Testing Strategies for Metabolite-Mediated Neurotoxicity
Abstract
1. Introduction
2. Results and Discussion
2.1. Parent/Metabolite Pairs to Explore Metabolite-Mediated Toxicity
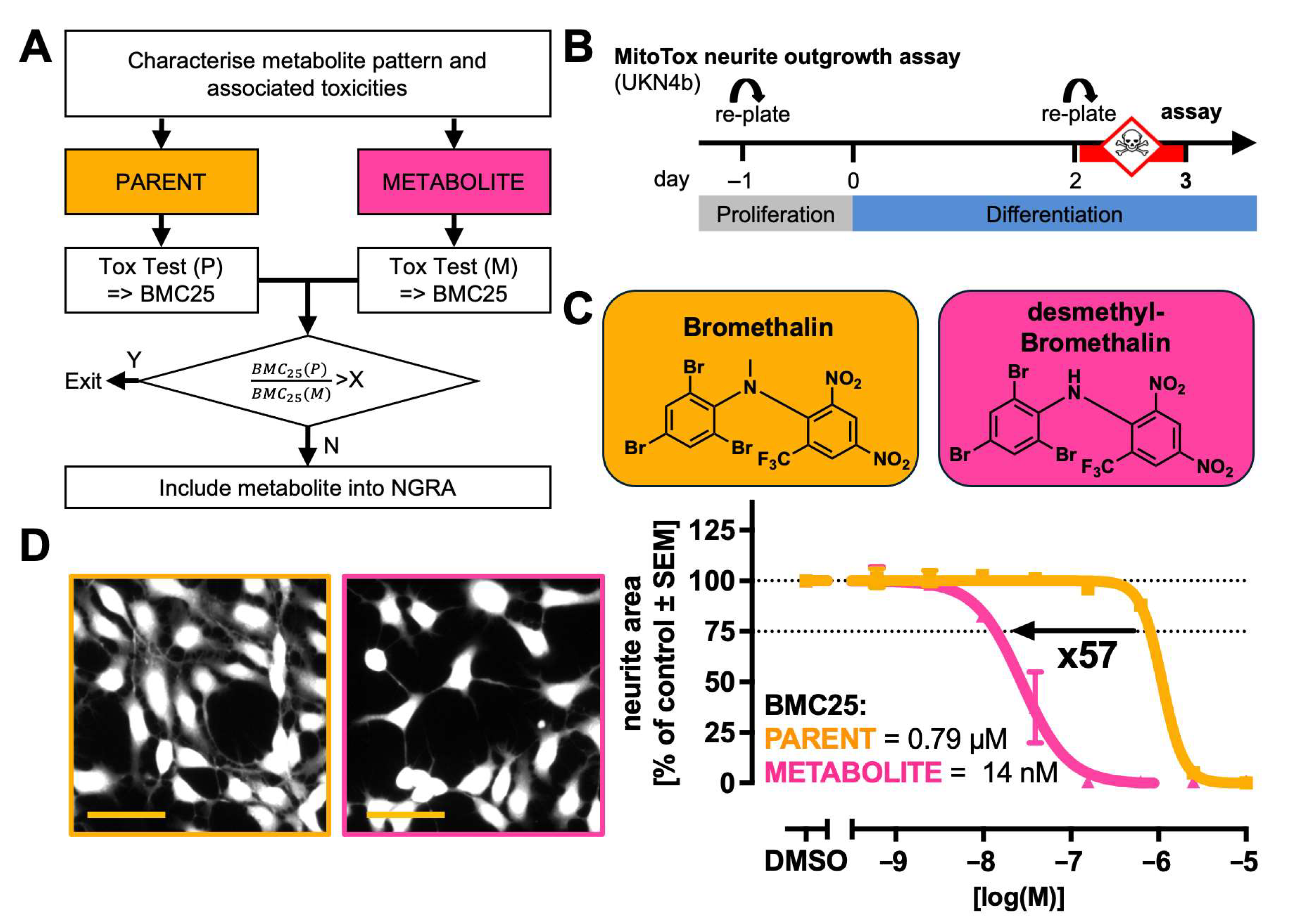
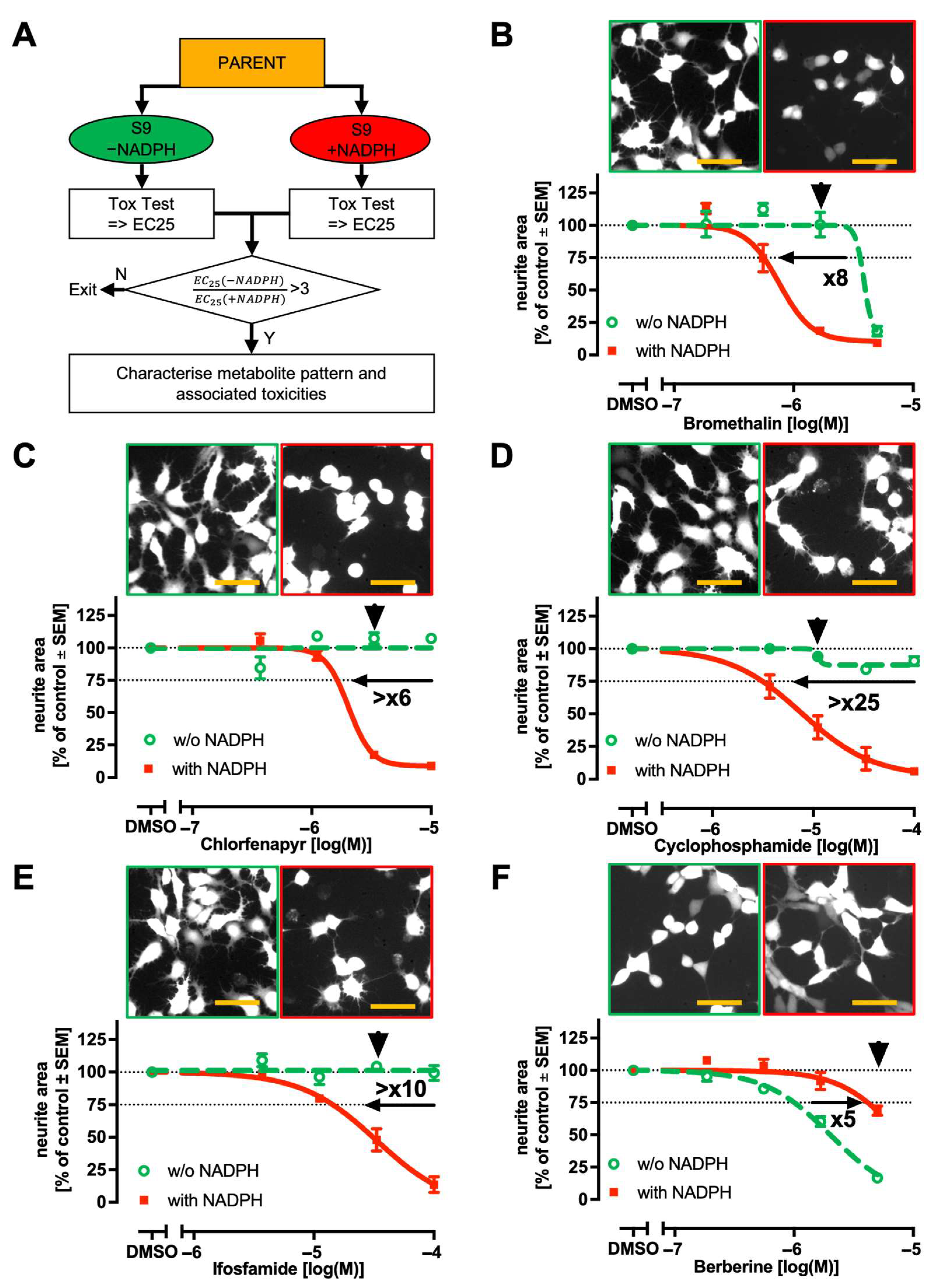
2.2. Strategy to Compare Potency of Metabolites to That of Their Cognate Parents
| Parent/CAS MW in g/mol | Comment | Free Fraction [%] | Metabolite of Concern CAS/MW in g/mol | Free fraction [%] | Literature Ref. | ||
|---|---|---|---|---|---|---|---|
| Plasma | Medium | Plasma | Medium | ||||
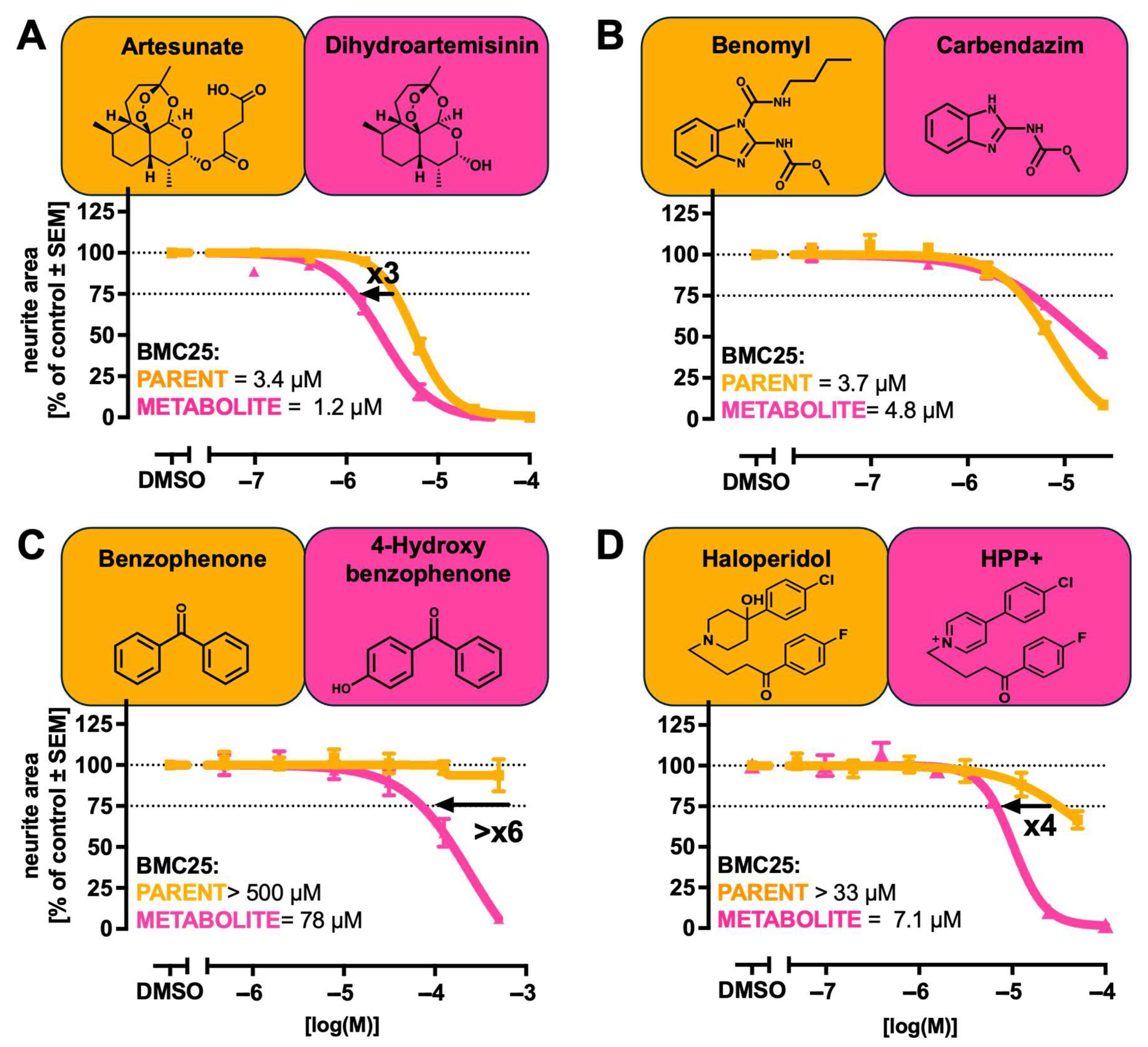
| Artesunate CAS: 88495-63-0 MW: 384.4 | Prodrug (human), which induces oxidative stress | 3.1 | 98.2 | Dihydroartemisinin CAS: 71939-50-9 MW: 284.4 | 14.9 | 99.7 | [54,55] |
| Benomyl CAS: 17804-35-2 MW: 290.3 | Pesticide; cytoskeletal toxicant | 14.1 | 99.6 | Carbendazim CAS: 10605-21-7 MW: 191.2 | 75.0 | 100.0 | [3,56,57] |
| Benzophenone CAS: 119-61-9 MW: 182.2 | Preservative, metabolite is mitochondrial toxicant | 93.0 | 99.4 | 4-Hydroxybenzophenone CAS: 1137-42-4 MW: 198.2 | 6.8 | 99.2 | [58,59,60] |
| Bromethalin CAS: 63333-35-7 MW: 577.9 | Pro-pesticide, metabolite is mitochondrial toxicant | 0.3 | 83.2 | Desmethyl-bromethalin CAS: 57729-86-9 MW: 563.9 | 0.1 | 62.6 | [61,62] |
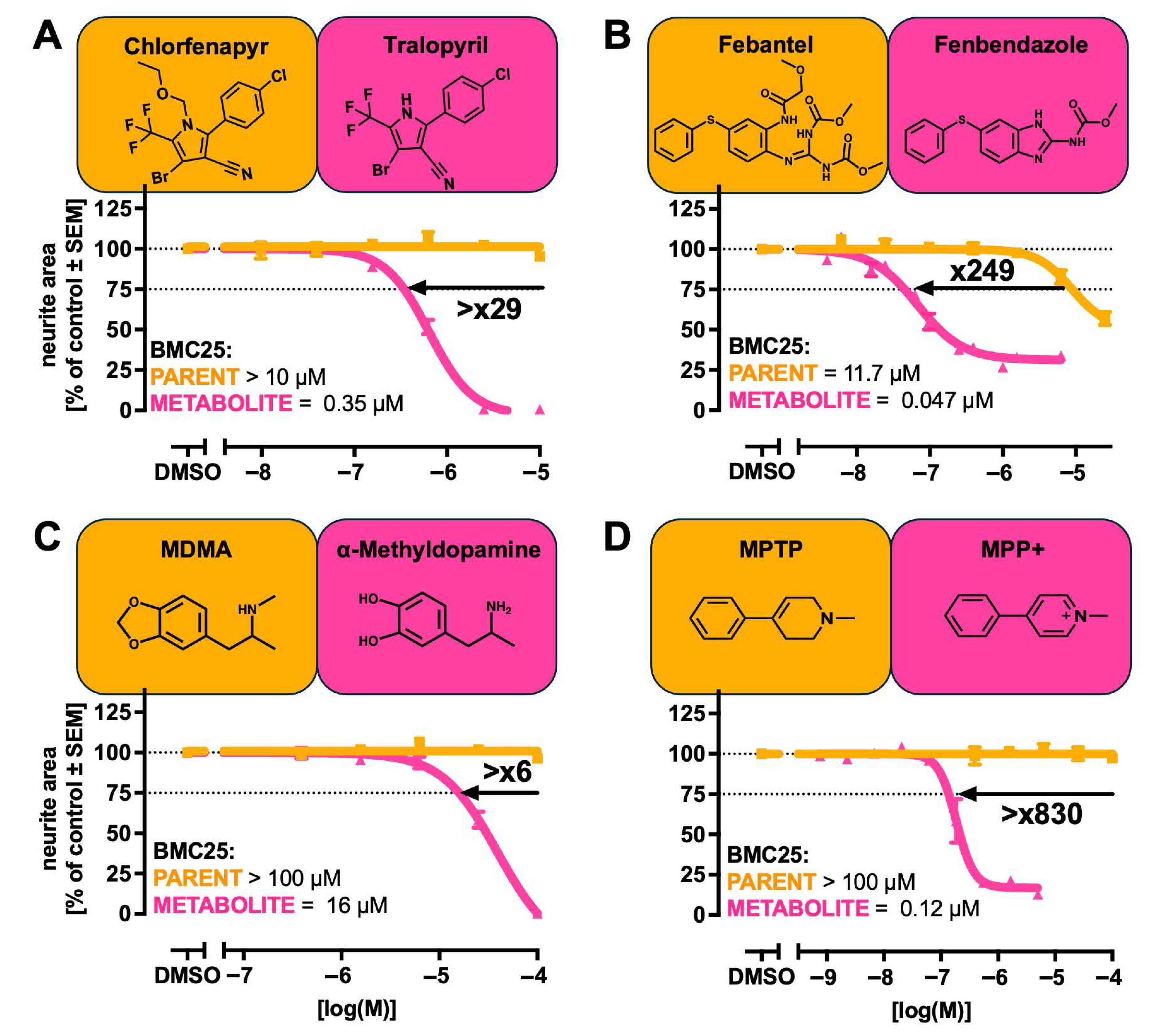
| Chlorfenapyr CAS: 122453-73-0 MW: 407.6 | Pro-pesticide, metabolite is mitochondrial toxicant | 1.6 | 96.4 | Tralopyril CAS: 122454-29-9 MW: 384.4 | 3.5 | 98.4 | [61,63] |
| Febantel CAS: 58306-30-2 MW: 446.5 | Prodrug (veterinary); which is used as cytoskeletal toxicant. | 6.0 | 99.1 | Fenbendazole CAS: 43210-67-9 MW: 299.3 | 75.0 | 100.0 | [64,65] |
| Haloperidol CAS: 52-86-8 MW: 375.9 | Drug (human), metabolite is reported neurotoxicant | 24.5 | 99.8 | HPP+ CAS: 125785-69-5 MW: 354.8 | 52.7 | 99.9 | [66,67] |
| MDMA CAS: 64057-70-1 MW: 193.3 | Recreational drug, metabolite reported as neurotoxicant | 61.7 | 100.0 | α-Methyldopamine CAS: 555-64-6 MW: 167.2 | 88.8 | 100.0 | [68] |
| MPTP CAS: 28289-54-5 MW: 173.3 | Recreational drug impurity, metabolite reported as neurotoxicant | 28.6 | 99.9 | MPP+ CAS: 48134-75-4 MW: 170.3 | 91.1 | 100.0 | [25,69] |
2.3. Comparison of Parent and Metabolite Potencies in Exemplary Assays
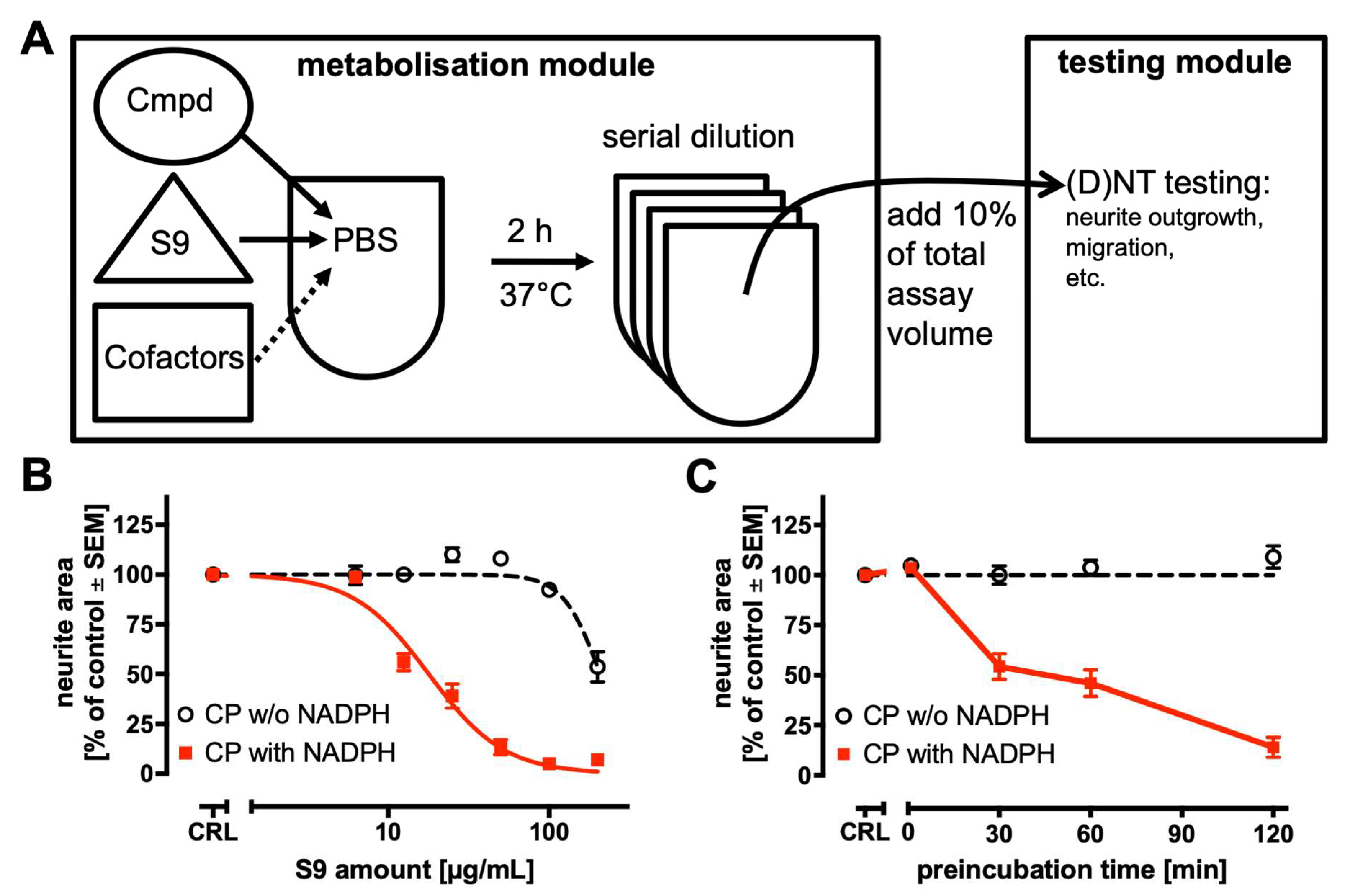
2.4. Strategy to Incorporate Metabolic Competence into In Vitro DNT Testing Methods
2.5. Effect of NADPH-Dependent Metabolism on Neurotoxicity of Selected Compounds
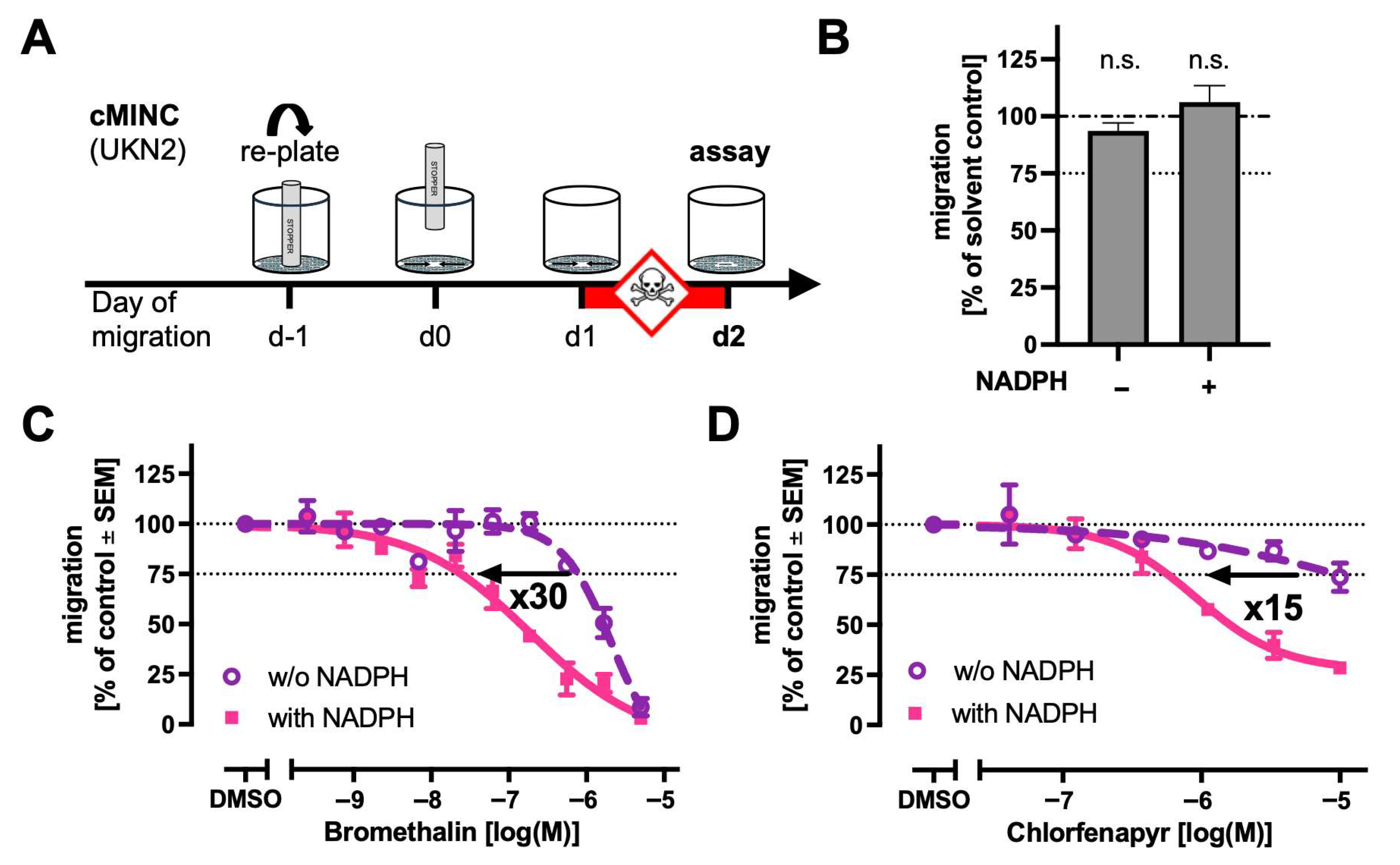
2.6. Transfer of Metabolisation Module to Neural Crest Cell Migration Assay (cMINC)
3. Materials and Methods
3.1. Materials
3.2. MitoMet Neurite Outgrowth Assay
3.3. PeriTox Neurite Outgrowth Assay
3.4. Image Acquisition and Quantification in Neurite Assays
3.5. Neural Crest Cell Migration Assay (cMINC)
3.6. Data Analysis: Curve Fitting and Deriving BMC and EC Values
3.7. Preincubation of Compound with S9
3.8. Microsomal Stability Assay
3.9. Liquid Chromatography–Tandem Mass Spectrometry Analysis
3.10. Prediction of Protein Binding
4. Conclusions and Outlook
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BMC | Benchmark Concentration |
| BMR | Benchmark Response |
| cMINC | Neural Crest Cell Migration Assay |
| CYP | Cytochrome P450 (enzyme family) |
| DNT | Developmental Neurotoxicity |
| DNT-IVB | Developmental Neurotoxicity In Vitro Battery |
| DoM | Day of Migration |
| EC | Effect Concentration |
| FMO | Flavin-containing Monooxygenase |
| G6P | Glucose-6-phosphate |
| HLM | Human Liver Microsomes |
| HPP+ | Haloperidol Pyridinium Ion |
| KNDP | Key Neurodevelopmental Processes |
