The Generation of iPSCs Expressing Interferon-Beta Under Doxycycline-Inducible Control
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Cultivation of iPSCs
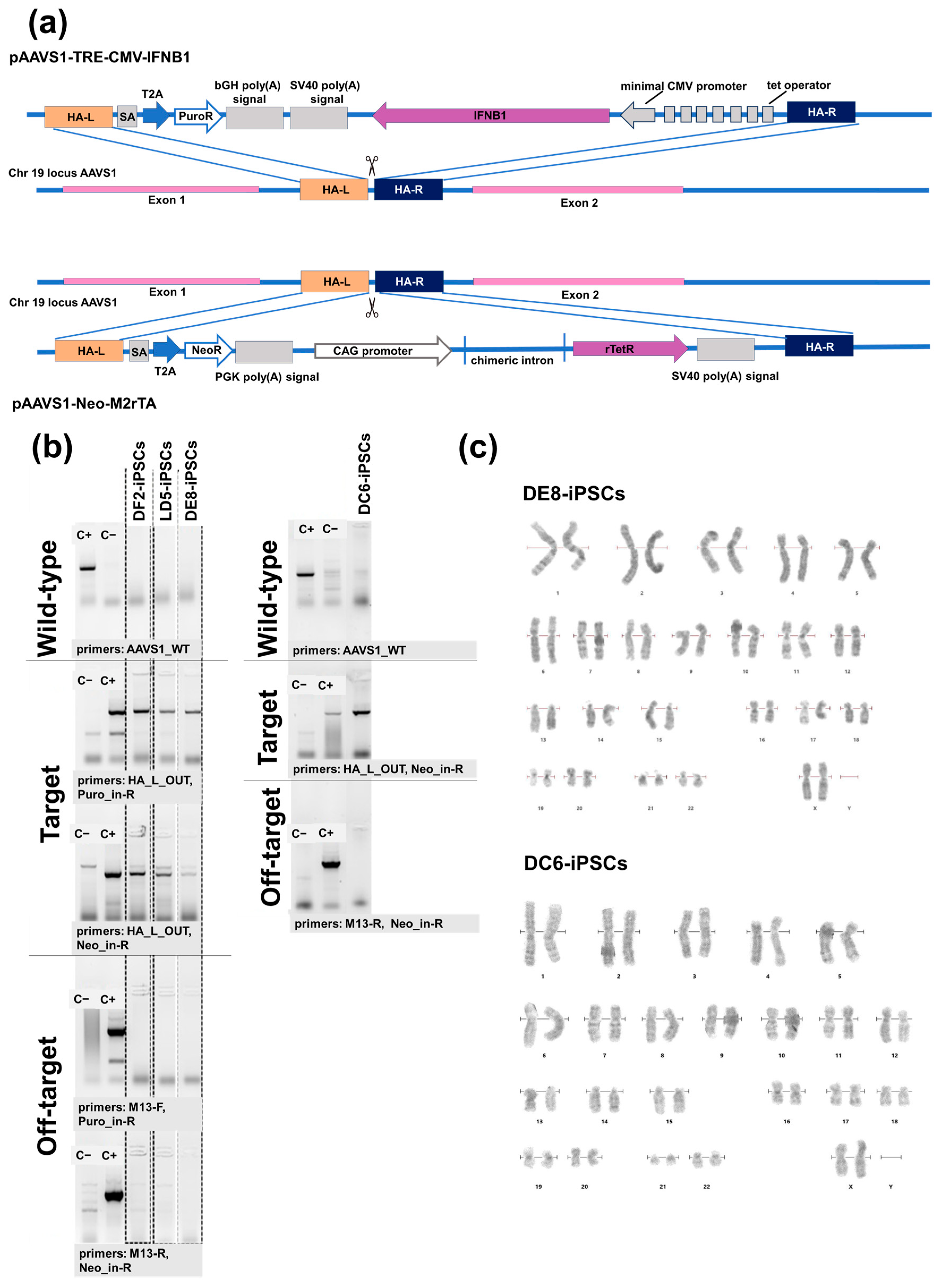
4.2. Obtaining iPSCs with IFN-β Overexpression
4.3. Karyotype Analysis
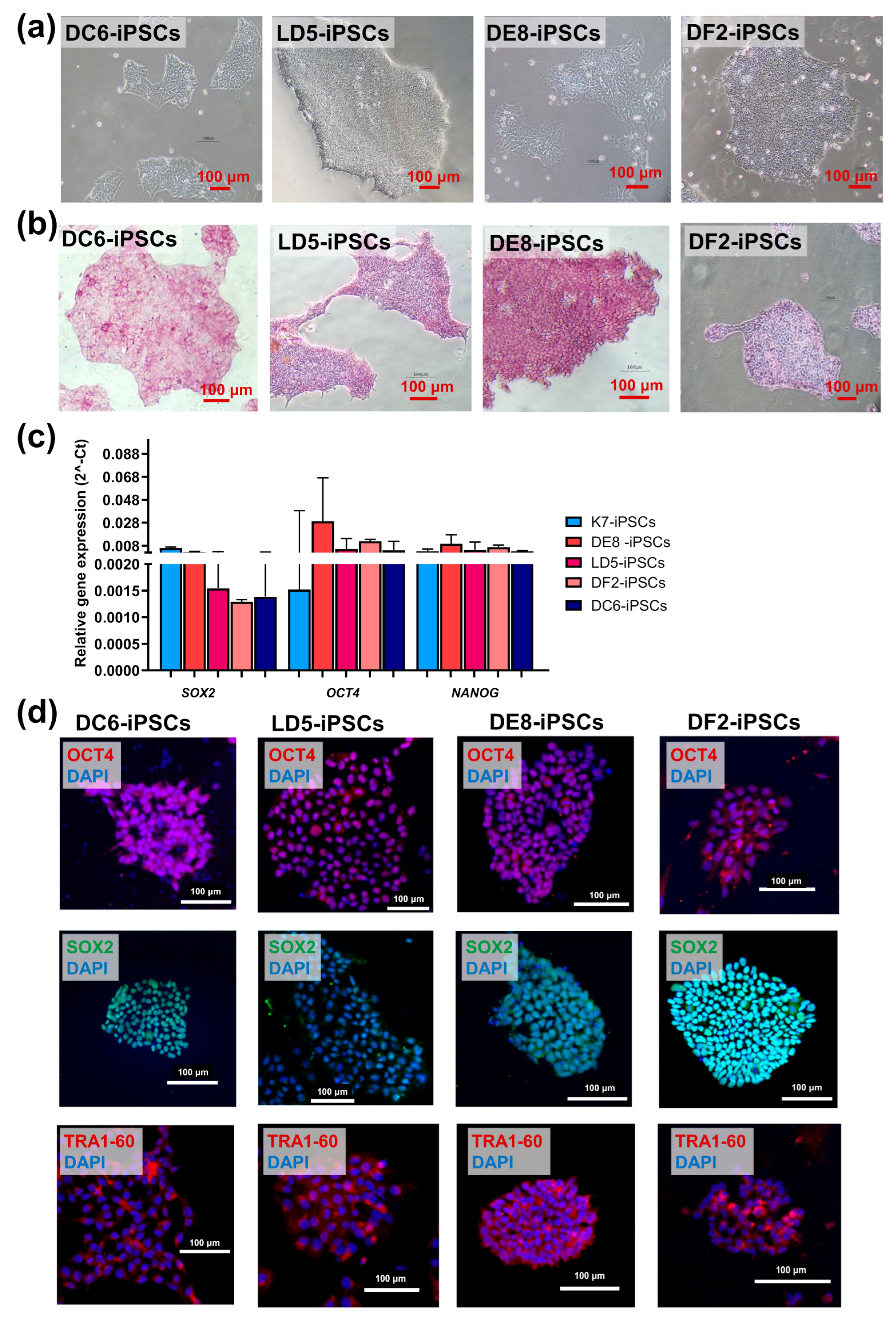
4.4. Alkaline Phosphatase Staining
4.5. Mycoplasma Detection
| Target | Forward/Reverse Primer (5′-3′) |
|---|---|
| OCT4 | GGGTTCTATTTGGGAAGGTATT/GGGTTTCTGCTTTGCATATCT |
| NANOG | GAAATACCTCAGCCTCCAGC/TGCCACCTCTTAGATTTCATTC |
| SOX2 | CTCGCAGACCTACATGAACG/GAGCCAAGAGCCATGCC |
| IFNB1 | GAACTTTGACATCCCTGAGGAG/CCAGTGCTAGATGAATCTTGTCTG |
| GATA4 | GCGGTGCTTCCAGCAACTCCA/GACATCGCACTGACTGAGAACG |
| SOX17 | ACGCTTTCATGGTGTGGGCTAAG/GTCAGCGCCTTCCACGACTT |
| MSX1 | GCGCCAAGGCAAAGAGAC/CGCCGAGAGGGAAGGAG |
| HAND1 | CAAGGATGCACAGTCTGGCGAT/GCAGGAGGAAAACCTTCGTGCT |
| LHX2 | ACGCCAAGGACTTGAAGCAGCT/TTTCCTGCCGTAAGAGGTTGCG |
| SOX1 | GAGTGGAAGGTCATGTCCGAGG/CCTTCTTGAGCAGCGTCTTGGT |
| RPL27 | GGACGCAAAGCTGTCATC/CAATTCCAGCCACCAGAG |
| RPS29 | GCTGTACTGGAGCCACCC/GGCGGCACATATTGAGG |
| Wild-type AAVS1 locus, without target insert (AAVS1_WT) | CTCTGGCTCCATCGTAAGCAA/CCCAAAGTACCCCGTCTCCC |
| M2rtTA plasmid insert position determination | HA_L-OUT CCGGACCACTTTGAGCTCTAC |
| Neo_in-R GCCCAGTCATAGCCGAATAG | |
| INFB1 plasmid insert position determination | HA_L-OUT CCGGACCACTTTGAGCTCTAC |
| Puro_in-R AGGCGCACCGTGGGCTTGTAC | |
| Determination of the presence of non-target inserts of M2rtTA plasmid in the genome | M13-R CAGGAAACAGCTATGAC |
| Neo_in-R GCCCAGTCATAGCCGAATAG | |
| Determination of the presence of non-target inserts of INFB1 plasmid in the genome | M13-F GTAAAACGACGGCCAGT |
| Puro_in-R AGGCGCACCGTGGGCTTGTAC | |
