Genome-Wide Identification and Analysis of bZIP Transcription Factors in Coptis chinensis Reveals Their Regulatory Roles in Stress Responses
Abstract
1. Introduction
2. Results
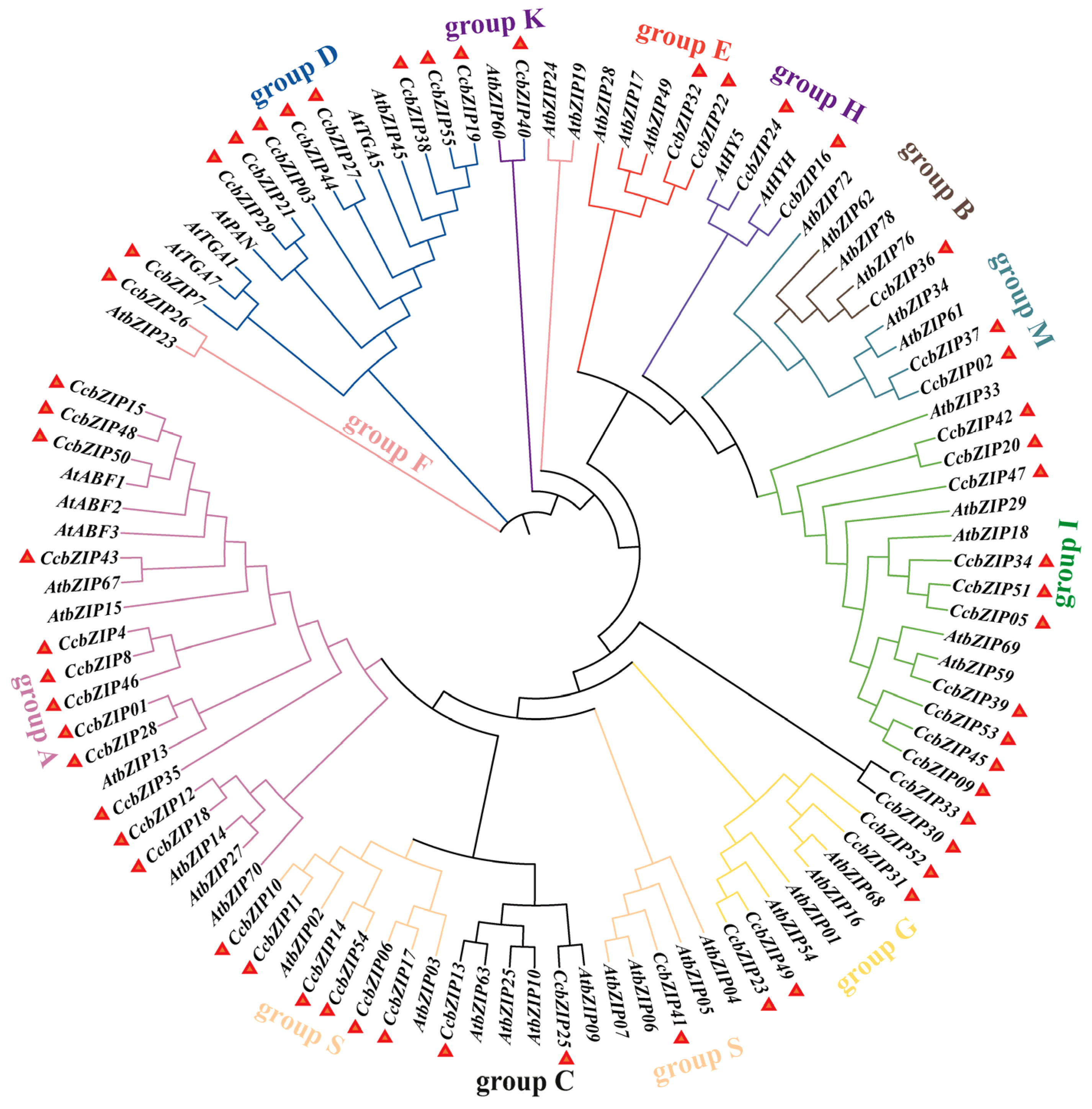
2.1. Genome-Wide Identification, Classification and Phylogenetic Analysis of CcbZIP Genes
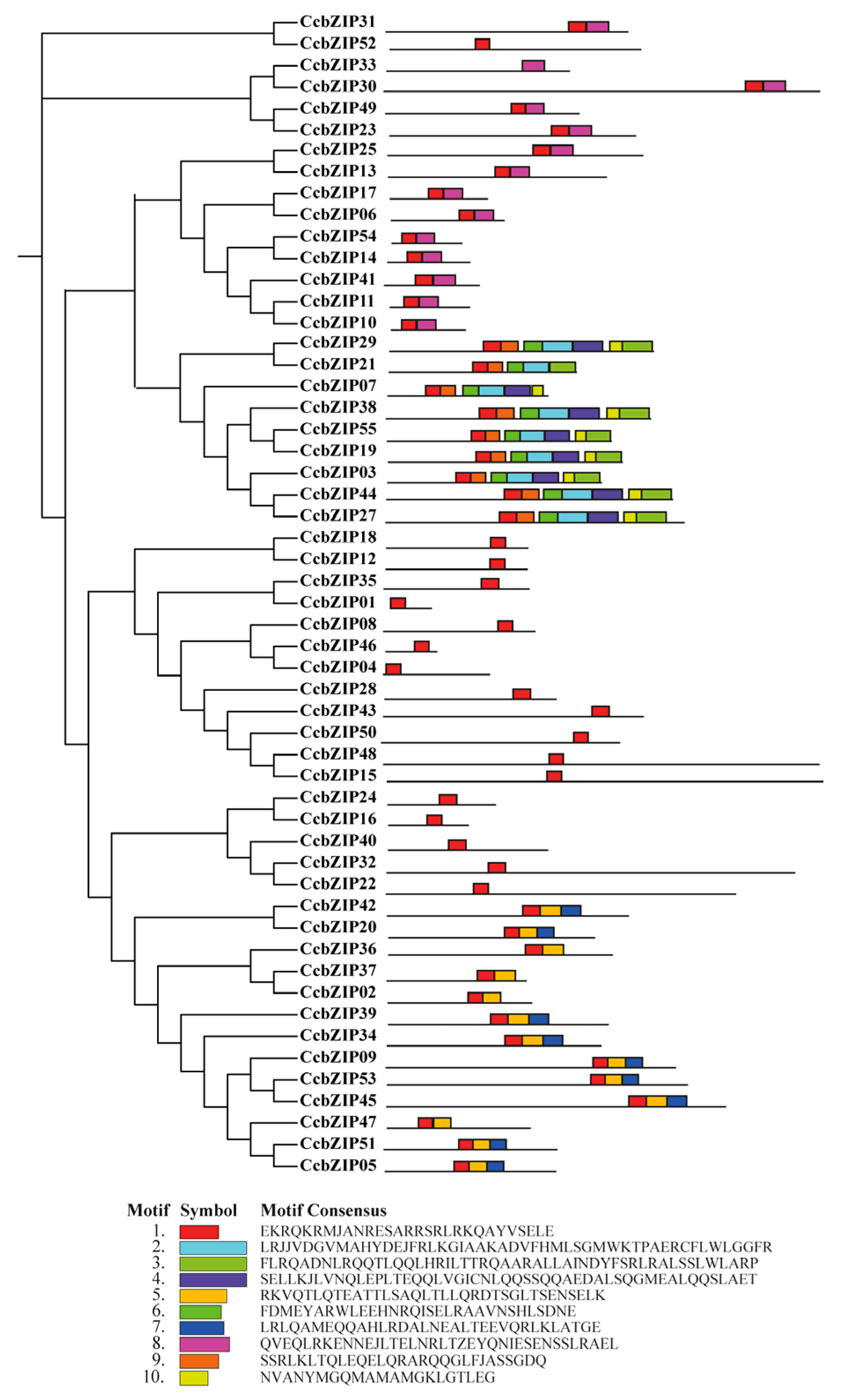
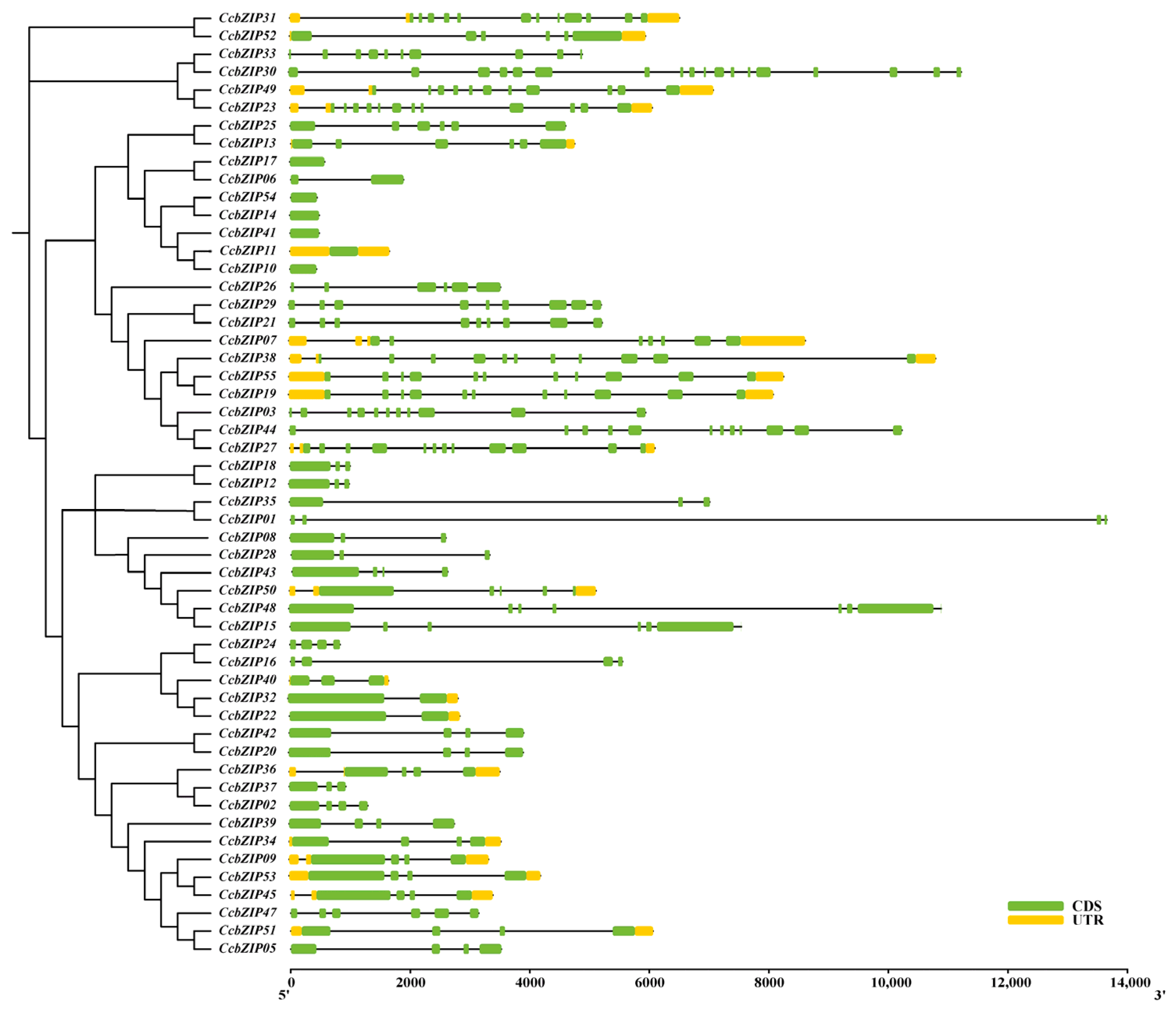
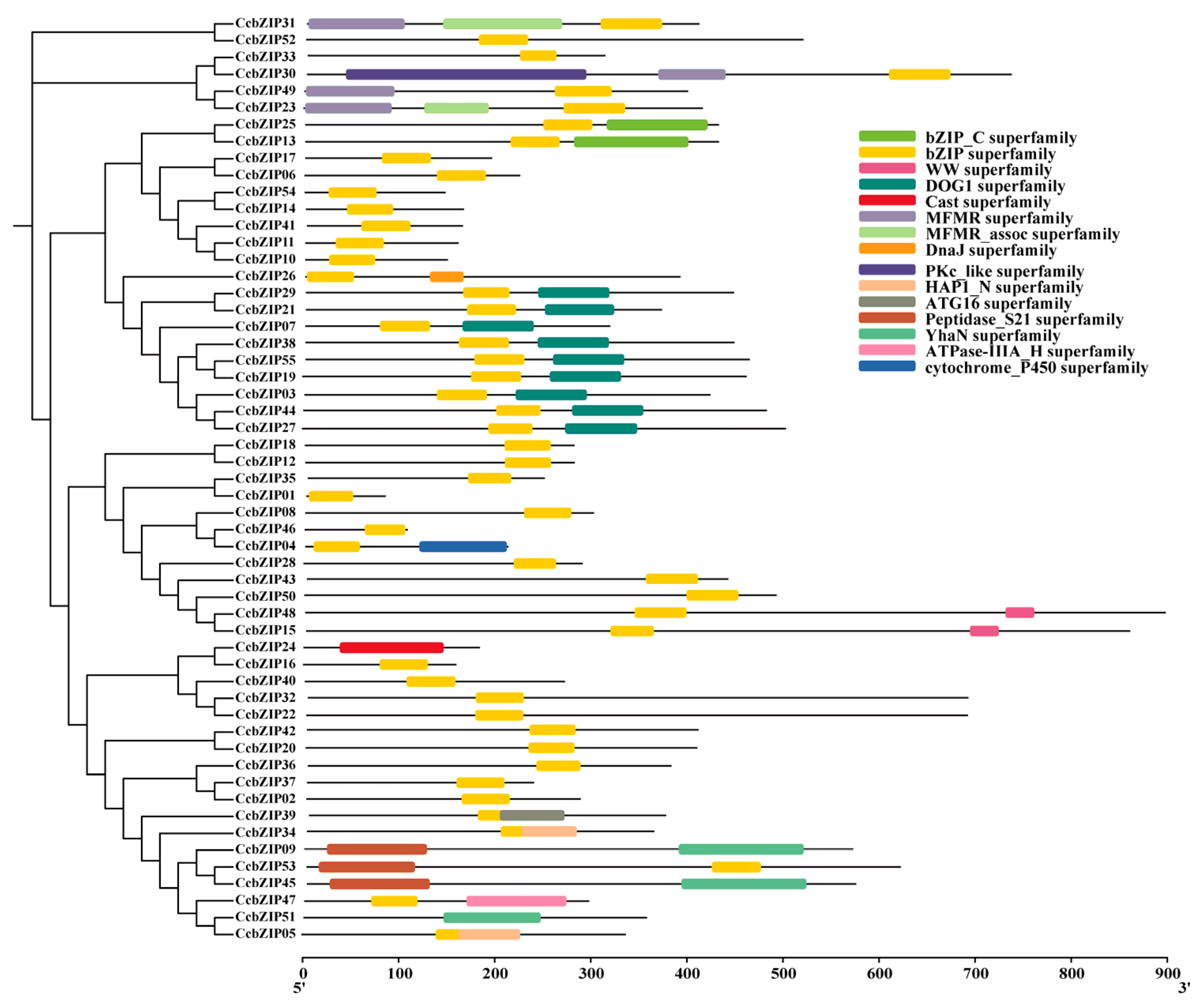
2.2. Gene Structure and Conserved Domain Analysis
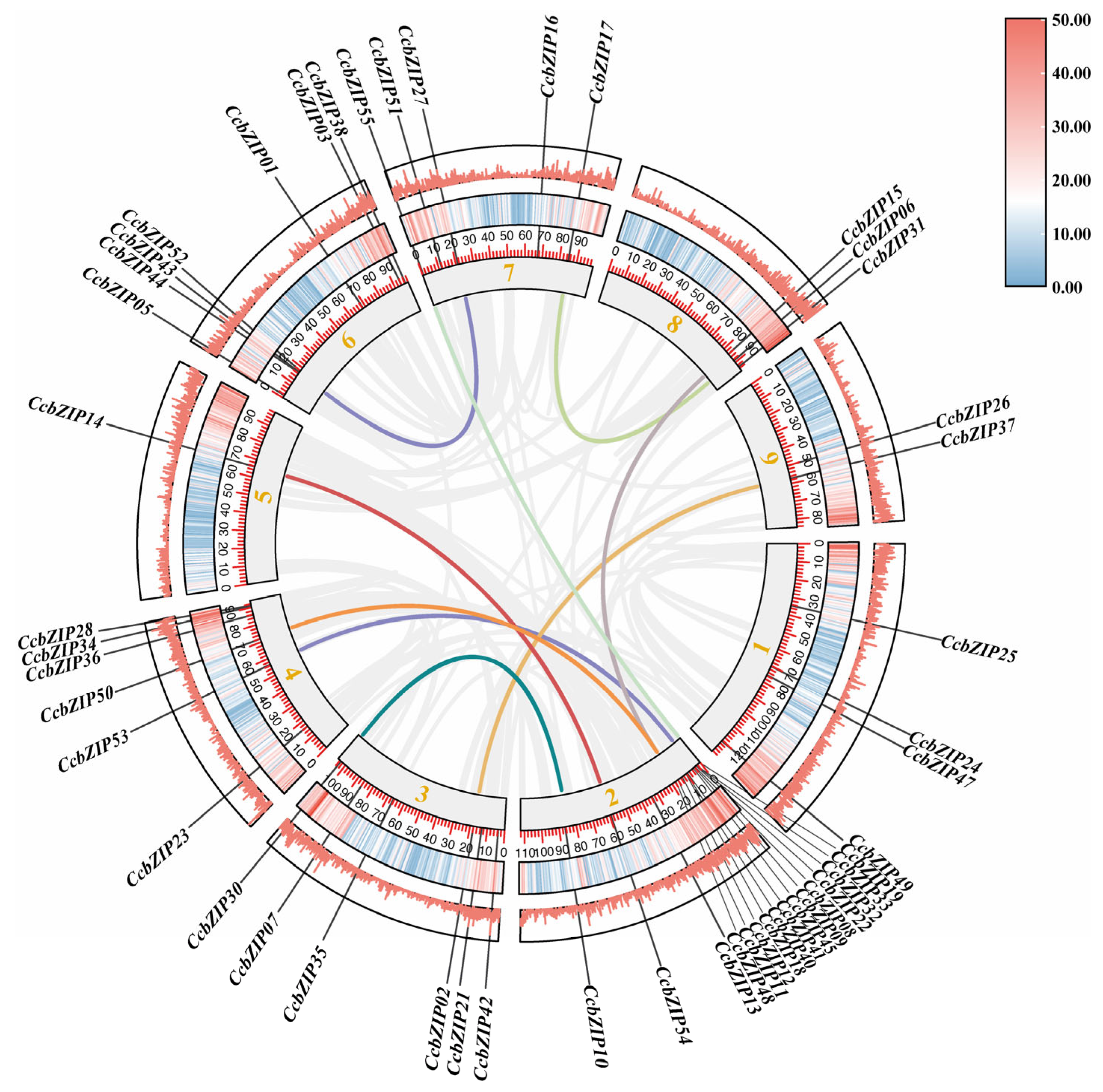
2.3. Chromosomal Distribution, Gene Duplications and Synteny Analysis of CcbZIPs
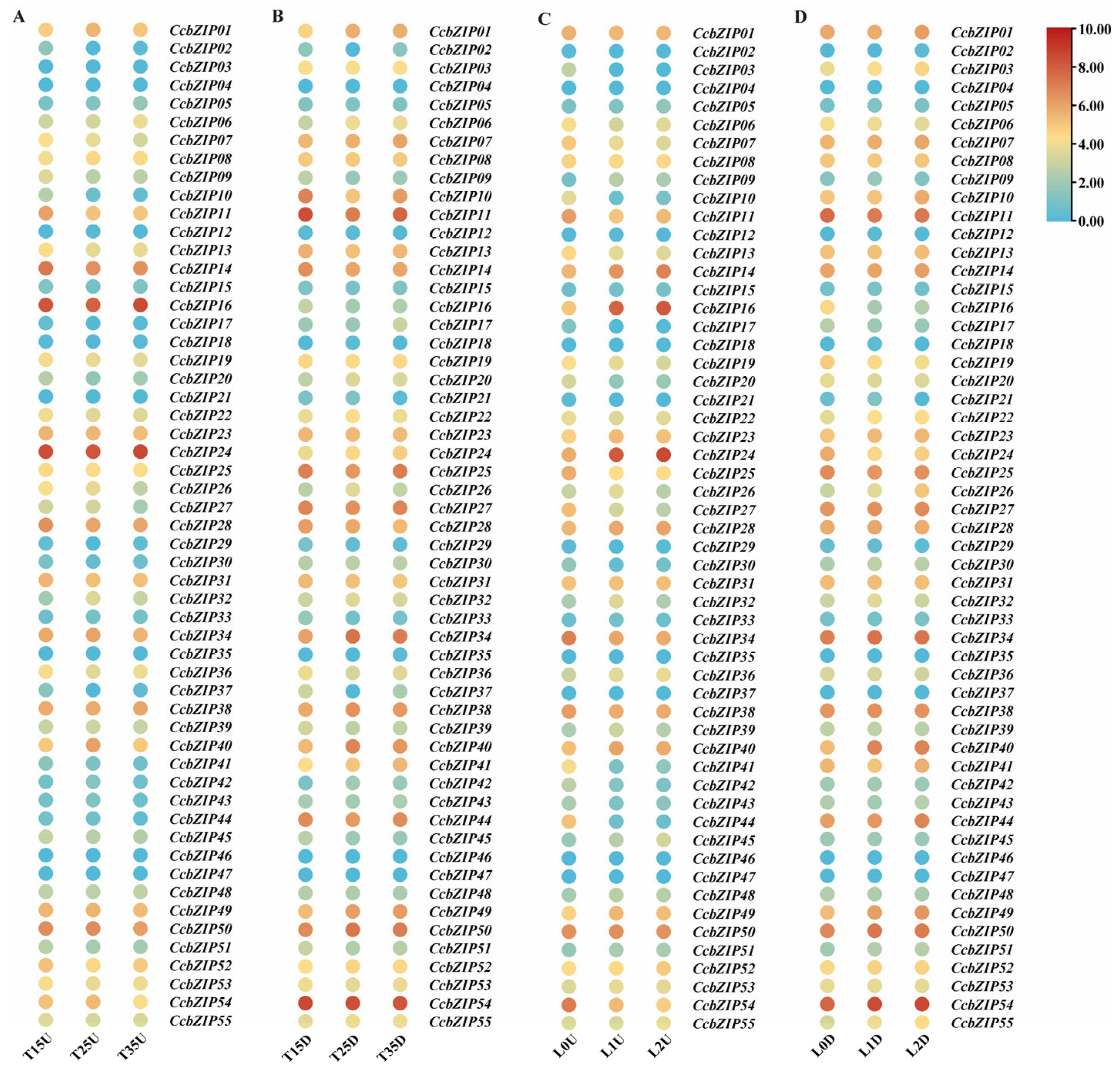
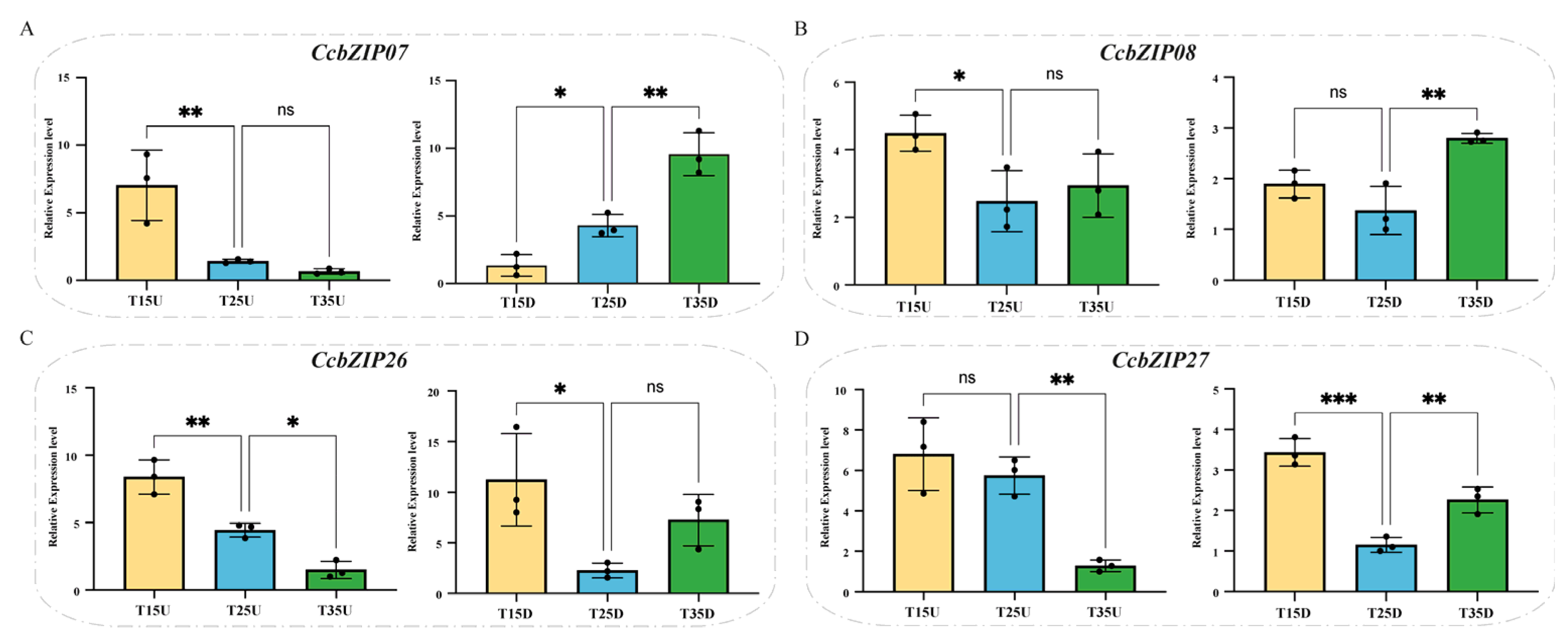
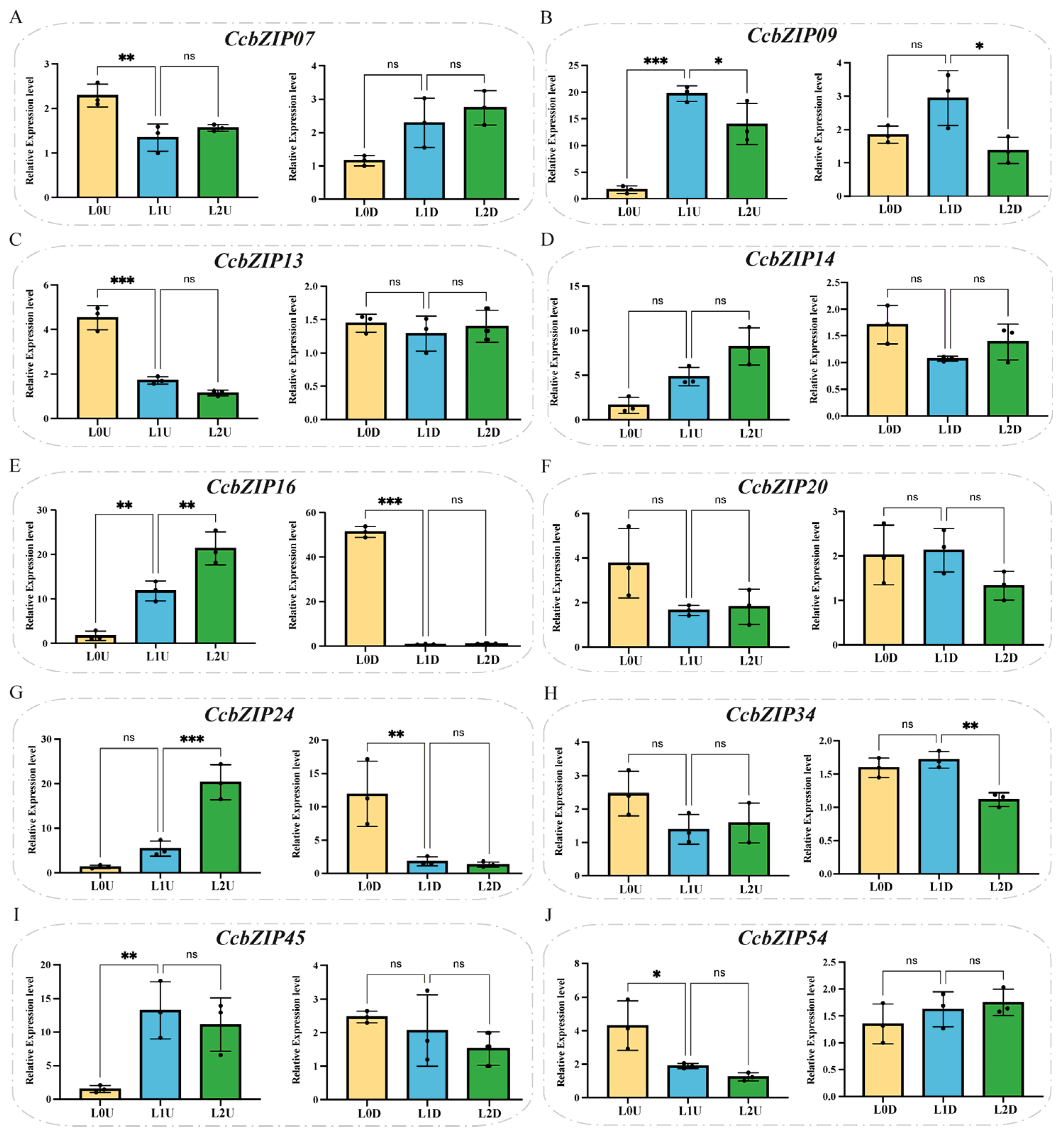
2.4. Expression Patterns of CcbZIP Genes Under Different Temperature and Light Conditions
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Stress Treatments
4.2. Identification and Sequence Analysis of CcbZIP Genes
4.3. Phylogenetic, Gene Structure, Conserved Motif Analysis, Chromosomal Distribution and Synteny Analysis
4.4. RNA Extraction, Transcriptome Sequencing, and qRT-PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| bZIP | basic leucine zipper |
| qRT-PCR | Quantitative Real-time PCR |
| ABA | abscisic acid |
| HMM | Hidden Markov Model |
| CDD | Conserved Domain Database |
References
- Dröge-Laser, W.; Snoek, B.L.; Snel, B.; Weiste, C. The Arabidopsis bZIP transcription factor family—An update. Curr. Opin. Plant Biol. 2018, 45, 36–49. [Google Scholar] [CrossRef]
- Weidemüller, P.; Kholmatov, M.; Petsalaki, E.; Zaugg, J.B. Transcription factors: Bridge between cell signaling and gene regulation. Proteomics 2021, 21, 23–24. [Google Scholar] [CrossRef] [PubMed]
- Jeung, D.; Li, X.; Cho, Y. New Roles of bZIP-Containing Membrane-Bound Transcription Factors in Chromatin Tethering and Karyoptosis. Int. J. Mol. Sci. 2025, 26, 10896. [Google Scholar] [CrossRef]
- Liu, Q.; Fang, H.; Zhou, H.; Wang, X.; Hou, Z. Identification and Characterization of bZIP Gene Family Combined Transcriptome Analysis Revealed Their Functional Roles on Abiotic Stress and Anthocyanin Biosynthesis in Mulberry (Morus alba). Horticulturae 2025, 11, 694. [Google Scholar] [CrossRef]
- Pérez-Rodríguez, P.; Riaño-Pachón, D.M.; Corrêa, L.G.G.; Rensing, S.A.; Kersten, B.; Mueller-Roeber, B. PlnTFDB: Updated content and new features of the plant transcription factor database. Nucleic Acids Res. 2010, 38, D822–D827. [Google Scholar] [CrossRef]
