The NAC Transcription Factors CjNAC43 and CjNAC54 Act as Positive Regulators of Leaf Senescence in Clerodendrum japonicum
Abstract
1. Introduction
2. Results
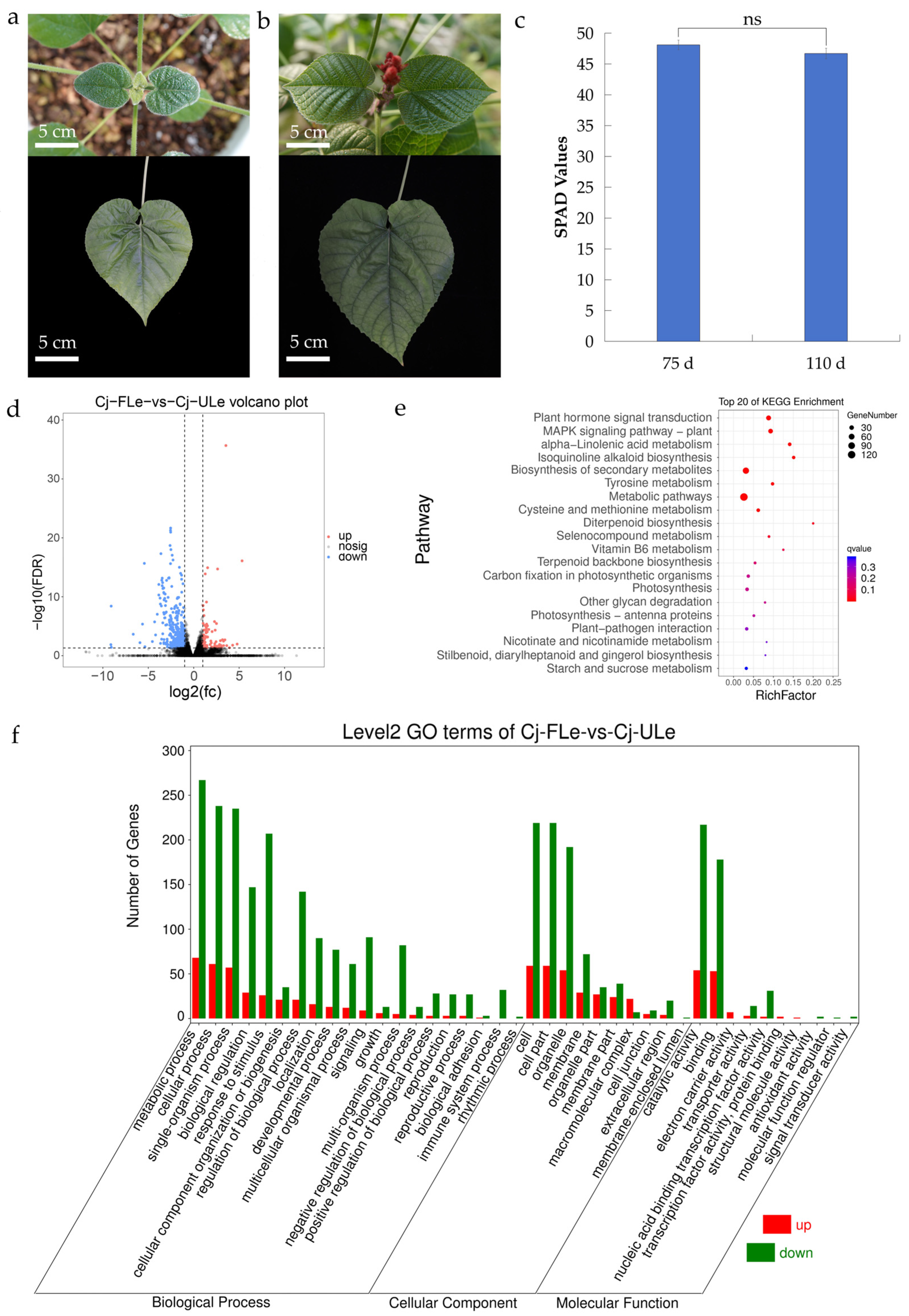
2.1. Transcriptomic Analysis and Selection of Differentially Expressed Genes
2.2. Screening for NAC Genes Associated with Leaf Senescence Among DEGs
2.3. Over-Expression of CjNAC43 and CjNAC54 Accelerates Leaf Senescence in Arabidopsis
2.4. CjNAC43 and CjNAC54 Mediates Leaf Senescence Induced by ABA Treatment and Darkness
2.5. Suppression of CjNAC43 and CjNAC54 Expression Results in Delayed Leaf Senescence
3. Discussion
4. Materials and Methods
4.1. Plant Meterial and Transcriptome Sequencing
4.2. Differential Gene Analysis
4.3. Identification of NAC Family
4.4. Relative Expression Measurement
4.5. Measurement of Physiological Parameters
4.6. Phylogenetic Analysis
4.7. Construction of Overexpression Vector
4.8. Generation of Transgenic Plants
4.9. Phenotypic and Functional Analysis of Overexpressing A. thaliana Lines
4.10. Abiotic Stress Treatment
4.11. VIGS System Construction and Functional Verification
4.12. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| ET | Ethylene |
| SGR | STAY-GREEN |
| PAO | PHEOPHORBIDE A OXYGENASE |
| SAG | senescence-associated genes |
| ACT | ACTIN |
| NAC | NAM/ATAF/CUC |
| TF | Transcription factors |
| VIGS | Virus-induced gene silencing |
| SPAD | Soil and Plant Analyzer Development |
| GC | Guanine-cytosine |
| FPKM | Fragments Per Kilobase of exon model per Million mapped fragments |
| ATP | Adenosine triphosphate |
| DEG | Differential expressed genes |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| GO | Gene Ontology |
| PDS | Phytoene desaturase |
References
- Zhang, Y.; Wang, H.-L.; Li, Z.; Guo, H. Genetic network between leaf senescence and plant immunity: Crucial regulatory nodes and new insights. Plants 2020, 9, 495. [Google Scholar] [CrossRef]
- Distelfeld, A.; Avni, R.; Fischer, A.M. Senescence, nutrient remobilization, and yield in wheat and barley. J. Exp. Bot. 2014, 65, 3783–3798. [Google Scholar] [CrossRef] [PubMed]
- Thomas, H. Senescence, ageing and death of the whole plant. New Phytol. 2013, 197, 696–711. [Google Scholar] [CrossRef] [PubMed]
- Doan, P.P.T.; Vuong, H.H.; Kim, J. Genetic foundation of leaf senescence: Insights from natural and cultivated plant diversity. Plants 2024, 13, 3405. [Google Scholar] [CrossRef]
- Woo, H.R.; Kim, H.J.; Nam, H.G.; Lim, P.O. Plant leaf senescence and death–regulation by multiple layers of control and implications for aging in general. J. Cell Sci. 2013, 126, 4823–4833. [Google Scholar] [CrossRef] [PubMed]