| Detection of mycoplasma, ribosomal 16S RNA gene | GGGAGCAAACAGGATTAGATACCCT/TGCACCATCTGTCACTCTGTTAACCTC |
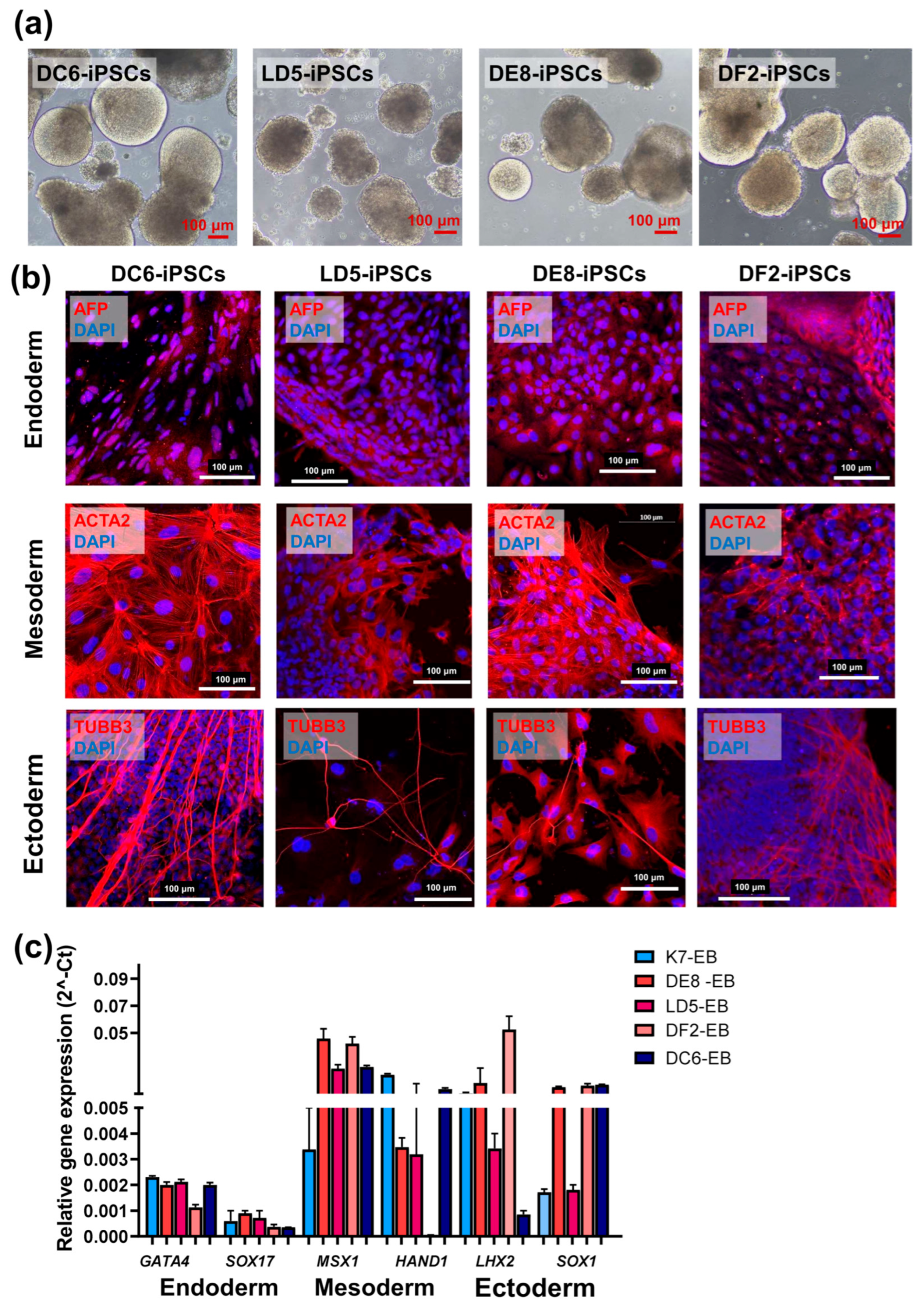
4.6. Spontaneous In Vitro Differentiation
4.7. Immunofluorescence Staining
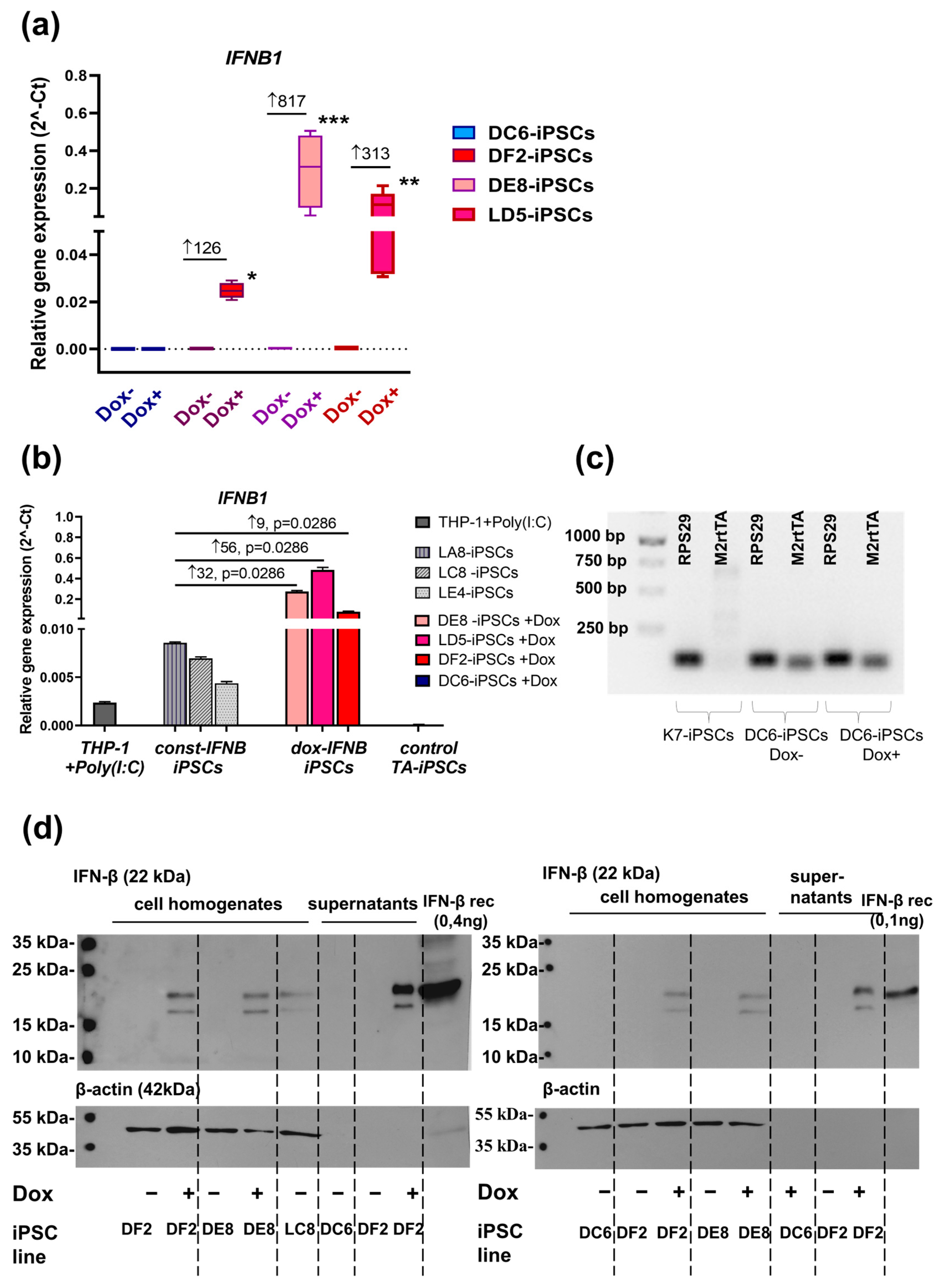
4.8. Reverse Transcription PCR
4.9. Western Blotting
4.10. Cultivation of THP-1
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AFP | alpha-fetoprotein |
| ACTA2 | actin alpha 2, smooth muscle actin |
| Const-IFNB-iPSCs | iPSC cell lines with constitutively overexpression of interferon-beta |
| DAPI | 4′,6-diamidino-2-phenylindol |
| Dox | doxycycline hylate |
| Dox-IFNB-iPSCs | iPSCs with doxycycline- inducible gene overexpression |
| EBs | embryoid bodies |
| FCS | fetal calf serum |
| FGF-2 | fibroblast growth factor-2 |
| IFN-I | type I interferon |
| IFNB1 | interferon-beta 1 |
| IFN-β | interferon-beta 1 protein |
| IFNB-iPSCs | iPSC cell lines with doxycycline-inducible overexpression of interferon-beta |
| iPSCs | induced pluripotent stem cells |
| IRF7 | interferon regulatory factor 7 |
| K7-iPSCs | K7-4Lf iPSC line (alternative name ICGi022-A)-parent iPSC |
| MEFs | mouse embryo fibroblasts |
| M2rtTAf | tetracycline-controlled transactivator protein |
| Poly (I: C) | polyinosinic-polycytidylic acid |
| TA-iPSCs | iPSC cell lines with expression of TA |
| TUBB3 | tubulin beta 3. |
References
- Hockemeyer, D.; Jaenisch, R. Induced Pluripotent Stem Cells Meet Genome Editing. Cell Stem Cell 2016, 18, 573–586. [Google Scholar] [CrossRef]
- Shi, Y.; Inoue, H.; Wu, J.C.; Yamanaka, S. Induced Pluripotent Stem Cell Technology: A Decade of Progress. Nat. Rev. Drug Discov. 2017, 16, 115–130. [Google Scholar] [CrossRef]