| LUHMES | Foetal Human Mesencephalic Cell Line |
| MDMA | Methylenedioxymethamphetamine |
| MitoMet | Mitochondrial Toxicity Assay Variant of NeuriTox |
| MPP+ | 1-methyl-4-phenylpyridinium |
| MPTP | 1-methyl-4-phenyl-tetrahydropyridine |
| NAM | New Approach Methodology |
| NCC | Neural Crest Cell |
| NGRA | Next-Generation Risk Assessment |
| S9 | Post-Mitochondrial Supernatant Fraction |
References
- Dent, M.; Amaral, R.T.; Da Silva, P.A.; Ansell, J.; Boisleve, F.; Hatao, M.; Hirose, A.; Kasai, Y.; Kern, P.; Kreiling, R.; et al. Principles Underpinning the Use of New Methodologies in the Risk Assessment of Cosmetic Ingredients. Comput. Toxicol. 2018, 7, 20–26. [Google Scholar] [CrossRef]
- Pallocca, G.; Moné, M.J.; Kamp, H.; Luijten, M.; van de Water, B.; Leist, M. Next-Generation Risk Assessment of Chemicals—Rolling Out a Human-Centric Testing Strategy to Drive 3R Implementation: The RISK-HUNT3R Project Perspective. ALTEX 2022, 39, 419–426. [Google Scholar] [CrossRef]
- Celardo, I.; Aschner, M.; Ashton, R.S.; Carstens, K.E.; Cediel-Ulloa, A.; Cöllen, E.; Crofton, K.M.; Debad, S.J.; Dreser, N.; Fitzpatrick, S.; et al. Developmental Neurotoxicity (DNT): A Call for Implementation of New Approach Methodologies for Regulatory Purposes: Summary of the 5th International Conference on DNT Testing. ALTEX 2025, 42, 323–349. [Google Scholar] [CrossRef]
- Fura, A. Role of Pharmacologically Active Metabolites in Drug Discovery and Development. Drug Discov. Today 2006, 11, 133–142. [Google Scholar] [CrossRef]
- Park, B.K.; Pirmohamed, M.; Kitteringham, N.R. The Role of Cytochrome P450 Enzymes in Hepatic and Extrahepatic Human Drug Toxicity. Pharmacol. Ther. 1995, 68, 385–424. [Google Scholar] [CrossRef]
- Rendic, S.P.; Guengerich, F.P. Human Family 1–4 Cytochrome P450 Enzymes Involved in the Metabolic Activation of Xenobiotic and Physiological Chemicals: An Update; Springer: Berlin/Heidelberg, Germany, 2021; Volume 95, ISBN 0123456789. [Google Scholar]
- Philip, P.A.; Ali-Sadat, S.; Doehmer, J.; Kocarek, T.; Akhtar, A.; Lu, H.; Chan, K.K. Use of V79 Cells with Stably Transfected Cytochrome P450 CDNAs in Studying the Metabolism and Effects of Cytotoxic Drugs. Cancer Chemother. Pharmacol. 1999, 43, 59–67. [Google Scholar] [CrossRef]
- Tolosa, L.; Gómez-Lechón, M.J.; Pérez-Cataldo, G.; Castell, J.V.; Donato, M.T. HepG2 Cells Simultaneously Expressing Five P450 Enzymes for the Screening of Hepatotoxicity: Identification of Bioactivable Drugs and the Potential Mechanism of Toxicity Involved. Arch. Toxicol. 2013, 87, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Amitai, G.; Moorad, D.; Adani, R.; Doctor, B.P. Inhibition of Acetylcholinesterase and Butyrylcholinesterase by Chlorpyrifos-Oxon. Biochem. Pharmacol. 1998, 56, 293–299. [Google Scholar] [CrossRef]
- Kasteel, E.E.J.; Nijmeijer, S.M.; Darney, K.; Lautz, L.S.; Dorne, J.L.C.M.; Kramer, N.I.; Westerink, R.H.S. Acetylcholinesterase Inhibition in Electric Eel and Human Donor Blood: An in Vitro Approach to Investigate Interspecies Differences and Human Variability in Toxicodynamics. Arch. Toxicol. 2020, 94, 4055–4065. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhao, J.; Huang, R.; Travers, J.; Klumpp-Thomas, C.; Yu, W.; Mackerell, A.D.; Sakamuru, S.; Ooka, M.; Xue, F.; et al. Profiling the Tox21 Chemical Collection for Acetylcholinesterase Inhibition. Environ. Health Perspect. 2021, 129, 047008. [Google Scholar] [CrossRef]
- Bal-Price, A.; Hogberg, H.T.; Crofton, K.M.; Daneshian, M.; FitzGerald, R.E.; Fritsche, E.; Heinonen, T.; Bennekou, S.H.; Klima, S.; Piersma, A.H.; et al. Recommendation on Test Readiness Criteria for New Approach Methods in Toxicology: Exemplified for Developmental Neurotoxicity. ALTEX 2018, 35, 306–352. [Google Scholar] [CrossRef]