- Avila-Ospina, L.; Moison, M.; Yoshimoto, K.; Masclaux-Daubresse, C. Autophagy, plant senescence, and nutrient recycling. J. Exp. Bot. 2014, 65, 3799–3811. [Google Scholar] [CrossRef]
- Zhang, J.; Yu, W.; Wang, Y.; Huang, Z.; Liu, G. Effects of climate, soil, and leaf traits on nutrient resorption efficiency and proficiency of different plant functional types across arid and semiarid regions of northwest China. BMC Plant Biol. 2024, 24, 1093. [Google Scholar] [CrossRef]
- Breeze, E.; Harrison, E.; McHattie, S.; Hughes, L.; Hickman, R.; Hill, C.; Kiddle, S.; Kim, Y.-S.; Penfold, C.A.; Jenkins, D. High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 2011, 23, 873–894. [Google Scholar] [CrossRef]
- Masclaux, C.; Valadier, M.-H.; Brugière, N.; Morot-Gaudry, J.-F.; Hirel, B. Characterization of the sink/source transition in tobacco (Nicotiana tabacum L.) shoots in relation to nitrogen management and leaf senescence. Planta 2000, 211, 510–518. [Google Scholar] [CrossRef]
- Watanabe, M.; Balazadeh, S.; Tohge, T.; Erban, A.; Giavalisco, P.; Kopka, J.; Mueller-Roeber, B.; Fernie, A.R.; Hoefgen, R. Comprehensive dissection of spatiotemporal metabolic shifts in primary, secondary, and lipid metabolism during developmental senescence in Arabidopsis. Plant Physiol. 2013, 162, 1290–1310. [Google Scholar] [CrossRef]
- Gan, S.; Amasino, R.M. Making sense of senescence (molecular genetic regulation and manipulation of leaf senescence). Plant Physiol. 1997, 113, 313. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Ren, G.; Zhang, K.; Li, Z.; Miao, Y.; Guo, H. Leaf senescence: Progression, regulation, and application. Mol. Hortic. 2021, 1, 5. [Google Scholar] [CrossRef]
- Cao, J.; Liu, H.; Tan, S.; Li, Z. Transcription factors-regulated leaf senescence: Current knowledge, challenges and approaches. Int. J. Mol. Sci. 2023, 24, 9245. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.; Sha, Y.; Sun, L.; Li, Z. Abiotic stress-induced leaf senescence: Regulatory mechanisms and application. Int. J. Mol. Sci. 2023, 24, 11996. [Google Scholar] [CrossRef]
- Zhao, W.; Zhao, H.; Wang, H.; He, Y. Research progress on the relationship between leaf senescence and quality, yield and stress resistance in horticultural plants. Front. Plant Sci. 2022, 13, 1044500. [Google Scholar] [CrossRef]
- Sakuraba, Y.; Schelbert, S.; Park, S.-Y.; Han, S.-H.; Lee, B.-D.; Andres, C.B.; Kessler, F.; Hörtensteiner, S.; Paek, N.-C. STAY-GREEN and chlorophyll catabolic enzymes interact at light-harvesting complex II for chlorophyll detoxification during leaf senescence in Arabidopsis. Plant Cell 2012, 24, 507–518. [Google Scholar] [CrossRef]