- Guo, Z.; Dzinyela, R.; Yang, L.; Hwarari, D. bZIP Transcription Factors: Structure, Modification, Abiotic Stress Responses and Application in Plant Improvement. Plants 2024, 13, 2058. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Wen, Y.; Sun, W.; Ma, Z.; Huang, L.; Wu, Q.; Tang, Z.; Bu, T.; Li, C.; Chen, H. Genome-wide identification, phylogeny, evolutionary expansion and expression analyses of bZIP transcription factor family in tartaty buckwheat. BMC Genom. 2019, 20, 483. [Google Scholar] [CrossRef]
- Li, H.; Li, L.; ShangGuan, G.; Jia, C.; Deng, S.; Noman, M.; Liu, Y.; Guo, Y.; Han, L.; Zhang, X.; et al. Genome-wide identification and expression analysis of bZIP gene family in Carthamus tinctorius L. Sci. Rep. 2020, 10, 15521. [Google Scholar] [CrossRef]
- Hou, H.; Kong, X.; Zhou, Y.; Yin, C.; Jiang, Y.; Qu, H.; Li, T. Genome-wide identification and characterization of bZIP transcription factors in relation to litchi (Litchi chinensis Sonn.) fruit ripening and postharvest storage. Int. J. Biol. Macromol. 2022, 222, 2176–2189. [Google Scholar] [CrossRef]
- Tian, X.; Jiang, Q.; Jia, Z.; Fang, Y.; Wang, Z.; Wang, J. Identification of TabZIP family members with possible roles in the response to auxin in wheat roots. Phytochemistry 2022, 196, 113103. [Google Scholar] [CrossRef] [PubMed]
- E, Z.; Zhang, Y.; Zhou, J.; Wang, L. Mini review roles of the bZIP gene family in rice. Genet. Mol. Res. 2014, 13, 3025–3036. [Google Scholar] [CrossRef]
- Cao, L.; Lu, X.; Zhang, P.; Wang, G.; Wei, L.; Wang, T. Systematic Analysis of Differentially Expressed Maize ZmbZIP Genes between Drought and Rewatering Transcriptome Reveals bZIP Family Members Involved in Abiotic Stress Responses. Int. J. Mol. Sci. 2019, 20, 4103. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Fu, F.; Zhang, H.; Song, F. Genome-wide systematic characterization of the bZIP transcriptional factor family in tomato (Solanum lycopersicum L.). BMC Genom. 2015, 16, 771. [Google Scholar] [CrossRef]
- Kumar, P.; Sharma, D.; Verma, S.K.; Halterman, D.; Kumar, A. Genome-wide identification and expression profiling of basic leucine zipper transcription factors following abiotic stresses in potato (Solanum tuberosum L.). PLoS ONE 2021, 16, e247864. [Google Scholar] [CrossRef]
