The Prognostic Value of CIP2A and Its Association with CD31, E-Cadherin, and pAMPK in Lung Cancer
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| Bcl-2 | B-cell lymphoma 2 |
| CD31 | cluster of differentiation 31 |
| CIP2A | cancerous inhibitor of PP2A |
| EGFR | epidermal growth factor receptor |
| E-cadherin | epithelial cadherin |
| LDCT | low-dose computed tomography |
| N-cadherin | neural cadherin |
| NSCLC | non-small-cell lung cancer |
| OR | odds ratio |
| pAkt | phosphorylated protein kinase B |
| pAMPK | phosphorylated AMP-activated protein kinase |
| pErk | phosphorylated extracellular signal-regulated kinase |
| pStat3 | phosphorylated signal transducer and activator of transcription 3 |
| PKM2 | pyruvate kinase M2 |
| PP2A | protein phosphatase 2A |
| TIMP1 | tissue inhibitor of metalloproteinases 1 |
| TP53 | tumor protein p53 |
| UTR | untranslated region |
References
- Nooreldeen, R.; Bach, H. Current and Future Development in Lung Cancer Diagnosis. Int. J. Mol. Sci. 2021, 22, 8661. [Google Scholar] [CrossRef]
- Smolarz, B.; Lukasiewicz, H.; Samulak, D.; Piekarska, E.; Kolacinski, R.; Romanowicz, H. Lung Cancer-Epidemiology, Pathogenesis, Treatment and Molecular Aspect (Review of Literature). Int. J. Mol. Sci. 2025, 26, 2049. [Google Scholar] [CrossRef]
- Oliver, A.L. Lung Cancer: Epidemiology and Screening. Surg. Clin. N. Am. 2022, 102, 335–344. [Google Scholar] [CrossRef]
- Kuo, C.N.; Liao, Y.M.; Kuo, L.N.; Tsai, H.J.; Chang, W.C.; Yen, Y. Cancers in Taiwan: Practical insight from epidemiology, treatments, biomarkers, and cost. J. Formos. Med. Assoc. 2020, 119, 1731–1741. [Google Scholar] [CrossRef]
- Wang, Y.T.; Hu, K.R.; Zhao, J.; Ai, F.L.; Shi, Y.L.; Wang, X.W.; Yang, W.Y.; Wang, J.X.; Ai, L.M.; Wan, X. The Association between Exposure to Second-Hand Smoke and Disease in the Chinese Population: A Systematic Review and Meta-Analysis. Biomed. Environ. Sci. 2023, 36, 24–37. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Mahendran, R.; Yu, P.; Xu, R.; Yu, W.; Godellawattage, S.; Li, S.; Guo, Y. Health Effects of Long-Term Exposure to Ambient PM(2.5) in Asia-Pacific: A Systematic Review of Cohort Studies. Curr. Environ. Health Rep. 2022, 9, 130–151. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.H.; Liang, K.H.; Huang, H.C.; Shen, C.I.; Chiang, C.L.; Wang, M.L.; Chiou, S.H.; Chen, Y.M. State-of-the-Art Molecular Oncology of Lung Cancer in Taiwan. Int. J. Mol. Sci. 2022, 23, 7037. [Google Scholar] [CrossRef] [PubMed]
