Genome-Wide Identification and Expression Analysis of the Ginkgo biloba B-Box Gene Family in Response to Hormone Treatments, Flavonoid Levels, and Water Stress
Abstract
1. Introduction
2. Results
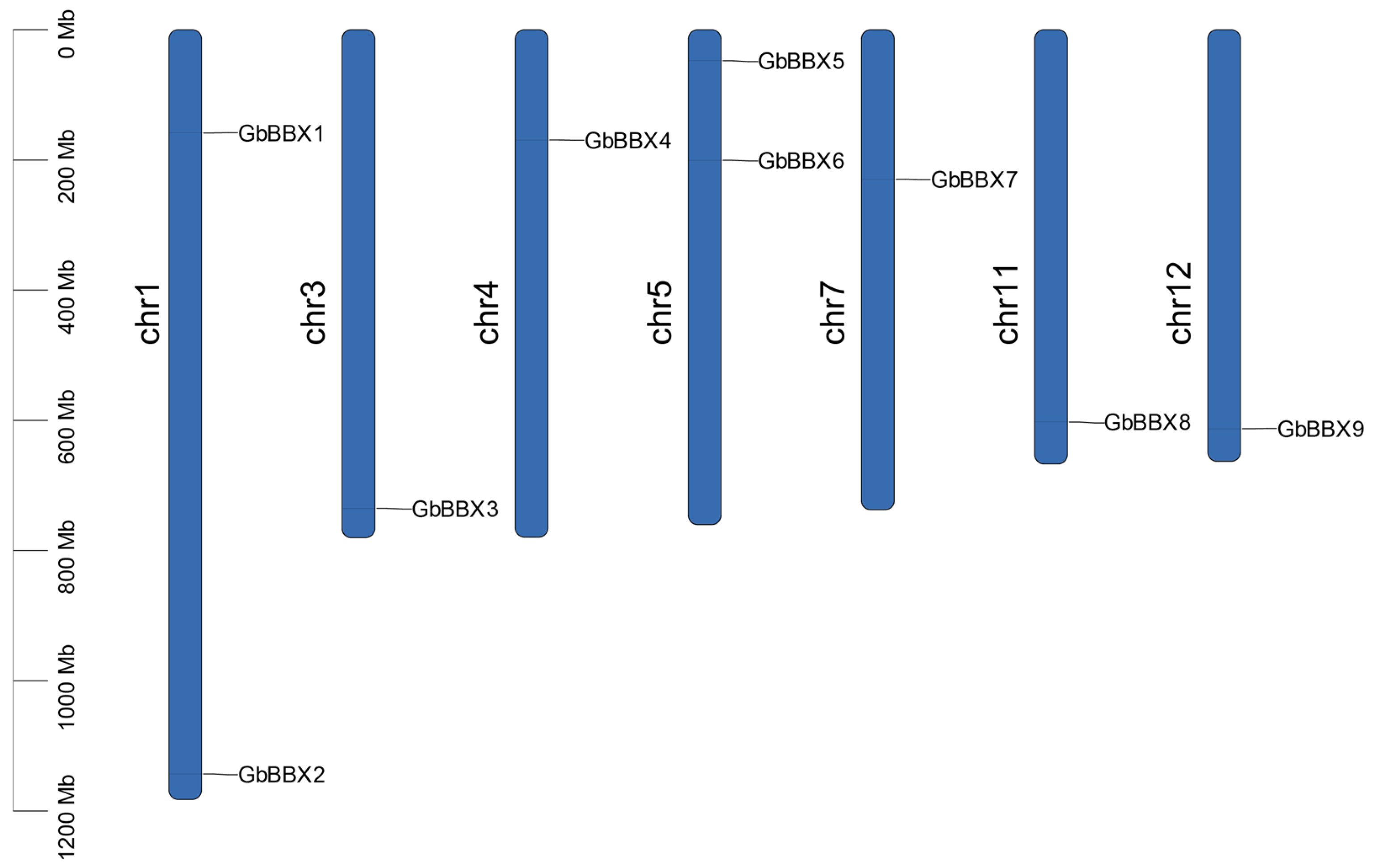
2.1. Whole-Genome Identification of the GbBBX Genes in G. biloba
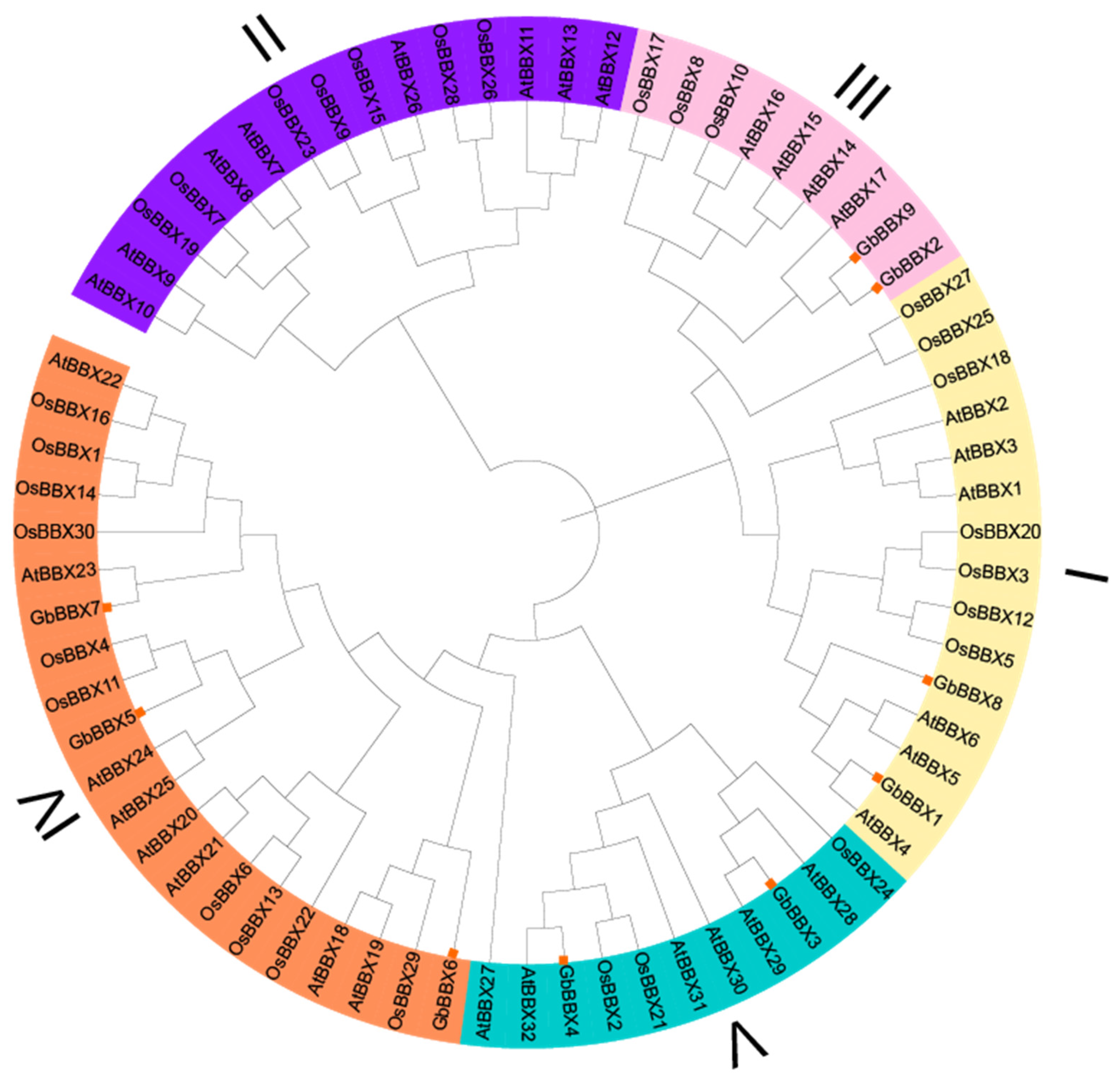
2.2. Phylogenetic Analysis of GbBBX Genes
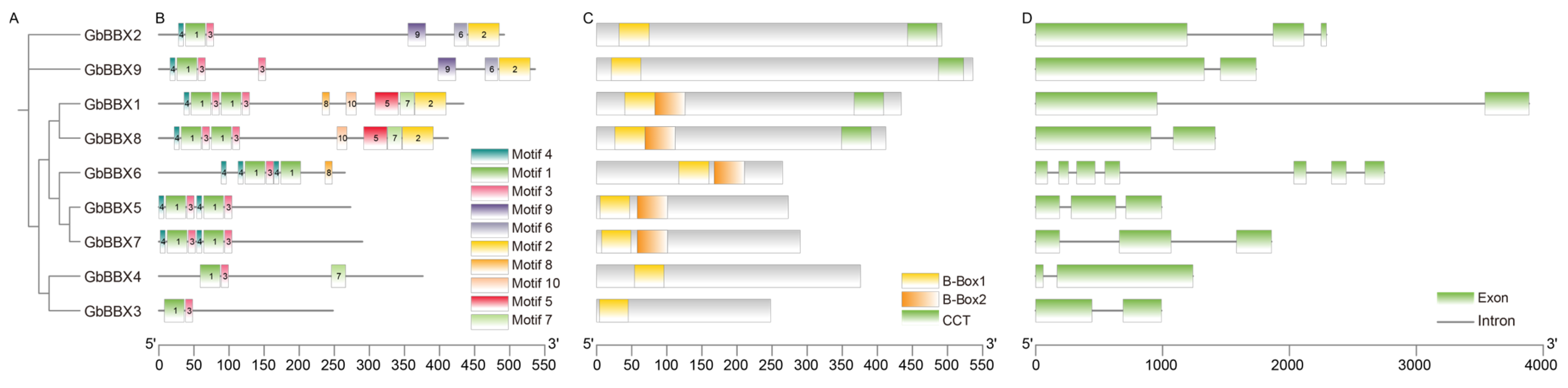
2.3. Motif, Conserved Domain, and Gene Structure Analysis of GbBBXs
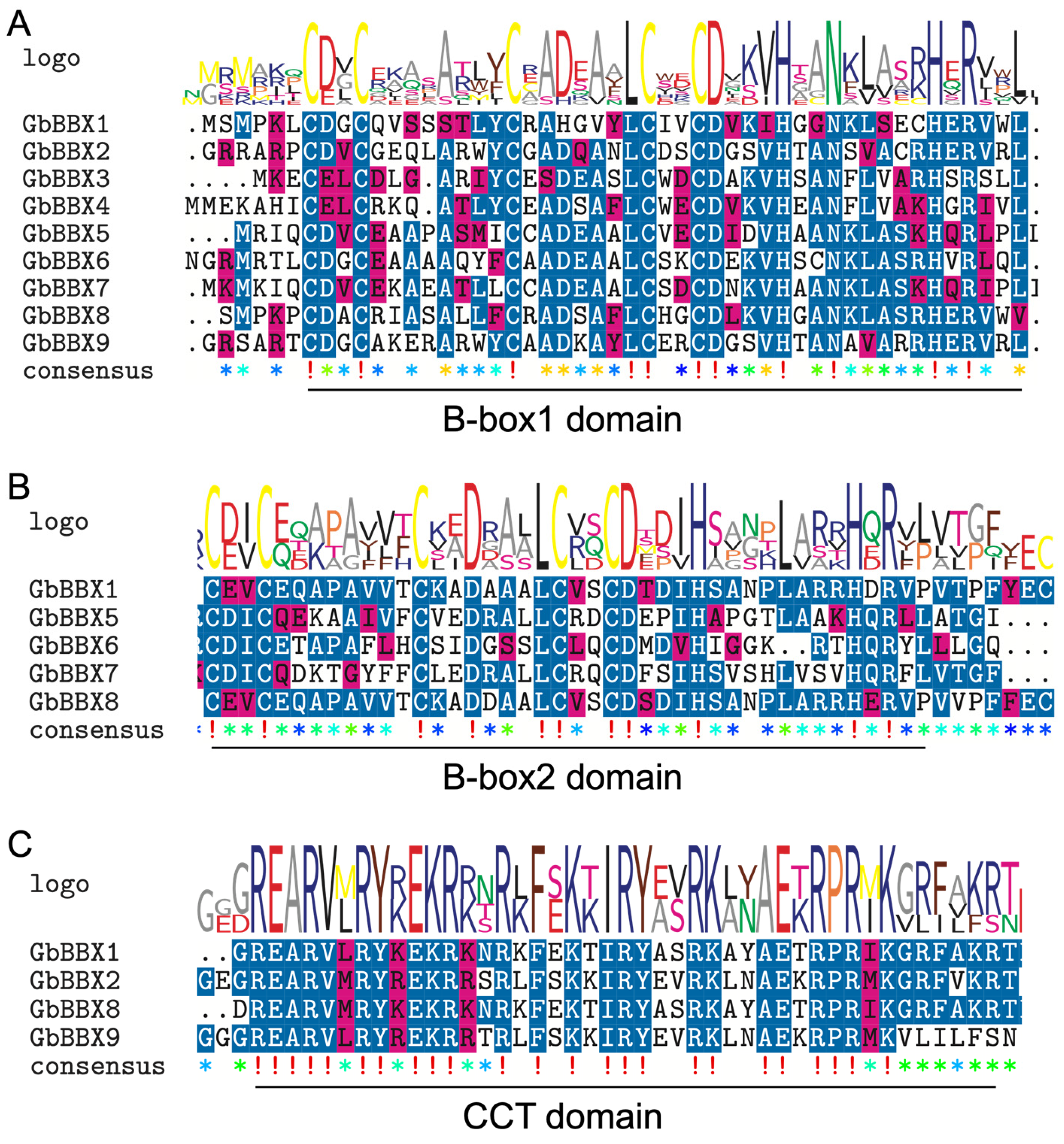
2.4. Multiple Sequence Alignment Analysis
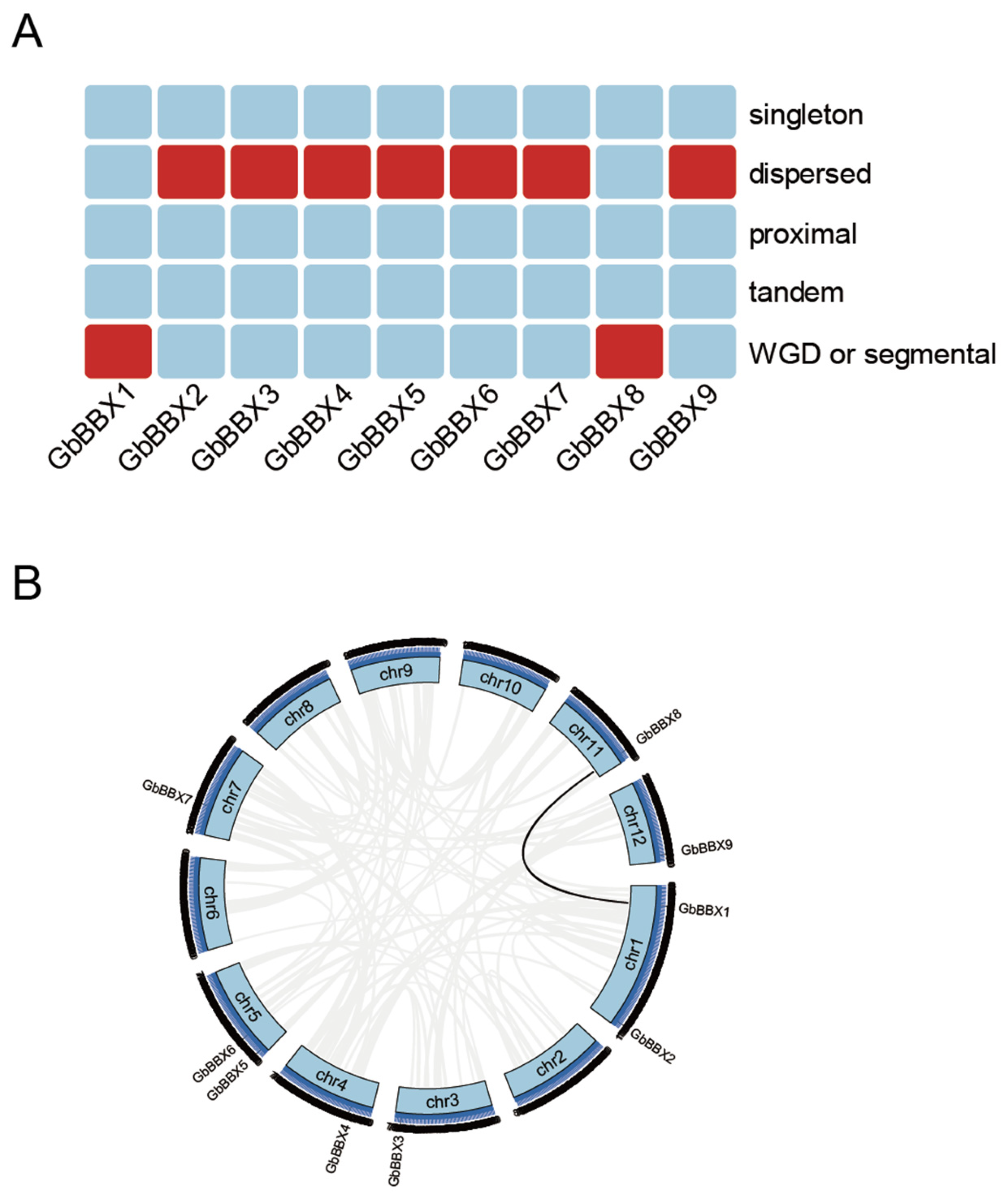
2.5. The Expansion and Collinearity Analysis of the GbBBX Gene Family
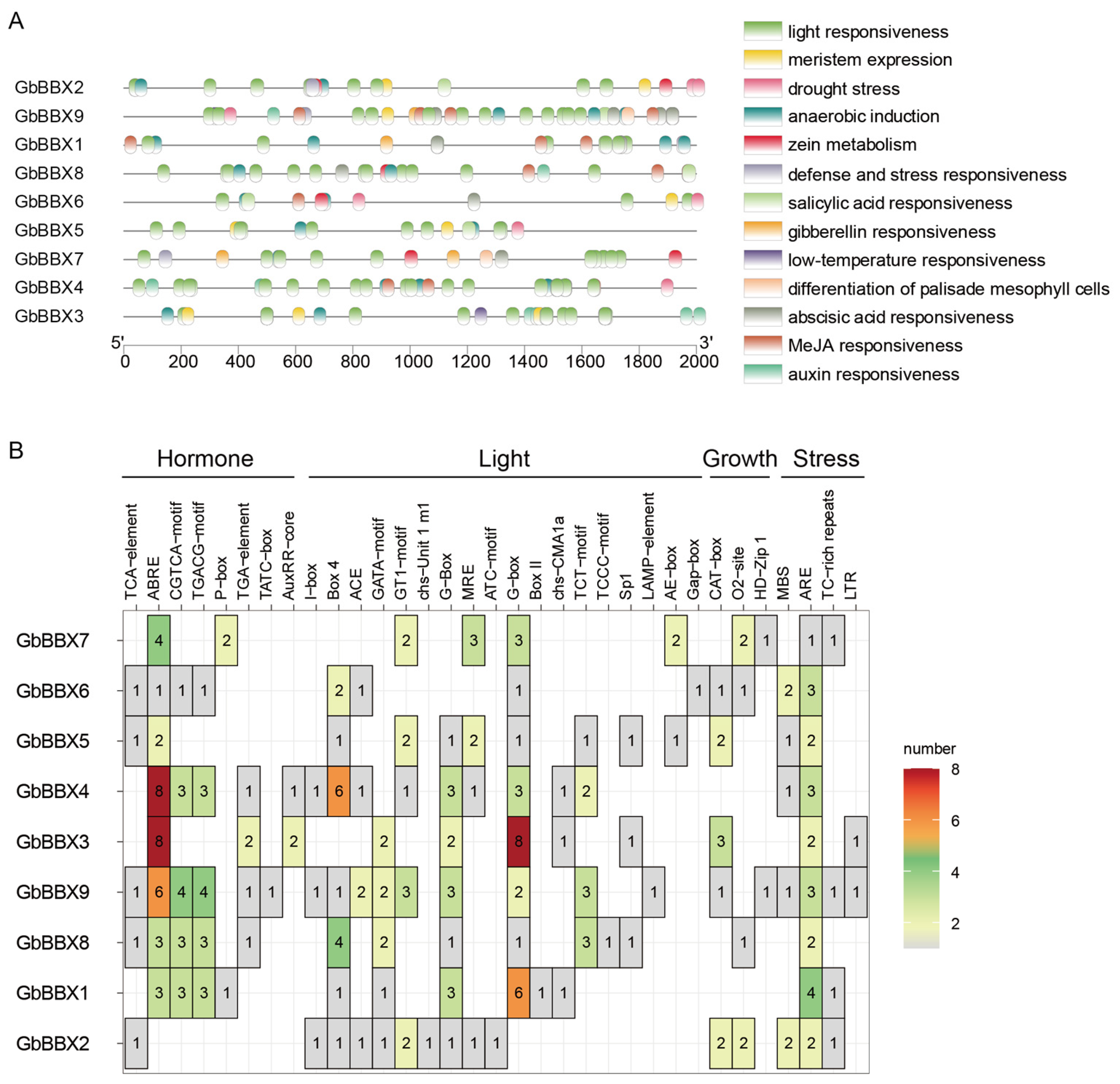
2.6. Cis-Acting Elements Analysis in the Promoter of GbBBX Genes
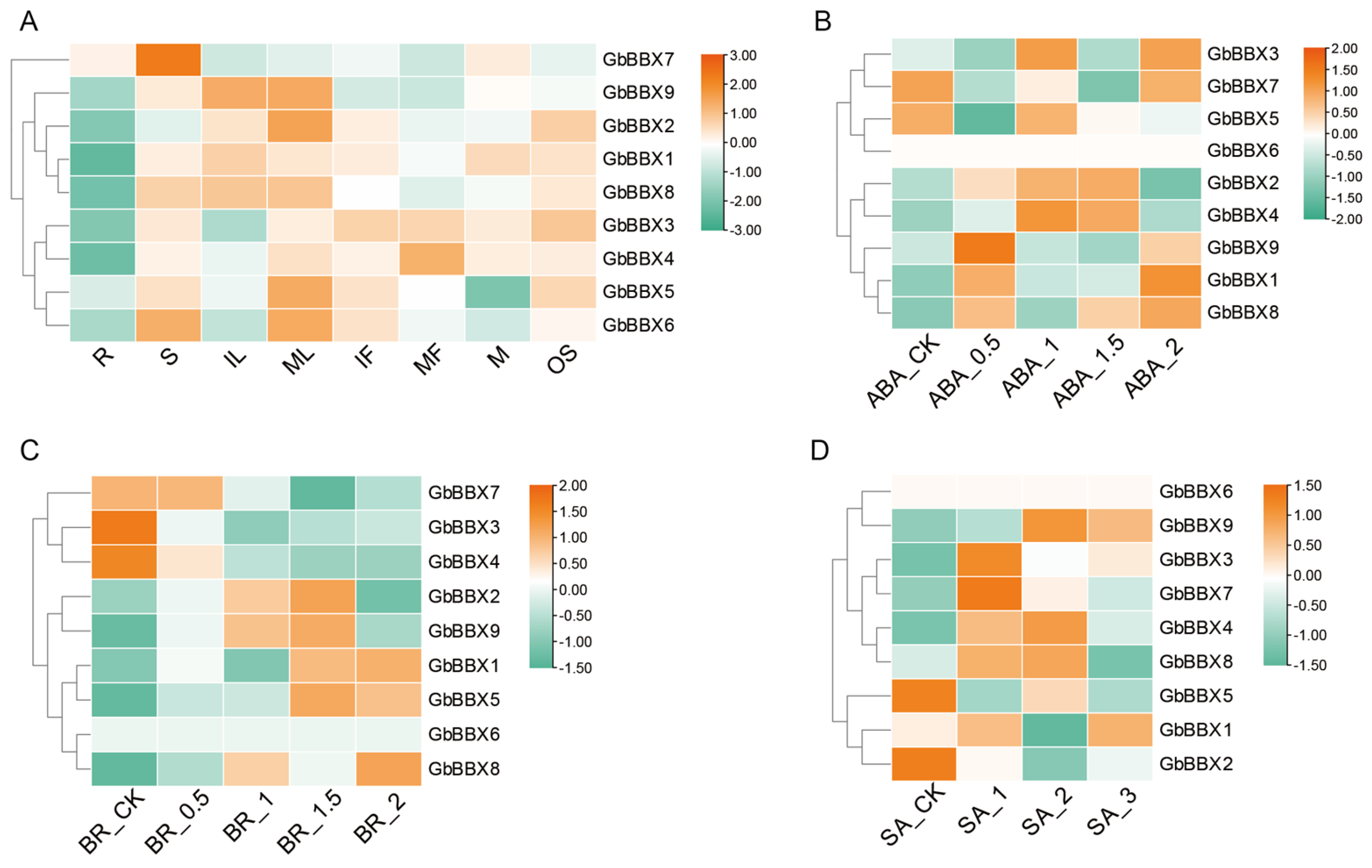
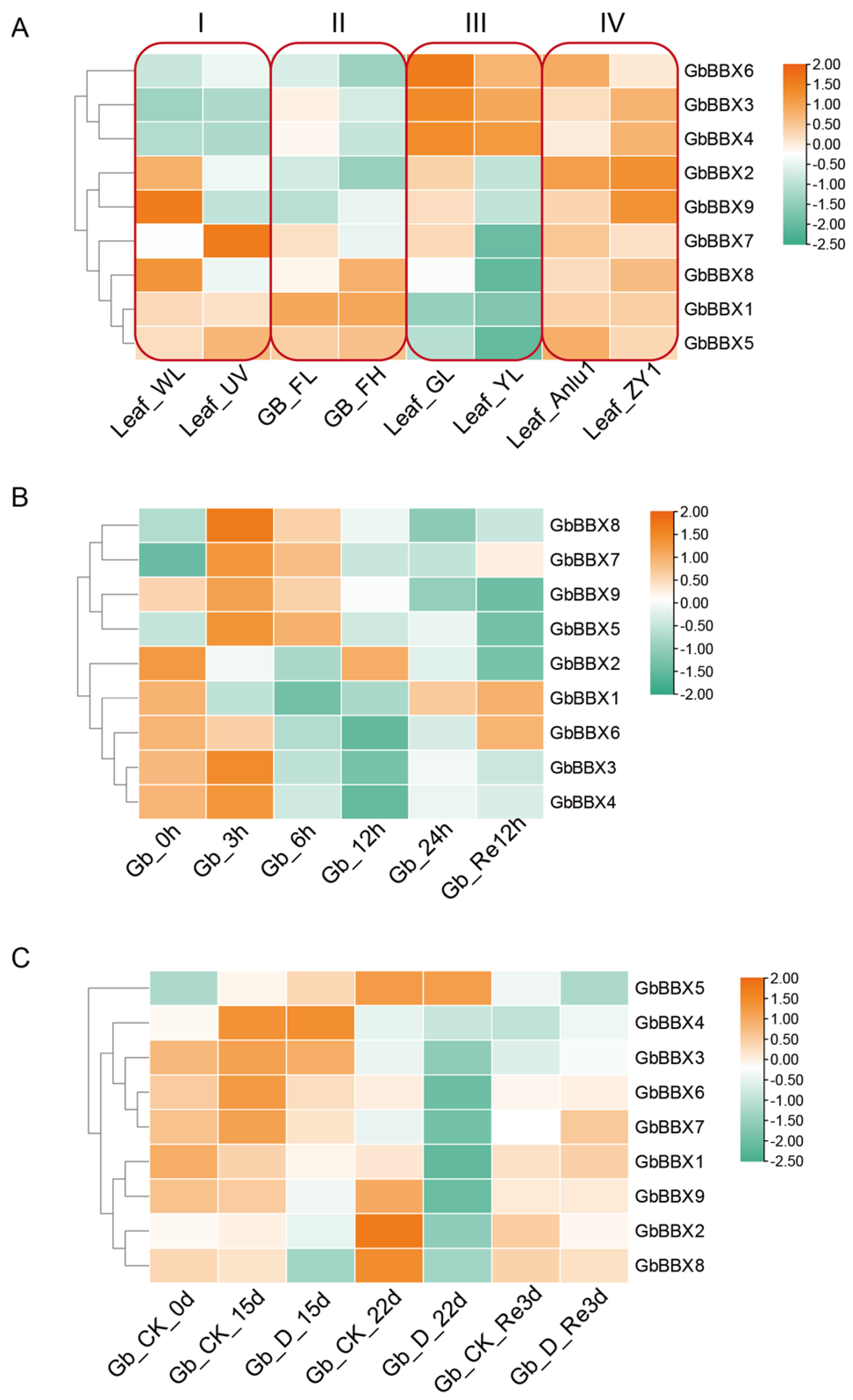
2.7. Expression Profiles of GbBBX Genes in Various Organs, Hormones, Low- and High-Flavonoid Leaves, and Water Stress in G. biloba
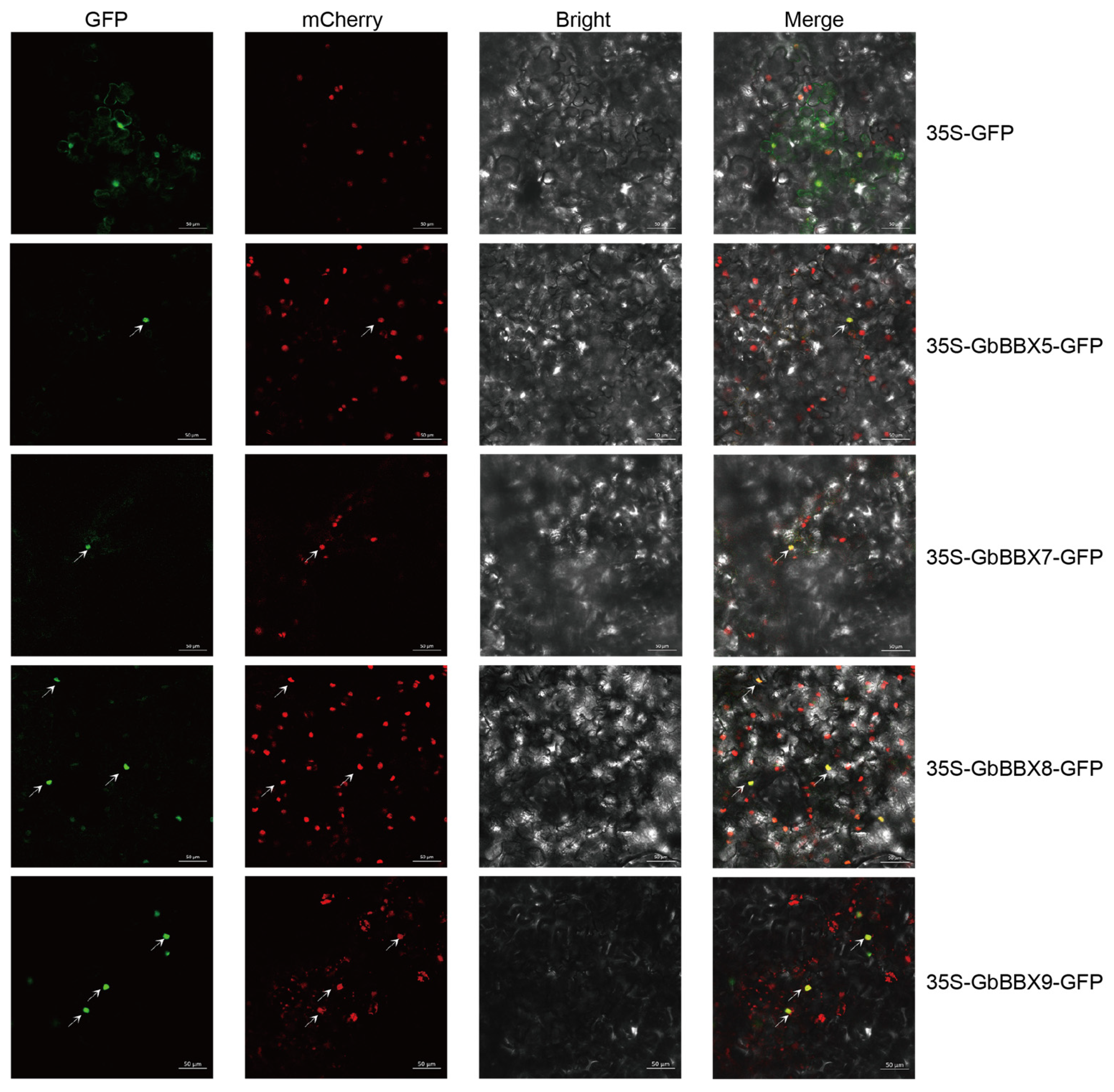
2.8. Quantitative Real-Time PCR (qRT-PCR) Analysis and Subcellular Localization Assay of the GbBBX Genes
3. Discussion
3.1. Characteristics and Evolution of BBX Genes in G. biloba
3.2. Expression Patterns and Potential Functions of the GbBBX Genes
4. Materials and Methods
4.1. Whole-Genome Identification of B-Box Genes in G. biloba
4.2. Physicochemical Properties, Subcellular Localization Prediction, and Chromosomal Location Analysis of BBX Genes
4.3. Phylogenetic Analysis
4.4. Motif, Conserved Domain, Gene Structure, and Cis-Elements Analysis
4.5. Gene Duplication and Syntenic Analysis of the GbBBX Gene Family
4.6. RNA-Seq and Quantitative Real-Time PCR (qRT-PCR) Based Expression Analysis of GbBBX Genes
4.7. Subcellular Localization Analysis
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef]
- Manna, M.; Thakur, T.; Chirom, O.; Mandlik, R.; Deshmukh, R.; Salvi, P. Transcription factors as key molecular target to strengthen the drought stress tolerance in plants. Physiol. Plant 2021, 172, 847–868. [Google Scholar] [CrossRef]
- Gangappa, S.N.; Botto, J.F. The BBX family of plant transcription factors. Trends Plant Sci. 2014, 19, 460–470. [Google Scholar] [CrossRef]
- Khanna, R.; Kronmiller, B.; Maszle, D.R.; Coupland, G.; Holm, M.; Mizuno, T.; Wu, S.H. The Arabidopsis B-Box Zinc Finger Family. Plant Cell 2009, 21, 3416–3420. [Google Scholar] [CrossRef]
- Lau, K.; Podolec, R.; Chappuis, R.; Ulm, R.; Hothorn, M. Plant photoreceptors and their signaling components compete for COP1 binding via VP peptide motifs. EMBO J. 2019, 38, e102140. [Google Scholar] [CrossRef]