- Cerneckis, J.; Cai, H.; Shi, Y. Induced Pluripotent Stem Cells (iPSCs): Molecular Mechanisms of Induction and Applications. Signal Transduct. Target. Ther. 2024, 9, 112. [Google Scholar] [CrossRef]
- Nakao, S.; Ihara, D.; Hasegawa, K.; Kawamura, T. Applications for Induced Pluripotent Stem Cells in Disease Modelling and Drug Development for Heart Diseases. Eur. Cardiol. Rev. 2020, 15, e02. [Google Scholar] [CrossRef] [PubMed]
- Ávila-González, D.; Gidi-Grenat, M.Á.; García-López, G.; Martínez-Juárez, A.; Molina-Hernández, A.; Portillo, W.; Díaz-Martínez, N.E.; Díaz, N.F. Pluripotent Stem Cells as a Model for Human Embryogenesis. Cells 2023, 12, 1192. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Wang, D.; Wang, B.; Guo, X.; Wang, X.; Wang, J.; Li, Y.; An, G.; Ren, Q. Generation of Heterozygous KCNA2 Knockout Induced Pluripotent Stem Cell Line. Stem Cell Res. 2024, 81, 103596. [Google Scholar] [CrossRef]
- Busley, A.V.; Cyganek, L. Generation of a Genetically-Modified Induced Pluripotent Stem Cell Line Harboring a Noonan Syndrome-Associated Gene Variant MRAS p. G23V. Stem Cell Res. 2023, 69, 103108. [Google Scholar] [CrossRef]
- Lyu, H.; Haag, C.; Schwarz, N.; Löffler, H.; Mau-Holzmann, U.A.; Lerche, H.; Rosa, F. Generation of an Induced Pluripotent Stem Cell (iPSC) Line Carrying a KCNA2 Homozygous (p. Arg294His, R294H) Mutation Related to Hereditary Spastic Paraplegia. Stem Cell Res. 2025, 86, 103737. [Google Scholar] [CrossRef]
- Liu, J.; Chen, W.; Zhao, Z.; Xu, H.H. Effect of NELL1 Gene Overexpression in iPSC-MSCs Seeded on Calcium Phosphate Cement. Acta Biomater. 2014, 10, 5128–5138. [Google Scholar] [CrossRef][Green Version]