- Sodeifian, F.; Kian, N.; Atefi, A.; Naserghandi, A.; Zangiabadian, M.; Sadeghzade, S.; Namakin, K.; Seghatoleslami, Z.S.; D’Ambrosio, L.; Nasiri, M.J.; et al. Pulmonary Tuberculosis and Risk of Lung Cancer: A Systematic Review and Meta-Analysis. Respiration 2025, 104, 360–376. [Google Scholar] [CrossRef]
- Luo, Y.H.; Chiu, C.H.; Scott Kuo, C.H.; Chou, T.Y.; Yeh, Y.C.; Hsu, H.S.; Yen, S.H.; Wu, Y.H.; Yang, J.C.; Liao, B.C.; et al. Lung Cancer in Republic of China. J. Thorac. Oncol. 2021, 16, 519–527. [Google Scholar] [CrossRef]
- Lam, D.C.; Liam, C.K.; Andarini, S.; Park, S.; Tan, D.S.W.; Singh, N.; Jang, S.H.; Vardhanabhuti, V.; Ramos, A.B.; Nakayama, T.; et al. Lung Cancer Screening in Asia: An Expert Consensus Report. J. Thorac. Oncol. 2023, 18, 1303–1322. [Google Scholar] [CrossRef]
- Oezkan, F.; Seweryn, M.; Shukuya, T.; Owen, D.H. Improvement in Survival for Patients With Lung Cancer in Taiwan: Implications and Call to Action. J. Thorac. Oncol. 2023, 18, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Ho, P.J.; Tan, I.B.; Chong, D.Q.; Khor, C.C.; Yuan, J.M.; Koh, W.P.; Dorajoo, R.; Li, J. Polygenic risk scores for the prediction of common cancers in East Asians: A population-based prospective cohort study. eLife 2023, 12, e82608. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, R. Breakthrough Biomarkers in Lung Cancer: Pioneering Early Detection and Precision Treatment Strategies. Zhongguo Ying Yong Sheng Li Xue Za Zhi 2024, 40, e20240034. [Google Scholar] [CrossRef] [PubMed]
- Dantas, E.; Murthy, A.; Ahmed, T.; Ahmed, M.; Ramsamooj, S.; Hurd, M.A.; Lam, T.; Malbari, M.; Agrusa, C.; Elemento, O.; et al. TIMP1 is an early biomarker for detection and prognosis of lung cancer. Clin. Transl. Med. 2023, 13, e1391. [Google Scholar] [CrossRef]
- Pu, J.; Ai, T.; Weng, W.; Wang, L.; Yang, Y.; Ma, L.; Hu, Z.; Meng, X. TJP1, a Membrane-Expressed Protein, is a Potential Therapeutic and Prognostic Target for Lung Cancer. Technol. Cancer Res. Treat. 2022, 21, 15330338221106855. [Google Scholar] [CrossRef]
- Lin, X.; Lin, K.; Lin, C.; Liu, T.; Ba, M.; Tang, Y.; Wang, J.; Zhou, L.; Wang, J.; Xiao, C. Prognostic and clinicopathological value of PD-L2 in lung cancer: A meta-analysis. Int. Immunopharmacol. 2021, 91, 107280. [Google Scholar] [CrossRef]