- Putterill, J.; Robson, F.; Lee, K.; Simon, R.; Coupland, G. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 1995, 80, 847–857. [Google Scholar] [CrossRef]
- Huang, J.; Zhao, X.; Weng, X.; Wang, L.; Xie, W. The Rice B-Box Zinc Finger Gene Family: Genomic Identification, Characterization, Expression Profiling and Diurnal Analysis. PLoS ONE 2012, 7, e48242. [Google Scholar] [CrossRef]
- Chu, Z.; Wang, X.; Li, Y.; Yu, H.; Li, J.; Lu, Y.; Li, H.; Ouyang, B. Genomic Organization, Phylogenetic and Expression Analysis of the B-BOX Gene Family in Tomato. Front. Plant Sci. 2016, 7, 1552. [Google Scholar] [CrossRef]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jiao, C.; Jin, Q.; Lin, Y.; Cai, Y. B-BOX genes: Genome-wide identification, evolution and their contribution to pollen growth in pear (Pyrus bretschneideri Rehd.). BMC Plant Biol. 2017, 17, 156. [Google Scholar] [CrossRef]
- Shalmani, A.; Fan, S.; Jia, P.; Li, G.; Muhammad, I.; Li, Y.; Sharif, R.; Dong, F.; Zuo, X.; Li, K.; et al. Genome Identification of B-BOX Gene Family Members in Seven Rosaceae Species and Their Expression Analysis in Response to Flower Induction in Malus domestica. Molecules 2018, 23, 1763. [Google Scholar] [CrossRef]
- Cao, Y.; Meng, D.; Han, Y.; Chen, T.; Jiao, C.; Chen, Y.; Jin, Q.; Cai, Y. Comparative analysis of B-BOX genes and their expression pattern analysis under various treatments in Dendrobium officinale. BMC Plant Biol. 2019, 19, 245. [Google Scholar] [CrossRef]
- Feng, Z.; Li, M.; Li, Y.; Yang, X.; Wei, H.; Fu, X.; Ma, L.; Lu, J.; Wang, H.; Yu, S. Comprehensive identification and expression analysis of B-Box genes in cotton. BMC Genom. 2021, 22, 439. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Liu, Y.; Li, X.; Wang, G.; Zhou, Q.; Chen, Q.; Li, J.; Wang, X.; Tang, H. An Evolutionary Analysis of B-Box Transcription Factors in Strawberry Reveals the Role of FaBBx28c1 in the Regulation of Flowering Time. Int. J. Mol. Sci. 2021, 22, 1766. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Tang, R.; Zhang, Y.; Liu, X.; Gao, Y.; Dai, Z.; Li, S.; Wu, B.; Wang, L. Genome-wide identification of B-box proteins and VvBBX44 involved in light-induced anthocyanin biosynthesis in grape (Vitis vinifera L.). Planta 2021, 253, 114. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, L.; Ji, M.; Wu, Y.; Zhang, S.; Zhu, Y.; Yao, J.; Li, Z.; Gao, H.; Wang, X. Genome-wide identification and expression analysis of the B-box transcription factor gene family in grapevine (Vitis vinifera L.). BMC Genom. 2021, 22, 221. [Google Scholar] [CrossRef]