- Zhou, J.; Cui, B.; Wang, X.; Wang, H.; Zheng, J.; Guo, F.; Sun, Y.; Fan, H.; Shen, J.; Su, J.; et al. Overexpression of KCNJ2 Enhances Maturation of Human-Induced Pluripotent Stem Cell-Derived Cardiomyocytes. Stem Cell Res. Ther. 2023, 14, 92. [Google Scholar] [CrossRef] [PubMed]
- Sheveleva, O.; Protasova, E.; Nenasheva, T.; Butorina, N.; Melnikova, V.; Gerasimova, T.; Sakovnich, O.; Kurinov, A.; Grigor’eva, E.; Medvedev, S. A Model of iPSC-Derived Macrophages with TNFAIP3 Overexpression Reveals the Peculiarities of TNFAIP3 Protein Expression and Function in Human Macrophages. Int. J. Mol. Sci. 2023, 24, 12868. [Google Scholar] [CrossRef]
- Shin, A.; Park, S.A.; Kim, J.Y.; Baek, E.J. TAL1 Overexpression in Induced Pluripotent Stem Cells Promotes the Formation of Hematopoietic Cell-Forming Complexes but Inhibits Enucleation in Vitro. Front. Cell Dev. Biol. 2025, 13, 1474631. [Google Scholar] [CrossRef]
- McNab, F.; Mayer-Barber, K.; Sher, A.; Wack, A.; O’garra, A. Type I Interferons in Infectious Disease. Nat. Rev. Immunol. 2015, 15, 87–103. [Google Scholar] [CrossRef] [PubMed]
- Boxx, G.M.; Cheng, G. The Roles of Type I Interferon in Bacterial Infection. Cell Host Microbe 2016, 19, 760–769. [Google Scholar] [CrossRef]
- Kovarik, P.; Castiglia, V.; Ivin, M.; Ebner, F. Type I Interferons in Bacterial Infections: A Balancing Act. Front. Immunol. 2016, 7, 652. [Google Scholar] [CrossRef] [PubMed]
- Kretschmer, S.; Lee-Kirsch, M.A. Type I Interferon-Mediated Autoinflammation and Autoimmunity. Curr. Opin. Immunol. 2017, 49, 96–102. [Google Scholar] [CrossRef]
- Yu, R.; Zhu, B.; Chen, D. Type I Interferon-Mediated Tumor Immunity and Its Role in Immunotherapy. Cell. Mol. Life Sci. 2022, 79, 191. [Google Scholar] [CrossRef] [PubMed]