- Kadara, H.; Behrens, C.; Yuan, P.; Solis, L.; Liu, D.; Gu, X.; Minna, J.D.; Lee, J.J.; Kim, E.; Hong, W.K.; et al. A five-gene and corresponding protein signature for stage-I lung adenocarcinoma prognosis. Clin. Cancer Res. 2011, 17, 1490–1501. [Google Scholar] [CrossRef]
- Ferrigno, D.; Buccheri, G.; Camilla, T. Prognosis and lung-cancer—The contribution of plasma-proteins. Oncol. Rep. 1995, 2, 637–641. [Google Scholar] [CrossRef]
- Lu, Y.F.; Chang, Y.H.; Chen, Y.J.; Hsieh, M.S.; Lin, M.W.; Hsu, H.H.; Han, C.L.; Chen, Y.J.; Yu, S.L.; Chen, J.S.; et al. Proteomic profiling of tumor microenvironment and prognosis risk prediction in stage I lung adenocarcinoma. Lung Cancer 2024, 191, 107791. [Google Scholar] [CrossRef]
- Chen, B.; Hu, H.; Chen, X. From Basic Science to Clinical Practice: The Role of Cancerous Inhibitor of Protein Phosphatase 2A (CIP2A)/p90 in Cancer. Front. Genet. 2023, 14, 1110656. [Google Scholar] [CrossRef]
- Nagelli, S.; Westermarck, J. CIP2A coordinates phosphosignaling, mitosis, and the DNA damage response. Trends Cancer 2024, 10, 52–64. [Google Scholar] [CrossRef]
- Liang, L.J.; Yang, F.Y.; Wang, D.; Zhang, Y.F.; Yu, H.; Wang, Z.; Sun, B.B.; Liu, Y.T.; Wang, G.Z.; Zhou, G.B. CIP2A induces PKM2 tetramer formation and oxidative phosphorylation in non-small cell lung cancer. Cell Discov. 2024, 10, 13. [Google Scholar] [CrossRef] [PubMed]
- Tarek, M.M.; Yahia, A.; El-Nakib, M.M.; Elhefnawi, M. Integrative assessment of CIP2A overexpression and mutational effects in human malignancies identifies possible deleterious variants. Comput. Biol. Med. 2021, 139, 104986. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Fan, S.; Wang, J.; Liang, Y.; Li, P.; Lv, X.; Sun, Y.; Wang, Q.; Liu, H.; Zhang, C.; et al. Cip2a induces arginine biosynthesis and promotes tumor progression in non-small cell lung cancer. Mol. Carcinog. 2023, 62, 561–572. [Google Scholar] [CrossRef]
- Wei, L.; Qu, W.; Sun, J.; Wang, X.; Lv, L.; Xie, L.; Song, X. Knockdown of cancerous inhibitor of protein phosphatase 2A may sensitize NSCLC cells to cisplatin. Cancer Gene Ther. 2014, 21, 194–199. [Google Scholar] [CrossRef]
- Wang, C.Y.; Chao, T.T.; Chang, F.Y.; Chen, Y.L.; Tsai, Y.T.; Lin, H.I.; Huang, Y.C.; Shiau, C.W.; Yu, C.J.; Chen, K.F. CIP2A mediates erlotinib-induced apoptosis in non-small cell lung cancer cells without EGFR mutation. Lung Cancer 2014, 85, 152–160. [Google Scholar] [CrossRef]
