Targeting Kupffer Cell Enolase 1 Attenuates Liver Inflammation and Injury in Hemorrhagic Shock
Abstract
1. Introduction
2. Results
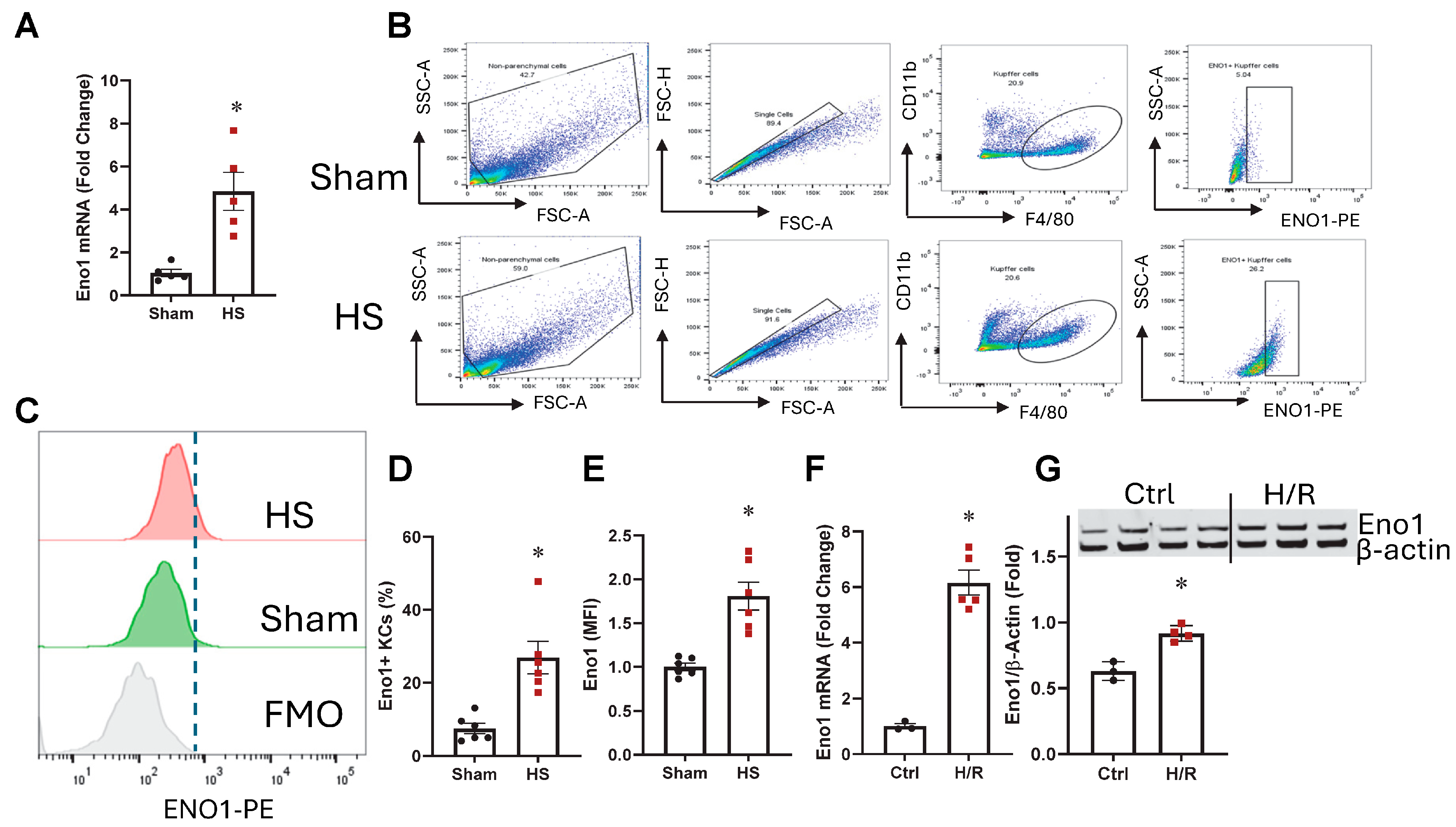
2.1. ENO1 Was Increased in KCs from HS Mice
2.2. ENO1 Was Upregulated in Kupffer Cells After H/R
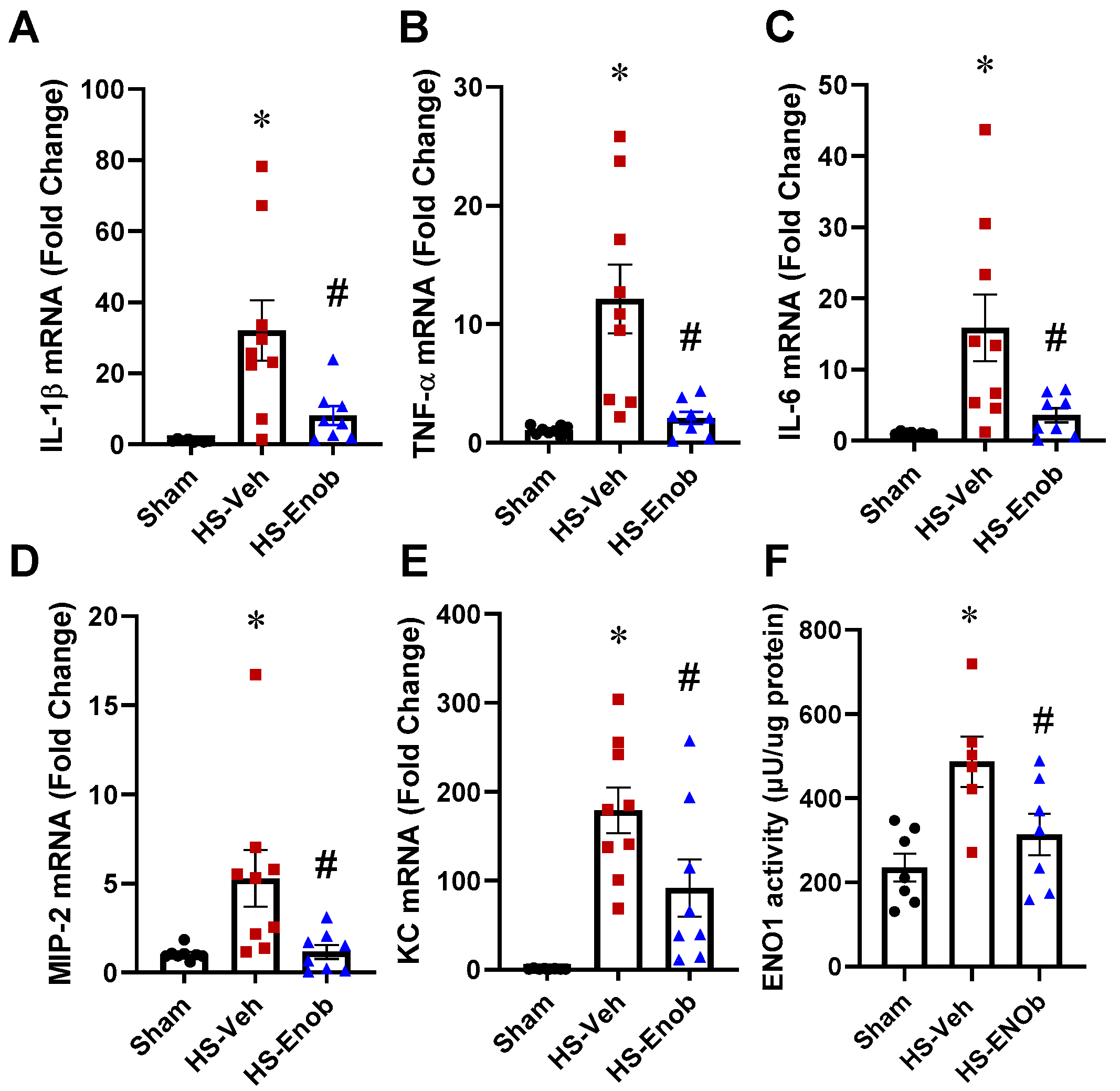
2.3. ENOblock Alleviates H/R-Induced Inflammatory Injury