- Ma, R.; Chen, J.; Huang, B.; Huang, Z.; Zhang, Z. The BBX gene family in Moso bamboo (Phyllostachys edulis): Identification, characterization and expression profiles. BMC Genom. 2021, 22, 533. [Google Scholar] [CrossRef]
- Zhao, J.; Li, H.; Huang, J.; Shi, T.; Meng, Z.; Chen, Q.; Deng, J. Genome-wide analysis of BBX gene family in Tartary buckwheat (Fagopyrum tataricum). PeerJ 2021, 9, e11939. [Google Scholar] [CrossRef]
- Nian, L.; Zhang, X.; Liu, X.; Li, X.; Liu, X.; Yang, Y.; Haider, F.U.; Zhu, X.; Ma, B.; Mao, Z.; et al. Characterization of B-box family genes and their expression profiles under abiotic stresses in the Melilotus albus. Front. Plant Sci. 2022, 13, 990929. [Google Scholar] [CrossRef]
- Obel, H.O.; Cheng, C.; Li, Y.; Tian, Z.; Njogu, M.K.; Li, J.; Lou, Q.; Yu, X.; Yang, Z.; Ogweno, J.O.; et al. Genome-Wide Identification of the B-Box Gene Family and Expression Analysis Suggests Their Potential Role in Photoperiod-Mediated beta-Carotene Accumulation in the Endocarp of Cucumber (Cucumis sativus L.) Fruit. Genes 2022, 13, 658. [Google Scholar] [CrossRef]
- Shan, B.; Bao, G.; Shi, T.; Zhai, L.; Bian, S.; Li, X. Genome-wide identification of BBX gene family and their expression patterns under salt stress in soybean. BMC Genom. 2022, 23, 820. [Google Scholar] [CrossRef]
- He, W.; Liu, H.; Li, Y.; Wu, Z.; Xie, Y.; Yan, X.; Wang, X.; Miao, Q.; Chen, T.; Rahman, S.-u.; et al. Genome-wide characterization of B-box gene family in Artemisia annua L. and its potential role in the regulation of artemisinin biosynthesis. Ind. Crops Prod. 2023, 199, 116736. [Google Scholar] [CrossRef]
- Li, Y.; Tong, Y.; Ye, J.; Zhang, C.; Li, B.; Hu, S.; Xue, X.; Tian, Q.; Wang, Y.; Li, L.; et al. Genome-Wide Characterization of B-Box Gene Family in Salvia miltiorrhiza. Int. J. Mol. Sci. 2023, 24, 2146. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Meng, Z.; He, H.; Du, P.; Dijkwel, P.P.; Shi, S.; Li, H.; Xie, Q. Genome-Wide Analysis of BBX Gene Family in Three Medicago Species Provides Insights into Expression Patterns under Hormonal and Salt Stresses. Int. J. Mol. Sci. 2024, 25, 5778. [Google Scholar] [CrossRef] [PubMed]
- Yadav, A.; Ravindran, N.; Singh, D.; Rahul, P.V.; Datta, S. Role of Arabidopsis BBX proteins in light signaling. J. Plant Biochem. Biotechnol. 2020, 29, 623–635. [Google Scholar] [CrossRef]