- Saafan, H.; Alahdab, A.; Michelet, R.; Gohlke, L.; Ziemann, J.; Holdenrieder, S.; McLaughlin, K.M.; Wass, M.N.; Cinatl, J., Jr.; Michaelis, M.; et al. Constitutive Cell Proliferation Regulating Inhibitor of Protein Phosphatase 2A (CIP2A) Mediates Drug Resistance to Erlotinib in an EGFR Activating Mutated NSCLC Cell Line. Cells 2021, 10, 716. [Google Scholar] [CrossRef]
- Cao, S.; Wang, Y.; Li, J.; Ling, X.; Zhang, Y.; Zhou, Y.; Zhong, H. Prognostic Implication of the Expression Level of PECAM-1 in Non-small Cell Lung Cancer. Front. Oncol. 2021, 11, 587744. [Google Scholar] [CrossRef]
- Cavazzoni, A.; Digiacomo, G.; Volta, F.; Alfieri, R.; Giovannetti, E.; Gnetti, L.; Bellini, L.; Galetti, M.; Fumarola, C.; Xu, G.; et al. PD-L1 overexpression induces STAT signaling and promotes the secretion of pro-angiogenic cytokines in non-small cell lung cancer (NSCLC). Lung Cancer 2024, 187, 107438. [Google Scholar] [CrossRef]
- Zhong, Z.T.; Wang, X.Y.; Pan, Y.; Zhou, K.; Chen, J.H.; Gao, Y.Q.; Dai, B.; Zhou, Z.L.; Wang, R.Q. AMPK: An energy sensor for non-small cell lung cancer progression and treatment. Pharmacol. Res. 2025, 212, 107592. [Google Scholar] [CrossRef]
- Che, J.; Wang, J.; Li, H.; Zhen, H.; Shang, K.; Yang, Y.; Cao, B. Decreased expression of Dlg5 is associated with a poor prognosis and epithelial-mesenchymal transition in squamous cell lung cancer. J. Thorac. Dis. 2021, 13, 3115–3125. [Google Scholar] [CrossRef]
- Hu, J.; Wang, L.; Guan, C. MiR-532-5p Suppresses Migration and Invasion of Lung Cancer Cells Through Inhibiting CCR4. Cancer Biother. Radiopharm. 2020, 35, 673–681. [Google Scholar] [CrossRef]
- Rezaei, M.; Mostafaei, S.; Aghaei, A.; Hosseini, N.; Darabi, H.; Nouri, M.; Etemadi, A.; Neill, A.O.; Nahand, J.S.; Mirzaei, H.; et al. The association between HPV gene expression, inflammatory agents and cellular genes involved in EMT in lung cancer tissue. BMC Cancer 2020, 20, 916. [Google Scholar] [CrossRef]
- Ji, S.; Guo, Y.; Ding, J.; Hong, W.; Yan, Z.; Cai, Z.; Yue, H.; Qiu, X.; Sang, N. Nontargeted Identification of Organic Components in Fine Particulate Matter Related to Lung Tumor Metastasis Based on an Adverse Outcome Pathway Strategy. Environ. Sci. Technol. 2024, 58, 4083–4091. [Google Scholar] [CrossRef] [PubMed]