2.4. ENOblock Alleviates HS-Induced Liver Inflammation and ENO1 Activity
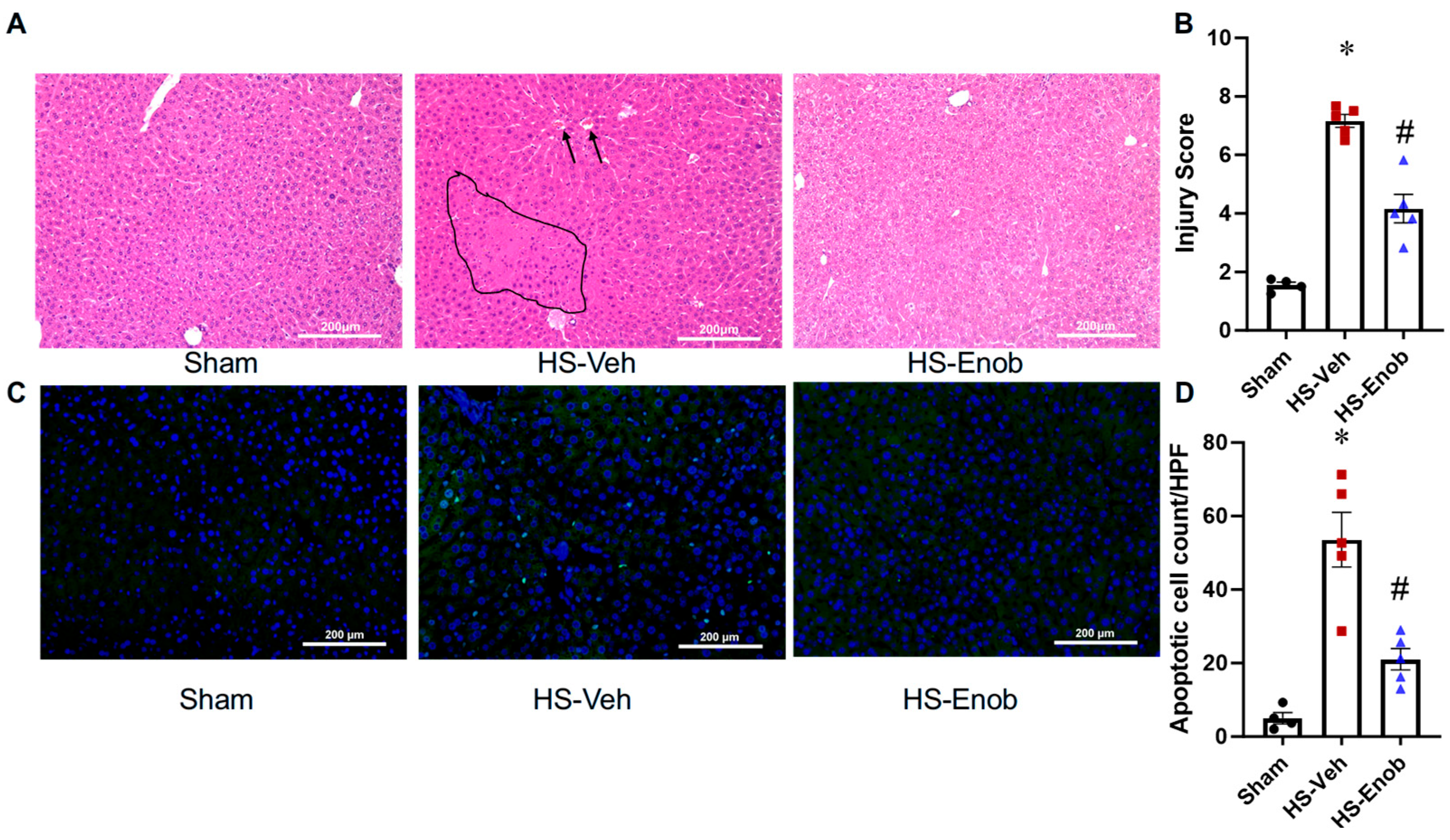
2.5. ENOblock Mitigates Liver Injury in HS
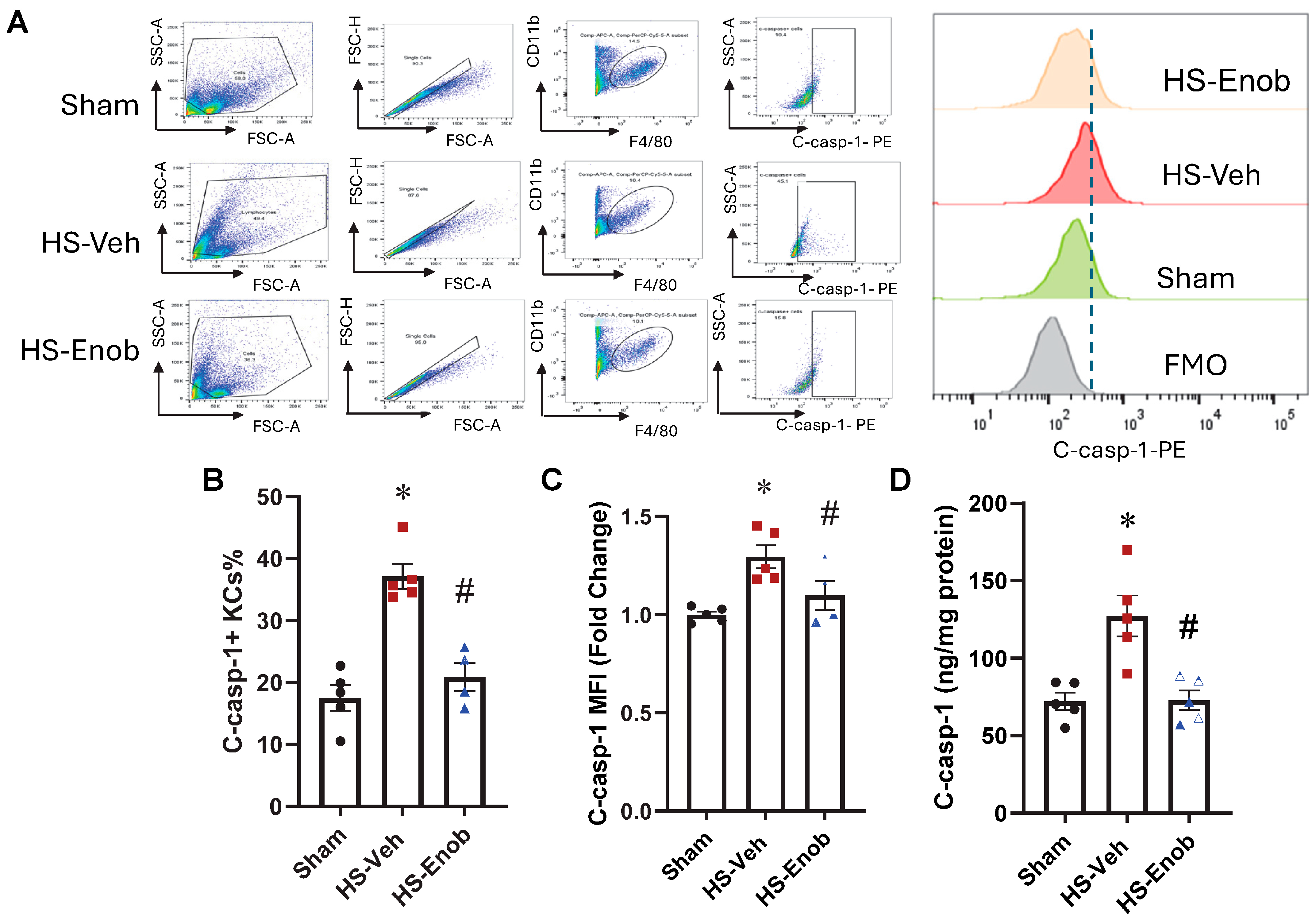
2.6. ENOblock Attenuates C-caspase-1 in KCs After HS
3. Discussion
4. Materials and Methods
4.1. Enolase1 Inhibitor
4.2. Experimental Animals
4.3. Mouse Model of Hemorrhagic Shock
4.4. Kupffer Cell Isolation
4.5. Hypoxia/Reoxygenation In Vitro