- Smirnova, L.; Hogberg, H.T.; Leist, M.; Hartung, T. Revolutionizing Developmental Neurotoxicity Testing—A Journey from Animal Models to Advanced In Vitro Systems. ALTEX 2024, 41, 152–178. [Google Scholar] [CrossRef] [PubMed]
- Blum, J.; Masjosthusmann, S.; Bartmann, K.; Bendt, F.; Dolde, X.; Dönmez, A.; Förster, N.; Holzer, A.K.; Hübenthal, U.; Keßel, H.E.; et al. Establishment of a Human Cell-Based in Vitro Battery to Assess Developmental Neurotoxicity Hazard of Chemicals. Chemosphere 2023, 311, 137035. [Google Scholar] [CrossRef] [PubMed]
- Aschner, M. Reference compounds for alternative test methods to indicate developmental neurotoxicity (DNT) potential of chemicals: Example lists and criteria for their selection and use. ALTEX-Altern. Anim. Exp. 2017, 34, 49–74. [Google Scholar] [CrossRef]
- Bal-Price, A.; Crofton, K.M.; Leist, M.; Allen, S.; Arand, M.; Buetler, T.; Delrue, N.; FitzGerald, R.E.; Hartung, T.; Heinonen, T.; et al. International STakeholder NETwork (ISTNET): Creating a Developmental Neurotoxicity (DNT) Testing Road Map for Regulatory Purposes. Arch. Toxicol. 2015, 89, 269–287. [Google Scholar] [CrossRef]
- Krug, A.K.; Balmer, N.V.; Matt, F.; Schönenberger, F.; Merhof, D.; Leist, M. Evaluation of a Human Neurite Growth Assay as Specific Screen for Developmental Neurotoxicants. Arch. Toxicol. 2013, 87, 2215–2231. [Google Scholar] [CrossRef]
- Delp, J.; Gutbier, S.; Klima, S.; Hoelting, L.; Pinto-Gil, K.; Hsieh, J.H.; Aichem, M.; Klein, K.; Schreiber, F.; Tice, R.R.; et al. A High-Throughput Approach to Identify Specific Neurotoxicants/Developmental Toxicants in Human Neuronal Cell Function Assays. ALTEX 2018, 35, 235–253. [Google Scholar] [CrossRef]
- Delp, J.; Funke, M.; Rudolf, F.; Cediel, A.; Bennekou, S.H.; van der Stel, W.; Carta, G.; Jennings, P.; Toma, C.; Gardner, I.; et al. Development of a Neurotoxicity Assay That Is Tuned to Detect Mitochondrial Toxicants. Arch. Toxicol. 2019, 93, 1585–1608. [Google Scholar] [CrossRef]
- Holzer, A.K.; Suciu, I.; Karreman, C.; Goj, T.; Leist, M. Specific Attenuation of Purinergic Signaling during Bortezomib-Induced Peripheral Neuropathy In Vitro. Int. J. Mol. Sci. 2022, 23, 3734. [Google Scholar] [CrossRef] [PubMed]
- Nyffeler, J.; Dolde, X.; Krebs, A.; Pinto-Gil, K.; Pastor, M.; Behl, M.; Waldmann, T.; Leist, M. Combination of Multiple Neural Crest Migration Assays to Identify Environmental Toxicants from a Proof-of-Concept Chemical Library. Arch. Toxicol. 2017, 91, 3613–3632. [Google Scholar] [CrossRef]
- Magel, V.; Blum, J.; Dolde, X.; Leisner, H.; Grillberger, K.; Khalidi, H.; Gardner, I.; Ecker, G.F.; Pallocca, G.; Dreser, N.; et al. Inhibition of Neural Crest Cell Migration by Strobilurin Fungicides and Other Mitochondrial Toxicants. Cells 2024, 13, 2057. [Google Scholar] [CrossRef] [PubMed]
- Lambrecht, R.; Jansen, J.; Rudolf, F.; El-Mesery, M.; Caporali, S.; Amelio, I.; Stengel, F.; Brunner, T. Drug-Induced Oxidative Stress Actively Prevents Caspase Activation and Hepatocyte Apoptosis. Cell Death Dis. 2024, 15, 659. [Google Scholar] [CrossRef]
- Capinha, L.; Jennings, P.; Commandeur, J.N.M. Exposure to Cis- and Trans-Regioisomers of S-(1,2-Dichlorovinyl)-L-Cysteine and S-(1,2-Dichlorovinyl)-Glutathione Result in Quantitatively and Qualitatively Different Cellular Effects in RPTEC/TERT1 Cells. Toxicol. Lett. 2023, 383, 75–88. [Google Scholar] [CrossRef]