- Luque, M.; Cristobal, I.; Sanz-Alvarez, M.; Santos, A.; Zazo, S.; Eroles, P.; Arpi, O.; Rovira, A.; Albanell, J.; Madoz-Gurpide, J.; et al. CIP2A as a Key Regulator for AKT Phosphorylation Has Partial Impact Determining Clinical Outcome in Breast Cancer. J. Clin. Med. 2022, 11, 1610. [Google Scholar] [CrossRef] [PubMed]
- Andriani, L.; Ling, Y.X.; Yang, S.Y.; Zhao, Q.; Ma, X.Y.; Huang, M.Y.; Zhang, Y.L.; Zhang, F.L.; Li, D.Q.; Shao, Z.M. Sideroflexin-1 promotes progression and sensitivity to lapatinib in triple-negative breast cancer by inhibiting TOLLIP-mediated autophagic degradation of CIP2A. Cancer Lett. 2024, 597, 217008. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Xu, C.; Peng, L.; Ma, L.; Du, M. cAMP/PKA signaling promotes AKT deactivation by reducing CIP2A expression, thereby facilitating decidualization. Mol. Cell. Endocrinol. 2023, 571, 111946. [Google Scholar] [CrossRef]
- Chen, L.; Xu, H.; Liu, R.; Yao, Z.; Xie, Q.; Zhang, X. Circular RNA Vav3 mediated ALV-J inhibition of autophagy by modulating the gga-miR-375/CIP2A axis and activating AKT. Poult. Sci. 2025, 104, 104923. [Google Scholar] [CrossRef]
- Fang, D.; Ou, X.; Sun, K.; Zhou, X.; Li, Y.; Shi, P.; Zhao, Z.; He, Y.; Peng, J.; Xu, J. m6A modification-mediated lncRNA TP53TG1 inhibits gastric cancer progression by regulating CIP2A stability. Cancer Sci. 2022, 113, 4135–4150. [Google Scholar] [CrossRef]
- Gao, D.; Nyalali, A.M.K.; Hou, Y.; Xu, Y.; Zhou, J.; Zhao, W.; Huang, B.; Li, F. 2,5-Dimethyl Celecoxib Inhibits Proliferation and Cell Cycle and Induces Apoptosis in Glioblastoma by Suppressing CIP2A/PP2A/Akt Signaling Axis. J. Mol. Neurosci. 2021, 71, 1703–1713. [Google Scholar] [CrossRef]
- Dai, S.; Wang, C.; Zhao, X.; Ma, C.; Fu, K.; Liu, Y.; Peng, C.; Li, Y. Cucurbitacin B: A review of its pharmacology, toxicity, and pharmacokinetics. Pharmacol. Res. 2023, 187, 106587. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yang, S.; Wang, J.; Jiang, Y. Blockade of AMPK-Mediated cAMP-PKA-CREB/ATF1 Signaling Synergizes with Aspirin to Inhibit Hepatocellular Carcinoma. Cancers 2021, 13, 1738. [Google Scholar] [CrossRef] [PubMed]
- Chao, T.T.; Wang, C.Y.; Lai, C.C.; Chen, Y.L.; Tsai, Y.T.; Chen, P.T.; Lin, H.I.; Huang, Y.C.; Shiau, C.W.; Yu, C.J.; et al. TD-19, an erlotinib derivative, induces epidermal growth factor receptor wild-type nonsmall-cell lung cancer apoptosis through CIP2A-mediated pathway. J. Pharmacol. Exp. Ther. 2014, 351, 352–358, Erratum in J. Pharmacol. Exp. Ther. 2018, 364, 26–27. [Google Scholar] [CrossRef] [PubMed]