4.6. Enolase Activity Assay
4.7. Flow Cytometry
4.8. Western Blot
4.9. Cytokines and C-caspase-1 Measurement
4.10. Real-Time Quantitative PCR
- Enolase1, forward 5′-CTGGCCAAGTACAATCAGATCCTC-3′ and reverse 5′-GGATCTCCGGTCCATGCTTTA-3′
- interleukin (IL)-6, forward 5′-CCGGAGAGGAGACTTCACAG-3′ and reverse 5′-CAGAATTGCCATTGCACAAC-3′;
- IL-1β, forward 5′-CAGGATGAGGACATGAGCACC-3′, and reverse 5′-CTCGCAGACTCAAACTCCAC-3′
- TNF-α, forward 5′-AGACCCTCACACTCAGATCATCTTC-3′ and reverse 5′-TTGCTACGACGTGGGCTACA-3′;
- KC, forward 5′-GCTGGGATTCACCTCAAGAA-3′ and reverse 5′-ACAGGTGCCATCAGAGCAGT-3′;
- macrophage inflammatory protein (MIP)-2, forward 5′-CATCCAGAGCTTGAGTGTGA-3′ and reverse 5′-CTTTGGTTCTTCCGTTGAGG-3′;
- β-actin, forward 5′-CGTGAAAAGATGACCCAGATCA-3′ and reverse 5′-TGGTACGACCAGAGGCATACAG-3′.