- Wang, P.; Richter, A.S.; Kleeberg, J.R.; Geimer, S.; Grimm, B. Post-translational coordination of chlorophyll biosynthesis and breakdown by BCMs maintains chlorophyll homeostasis during leaf development. Nat. Commun. 2020, 11, 1254. [Google Scholar] [CrossRef]
- Dey, D.; Dhar, D.; Fortunato, H.; Obata, D.; Tanaka, A.; Tanaka, R.; Basu, S.; Ito, H. Insights into the structure and function of the rate-limiting enzyme of chlorophyll degradation through analysis of a bacterial Mg-dechelatase homolog. Comput. Struct. Biotechnol. J. 2021, 19, 5333–5347. [Google Scholar] [CrossRef]
- Zhang, K.; Gan, S.-S. An abscisic acid-AtNAP transcription factor-SAG113 protein phosphatase 2C regulatory chain for controlling dehydration in senescing Arabidopsis leaves. Plant Physiol. 2012, 158, 961–969. [Google Scholar] [CrossRef] [PubMed]
- Keech, O.; Pesquet, E.; Gutierrez, L.; Ahad, A.; Bellini, C.; Smith, S.M.; Gardeström, P. Leaf senescence is accompanied by an early disruption of the microtubule network in Arabidopsis. Plant Physiol. 2010, 154, 1710–1720. [Google Scholar] [CrossRef]
- Woo, H.R.; Kim, H.J.; Lim, P.O.; Nam, H.G. Leaf senescence: Systems and dynamics aspects. Annu. Rev. Plant Biol. 2019, 70, 347–376. [Google Scholar] [CrossRef]
- Kim, H.J.; Nam, H.G.; Lim, P.O. Regulatory network of NAC transcription factors in leaf senescence. Curr. Opin. Plant Biol. 2016, 33, 48–56. [Google Scholar] [CrossRef]
- Balazadeh, S.; Riaño-Pachón, D.; Mueller-Roeber, B. Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Biol. 2008, 10, 63–75. [Google Scholar] [CrossRef]
- Jin, J.; Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2016, 45, D1040–D1045. [Google Scholar] [CrossRef]
- Chen, Y.; Xia, P. NAC transcription factors as biological macromolecules responded to abiotic stress: A comprehensive review. Int. J. Biol. Macromol. 2025, 308, 142400. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Zhao, Y.; Wang, W.; Liu, X.; Shi, W.; Zhang, D.; Chen, J.; Ma, S.; Sun, Q.; Wang, T. Transcriptome analysis in contrasting maize inbred lines and functional analysis of five maize NAC genes under drought stress treatment. Front. Plant Sci. 2023, 13, 1097719. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Liu, X.; Gan, S.-S. The ABA–AtNAP–SAG113 PP2C module regulates leaf senescence by dephoshorylating SAG114 SnRK3. 25 in Arabidopsis. Mol. Hortic. 2023, 3, 22. [Google Scholar] [CrossRef]