- Feng, F.F.; Cheng, P.; Sun, C.; Wang, H.; Wang, W. Inhibitory effects of polyphyllins I and VII on human cisplatin-resistant NSCLC via p53 upregulation and CIP2A/AKT/mTOR signaling axis inhibition. Chin. J. Nat. Med. 2019, 17, 768–777. [Google Scholar] [CrossRef]
- Feng, F.; Cheng, P.; Wang, C.; Wang, Y.; Wang, W. Polyphyllin I and VII potentiate the chemosensitivity of A549/DDP cells to cisplatin by enhancing apoptosis, reversing EMT and suppressing the CIP2A/AKT/mTOR signaling axis. Oncol. Lett. 2019, 18, 5428–5436. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Z.; Wang, F.; Liu, J.; Zhang, Y.; Cao, J.; Huang, G.; Ma, L. ent-8(14),15-Pimaradiene-2beta,19-diol, a diterpene from Aleuritopteris albofusca, inhibits growth and induces protective autophagy in hepatocellular carcinoma cells. Naunyn-Schmiedeberg’s Arch. Pharmacol. 2024, 397, 6867–6878. [Google Scholar] [CrossRef]
- Zhao, S.; Gao, X.; Zang, S.; Li, Y.; Feng, X.; Yuan, X. MicroRNA-383-5p acts as a prognostic marker and inhibitor of cell proliferation in lung adenocarcinoma by cancerous inhibitor of protein phosphatase 2A. Oncol. Lett. 2017, 14, 3573–3579. [Google Scholar] [CrossRef]
- Chao, T.T.; Wang, C.Y.; Chen, Y.L.; Lai, C.C.; Chang, F.Y.; Tsai, Y.T.; Chao, C.H.; Shiau, C.W.; Huang, Y.C.; Yu, C.J.; et al. Afatinib induces apoptosis in NSCLC without EGFR mutation through Elk-1-mediated suppression of CIP2A. Oncotarget 2015, 6, 2164–2179. [Google Scholar] [CrossRef]
- Hsu, C.C.; Peng, D.; Cai, Z.; Lin, H.K. AMPK signaling and its targeting in cancer progression and treatment. Semin. Cancer Biol. 2022, 85, 52–68. [Google Scholar] [CrossRef]
- Sahu, M.; Paliwal, T.; Jain, S.; Verma, K.; Chakraborty, D.; Jaiswal, S.; Dwivedi, J.; Sharma, S. Multifaceted Therapeutic Impacts of Cucurbitacin B: Recent Evidences From Preclinical Studies. Phytother. Res. 2025, 39, 1966–1995. [Google Scholar] [CrossRef]
- Ashrafizadeh, M.; Mirzaei, S.; Hushmandi, K.; Rahmanian, V.; Zabolian, A.; Raei, M.; Farahani, M.V.; Goharrizi, M.; Khan, H.; Zarrabi, A.; et al. Therapeutic potential of AMPK signaling targeting in lung cancer: Advances, challenges and future prospects. Life Sci. 2021, 278, 119649. [Google Scholar] [CrossRef]