- Liu, W.; Wang, T.; Wang, Y.; Liang, X.; Han, J.; Han, D. MbMYBC1, a M. baccata MYB transcription factor, contribute to cold and drought stress tolerance in transgenic Arabidopsis. Front. Plant Sci. 2023, 14, 1446. [Google Scholar] [CrossRef]
- Duan, Y.; Han, J.; Guo, B.; Zhao, W.; Zhou, S.; Zhou, C.; Zhang, L.; Li, X.; Han, D. MbICE1 Confers Drought and Cold Tolerance through Up-Regulating Antioxidant Capacity and Stress-Resistant Genes in Arabidopsis thaliana. Int. J. Mol. Sci. 2022, 23, 6072. [Google Scholar] [CrossRef]
- Li, W.; Li, P.; Chen, H.; Zhong, J.; Liang, X.; Wei, Y.; Zhang, L.; Wang, H.; Han, D. Overexpression of a Fragaria vesca 1R-MYB Transcription Factor Gene (FvMYB114) Increases Salt and Cold Tolerance in Arabidopsis thaliana. Int. J. Mol. Sci. 2023, 24, 5261. [Google Scholar] [CrossRef]
- Li, W.; Wei, Y.; Zhang, L.; Wang, Y.; Song, P.; Li, X.; Han, D. FvMYB44, a Strawberry R2R3-MYB Transcription Factor, Improved Salt and Cold Stress Tolerance in Transgenic Arabidopsis. Agronomy 2023, 13, 1051. [Google Scholar] [CrossRef]
- Wang, D.; Chen, Q.; Chen, W.; Liu, X.; Xia, Y.; Guo, Q.; Jing, D.; Liang, G. A WRKY Transcription Factor, EjWRKY17, from Eriobotrya japonica Enhances Drought Tolerance in Transgenic Arabidopsis. Int. J. Mol. Sci. 2021, 22, 5593. [Google Scholar] [CrossRef]
- Talar, U.; Kielbowicz-Matuk, A. Beyond Arabidopsis: BBX Regulators in Crop Plants. Int. J. Mol. Sci. 2021, 22, 2906. [Google Scholar] [CrossRef]
- Shalmani, A.; Jing, X.-Q.; Shi, Y.; Muhammad, I.; Zhou, M.-R.; Wei, X.-Y.; Chen, Q.-Q.; Li, W.-Q.; Liu, W.-T.; Chen, K.-M. Characterization of B-BOX gene family and their expression profiles under hormonal, abiotic and metal stresses in Poaceae plants. BMC Genom. 2019, 20, 27. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, Q.; Zhai, H.; Gao, S.; Yang, L.; Wang, Z.; Xu, Y.; Huo, J.; Ren, Z.; Zhao, N.; et al. IbBBX24 Promotes the Jasmonic Acid Pathway and Enhances Fusarium Wilt Resistance in Sweet Potato. Plant Cell 2020, 32, 1102–1123. [Google Scholar] [CrossRef]
- An, J.P.; Wang, X.F.; Zhang, X.W.; Bi, S.Q.; You, C.X.; Hao, Y.J. MdBBX22 regulates UV-B-induced anthocyanin biosynthesis through regulating the function of MdHY5 and is targeted by MdBT2 for 26S proteasome-mediated degradation. Plant Biotechnol. J. 2019, 17, 2231–2233. [Google Scholar] [CrossRef]