- Schildknecht, S.; Pape, R.; Meiser, J.; Karreman, C.; Strittmatter, T.; Odermatt, M.; Cirri, E.; Friemel, A.; Ringwald, M.; Pasquarelli, N.; et al. Preferential Extracellular Generation of the Active Parkinsonian Toxin MPP+ by Transporter-Independent Export of the Intermediate MPDP+. Antioxid. Redox Signal. 2015, 23, 1001–1016. [Google Scholar] [CrossRef]
- Schmuck, G.; Ahr, H.J.; Mihail, F.; Stahl, B.; Kayser, M. Effects of the Dithiocarbamate Fungicide Propineb in Primary Neuronal Cell Cultures and Skeletal Muscle Cells of the Rat. Arch. Toxicol. 2002, 76, 414–422. [Google Scholar] [CrossRef]
- Pinto, C.L.; Mansouri, K.; Judson, R.; Browne, P. Prediction of Estrogenic Bioactivity of Environmental Chemical Metabolites. Chem. Res. Toxicol. 2016, 29, 1410–1427. [Google Scholar] [CrossRef] [PubMed]
- Paris, F.; Balaguer, P.; Térouanne, B.; Servant, N.; Lacoste, C.; Cravedi, J.P.; Nicolas, J.C.; Sultan, C. Phenylphenols, Biphenols, Bisphenol-A and 4-Tert-Octylphenol Exhibit α and β Estrogen Activities and Antiandrogen Activity in Reporter Cell Lines. Mol. Cell. Endocrinol. 2002, 193, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Koenig, L.; Ramme, A.P.; Faust, D.; Mayer, M.; Flötke, T.; Gerhartl, A.; Brachner, A.; Neuhaus, W.; Appelt-Menzel, A.; Metzger, M.; et al. A Human Stem Cell-Derived Brain-Liver Chip for Assessing Blood-Brain-Barrier Permeation of Pharmaceutical Drugs. Cells 2022, 11, 3295. [Google Scholar] [CrossRef]
- Oleaga, C.; Bernabini, C.; Smith, A.S.T.; Srinivasan, B.; Jackson, M.; McLamb, W.; Platt, V.; Bridges, R.; Cai, Y.; Santhanam, N.; et al. Multi-Organ Toxicity Demonstration in a Functional Human in Vitro System Composed of Four Organs. Sci. Rep. 2016, 6, 20030. [Google Scholar] [CrossRef]
- Materne, E.M.; Ramme, A.P.; Terrasso, A.P.; Serra, M.; Alves, P.M.; Brito, C.; Sakharov, D.A.; Tonevitsky, A.G.; Lauster, R.; Marx, U. A Multi-Organ Chip Co-Culture of Neurospheres and Liver Equivalents for Long-Term Substance Testing. J. Biotechnol. 2015, 205, 36–46. [Google Scholar] [CrossRef]
- Bruinink, A.; Yu, D.; Maier, P. Short-Term Assay for the Identification of Neurotoxic Compounds and Their Liver Derived Stable Metabolites. Toxicol. Vitr. 2002, 16, 717–724. [Google Scholar] [CrossRef]
- Bruinink, A.; Maier, P. Identification of Neurotoxic Chemicals in Cell Cultures. ALTEX 2007, 24, 22–25. [Google Scholar] [PubMed]
- Yu, K.-N.; Kang, S.-Y.; Hong, S.; Lee, M.-Y. High-Throughput Metabolism-Induced Toxicity Assays Demonstrated on a 384-Pillar Plate. Arch. Toxicol. 2018, 92, 2501–2516. [Google Scholar] [CrossRef]
- DeGroot, D.E.; Swank, A.; Thomas, R.S.; Strynar, M.; Lee, M.-Y.; Carmichael, P.L.; Simmons, S.O. MRNA Transfection Retrofits Cell-Based Assays with Xenobiotic Metabolism. J. Pharmacol. Toxicol. Methods 2018, 92, 77–94. [Google Scholar] [CrossRef]
- Mollergues, J.; Van Vugt-Lussenburg, B.; Kirchnawy, C.; Bandi, R.A.; Van Der Lee, R.B.; Marin-Kuan, M.; Schilter, B.; Fussell, K.C. Incorporation of a Metabolizing System in Biodetection Assays for Endocrine Active Substances. ALTEX 2017, 34, 389–398. [Google Scholar] [CrossRef] [PubMed]
- van Vugt-Lussenburg, B.M.A.; van der Lee, R.B.; Man, H.Y.; Middelhof, I.; Brouwer, A.; Besselink, H.; van der Burg, B. Incorporation of Metabolic Enzymes to Improve Predictivity of Reporter Gene Assay Results for Estrogenic and Anti-Androgenic Activity. Reprod. Toxicol. 2018, 75, 40–48. [Google Scholar] [CrossRef]
- Coecke, S.; Ahr, H.; Blaauboer, B.J.; Bremer, S.; Casati, S.; Castell, J.; Combes, R.; Corvi, R.; Crespi, C.L.; Cunningham, M.L.; et al. Metabolism: A Bottleneck in in Vitro Toxicological Test Development. ATLA Altern. Lab. Anim. 2006, 34, 49–84. [Google Scholar] [CrossRef] [PubMed]