- Hu, W.; Wang, L.; Tie, W.; Yan, Y.; Ding, Z.; Liu, J.; Li, M.; Peng, M.; Xu, B.; Jin, Z. Genome-wide analyses of the bZIP family reveal their involvement in the development, ripening and abiotic stress response in banana. Sci. Rep. 2016, 6, 30203. [Google Scholar] [CrossRef]
- Baloglu, M.C.; Eldem, V.; Hajyzadeh, M.; Unver, T. Genome-wide analysis of the bZIP transcription factors in cucumber. PLoS ONE 2014, 9, e96014. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, Y.; Zhou, R.; Dossa, K.; Yu, J.; Li, D.; Liu, A.; Mmadi, M.A.; Zhang, X.; You, J. Identification and characterization of the bZIP transcription factor family and its expression in response to abiotic stresses in sesame. PLoS ONE 2018, 13, e200850. [Google Scholar] [CrossRef]
- Wang, S.; Zhang, X.; Li, B.; Zhao, X.; Shen, Y.; Yuan, Z. Genome-wide identification and characterization of bZIP gene family and cloning of candidate genes for anthocyanin biosynthesis in pomegranate (Punica granatum). BMC Plant Biol. 2022, 22, 170. [Google Scholar] [CrossRef]
- Liu, C.; Wu, Y.; Wang, X. bZIP transcription factor OsbZIP52/RISBZ5: A potential negative regulator of cold and drought stress response in rice. Planta 2012, 235, 1157–1169. [Google Scholar] [CrossRef] [PubMed]
- Shu, G.; Tang, Y.; Yuan, M.; Wei, N.; Zhang, F.; Yang, C.; Lan, X.; Chen, M.; Tang, K.; Xiang, L.; et al. Molecular insights into AabZIP1-mediated regulation on artemisinin biosynthesis and drought tolerance in Artemisia annua. Acta Pharm. Sin. B 2022, 12, 1500–1513. [Google Scholar] [CrossRef] [PubMed]
- Shen, Q.; Huang, H.; Zhao, Y.; Xie, L.; He, Q.; Zhong, Y.; Wang, Y.; Wang, Y.; Tang, K. The Transcription Factor Aabzip9 Positively Regulates the Biosynthesis of Artemisinin in Artemisia annua. Front. Plant Sci. 2019, 10, 1294. [Google Scholar] [CrossRef]
- Lv, Z.; Guo, Z.; Zhang, L.; Zhang, F.; Jiang, W.; Shen, Q.; Fu, X.; Yan, T.; Shi, P.; Hao, X.; et al. Interaction of bZIP transcription factor TGA6 with salicylic acid signaling modulates artemisinin biosynthesis in Artemisia annua. J. Exp. Bot. 2019, 70, 3969–3979. [Google Scholar] [CrossRef]
- Wu, Z.; Li, R.; Sun, M.; Hu, X.; Xiao, M.; Hu, Z.; Jiao, P.; Pu, S.; Zhai, J.; Zhang, J. Current advances of Carthamus tinctorius L.: A review of its application and molecular regulation of flavonoid biosynthesis. Med. Plant Biol. 2024, 3, e004. [Google Scholar] [CrossRef]