- Snell, L.M.; McGaha, T.L.; Brooks, D.G. Type I Interferon in Chronic Virus Infection and Cancer. Trends Immunol. 2017, 38, 542–557. [Google Scholar] [CrossRef]
- Boukhaled, G.M.; Harding, S.; Brooks, D.G. Opposing Roles of Type I Interferons in Cancer Immunity. Annu. Rev. Pathol. Mech. Dis. 2021, 16, 167–198. [Google Scholar] [CrossRef]
- Li, Q.; Tan, F.; Wang, Y.; Liu, X.; Kong, X.; Meng, J.; Yang, L.; Cen, S. The Gamble between Oncolytic Virus Therapy and IFN. Front. Immunol. 2022, 13, 971674. [Google Scholar] [CrossRef]
- Plosker, G.L. Interferon-β-1b: A Review of Its Use in Multiple Sclerosis. CNS Drugs 2011, 25, 67–88. [Google Scholar] [CrossRef]
- Huang, L.; Li, L.; Lemos, H.; Chandler, P.R.; Pacholczyk, G.; Baban, B.; Barber, G.N.; Hayakawa, Y.; McGaha, T.L.; Ravishankar, B. Cutting Edge: DNA Sensing via the STING Adaptor in Myeloid Dendritic Cells Induces Potent Tolerogenic Responses. J. Immunol. 2013, 191, 3509–3513. [Google Scholar] [CrossRef]
- Zitvogel, L.; Galluzzi, L.; Kepp, O.; Smyth, M.J.; Kroemer, G. Type I Interferons in Anticancer Immunity. Nat. Rev. Immunol. 2015, 15, 405–414. [Google Scholar] [CrossRef] [PubMed]
- Kirkeby, A.; Main, H.; Carpenter, M. Pluripotent Stem-Cell-Derived Therapies in Clinical Trial: A 2025 Update. Cell Stem Cell 2025, 32, 10–37. [Google Scholar] [CrossRef] [PubMed]
- Eggenberger, J.; Blanco-Melo, D.; Panis, M.; Brennand, K.J.; tenOever, B.R. Type I Interferon Response Impairs Differentiation Potential of Pluripotent Stem Cells. Proc. Natl. Acad. Sci. USA 2019, 116, 1384–1393. [Google Scholar] [CrossRef] [PubMed]
- Sheveleva, O.; Protasova, E.; Grigor’eva, E.; Butorina, N.; Kuziaeva, V.; Antonov, D.; Melnikova, V.; Medvedev, S.; Lyadova, I. The Generation of Genetically Engineered Human Induced Pluripotent Stem Cells Overexpressing IFN-β for Future Experimental and Clinically Oriented Studies. Int. J. Mol. Sci. 2024, 25, 12456. [Google Scholar] [CrossRef]