- Qiu, K.; Li, Z.; Yang, Z.; Chen, J.; Wu, S.; Zhu, X.; Gao, S.; Gao, J.; Ren, G.; Kuai, B. EIN3 and ORE1 accelerate degreening during ethylene-mediated leaf senescence by directly activating chlorophyll catabolic genes in Arabidopsis. PLoS Genet. 2015, 11, e1005399. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Huang, W.; Liu, L.; Chen, T.; Zhou, F.; Lin, Y. Identification and functional characterization of a rice NAC gene involved in the regulation of leaf senescence. BMC Plant Biol. 2013, 13, 132. [Google Scholar] [CrossRef]
- Sun, L.; Xu, H.; Song, J.; Yang, X.; Wang, X.; Liu, H.; Pang, M.; Hu, Y.; Yang, Q.; Ning, X. OsNAC103, a NAC transcription factor, positively regulates leaf senescence and plant architecture in rice. Rice 2024, 17, 15. [Google Scholar] [CrossRef]
- Yang, Z.; Wang, C.; Qiu, K.; Chen, H.; Li, Z.; Li, X.; Song, J.; Wang, X.; Gao, J.; Kuai, B. The transcription factor ZmNAC126 accelerates leaf senescence downstream of the ethylene signalling pathway in maize. Plant Cell Environ. 2020, 43, 2287–2300. [Google Scholar] [CrossRef]
- Yang, J.; Worley, E.; Udvardi, M. A NAP-AAO3 regulatory module promotes chlorophyll degradation via ABA biosynthesis in Arabidopsis leaves. Plant Cell 2014, 26, 4862–4874. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Huang, G.; Zhang, Q.; Wang, X.; Li, C.; Tao, Y.; Zhang, S.; Lai, J.; Yang, C.; Wang, Y. The LEA protein, ABR, is regulated by ABI5 and involved in dark-induced leaf senescence in Arabidopsis thaliana. Plant Sci. 2016, 247, 93–103. [Google Scholar] [CrossRef]
- Fernandez, P.; Moschen, S.; Paniego, N.; Heinz, R.A. Functional approaches to study leaf senescence in sunflower. In Senescence; InTech Open Access Publisher: Rijeka, Croatia, 2012; pp. 69–88. [Google Scholar]
- Trupkin, S.A.; Astigueta, F.H.; Baigorria, A.H.; García, M.N.; Delfosse, V.C.; González, S.A.; de la Torre, M.C.P.; Moschen, S.; Lía, V.V.; Fernández, P. Identification and expression analysis of NAC transcription factors potentially involved in leaf and petal senescence in Petunia hybrida. Plant Sci. 2019, 287, 110195. [Google Scholar] [CrossRef]
- Bengoa Luoni, S.A.; Cenci, A.; Moschen, S.; Nicosia, S.; Radonic, L.M.; Sabio y García, J.V.; Langlade, N.B.; Vile, D.; Rovere, C.V.; Fernandez, P. Genome-wide and comparative phylogenetic analysis of senescence-associated NAC transcription factors in sunflower (Helianthus annuus). BMC Genom. 2021, 22, 893. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, J.; Zhang, T.; Wang, G.; Han, Z.; Meng, Y.; Bi, J.; Ren, Y.; Yang, M. Research progress of NAC transcription factors in woody plants. Front. Plant Sci. 2025, 16, 1592898. [Google Scholar] [CrossRef]