- Chen, J.; Chen, M.; Li, H.; Tan, R.; Zhang, J.; Jones, J.A.; Zhao, S. An ABA-responsive transcription factor SmbZIP38 positively regulates salvianolic acids biosynthesis and stress-associated morphogenesis of Salvia miltiorrhiza. Hortic. Plant J. 2025; in press. [Google Scholar] [CrossRef]
- Sun, T.; Hu, C.; Zhu, C.; Cai, J.; Wang, G.; Xia, X.; Shi, K.; Zhou, Y.; Yu, J.; Foyer, C.H.; et al. HY5 integrates light and electrical signaling to trigger a jasmonate burst for nematode defense in tomato. Nat. Commun. 2025, 16, 8750. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wei, J.; Li, S.; Li, J.; Cao, H.; Huang, D.; Zhang, D.; Zhang, Z.; Gao, T.; Zhang, Y.; et al. MdHY5 positively regulates cold tolerance in apple by integrating the auxin and abscisic acid pathways. New Phytol. 2025, 246, 2155–2173. [Google Scholar] [CrossRef] [PubMed]
- Bai, M.; Yang, Y.; Gao, Y.; Xu, M.; Zhang, Q.; Liu, S.; Lu, J.; Wang, J.; Wang, C.; Feng, L. RrMYB2 Regulates Drought Stress via RrJMJ12-Dependent Epigenetic Modification in Rosa rugosa. Plant Biotechnol J. 2025. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, B.; Shu, S.; Li, Z.; Song, C.; Liu, D.; Niu, Y.; Liu, J.; Zhang, J.; Liu, H.; et al. Analysis of the Coptis chinensis genome reveals the diversification of protoberberine-type alkaloids. Nat. Commun. 2021, 12, 3276. [Google Scholar] [CrossRef]
- Li, C.; Jiao, M.; Zhao, X.; Ma, J.; Cui, Y.; Kou, X.; Long, Y.; Xing, Z. bZIP transcription factor responds to changes in light quality and affects saponins synthesis in Eleutherococcus senticosus. Int. J. Biol. Macromol. 2024, 279, 135273. [Google Scholar] [CrossRef]
- Zheng, Q.; Yang, Q.; Yang, D.; Hong, L.; Zhu, H.; Sui, N.; Wang, H. Enhanced carbon backbone supply via the mevalonate pathway confers sesquiterpene accumulation in Atractylodes macrocephala rhizomes under low temperature. Ind. Crops Prod. 2025, 236, 122127. [Google Scholar] [CrossRef]
- Marathe, S.; Grotewold, E.; Otegui, M.S. Should I stay or should I go? Trafficking of plant extra-nuclear transcription factors. Plant Cell 2024, 36, 1524–1539. [Google Scholar] [CrossRef]
- Cheng, L.; Han, Q.; Hao, Y.; Qiao, Z.; Li, M.; Liu, D.; Yin, H.; Li, T.; Long, W.; Luo, S.; et al. Genome assembly of Stewartia sinensis reveals origin and evolution of orphan genes in Theaceae. Commun. Biol. 2025, 8, 354. [Google Scholar] [CrossRef]