- An, J.-P.; Wang, X.-F.; Espley, R.V.; Lin-Wang, K.; Bi, S.-Q.; You, C.-X.; Hao, Y.-J. An Apple B-Box Protein MdBBX37 Modulates Anthocyanin Biosynthesis and Hypocotyl Elongation Synergistically with MdMYBs and MdHYS. Plant Cell Physiol. 2020, 61, 130–143. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Saito, T.; Honda, C.; Hatsuyama, Y.; Ito, A.; Moriguchi, T. An apple B-box protein, MdCOL11, is involved in UV-B- and temperature-induced anthocyanin biosynthesis. Planta 2014, 240, 1051–1062. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Tao, R.; Yin, L.; Ni, J.; Yang, Q.; Yan, X.; Yang, F.; Guo, X.; Li, H.; Teng, Y. Two B-box proteins, PpBBX18 and PpBBX21, antagonistically regulate anthocyanin biosynthesis via competitive association with Pyrus pyrifolia ELONGATED HYPOCOTYL 5 in the peel of pear fruit. Plant J. 2019, 100, 1208–1223. [Google Scholar] [CrossRef] [PubMed]
- Bai, S.; Tao, R.; Tang, Y.; Yin, L.; Ma, Y.; Ni, J.; Yan, X.; Yang, Q.; Wu, Z.; Zeng, Y.; et al. BBX16, a B-box protein, positively regulates light-induced anthocyanin accumulation by activating MYB10 in red pear. Plant Biotechnol. J. 2019, 17, 1985–1997. [Google Scholar] [CrossRef]
- Zheng, X.-T.; Yu, Z.-C.; Tang, J.-W.; Cai, M.-L.; Chen, Y.-L.; Yang, C.-W.; Chow, W.S.; Peng, C.-L. The major photoprotective role of anthocyanins in leaves of Arabidopsis thaliana under long-term high light treatment: Antioxidant or light attenuator? Photosynth. Res. 2021, 149, 25–40. [Google Scholar] [CrossRef]
- Nagaoka, S.; Takano, T. Salt tolerance-related protein STO binds to a Myb transcription factor homologue and confers salt tolerance in Arabidopsis. J. Exp. Bot. 2003, 54, 2231–2237. [Google Scholar] [CrossRef]
- Liu, X.; Li, R.; Dai, Y.; Yuan, L.; Sun, Q.; Zhang, S.; Wang, X. A B-box zinc finger protein, MdBBX10, enhanced salt and drought stresses tolerance in Arabidopsis. Plant Mol. Biol. 2019, 99, 437–447. [Google Scholar] [CrossRef]
- Mbambalala, N.; Panda, S.K.; van der Vyver, C. Overexpression of AtBBX29 Improves Drought Tolerance by Maintaining Photosynthesis and Enhancing the Antioxidant and Osmolyte Capacity of Sugarcane Plants. Plant Mol. Biol. Rep. 2021, 39, 419–433. [Google Scholar] [CrossRef]
- Liu, J.; Shen, J.; Xu, Y.; Li, X.; Xiao, J.; Xiong, L. Ghd2, a CONSTANS-like gene, confers drought sensitivity through regulation of senescence in rice. J. Exp. Bot. 2016, 67, 5785–5798. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Chen, C.; Xu, M.; Wang, G.; Xu, L.A.; Wu, Y. Overexpression of Ginkgo BBX25 enhances salt tolerance in Transgenic Populus. Plant Physiol. Biochem. 2021, 167, 946–954. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, X.; Wang, G.; Cui, P.; Wu, S.; Ai, C.; Hu, N.; Li, A.; He, B.; Shao, X.; et al. The nearly complete genome of Ginkgo biloba illuminates gymnosperm evolution. Nat. Plants 2021, 7, 748–756. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Cheng, S.; Zhou, X.; Chen, Z.; Kim, S.U.; Tan, J.; Zheng, J.; Xu, F.; Zhang, W.; Liao, Y.; et al. A global survey of full-length transcriptome of Ginkgo biloba reveals transcript variants involved in flavonoid biosynthesis. Ind. Crops Prod. 2019, 139, 111547. [Google Scholar] [CrossRef]
