c | 50.6 ± 7.4 a,b,c |
| AK-5 | 3.7 ± 0.1 a,b | 69.2 ± 0.9 a,b,c,d | 20.8 ± 2.5 a | 24.5 ± 1.0 a,b,c | 35.5 ± 1.9 a,b,c |
| AK-6 | 3.7 ± 0.2 a,b | 64.2 ± 0.5 b,c,d,e | 10.4 ± 0.5 c,d | 25.1 ± 2.5 a,b,c | 36.1 ± 1.8 a,b,c |
| AK-7 | 3.6 ± 0.4 a,b | 60.2 ± 5.7 b,c,d,e | 9.9 ± 0.7 d | 24.3 ± 6.7 a,b,c | 33.2 ± 6.5 a,b,c |
| AK-8 | 4.1 ± 0.1 a,b | 69.4 ± 0.7 a,b,c,d | 19.6 ± 0.5 a,b,c | 27.7 ± 0.8 b,c | 44.0 ± 1.3 a,b,c |
| AK-9 | 3.3 ± 0.3 a,b | 55.9 ± 1.2 c,d,e | 11.2 ± 2.4 b,c,d | 22.5 ± 5.4 a,b,c | 27.0 ± 8.1 b,c |
| AK-10 | 3.9 ± 0.5 a,b | 56.3 ± 0.3 c,d,e | 17.7 ± 0.1 a,b,c,d | 26.4 ± 8.0 a,b,c | 44.0 ± 13.5 a,b,c |
| AK-11 | 4.0 ± 0.4 a,b | 73.1 ± 8.8 a,b,c | 12.6 ± 2.5 a,b,c,d | 27.9 ± 6.3 a,b,c | 48.3 ± 9.7 a,b,c |
| AK-12 | 3.8 ± 0.2 a,b | 75.5 ± 5.0 a,b | 20.3 ± 2.9 a,b | 25.8 ± 2.7 a | 34.3 ± 1.5 a,b,c |
| AK-13 | 4.5 ± 0.1 a | 77.4 ± 1.5 a,b | 19.1 ± 0.2 a,b,c,d | 31.4 ± 0.4 c | 61.3 ± 2.1 a |
| AK-14 | 3.0 ± 0.1 b | 51.1 ± 2.2 e | 11.9 ± 0.9 a,b,c,d | 20.1 ± 1.9 a,b,c | 23.2 ± 2.5 b,c |
| AK-15 | 3.4 ± 0.2 ab | 61.4 ± 2.6 b,c,d,e | 14.2 ± 0.2 a,b,c,d | 25.4 ± 3.5 a,b,c | 49.8 ± 3.4 a,b,c |
| AK-16 | 3.8 ± 0.3 a,b | 75.5 ± 4.9 a,b | 15.1 ± 0.9 a,b,c,d | 27.9 ± 6.8 b,c | 52.5 ± 13.4 a,b |
| AK-17 | 3.1 ± 0.3 b | 54.0 ± 5.3 d,e | 12.2 ± 1.0 a,b,c,d | 21.4 ± 4.5 c | 23.7 ± 7.2 b,c |
| AK-18 | 3.0 ± 0.1 b | 53.8 ± 2.3 d,e | 13.5 ± 0.1 a,b,c,d | 19.5 ± 2.0 a,b,c | 18.3 ± 1.5 c |
| AK-19 | 3.4 ± 0.1 a,b | 63.4 ± 0.5 b,c,d,e | 19.3 ± 3.1 a,b,c | 23.6 ± 1.3 a,b,c | 32.2 ± 3.8 a,b,c |
| Mean | 3.7 ± 0.2 | 65.2 ± 2.6 | 15.1 ± 1.4 | 25.5 ± 3.3 | 38.4 ± 4.9 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lotti, C.; Iovieno, P.; Centomani, I.; Marcotrigiano, A.R.; Fanelli, V.; Mimiola, G.; Summo, C.; Pavan, S.; Ricciardi, L. Genetic, Bio-Agronomic, and Nutritional Characterization of Kale (Brassica Oleracea L. var. Acephala) Diversity in Apulia, Southern Italy. Diversity 2018, 10, 25. https://doi.org/10.3390/d10020025
Lotti C, Iovieno P, Centomani I, Marcotrigiano AR, Fanelli V, Mimiola G, Summo C, Pavan S, Ricciardi L. Genetic, Bio-Agronomic, and Nutritional Characterization of Kale (Brassica Oleracea L. var. Acephala) Diversity in Apulia, Southern Italy. Diversity. 2018; 10(2):25. https://doi.org/10.3390/d10020025
Chicago/Turabian StyleLotti, Concetta, Paolo Iovieno, Isabella Centomani, Angelo R. Marcotrigiano, Valentina Fanelli, Giancarlo Mimiola, Carmine Summo, Stefano Pavan, and Luigi Ricciardi. 2018. "Genetic, Bio-Agronomic, and Nutritional Characterization of Kale (Brassica Oleracea L. var. Acephala) Diversity in Apulia, Southern Italy" Diversity 10, no. 2: 25. https://doi.org/10.3390/d10020025
APA StyleLotti, C., Iovieno, P., Centomani, I., Marcotrigiano, A. R., Fanelli, V., Mimiola, G., Summo, C., Pavan, S., & Ricciardi, L. (2018). Genetic, Bio-Agronomic, and Nutritional Characterization of Kale (Brassica Oleracea L. var. Acephala) Diversity in Apulia, Southern Italy. Diversity, 10(2), 25. https://doi.org/10.3390/d10020025