- Wang, H.-L.; Zhang, Y.; Wang, T.; Yang, Q.; Yang, Y.; Li, Z.; Li, B.; Wen, X.; Li, W.; Yin, W. An alternative splicing variant of PtRD26 delays leaf senescence by regulating multiple NAC transcription factors in Populus. Plant Cell 2021, 33, 1594–1614. [Google Scholar] [CrossRef]
- D’Incà, E.; Foresti, C.; Orduña, L.; Amato, A.; Vandelle, E.; Santiago, A.; Botton, A.; Cazzaniga, S.; Bertini, E.; Pezzotti, M. The transcription factor VviNAC60 regulates senescence-and ripening-related processes in grapevine. Plant Physiol. 2023, 192, 1928–1946. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Wang, L.-G.; Wang, Y.-H. Complete chloroplast genome sequence and phylogenetic analysis of Clerodendrum japonicum (Thunb.) Sweet (Ajugoideae, Lamiaceae). Mitochondrial DNA Part B 2021, 6, 2218–2220. [Google Scholar] [CrossRef]
- Rabha, D.; Sarma, H.; Borah, M.; Ghose, V.; Barman, D.; Rao, S.; Sarma, H.K.; Devi, R.; Boruah, D.C. Clerodendrum japonicum (Thunb.) Sweet leaf extract supported nanosilver particles: Characterization, antioxidant and antibacterial activity: Clerodendrum japonicum nanosilver particles: Antioxidant and antibacterial activity. Vietnam J. Sci. Technol. 2024, 62, 1065–1076. [Google Scholar] [CrossRef]
- Wu, Y.; Cui, Y.; Zhang, Y.; Zhang, X.; Zhou, J.; Wang, C.; Wang, H. MnO Nanoenzymes Synthesized from Clerodendrum japonicum Extract Mitigate Ferroptosis in Pulmonary Fibrosis Rats through Enhanced Antioxidant Surface Activity. Surf. Interfaces 2025, 69, 106740. [Google Scholar] [CrossRef]
- Wang, C.; Liao, G.; Duan, Y.; Su, L.; He, C.; Xu, M.; Wang, H. Identification of NAC Transcription Factors Associated with Leaf Senescence in Clerodendrum japonicum. Int. J. Mol. Sci. 2025, 26, 8846. [Google Scholar] [CrossRef]
- Alshareef, N.O.; Otterbach, S.L.; Allu, A.D.; Woo, Y.H.; de Werk, T.; Kamranfar, I.; Mueller-Roeber, B.; Tester, M.; Balazadeh, S.; Schmöckel, S.M. NAC transcription factors ATAF1 and ANAC055 affect the heat stress response in Arabidopsis. Sci. Rep. 2022, 12, 11264. [Google Scholar] [CrossRef] [PubMed]
- Eerapagula, R.; Singh, A.; Maurya, A.; Singh, R.; Singh, N.K.; Mahato, A.K. Genome-wide analysis of NAC transcription factors in grain amaranth reveals structural diversity and regulatory features. Sci. Rep. 2025, 15, 39968. [Google Scholar] [CrossRef] [PubMed]
- Garapati, P.; Xue, G.-P.; Munné-Bosch, S.; Balazadeh, S. Transcription factor ATAF1 in Arabidopsis promotes senescence by direct regulation of key chloroplast maintenance and senescence transcriptional cascades. Plant Physiol. 2015, 168, 1122–1139. [Google Scholar] [CrossRef]