- Guo, F.; Guo, J.; El-Kassaby, Y.A.; Wang, G. Genome-Wide Identification of Expansin Gene Family and Their Response under Hormone Exposure in Ginkgo biloba L. Int. J. Mol. Sci. 2023, 24, 5901. [Google Scholar] [CrossRef]
- Guo, F.; Xiong, W.; Guo, J.; Wang, G. Systematic Identification and Expression Analysis of the Auxin Response Factor (ARF) Gene Family in Ginkgo biloba L. Int. J. Mol. Sci. 2022, 23, 6754. [Google Scholar] [CrossRef]
- Zhao, B.; Wang, L.; Pang, S.; Jia, Z.; Wang, L.; Li, W.; Jin, B. UV-B promotes flavonoid synthesis in Ginkgo biloba leaves. Ind. Crops Prod. 2020, 151, 112483. [Google Scholar] [CrossRef]
- Wu, Y.; Guo, J.; Zhou, Q.; Xin, Y.; Wang, G.; Xu, L.-a. De novo transcriptome analysis revealed genes involved in flavonoid biosynthesis, transport and regulation in Ginkgo biloba. Ind. Crops Prod. 2018, 124, 226–235. [Google Scholar] [CrossRef]
- Wu, Y.; Guo, J.; Wang, T.; Cao, F.; Wang, G. Metabolomic and transcriptomic analyses of mutant yellow leaves provide insights into pigment synthesis and metabolism in Ginkgo biloba. BMC Genom. 2020, 21, 858. [Google Scholar] [CrossRef]
- Meng, J.; Wang, B.; He, G.; Wang, Y.; Tang, X.; Wang, S.; Ma, Y.; Fu, C.; Chai, G.; Zhou, G. Metabolomics Integrated with Transcriptomics Reveals Redirection of the Phenylpropanoids Metabolic Flux in Ginkgo biloba. J. Agric. Food Chem. 2019, 67, 3284–3291. [Google Scholar] [CrossRef]
- Ming, M.; Zhang, J.; Tang, J.; Zhang, J.; Fu, F.; Cao, F. Transcriptome Profiling Revealed ABA Signaling Pathway-Related Genes and Major Transcription Factors Involved in the Response to Water Shock and Rehydration in Ginkgo biloba. Forests 2024, 15, 1690. [Google Scholar] [CrossRef]
- Ming, M.; Zhang, J.; Zhang, J.; Tang, J.; Fu, F.; Cao, F. Transcriptome Profiling Identifies Plant Hormone Signaling Pathway-Related Genes and Transcription Factors in the Drought and Re-Watering Response of Ginkgo biloba. Plants 2024, 13, 2685. [Google Scholar] [CrossRef] [PubMed]