- Callander, R.D.; Mackay, J.M.; Clay, P.; Elcombe, C.R.; Elliott, B.M. Evaluation of Phenobarbital/β-Naphthoflavone as an Alternative S9-Induction Regime to Aroclor 1254 in the Rat for Use in in Vitro Genotoxicity Assays. Mutagenesis 1995, 10, 517–522. [Google Scholar] [CrossRef]
- Maron, D.M.; Ames, B.N. Revised Methods for the Salmonella Mutagenicity Test. Mutat. Res./Environ. Mutagen. Relat. Subj. 1983, 113, 173–215. [Google Scholar] [CrossRef] [PubMed]
- OECD. Test No. 471: Bacterial Reverse Mutation Test. 1997. Available online: https://www.oecd.org/en/publications/test-no-471-bacterial-reverse-mutation-test_9789264071247-en.html (accessed on 25 August 2025).
- Taxvig, C.; Olesen, P.T.; Nellemann, C. Use of External Metabolizing Systems When Testing for Endocrine Disruption in the T-Screen Assay. Toxicol. Appl. Pharmacol. 2011, 250, 263–269. [Google Scholar] [CrossRef]
- Deisenroth, C.; DeGroot, D.E.; Zurlinden, T.; Eicher, A.; McCord, J.; Lee, M.Y.; Carmichael, P.; Thomas, R.S. The Alginate Immobilization of Metabolic Enzymes Platform Retrofits an Estrogen Receptor Transactivation Assay with Metabolic Competence. Toxicol. Sci. 2020, 178, 281–301. [Google Scholar] [CrossRef]
- Hopperstad, K.; Deisenroth, C. Development of a Bioprinter-Based Method for Incorporating Metabolic Competence into High-Throughput in Vitro Assays. Front. Toxicol. 2023, 5, 1196245. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, G.A.; Frömbling, S.; Joseph, N.; Bode, K.; Bicker, G.; Stern, M. An Intact Insect Embryo for Developmental Neurotoxicity Testing of Directed Axonal Elongation. ALTEX 2019, 36, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Escher, B.I.; Scholz, S.; Schlichting, R. Inhibition of Neurite Outgrowth and Enhanced Effects Compared to Baseline Toxicity in SH-SY5Y Cells. Arch. Toxicol. 2022, 96, 1039–1053. [Google Scholar] [CrossRef] [PubMed]
- Pamies, D.; Block, K.; Lau, P.; Gribaldo, L.; Pardo, C.A.; Barreras, P.; Smirnova, L.; Wiersma, D.; Zhao, L.; Harris, G.; et al. Rotenone Exerts Developmental Neurotoxicity in a Human Brain Spheroid Model. Toxicol. Appl. Pharmacol. 2018, 354, 101–114. [Google Scholar] [CrossRef]
- Flaskos, J.; Nikolaidis, E.; Harris, W.; Sachana, M.; Hargreaves, A.J. Effects of Sub-Lethal Neurite Outgrowth Inhibitory Concentrations of Chlorpyrifos Oxon on Cytoskeletal Proteins and Acetylcholinesterase in Differentiating N2a Cells. Toxicol. Appl. Pharmacol. 2011, 256, 330–336. [Google Scholar] [CrossRef]
- Liu, K.H.; Lee, C.M.; Singer, G.; Bais, P.; Castellanos, F.; Woodworth, M.H.; Ziegler, T.R.; Kraft, C.S.; Miller, G.W.; Li, S.; et al. Large Scale Enzyme Based Xenobiotic Identification for Exposomics. Nat. Commun. 2021, 12, 5418. [Google Scholar] [CrossRef]
- Jamnik, T.; Flasch, M.; Braun, D.; Fareed, Y.; Wasinger, D.; Seki, D.; Berry, D.; Berger, A.; Wisgrill, L.; Warth, B. Next-Generation Biomonitoring of the Early-Life Chemical Exposome in Neonatal and Infant Development. Nat. Commun. 2022, 13, 2653. [Google Scholar] [CrossRef]
- Tyzack, J.D.; Kirchmair, J. Computational Methods and Tools to Predict Cytochrome P450 Metabolism for Drug Discovery. Chem. Biol. Drug Des. 2019, 93, 377–386. [Google Scholar] [CrossRef]
- Kirchmair, J.; Göller, A.H.; Lang, D.; Kunze, J.; Testa, B.; Wilson, I.D.; Glen, R.C.; Schneider, G. Predicting Drug Metabolism: Experiment and/or Computation? Nat. Rev. Drug Discov. 2015, 14, 387–404. [Google Scholar] [CrossRef]
- Jin, X.; Pybus, B.S.; Marcsisin, S.R.; Logan, T.; Luong, T.L.; Sousa, J.; Matlock, N.; Collazo, V.; Asher, C.; Carroll, D.; et al. An LC-MS Based Study of the Metabolic Profile of Primaquine, an 8-Aminoquinoline Antiparasitic Drug, with an in Vitro Primary Human Hepatocyte Culture Model. Eur. J. Drug Metab. Pharmacokinet. 2014, 39, 139–146. [Google Scholar] [CrossRef]