- Malakhova, A.A.; Grigor’eva, E.V.; Pavlova, S.V.; Malankhanova, T.B.; Valetdinova, K.R.; Vyatkin, Y.V.; Khabarova, E.A.; Rzaev, J.A.; Zakian, S.M.; Medvedev, S.P. Generation of Induced Pluripotent Stem Cell Lines ICGi021-A and ICGi022-A from Peripheral Blood Mononuclear Cells of Two Healthy Individuals from Siberian Population. Stem Cell Res. 2020, 48, 101952. [Google Scholar] [CrossRef]
- Pavlova, S.V.; Valetdinova, K.R.; Malankhanova, T.B.; Polivtsev, D.E.; Malahova, A.A.; Grigor’eva, E.V.; Shevchenko, A.I.; Zakian, S.M.; Medvedev, S.P. Transgenic Lines of Human Induced Pluripotent Stem Cells ICGi022-A-6 and ICGi022-A-7 with Doxycycline-Inducible Variants of Programmable Nuclease AsCas12a. Russ. J. Dev. Biol. 2023, 54, 374–386. [Google Scholar] [CrossRef]
- Ustyantseva, E.; Pavlova, S.V.; Malakhova, A.A.; Ustyantsev, K.; Zakian, S.M.; Medvedev, S.P. Oxidative Stress Monitoring in iPSC-Derived Motor Neurons Using Genetically Encoded Biosensors of H2O2. Sci. Rep. 2022, 12, 8928. [Google Scholar] [CrossRef]
- Blank, T.; Prinz, M. Type I Interferon Pathway in CNS Homeostasis and Neurological Disorders. Glia 2017, 65, 1397–1406. [Google Scholar] [CrossRef]
- Tan, P.H.; Ji, J.; Yeh, C.C.; Ji, R.R. Interferons in Pain and Infections: Emerging Roles in Neuro-Immune and Neuro-Glial Interactions. Front. Immunol. 2021, 12, 783725. [Google Scholar] [CrossRef] [PubMed]
- Tresse, E.; Riera-Ponsati, L.; Jaberi, E.; Sew, W.Q.G.; Ruscher, K.; Issazadeh-Navikas, S. IFN-β Rescues Neurodegeneration by Regulating Mitochondrial Fission via STAT5, PGAM5, and Drp1. EMBO J. 2021, 40, e106868. [Google Scholar] [CrossRef]