- Wang, R.; Hu, S.; Jiang, Y.; Zhou, L.; Liu, Q.; He, J. Temporal transcriptome and genome-wide identification of lily jasmonate ZIM-domain (JAZ) protein gene family reveals the roles of LfJAZ3 and LfJAZ12 in bulb development in Lilium fargesii. Int. J. Biol. Macromol. 2025, 308, 142656. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Xu, Z.; Bai, S. OsFON879, an orphan gene, regulates floral organ homeostasis in rice. Plant Biotechnol. J. 2025, 23, 2888–2890. [Google Scholar] [CrossRef]
- Chen, D.-X.; Pan, Y.; Wang, Y.; Cui, Y.-Z.; Zhang, Y.-J.; Mo, R.-Y.; Wu, X.-L.; Tan, J.; Zhang, J.; Guo, L.-A.; et al. The chromosome-level reference genome of Coptis chinensis provides insights into genomic evolution and berberine biosynthesis. Hortic. Res. 2021, 8, 121. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Sharma, M.; Gahlaut, V.; Nagaraju, M.; Chaudhary, S.; Kumar, A.; Tyagi, P.; Gajula, M.P.; Singh, K.P. Genome-wide identification, characterization, and expression profiling of SPX gene family in wheat. Int. J. Biol. Macromol. 2019, 140, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Lin, X.; Le, Z.; Wang, M.; Qin, X.; Zeng, L.; Qian, Y.; Shu, G.; Chen, M.; Lan, X.; et al. Chemical Profiling of Monoterpenes and Genome-Wide Discovery of Monoterpene Synthases in Artemisia annua. Horticulturae 2025, 11, 1083. [Google Scholar] [CrossRef]

| Gene Name | Gene ID | Chr | Chromosome Location | Gene Length (bp) | ORF Length (aa) | Deduced Protein | Subcellular Location | |||
|---|---|---|---|---|---|---|---|---|---|---|
| Size (aa) | MW (kDa) | PI | GRAVY | |||||||
| CcbZIP01 | evm.model.Scaffold_63.37 | 6 | 68351006–68364660 | 13,654 | 242 | 81 | 9563.17 | 11.44 | −0.932 | Nuclear |
| CcbZIP02 | evm.model.Scaffold_124.235 | 3 | 18006083–18007384 | 1301 | 848 | 283 | 31,878.16 | 6.37 | −0.913 | Nuclear |
| CcbZIP03 | evm.model.Scaffold_123.165 | 6 | 93895910–93901866 | 5956 | 1252 | 420 | 47,214.42 | 6.69 | −0.541 | Nuclear |
| CcbZIP04 | evm.model.Scaffold_65.11 | 3 | 22686753–22693929 | 4713 | 625 | 209 | 24,202.92 | 9.65 | −0.506 | Cytoplasmic |
| CcbZIP05 | evm.model.Scaffold_56.68 | 6 | 1542536–1546055 | 3519 | 1004 | 335 | 37,011.98 | 5.84 | −0.704 | Nuclear |
| CcbZIP06 | evm.model.Scaffold_128.187 | 8 | 88687098–88688981 | 1883 | 667 | 222 | 25,630.64 | 8.57 | −0.919 | Nuclear |
| CcbZIP07 | evm.model.Scaffold_175.551 | 3 | 85971174–85979810 | 8636 | 941 | 315 | 35,617.36 | 6.64 | −0.49 | Nuclear |