- Li, X.Q.; Björkman, A.; Andersson, T.B.; Gustafsson, L.L.; Masimirembwa, C.M. Identification of Human Cytochrome P450s That Metabolise Anti-Parasitic Drugs and Predictions of in Vivo Drug Hepatic Clearance from in Vitro Data. Eur. J. Clin. Pharmacol. 2003, 59, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.G.; Peggins, J.O.; Fleckenstein, L.L.; Masonic, K.; Heiffer, M.H.; Brewer, T.G. The Pharmacokinetics and Bioavailability of Dihydroartemisinin, Arteether, Artemether, Artesunic Acid and Artelinic Acid in Rats. J. Pharm. Pharmacol. 1998, 50, 173–182. [Google Scholar] [CrossRef]
- Lim, J.; Miller, M.G. The Role of the Benomyl Metabolite Carbendazim in Benomyl-Induced Testicular Toxicity. Toxicol. Appl. Pharmacol. 1997, 142, 401–410. [Google Scholar] [CrossRef] [PubMed]
- Gardiner, J.A.; Kirkland, J.J.; Klopping, H.L.; Sherman, H. Fate of Benomyl in Animals. J. Agric. Food Chem. 1974, 22, 419–427. [Google Scholar] [CrossRef]
- Krause, M.; Frederiksen, H.; Sundberg, K.; Jørgensen, F.S.; Jensen, L.N.; Nørgaard, P.; Jørgensen, C.; Ertberg, P.; Juul, A.; Drzewiecki, K.T.; et al. Presence of Benzophenones Commonly Used as UV Filters and Absorbers in Paired Maternal and Fetal Samples. Environ. Int. 2018, 110, 51–60. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Suzuki, T.; Tayama, S. Metabolism and Toxicity of Benzophenone in Isolated Rat Hepatocytes and Estrogenic Activity of Its Metabolites in MCF-7 Cells. Toxicology 2000, 156, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Attene-Ramos, M.S.; Huang, R.; Michael, S.; Witt, K.L.; Richard, A.; Tice, R.R.; Simeonov, A.; Austin, C.P.; Xia, M. Profiling of the Tox21 Chemical Collection for Mitochondrial Function to Identify Compounds That Acutely Decrease Mitochondrial Membrane Potential. Environ. Health Perspect. 2015, 123, 49–56. [Google Scholar] [CrossRef]
- Hollingworth, R.M.; Kurihara, N.; Miyamoto, J.; Otto, S.; Paulson, G.D. Detection and Significance of Active Metabolites of Agrochemicals and Related Xenobiotics in Animals (Technical Report). Pure Appl. Chem. 1995, 67, 1487–1532. [Google Scholar] [CrossRef]
- van Lier, R.B.L.; Cherry, L.D. The Toxicity and Mechanism of Action of Bromethalin: A New Single-Feeding Rodenticide. Fundam. Appl. Toxicol. 1988, 11, 664–672. [Google Scholar] [CrossRef]
- Bellisai, G.; Bernasconi, G.; Binaglia, M.; Carrasco Cabrera, L.; Castellan, I.; Castoldi, A.F.; Chiusolo, A.; Chukwubike, K.; Crivellente, F.; Del Aguila, M.; et al. Targeted Review of Maximum Residue Levels (MRLs) for Chlorfenapyr. EFSA J. 2023, 21, e8444. [Google Scholar] [CrossRef]
- Beretta, C.; Fadini, L.; Stracciari, J.M.; Montesissa, C. In Vitro Febantel Transformation by Sheep and Cattle Ruminal Fluids and Metabolism by Hepatic Subcellular Fractions from Different Animal Species. Biochem. Pharmacol. 1987, 36, 3107–3114. [Google Scholar] [CrossRef] [PubMed]
- Mahajan, A.A.; Marathe, A.M.; Jarande, S.S.; Pissurlenkar, R.; Gawande, V.T. Characterization and Toxicity Evaluation of Degradation Products of Febantel. Future J. Pharm. Sci. 2020, 6, 127. [Google Scholar] [CrossRef]
- Kalgutkar, A.S.; Nguyen, H.T. Identification of an N-Methyl-4-Phenylpyridinium-like Metabolite of the Antidiarrheal Agent Loperamide in Human Liver Microsomes: Underlying Reason(s) for the Lack of Neurotoxicity despite the Bioactivation Event. Drug Metab. Dispos. 2004, 32, 943–952. [Google Scholar] [CrossRef] [PubMed]
- Usuki, E.; Pearce, R.; Parkinson, A.; Castagnoli, N. Studies on the Conversion of Haloperidol and Its Tetrahydropyridine Dehydration Product to Potentially Neurotoxic Pyridinium Metabolites by Human Liver Microsomes. Chem. Res. Toxicol. 1996, 9, 800–806. [Google Scholar] [CrossRef]