- Yuan, J.; Dong, X.; Yap, J.; Hu, J. The MAPK and AMPK signalings: Interplay and implication in targeted cancer therapy. J. Hematol. Oncol. 2020, 13, 113. [Google Scholar] [CrossRef]
- Keerthana, C.K.; Rayginia, T.P.; Shifana, S.C.; Anto, N.P.; Kalimuthu, K.; Isakov, N.; Anto, R.J. The role of AMPK in cancer metabolism and its impact on the immunomodulation of the tumor microenvironment. Front. Immunol. 2023, 14, 1114582. [Google Scholar] [CrossRef]
- Steinberg, G.R.; Hardie, D.G. New insights into activation and function of the AMPK. Nat. Rev. Mol. Cell Biol. 2023, 24, 255–272. [Google Scholar] [CrossRef]
- Cheng, C.Y.; Chen, Y.L.; Ho, H.; Huang, C.Y.; Chu, S.E.; Liang, Y.J. Prognostic Significance of DNAJB4 Expression in Gastric Cancer: Correlation with CD31, Caspase-3, and Tumor Progression. Diagnostics 2025, 15, 652. [Google Scholar] [CrossRef]
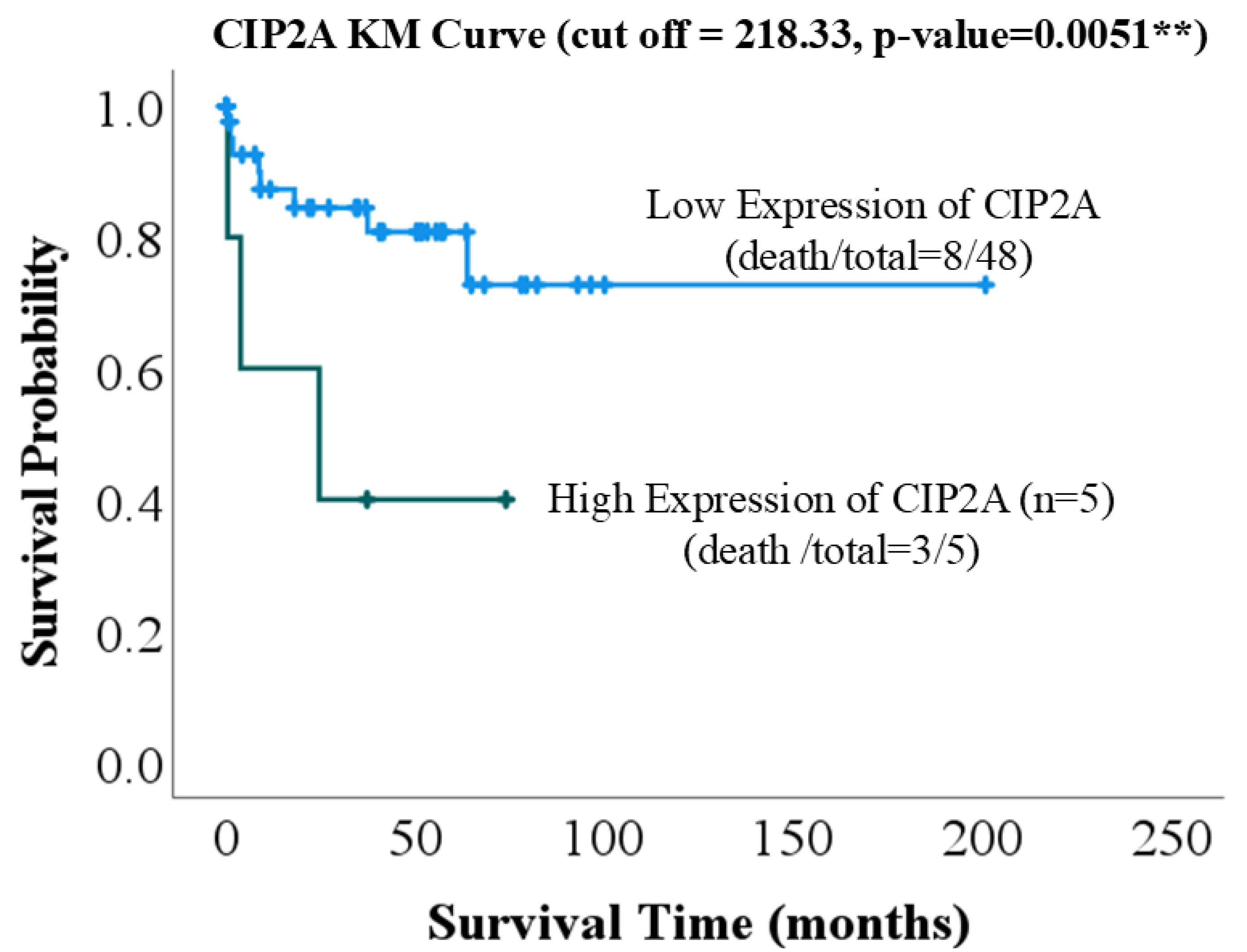
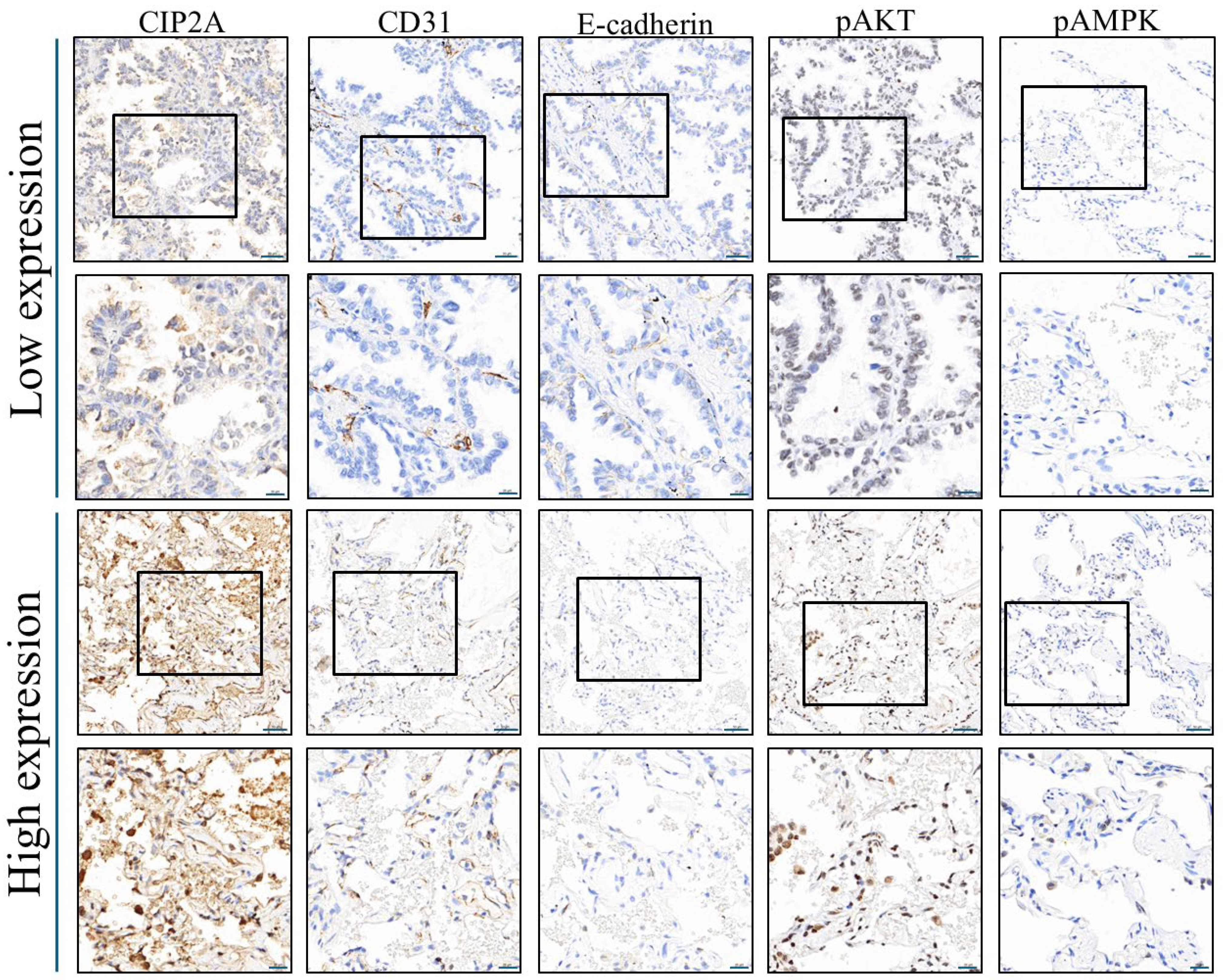
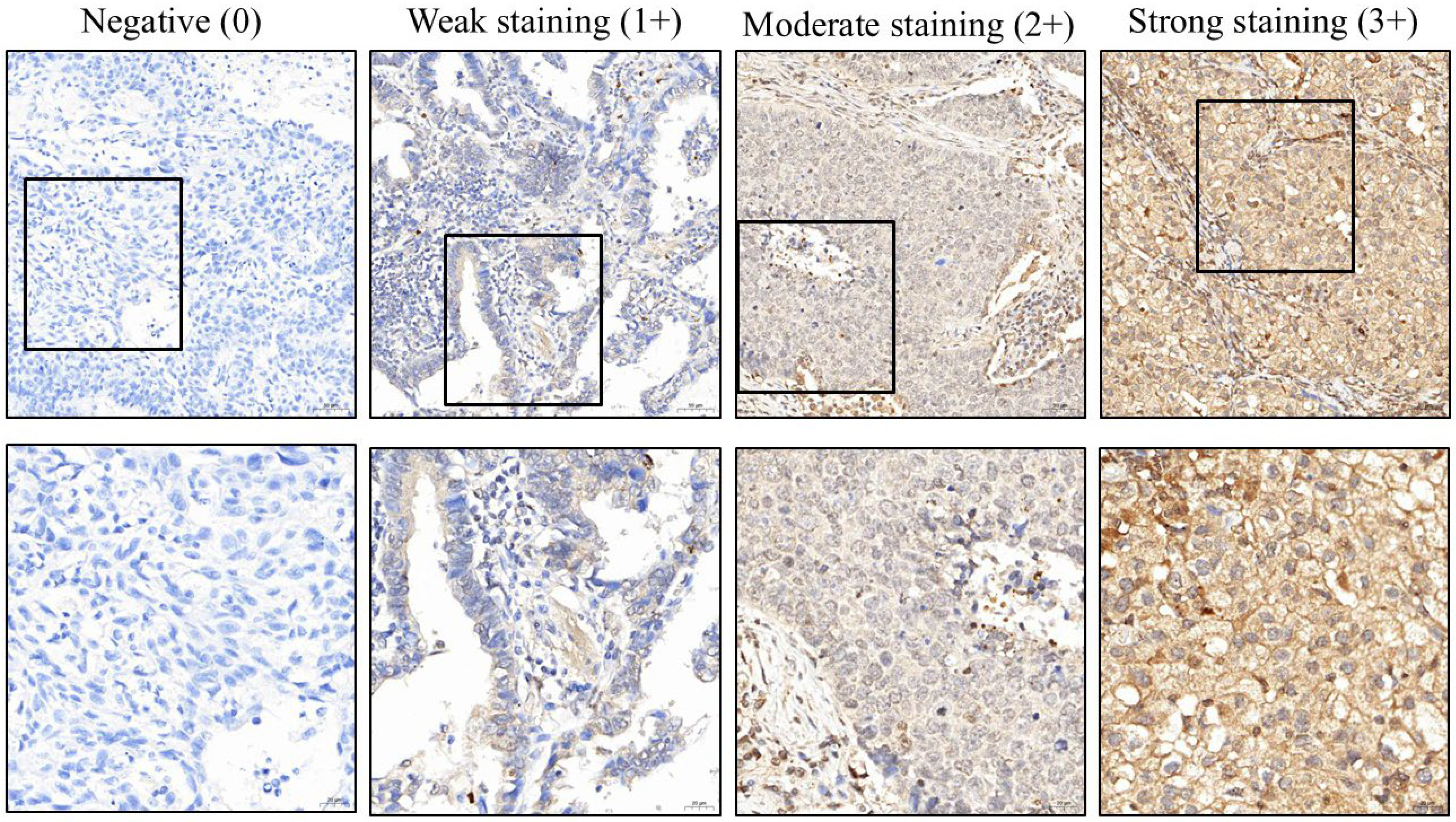
| Low Expression of CIP2A | High Expression of CIP2A | Total | p-Value | |
|---|---|---|---|---|
| Sex | 0.913 | |||
| Female | 30 (62.5%) | 3 (60.0%) | 33 (62.3%) | |
| Male | 18 (37.5%) | 2 (40.0%) | 20 (37.7%) | |
| Age | ||||
| Grading | 0.627 | |||
| Well | 5 (10.4%) | 0 (0.0%) | 5 (9.4%) | |
| Moderate | 29 (60.4%) | 4 (80.0%) | 33 (62.3%) | |
| Low | 14 (29.2%) | 1 (20.0%) | 15 (28.3%) | |
| Size | ||||
| Stage | 0.485 | |||
| I and II | 37 (77.1%) | 5 (100.0%) | 42 (79.2%) | |
| IIIa | 10 (20.8%) | 0 (0.0%) | 10 (18.9%) | |
| IIIb and IV | 1 (2.1%) | 0 (0.0%) | 1 (1.9%) |
| Low Expression of CIP2A | High Expression of CIP2A | Total | p-Value | |
|---|---|---|---|---|
| Caspase3 | 4.6 (3.8, 7.9) | 4.5 (3.4, 14.4) | 4.5 (3.8, 7.9) | 1.000 |
| Ki67 | 3.8 (1.1, 13.7) | 2.5 (0.2, 4.6) | 3.3 (1.1, 12.7) | 0.192 |
| CD31 | 23.2 (11.4, 45.6) | 111.0 (43.0, 153.6) | 29.3 (11.9, 54.6) | 0.004 ** |
| E-cad | 102.9 (100.7, 111.9) | 110.8 (104.3, 134.6) | 203.4 (101.1, 112.0) | 0.060 |
| N-cad | 4.4 (2.7, 8.3) | 2.7 (2.0, 3.9) | 4.26 (2.6, 8.1) | 0.069 |
| Fibronectin | 214.7 (158.3, 244.2) | 186.4 (97.8, 239.6) | 209.3 (154.2, 243.9) | 0.432 |
| pAkt | 46.6 (17.2, 95.8) | 116.4 (45.5, 159.6) | 50.2 (17.5, 102.2) | 0.181 |
| pErk | 3.5 (0.7, 22.0) | 3.2 (1.3, 92.2) | 3.2 (0.9, 21.6) | 0.605 |
| pStat3 | 1.6 (0.2, 11.5) | 3.3 (1.9, 8.9) | 1.9 (0.4, 11.1) | 0.468 |