| CcbZIP08 | evm.model.Scaffold_161.2 | 2 | 3556701–3557183 | 2609 | 894 | 298 | 33,482.61 | 7.8 | −0.881 | Nuclear |
| CcbZIP09 | evm.model.Scaffold_204.1 | 2 | 4085267–4090711 | 3335 | 1706 | 569 | 62,560.2 | 6.78 | −0.895 | Nuclear |
| CcbZIP10 | evm.model.Scaffold_84.32 | 2 | 86525851–86526291 | 440 | 440 | 146 | 16,747.67 | 8.02 | −0.784 | Nuclear |
| CcbZIP11 | evm.model.Scaffold_24.21 | 9 | 65115883–65116317 | 1659 | 473 | 157 | 17,965.1 | 6.29 | −0.817 | Nuclear |
| CcbZIP12 | evm.model.Scaffold_165.276 | 2 | 12928906–12929914 | 1008 | 834 | 278 | 30,923.83 | 8.43 | −0.915 | Nuclear |
| CcbZIP13 | evm.model.Scaffold_145.55 | 2 | 33619210–33623955 | 4745 | 1281 | 428 | 46,805.05 | 5.04 | −0.711 | Nuclear |
| CcbZIP14 | evm.model.Scaffold_20.387 | 5 | 65762806–65763294 | 488 | 488 | 162 | 18,185.72 | 6.75 | −0.466 | Nuclear |
| CcbZIP15 | evm.model.Scaffold_356.10 | 8 | 83877338–83884882 | 7544 | 2561 | 855 | 93,518.33 | 4.82 | −0.619 | Nuclear |
| CcbZIP16 | evm.model.Scaffold_62.65 | 7 | 67370827–67376377 | 5550 | 470 | 157 | 18,044.1 | 9.52 | −1.246 | Nuclear |
| CcbZIP17 | evm.model.Scaffold_27.71 | 7 | 85094226–85094804 | 578 | 578 | 192 | 22,328.99 | 6.06 | −0.759 | Nuclear |
| CcbZIP18 | evm.model.Scaffold_106.121 | 2 | 10277150–10278158 | 1008 | 834 | 278 | 30,923.83 | 8.43 | −0.915 | Nuclear |
| CcbZIP19 | evm.model.Scaffold_106.478 | 2 | 251653–259758 | 8105 | 1372 | 460 | 50,789.87 | 8.75 | −0.603 | Nuclear |
| CcbZIP20 | evm.model.Scaffold_191.13 | -- | 97202–101121 | 3919 | 1214 | 405 | 44,374.16 | 5.51 | −0.613 | Nuclear |
| CcbZIP21 | evm.model.Scaffold_252.40 | 3 | 13184165–13189401 | 5236 | 1098 | 368 | 42,110.74 | 6.14 | −0.498 | Nuclear |
| CcbZIP22 | evm.model.Scaffold_161.83 | 2 | 2983894–2986739 | 2845 | 2059 | 686 | 74,454.55 | 7.18 | −0.524 | Endoplasmic Reticulum |
| CcbZIP23 | evm.model.Scaffold_360.34 | 4 | 16145365–16151420 | 6055 | 1230 | 413 | 43,786.68 | 8.31 | −0.792 | Nuclear |
| CcbZIP24 | evm.model.Scaffold_46.103 | 1 | 70302836–70303670 | 834 | 542 | 181 | 20,387.9 | 6.99 | −0.809 | Nuclear |
| CcbZIP25 | evm.model.Scaffold_194.41 | 1 | 34592896–34597504 | 4608 | 1281 | 428 | 47,129.15 | 5.64 | −0.636 | Nuclear |
| CcbZIP26 | evm.model.Scaffold_38.133 | 9 | 50051894–50055400 | 3506 | 1161 | 388 | 44,472.64 | 9.13 | −0.609 | Nuclear |