- Costa, G.; Gołembiowska, K. Neurotoxicity of MDMA: Main Effects and Mechanisms. Exp. Neurol. 2022, 347, 113894. [Google Scholar] [CrossRef] [PubMed]
- Heikkila, R.E.; Manzino, L.; Cabbat, F.S.; Duvoisin, R.C. Protection against the Dopaminergic Neurotoxicity of 1-Methyl-4-Phenyl-1,2,5,6-Tetrahydropyridine by Monoamine Oxidase Inhibitors. Nature 1984, 311, 467–469. [Google Scholar] [CrossRef]
- Shao, Y.; Schiwy, A.; Glauch, L.; Henneberger, L.; König, M.; Mühlenbrink, M.; Xiao, H.; Thalmann, B.; Schlichting, R.; Hollert, H.; et al. Optimization of a Pre-Metabolization Procedure Using Rat Liver S9 and Cell-Extracted S9 in the Ames Fluctuation Test. Sci. Total Environ. 2020, 749, 141468. [Google Scholar] [CrossRef]
- de Jonge, M.E.; Huitema, A.D.R.; Rodenhuis, S.; Beijnen, J.H. Clinical Pharmacokinetics of Cyclophosphamide. Clin. Pharmacokinet. 2005, 44, 1135–1164. [Google Scholar] [CrossRef]
- Feng, X.; Wang, K.; Cao, S.; Ding, L.; Qiu, F. Pharmacokinetics and Excretion of Berberine and Its Nine Metabolites in Rats. Front. Pharmacol. 2020, 11, 594852. [Google Scholar] [CrossRef]
- Turner, N.; Li, J.; Gosby, A.; To, S.W.C.; Cheng, Z.; Miyoshi, H.; Taketo, M.M.; Cooney, G.J.; Kraegen, E.W.; James, D.E.; et al. A Mechanism for the Action of Berberine to Activate. Diabetes 2008, 57, 1414–1418. [Google Scholar] [CrossRef]
- Suciu, I.; Delp, J.; Gutbier, S.; Suess, J.; Henschke, L.; Celardo, I.; Mayer, T.U.; Amelio, I.; Leist, M. Definition of the Neurotoxicity-Associated Metabolic Signature Triggered by Berberine and Other Respiratory Chain Inhibitors. Antioxidants 2024, 13, 49. [Google Scholar] [CrossRef]
- Nyffeler, J.; Willis, C.; Lougee, R.; Richard, A.; Paul-Friedman, K.; Harrill, J.A. Bioactivity Screening of Environmental Chemicals Using Imaging-Based High-Throughput Phenotypic Profiling. Toxicol. Appl. Pharmacol. 2020, 389, 114876. [Google Scholar] [CrossRef]
- Alimohammadi, M.; Meyburg, B.; Ückert, A.; Holzer, A.; Leist, M. EFSA Pilot Project on New Approach Methodologies (NAMs) for Tebufenpyrad Risk Assessment. Part 2. Hazard Characterisation and Identification of the Reference Point. EFSA Support. Publ. 2023, 20, 7794E. [Google Scholar] [CrossRef]
- Hoelting, L.; Klima, S.; Karreman, C.; Grinberg, M.; Meisig, J.; Henry, M.; Rotshteyn, T.; Rahnenführer, J.; Blüthgen, N.; Sachinidis, A.; et al. Stem Cell-Derived Immature Human Dorsal Root Ganglia Neurons to Identify Peripheral Neurotoxicants. Stem Cells Transl. Med. 2016, 5, 476–487. [Google Scholar] [CrossRef]
- Krebs, A.; Nyffeler, J.; Karreman, C.; Kappenberg, F.; Mellert, J.; Pallocca, G.; Pastor, M.; Leist, M. Determination of Benchmark Concentrations and Their Statistical Uncertainty for Cytotoxicity Test Data and Functional In Vitro Assays. ALTEX 2020, 37, 155–163. [Google Scholar] [CrossRef]
- Lobell, M.; Sivarajah, V. In Silico Prediction of Aqueous Solubility, Human Plasma Protein Binding and Volume of Distribution of Compounds from Calculated PKa and AlogP98 Values. Mol. Divers. 2003, 7, 69–87. [Google Scholar] [CrossRef]
- Gardner, I.; Xu, M.; Han, C.; Wang, Y.; Jiao, X.; Jamei, M.; Khalidi, H.; Kilford, P.; Neuhoff, S.; Southall, R.; et al. Non-Specific Binding of Compounds in in Vitro Metabolism Assays: A Comparison of Microsomal and Hepatocyte Binding in Different Species and an Assessment of the Accuracy of Prediction Models. Xenobiotica 2022, 52, 943–956. [Google Scholar] [CrossRef] [PubMed]