| pAMPK | 8.3 (5.7, 11.0) | 3.8 (3.5, 7.3) | 8.2 (5.1, 10.4) | 0.018 * |
| Univariable | p-Value | |
|---|---|---|
| Sex | ||
| Female | Reference | |
| Male | 1.24 (0.33–4.60) | 0.750 |
| Age | 0.99 (0.94–1.05) | 0.792 |
| Grading | ||
| Well | Reference | |
| Moderate | 0.89 (0.08–9.44) | 0.922 |
| Low | 2.00 (0.17–22.95) | 0.578 |
| Size | 1.13 (0.84–1.51) | 0.428 |
| Stage | ||
| I and II | Reference | |
| IIIa | 0.31 (0.04–2.76) | 0.296 |
| IIIb and IV | 0.00 (0.00–) | 1.000 |
| Marker | ||
| Caspase3 | 1.03 (0.97–1.08) | 0.346 |
| Ki67 | 1.00 (0.99–1.02) | 0.652 |
| CD31 | 0.99 (0.97–1.01) | 0.274 |
| E-cad | 1.00 (0.93–1.08) | 0.997 |
| N-cad | 1.01 (0.98–1.04) | 0.538 |
| Fibronectin | 1.00 (0.99–1.01) | 0.748 |
| pAkt | 1.00 (0.99–1.01) | 0.956 |
| pErk | 1.00 (0.99–1.01) | 0.793 |
| pStat3 | 0.96 (0.87–1.05) | 0.356 |
| pAMPK | 0.99 (0.88–1.12) | 0.870 |
| Marker | Pearson Correlation | p-Value |
|---|---|---|
| Caspase3 | 0.104 | 0.461 |
| Ki67 | −0.017 | 0.902 |
| CD31 | 0.420 | 0.002 ** |
| E-cad | 0.372 | 0.006 ** |
| N-cad | −0.118 | 0.400 |
| Fibronectin | 0.155 | 0.268 |
| pAkt | 0.332 | 0.015 * |
| pErk | 0.165 | 0.236 |
| pStat3 | 0.116 | 0.407 |
| pAMPK | −0.474 | <0.001 *** |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, P.Y.; Shih, C.-Y.; Cheng, C.-Y.; Ho, H.; Chen, Y.-L.; Chang, C.-J. The Prognostic Value of CIP2A and Its Association with CD31, E-Cadherin, and pAMPK in Lung Cancer. Int. J. Mol. Sci. 2025, 26, 8362. https://doi.org/10.3390/ijms26178362
Lee PY, Shih C-Y, Cheng C-Y, Ho H, Chen Y-L, Chang C-J. The Prognostic Value of CIP2A and Its Association with CD31, E-Cadherin, and pAMPK in Lung Cancer. International Journal of Molecular Sciences. 2025; 26(17):8362. https://doi.org/10.3390/ijms26178362
Chicago/Turabian StyleLee, Peng Yu, Ching-Yu Shih, Chiao-Yin Cheng, Hua Ho, Yen-Lin Chen, and Chih-Jung Chang. 2025. "The Prognostic Value of CIP2A and Its Association with CD31, E-Cadherin, and pAMPK in Lung Cancer" International Journal of Molecular Sciences 26, no. 17: 8362. https://doi.org/10.3390/ijms26178362
APA StyleLee, P. Y., Shih, C.-Y., Cheng, C.-Y., Ho, H., Chen, Y.-L., & Chang, C.-J. (2025). The Prognostic Value of CIP2A and Its Association with CD31, E-Cadherin, and pAMPK in Lung Cancer. International Journal of Molecular Sciences, 26(17), 8362. https://doi.org/10.3390/ijms26178362