- Chia, T.Y.P.; Mueller, A.; Jung, C.; Mutasa-Goettgens, E.S. Sugar beet contains a large CONSTANS-LIKE gene family including a CO homologue that is independent of the early-bolting (B) gene locus. J. Exp. Bot. 2008, 59, 2735–2748. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Sun, M.; Brewer, L.; Tang, Z.; Nieuwenhuizen, N.; Cooney, J.; Xu, S.; Sheng, J.; Andre, C.; Xue, C.; et al. Allelic variation of BBX24 is a dominant determinant controlling red coloration and dwarfism in pear. Plant Biotechnol. J. 2024, 22, 1468–1490. [Google Scholar] [CrossRef] [PubMed]
- Panchy, N.; Lehti-Shiu, M.; Shiu, S.-H. Evolution of Gene Duplication in Plants. Plant Physiol. 2016, 171, 2294–2316. [Google Scholar] [CrossRef]
- Waadt, R.; Seller, C.A.; Hsu, P.K.; Takahashi, Y.; Munemasa, S.; Schroeder, J.I. Plant hormone regulation of abiotic stress responses. Nat. Rev. Mol. Cell Biol. 2022, 23, 680–694. [Google Scholar] [CrossRef]
- Liu, S.; Gu, X.; Jiang, Y.; Wang, L.; Xiao, N.; Chen, Y.; Jin, B.; Wang, L.; Li, W. UV-B promotes flavonoid biosynthesis in Ginkgo biloba by inducing the GbHY5-GbMYB1-GbFLS module. Hortic. Res. 2023, 10, uhad118. [Google Scholar] [CrossRef]
- Gao, X.-r.; Zhang, H.; Li, X.; Bai, Y.-w.; Peng, K.; Wang, Z.; Dai, Z.-r.; Bian, X.-f.; Zhang, Q.; Jia, L.-c.; et al. The B-box transcription factor IbBBX29 regulates leaf development and flavonoid biosynthesis in sweet potato. Plant Physiol. 2023, 191, 496–514. [Google Scholar] [CrossRef]
- Hrmova, M.; Hussain, S.S. Plant Transcription Factors Involved in Drought and Associated Stresses. Int. J. Mol. Sci. 2021, 22, 5662. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “One for All, All for One” Bioinformatics Platform for Biological Big-data Mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Qiu, D.; Bai, S.; Ma, J.; Zhang, L.; Shao, F.; Zhang, K.; Yang, Y.; Sun, T.; Huang, J.; Zhou, Y.; et al. The genome of Populus alba × Populus tremula var glandulosa clone 84K. DNA Res. 2019, 26, 423–431. [Google Scholar] [CrossRef]
- Liu, S.; Meng, Z.; Zhang, H.; Chu, Y.; Qiu, Y.; Jin, B.; Wang, L. Identification and characterization of thirteen gene families involved in flavonoid biosynthesis in Ginkgo biloba. Ind. Crops Prod. 2022, 188, 115576. [Google Scholar] [CrossRef]
- Zhou, X.; Wang, L.; Yan, J.; Ye, J.; Cheng, S.; Xu, F.; Wang, G.; Zhang, W.; Liao, Y.; Liu, X. Functional Characterization of the EMBRYONIC FLOWER 2 Gene Involved in Flowering in Ginkgo biloba. Front. Plant Sci. 2021, 12, 681166. [Google Scholar] [CrossRef]
- Wang, J.; Luo, Q.; Deng, J.; Liang, X.; Li, Y.; Wang, A.; Lin, T.; Liu, H.; Zhang, X.; Liu, Z.; et al. The G-protein beta subunit SlGB1 regulates tyramine-derived phenolamide metabolism for shoot apex growth and development in tomato. Plant Cell 2025, 37, koaf070. [Google Scholar] [CrossRef]