| CcbZIP27 | evm.model.Scaffold_203.105 | 7 | 21177099–21183198 | 6099 | 1494 | 501 | 55,899.55 | 6.25 | −0.588 | Nuclear |
| CcbZIP28 | evm.model.Scaffold_41.681 | 4 | 90980279–90983595 | 3316 | 864 | 288 | 31,318.5 | 4.78 | −0.837 | Nuclear |
| CcbZIP29 | evm.model.Scaffold_370.16 | -- | 123764–128992 | 5228 | 1323 | 443 | 50,180.74 | 5.22 | −0.403 | Nuclear |
| CcbZIP30 | evm.model.Scaffold_4.341 | 3 | 103306282–103317534 | 11,252 | 2178 | 731 | 81,352.76 | 5.28 | −0.526 | Nuclear |
| CcbZIP31 | evm.model.Scaffold_78.543 | 8 | 93164141–93170657 | 6516 | 1209 | 406 | 43,075.3 | 5.6 | −0.893 | Nuclear |
| CcbZIP32 | evm.model.Scaffold_226.12 | 2 | 2736440–2739282 | 2842 | 2056 | 685 | 74,336.37 | 7.18 | −0.52 | Cytoplasmic |
| CcbZIP33 | evm.model.Scaffold_226.30 | 2 | 2628293–2633196 | 4903 | 914 | 307 | 33,109.99 | 4.89 | −0.457 | Chloroplast |
| CcbZIP34 | evm.model.Scaffold_41.632 | 4 | 90521302–90524839 | 3537 | 1076 | 359 | 39,393.77 | 6.34 | −0.772 | Nuclear |
| CcbZIP35 | evm.model.Scaffold_187.200 | 3 | 66584201–66591229 | 7028 | 732 | 244 | 27,969.38 | 5.31 | −0.802 | Nuclear |
| CcbZIP36 | evm.model.Scaffold_282.29 | 4 | 85802034–85805554 | 3520 | 1127 | 376 | 42,433.66 | 6.78 | −1.045 | Nuclear |
| CcbZIP37 | evm.model.Scaffold_89.303 | 9 | 57192590–57193532 | 942 | 702 | 231 | 25,845.31 | 5.23 | −0.756 | Nuclear |
| CcbZIP38 | evm.model.Scaffold_81.149 | 6 | 96334268–96345074 | 10,806 | 1324 | 444 | 48,989.11 | 5.91 | −0.44 | Nuclear |
| CcbZIP39 | evm.model.Scaffold_371.5 | -- | 73269–76031 | 2762 | 1106 | 369 | 41,307.57 | 6.5 | −0.943 | Nuclear |
| CcbZIP40 | evm.model.Scaffold_72.208 | 2 | 9921150–9922803 | 1653 | 804 | 268 | 30,140.36 | 5.76 | −0.432 | Nuclear |
| CcbZIP41 | evm.model.Scaffold_72.137 | 2 | 9254146–9254628 | 482 | 482 | 160 | 18,879.41 | 11.04 | −0.831 | Nuclear |
| CcbZIP42 | evm.model.Scaffold_34.97 | 3 | 3647551–3651470 | 3919 | 1214 | 405 | 44,422.25 | 5.59 | −0.607 | Nuclear |
| CcbZIP43 | evm.model.Scaffold_67.62 | 6 | 16899934–16900026 | 2600 | 1307 | 436 | 47,732.88 | 8.54 | −0.918 | Nuclear |
| CcbZIP44 | evm.model.Scaffold_67.72 | 6 | 16703040–16703186 | 10,243 | 1431 | 480 | 54,036.08 | 8.88 | −0.579 | Nuclear |
| CcbZIP45 | evm.model.Scaffold_31.662 | 2 | 4956546–4959921 | 3375 | 1706 | 569 | 62,557.24 | 6.78 | −0.888 | Nuclear |
| CcbZIP46 | evm.model.Scaffold_11.13 | 8 | 67776316–67781705 | 1657 | 315 | 105 | 11,959.71 | 10.28 | −0.69 | Chloroplast |