- Podzimska-Sroka, D.; O’Shea, C.; Gregersen, P.L.; Skriver, K. NAC transcription factors in senescence: From molecular structure to function in crops. Plants 2015, 4, 412–448. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Rhoads, A.; Au, K.F. PacBio sequencing and its applications. Genom. Proteom. Bioinform. 2015, 13, 278–289. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Bu, D.; Luo, H.; Huo, P.; Wang, Z.; Zhang, S.; He, Z.; Wu, Y.; Zhao, L.; Liu, J.; Guo, J. KOBAS-i: Intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res. 2021, 49, W317–W325. [Google Scholar] [CrossRef]
- Stajich, J.E.; Block, D.; Boulez, K.; Brenner, S.E.; Chervitz, S.A.; Dagdigian, C.; Fuellen, G.; Gilbert, J.G.; Korf, I.; Lapp, H. The Bioperl toolkit: Perl modules for the life sciences. Genome Res. 2002, 12, 1611–1618. [Google Scholar] [CrossRef] [PubMed]
- Clarke, K.; Gorley, R. PRIMER: Getting Started with v6; PRIMER-E Ltd.: Plymouth, UK, 2005; Volume 931, p. 932. [Google Scholar]
- Woo, H.R.; Masclaux-Daubresse, C.; Lim, P.O. Plant Senescence: How Plants Know When and How to Die; Oxford University Press: Oxford, UK, 2018; pp. 715–718. [Google Scholar]
- Bajji, M.; Kinet, J.-M.; Lutts, S. The use of the electrolyte leakage method for assessing cell membrane stability as a water stress tolerance test in durum wheat. Plant Growth Regul. 2002, 36, 61–70. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]





Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Deng, Y.; Wang, C.; Huang, R.; Su, L.; He, C.; Xu, M.; Wang, H. The NAC Transcription Factors CjNAC43 and CjNAC54 Act as Positive Regulators of Leaf Senescence in Clerodendrum japonicum. Int. J. Mol. Sci. 2026, 27, 133. https://doi.org/10.3390/ijms27010133
Deng Y, Wang C, Huang R, Su L, He C, Xu M, Wang H. The NAC Transcription Factors CjNAC43 and CjNAC54 Act as Positive Regulators of Leaf Senescence in Clerodendrum japonicum. International Journal of Molecular Sciences. 2026; 27(1):133. https://doi.org/10.3390/ijms27010133
Chicago/Turabian StyleDeng, Yanwen, Congcong Wang, Rutao Huang, Lingye Su, Chunmei He, Mingfeng Xu, and Hongfeng Wang. 2026. "The NAC Transcription Factors CjNAC43 and CjNAC54 Act as Positive Regulators of Leaf Senescence in Clerodendrum japonicum" International Journal of Molecular Sciences 27, no. 1: 133. https://doi.org/10.3390/ijms27010133
APA StyleDeng, Y., Wang, C., Huang, R., Su, L., He, C., Xu, M., & Wang, H. (2026). The NAC Transcription Factors CjNAC43 and CjNAC54 Act as Positive Regulators of Leaf Senescence in Clerodendrum japonicum. International Journal of Molecular Sciences, 27(1), 133. https://doi.org/10.3390/ijms27010133