| Gene Name | Gene ID | Peptide (aa) | Molecular Weight MW (Da) | Theoretical pI | Grand Average of Hydropathicity (GRAVY) | Instability Index | Aliphatic Index | Prediction of Subcellular Localization |
|---|---|---|---|---|---|---|---|---|
| GbBBX1 | evm.model.chr1.520 | 434 | 48,292.50 | 5.19 | −0.690 | 41.33 | 62.67 | nuclear |
| GbBBX2 | evm.model.chr1.3215 | 492 | 54,652.64 | 5.08 | −0.684 | 44.93 | 63.82 | nuclear |
| GbBBX3 | evm.model.chr3.2337 | 248 | 26,808.51 | 4.84 | −0.598 | 53.98 | 67.62 | nuclear |
| GbBBX4 | evm.model.chr4.597 | 376 | 42,009.76 | 8.11 | −0.173 | 46.94 | 87.98 | chloroplast |
| GbBBX5 (GbBBX25) | evm.model.chr5.165 | 273 | 29,581.52 | 4.96 | −0.319 | 56.35 | 74.07 | nuclear |
| GbBBX6 | evm.model.chr5.546 | 265 | 29,541.79 | 7.83 | −0.271 | 45.33 | 79.13 | chloroplast |
| GbBBX7 | evm.model.chr7.750 | 290 | 31,887.14 | 5.65 | −0.434 | 51.32 | 75.38 | nuclear |
| GbBBX8 | evm.model.chr11.1562 | 412 | 45,117.02 | 5.80 | −0.286 | 46.13 | 72.18 | nuclear |
| GbBBX9 | evm.model.chr12.1654 | 536 | 60,029.39 | 4.95 | −0.746 | 47.55 | 66.08 | nuclear |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ming, M.; Yi, M.; Sun, K.; Zu, A.; Zhang, J.; Fu, F.; Cao, F.; Yang, X. Genome-Wide Identification and Expression Analysis of the Ginkgo biloba B-Box Gene Family in Response to Hormone Treatments, Flavonoid Levels, and Water Stress. Int. J. Mol. Sci. 2025, 26, 8427. https://doi.org/10.3390/ijms26178427
Ming M, Yi M, Sun K, Zu A, Zhang J, Fu F, Cao F, Yang X. Genome-Wide Identification and Expression Analysis of the Ginkgo biloba B-Box Gene Family in Response to Hormone Treatments, Flavonoid Levels, and Water Stress. International Journal of Molecular Sciences. 2025; 26(17):8427. https://doi.org/10.3390/ijms26178427
Chicago/Turabian StyleMing, Meiling, Mulin Yi, Kexin Sun, Anning Zu, Juan Zhang, Fangfang Fu, Fuliang Cao, and Xiaoming Yang. 2025. "Genome-Wide Identification and Expression Analysis of the Ginkgo biloba B-Box Gene Family in Response to Hormone Treatments, Flavonoid Levels, and Water Stress" International Journal of Molecular Sciences 26, no. 17: 8427. https://doi.org/10.3390/ijms26178427
APA StyleMing, M., Yi, M., Sun, K., Zu, A., Zhang, J., Fu, F., Cao, F., & Yang, X. (2025). Genome-Wide Identification and Expression Analysis of the Ginkgo biloba B-Box Gene Family in Response to Hormone Treatments, Flavonoid Levels, and Water Stress. International Journal of Molecular Sciences, 26(17), 8427. https://doi.org/10.3390/ijms26178427