- Heringa, M.B.; Schreurs, R.H.M.M.; Busser, F.; Van Der Saag, P.T.; Van Der Burg, B.; Hermens, J.L.M. Toward More Useful in Vitro Toxicity Data with Measured Free Concentrations. Environ. Sci. Technol. 2004, 38, 6263–6270. [Google Scholar] [CrossRef] [PubMed]
- Zanger, U.M.; Schwab, M. Cytochrome P450 Enzymes in Drug Metabolism: Regulation of Gene Expression, Enzyme Activities, and Impact of Genetic Variation. Pharmacol. Ther. 2013, 138, 103–141. [Google Scholar] [CrossRef]
- Rendic, S.; Guengerich, F.P. Survey of Human Oxidoreductases and Cytochrome P450 Enzymes Involved in the Metabolism of Xenobiotic and Natural Chemicals. Chem. Res. Toxicol. 2015, 28, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Faustman-Watts, E.M.; Greenaway, J.C.; Namkung, M.J.; Fantel, A.G.; Juchau, M.R. Teratogenicity in Vitro of Two Deacetylated Metabolites of N-Hydroxy-2-Acetylaminofluorene. Toxicol. Appl. Pharmacol. 1984, 76, 161–171. [Google Scholar] [CrossRef]
- Smith, T.J.; Hanna, P.E. N -Acetyltransferase Multiplicity and the Bioactivation of N -Arylhydroxamic Acids by Hamster Hepatic and Intestinal Enzymes. Carcinogenesis 1986, 7, 697–702. [Google Scholar] [CrossRef]
- Hammer, H.; Schmidt, F.; Marx-Stoelting, P.; Pötz, O.; Braeuning, A. Cross-Species Analysis of Hepatic Cytochrome P450 and Transport Protein Expression. Arch. Toxicol. 2020, 95, 117–133. [Google Scholar] [CrossRef]
- Cox, J.A.; Fellows, M.D.; Hashizume, T.; White, P.A. The Utility of Metabolic Activation Mixtures Containing Human Hepatic Post-Mitochondrial Supernatant (S9) for in Vitro Genetic Toxicity Assessment. Mutagenesis 2016, 31, 117–130. [Google Scholar] [CrossRef]
- Richardson, S.J.; Bai, A.; Kulkarni, A.A.; Moghaddam, M.F. Efficiency in Drug Discovery: Liver S9 Fraction Assay As a Screen for Metabolic Stability. Drug Metab. Lett. 2016, 10, 83–90. [Google Scholar] [CrossRef]
- Mishra, N.; Srivastava, R. Bacterial Worth in Genotoxicity Assessment Studies. J. Microbiol. Methods 2023, 215, 106860. [Google Scholar] [CrossRef]
- Brendt, J.; Lackmann, C.; Heger, S.; Velki, M.; Crawford, S.E.; Xiao, H.; Thalmann, B.; Schiwy, A.; Hollert, H. Using a High-Throughput Method in the Micronucleus Assay to Compare Animal-Free with Rat-Derived S9. Sci. Total Environ. 2021, 751, 142269. [Google Scholar] [CrossRef] [PubMed]
- Riley, R.; Roberts, P.; Coleman, M.; Kitteringham, N.; Park, B. Bioactivation of Dapsone to a Cytotoxic Metabolite: In Vitro Use of a Novel Two Compartment System Which Contains Human Tissues. Br. J. Clin. Pharmacol. 1990, 30, 417–426. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Suess, J.; Reinmoeller, M.; Magel, V.; Gukalova, B.; Liepinsh, E.; Gardner, I.; Dreser, N.; Holzer, A.-K.; Leist, M. Testing Strategies for Metabolite-Mediated Neurotoxicity. Int. J. Mol. Sci. 2025, 26, 8338. https://doi.org/10.3390/ijms26178338
Suess J, Reinmoeller M, Magel V, Gukalova B, Liepinsh E, Gardner I, Dreser N, Holzer A-K, Leist M. Testing Strategies for Metabolite-Mediated Neurotoxicity. International Journal of Molecular Sciences. 2025; 26(17):8338. https://doi.org/10.3390/ijms26178338
Chicago/Turabian StyleSuess, Julian, Moritz Reinmoeller, Viktoria Magel, Baiba Gukalova, Edgars Liepinsh, Iain Gardner, Nadine Dreser, Anna-Katharina Holzer, and Marcel Leist. 2025. "Testing Strategies for Metabolite-Mediated Neurotoxicity" International Journal of Molecular Sciences 26, no. 17: 8338. https://doi.org/10.3390/ijms26178338
APA StyleSuess, J., Reinmoeller, M., Magel, V., Gukalova, B., Liepinsh, E., Gardner, I., Dreser, N., Holzer, A.-K., & Leist, M. (2025). Testing Strategies for Metabolite-Mediated Neurotoxicity. International Journal of Molecular Sciences, 26(17), 8338. https://doi.org/10.3390/ijms26178338