- Wang, Q.; Yuan, S.; Wang, C.; Huang, D.; Zhang, M.; Zhan, Y.; Gao, F.; Shi, J.; Levey, A.I.; Shen, Y. Brain Derived β-Interferon Is a Potential Player in Alzheimer’s Disease Pathogenesis and Cognitive Impairment. Alz. Res. Ther. 2024, 16, 271. [Google Scholar] [CrossRef]
- Quan, P.; Li, X.; Si, Y.; Sun, L.; Ding, F.F.; Fan, Y.; Liu, H.; Wei, C.; Li, R.; Zhao, X.; et al. Single Cell Analysis Reveals the Roles and Regulatory Mechanisms of Type-I Interferons in Parkinson’s Disease. Cell Commun. Signal. 2024, 22, 212. [Google Scholar] [CrossRef] [PubMed]
- Jash, S.; Sharma, S. Viral Infections and Temporal Programming of Autism Spectrum Disorders in the Mother’s Womb. Front. Virol. 2022, 2, 863202. [Google Scholar] [CrossRef]
- Uribe, F.R.; González, V.P.; Kalergis, A.M.; Soto, J.A.; Bohmwald, K. Understanding the Neurotrophic Virus Mechanisms and Their Potential Effect on Systemic Lupus Erythematosus Development. Brain Sci. 2024, 14, 59. [Google Scholar] [CrossRef]
- Ning, S.; Pagano, J.S.; Barber, G.N. IRF7: Activation, Regulation, Modification and Function. Genes Immun. 2011, 12, 399–414. [Google Scholar] [CrossRef] [PubMed]
| Antibodies | Dilution | Manufacturer, Cat. № | |
|---|---|---|---|
| Pluripotency Markers | Rabbit monoclonal anti-OCT4 | 1:100 | Abcam, Cambridge, UK #ab19857 |
| Rabbit monoclonal anti-SOX2 | 1:200 | Abclonal, Woburn, MA, USA #A11501 | |
| Mouse monoclonal anti-TRA 1-60 | 1:100 | StemCell Techn., Vancouver, BC, Canada, #60064 | |
| Differentiation Markers | Mouse Monoclonal Anti-TUBB3 | 1:200 | Abclonal, Woburn, MA, USA A18132 |
| Rabbit Monoclonal Anti-ACTA2 | 1:200 | Abclonal, Woburn, MA, USA A17910 | |