| CcbZIP47 | evm.model.Scaffold_110.91 | 1 | 75592581–75595718 | 3137 | 879 | 294 | 33,138.64 | 6.83 | 0.023 | Plasma Membrane |
| CcbZIP48 | evm.model.Scaffold_110.382 | 2 | 17308158–17319069 | 10,911 | 2674 | 893 | 97,509.61 | 4.79 | −0.63 | Nuclear |
| CcbZIP49 | evm.model.Scaffold_1.8 | -- | 55368–59792 | 7079 | 1183 | 397 | 41,833.24 | 6.11 | −0.696 | Nuclear |
| CcbZIP50 | evm.model.Scaffold_109.640 | 4 | 71801476–71806600 | 5124 | 1465 | 489 | 53,474.72 | 9.41 | −0.791 | Nuclear |
| CcbZIP51 | evm.model.Scaffold_273.40 | 7 | 11663351–11669409 | 6058 | 1064 | 355 | 38,922.79 | 5.52 | −0.794 | Nuclear |
| CcbZIP52 | evm.model.Scaffold_232.8 | 6 | 17932846–17933243 | 5953 | 1542 | 515 | 57,476.6 | 8.13 | −0.797 | Nuclear |
| CcbZIP53 | evm.model.Scaffold_93.79 | 4 | 56509773–56510094 | 4200 | 1847 | 616 | 68,403.26 | 7.77 | −0.997 | Nuclear |
| CcbZIP54 | evm.model.Scaffold_15.169 | 2 | 59307068–59307502 | 434 | 434 | 144 | 16,569.83 | 5.22 | −0.576 | Nuclear |
| CcbZIP55 | evm.model.Scaffold_264.349 | 7 | 397066–405341 | 8275 | 1372 | 460 | 50,764.88 | 8.43 | −0.59 | Nuclear |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Wei, W.; Le, Z.; Guo, L.; Mo, R.; Wang, Y.; Pan, Y. Genome-Wide Identification and Analysis of bZIP Transcription Factors in Coptis chinensis Reveals Their Regulatory Roles in Stress Responses. Int. J. Mol. Sci. 2026, 27, 431. https://doi.org/10.3390/ijms27010431
Wei W, Le Z, Guo L, Mo R, Wang Y, Pan Y. Genome-Wide Identification and Analysis of bZIP Transcription Factors in Coptis chinensis Reveals Their Regulatory Roles in Stress Responses. International Journal of Molecular Sciences. 2026; 27(1):431. https://doi.org/10.3390/ijms27010431
Chicago/Turabian StyleWei, Wuke, Zijian Le, Lianan Guo, Rangyu Mo, Yu Wang, and Yuan Pan. 2026. "Genome-Wide Identification and Analysis of bZIP Transcription Factors in Coptis chinensis Reveals Their Regulatory Roles in Stress Responses" International Journal of Molecular Sciences 27, no. 1: 431. https://doi.org/10.3390/ijms27010431
APA StyleWei, W., Le, Z., Guo, L., Mo, R., Wang, Y., & Pan, Y. (2026). Genome-Wide Identification and Analysis of bZIP Transcription Factors in Coptis chinensis Reveals Their Regulatory Roles in Stress Responses. International Journal of Molecular Sciences, 27(1), 431. https://doi.org/10.3390/ijms27010431