| Rabbit Polyclonal anti-AFP | 1:200 | Huabio, Woburn, MA, USA, ER80601 | |
| Secondary antibodies | Goat anti-Rabbit IgG, Alexa Fluor 568 | 1:500 | Thermo Fisher Scientific, Waltham, MA, USA #A-11011 |
| Rabbit anti-Mouse IgG, Alexa Fluor 568 | 1:500 | Thermo Fisher Scientific, Waltham, MA, USA #A-11061 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheveleva, O.; Butorina, N.; Protasova, E.; Medvedev, S.; Grigor’eva, E.; Melnikova, V.; Kuziaeva, V.; Minzhenkova, M.; Tatarenko, Y.; Lyadova, I. The Generation of iPSCs Expressing Interferon-Beta Under Doxycycline-Inducible Control. Int. J. Mol. Sci. 2025, 26, 8376. https://doi.org/10.3390/ijms26178376
Sheveleva O, Butorina N, Protasova E, Medvedev S, Grigor’eva E, Melnikova V, Kuziaeva V, Minzhenkova M, Tatarenko Y, Lyadova I. The Generation of iPSCs Expressing Interferon-Beta Under Doxycycline-Inducible Control. International Journal of Molecular Sciences. 2025; 26(17):8376. https://doi.org/10.3390/ijms26178376
Chicago/Turabian StyleSheveleva, Olga, Nina Butorina, Elena Protasova, Sergey Medvedev, Elena Grigor’eva, Victoria Melnikova, Valeriia Kuziaeva, Marina Minzhenkova, Yana Tatarenko, and Irina Lyadova. 2025. "The Generation of iPSCs Expressing Interferon-Beta Under Doxycycline-Inducible Control" International Journal of Molecular Sciences 26, no. 17: 8376. https://doi.org/10.3390/ijms26178376
APA StyleSheveleva, O., Butorina, N., Protasova, E., Medvedev, S., Grigor’eva, E., Melnikova, V., Kuziaeva, V., Minzhenkova, M., Tatarenko, Y., & Lyadova, I. (2025). The Generation of iPSCs Expressing Interferon-Beta Under Doxycycline-Inducible Control. International Journal of Molecular Sciences, 26(17), 8376. https://doi.org/10.3390/ijms26178376