4.11. Liver Injury Histopathological Evaluation
4.12. Terminal Deoxynucleotidyl Transferase dUTP Nick End Labeling (TUNEL) Assay
4.13. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HS | Hemorrhagic shock |
| ENO | Enolase |
| KCs | Kupffer cells |
| H/R | Hypoxia/reoxygenation |
| mRNA | Messenger RNA |
| TUNEL | Terminal deoxynucleotidyl transferase dUTP nick end labeling |
| IL-1β | Interleukin-1beta |
| TNF-α | Tumor necrosis factor-alpha |
| IL-6 | Interleukin-6 |
| MAP | Mean arterial pressure |
| HBSS | Hank’s balanced salt solution |
| FBS | Fetal bovine serum |
| FACS | Fluorescence-activated cell sorting |
| PBS | Phosphate-buffered saline |
| PAGE | Polyacrylamide gel electrophoresis |
| TBS | Tris-buffered saline |
| ELISA | Enzyme-linked immunosorbent assay |
| BCP | 1-bromo-3-chloropropane |
| qPCR | Quantitative polymerase chain reaction |
| KC | Keratinocyte-derived cytokine |
| MIP-2 | Macrophage inflammatory protein-2 |
| H&E | Hematoxylin and eosin |
| SEM | Standard error of the mean |
| ANOVA | Analysis of variance |
| SNK | Student–Newman–Keuls |
References
- Latif, R.K.; Clifford, S.P.; Baker, J.A.; Lenhardt, R.; Haq, M.Z.; Huang, J.; Farah, I.; Businger, J.R. Traumatic hemorrhage and chain of survival. Scand. J. Trauma Resusc. Emerg. Med. 2023, 31, 25. [Google Scholar] [CrossRef]
- Gallimore, E. Clinical features and management of haemorrhagic shock. Nurs. Stand. 2015, 30, 51–58, quiz 60. [Google Scholar] [CrossRef]
- Gann, D.S.; Drucker, W.R. Hemorrhagic shock. J. Trauma Acute Care Surg. 2013, 75, 888–895. [Google Scholar] [CrossRef]
- Xie, Y.; Xu, M.; Deng, M.; Li, Z.; Wang, P.; Ren, S.; Guo, Y.; Ma, X.; Fan, J.; Billiar, T.R.; et al. Activation of Pregnane X Receptor Sensitizes Mice to Hemorrhagic Shock-Induced Liver Injury. Hepatology 2019, 70, 995–1010. [Google Scholar] [CrossRef] [PubMed]
- Wetzel, G.; Relja, B.; Klarner, A.; Henrich, D.; Dehne, N.; Brühne, B.; Lehnert, M.; Marzi, I. Myeloid knockout of HIF-1 α does not markedly affect hemorrhage/resuscitation-induced inflammation and hepatic injury. Mediat. Inflamm. 2014, 2014, 930419. [Google Scholar] [CrossRef] [PubMed]
- Qiao, G.; Wu, A.; Chen, X.; Tian, Y.; Lin, X. Enolase 1, a Moonlighting Protein, as a Potential Target for Cancer Treatment. Int. J. Biol. Sci. 2021, 17, 3981–3992. [Google Scholar] [CrossRef]
- Yuan, Z.; Hu, H.; Zhu, Y.; Zhang, W.; Fang, Q.; Qiao, T.; Ma, T.; Wang, M.; Huang, R.; Tang, Q.; et al. Colorectal cancer cell intrinsic fibroblast activation protein alpha binds to Enolase1 and activates NF-κB pathway to promote metastasis. Cell Death Dis. 2021, 12, 543. [Google Scholar] [CrossRef]
- Cho, H.; Lee, J.H.; Um, J.; Kim, S.; Kim, Y.; Kim, W.H.; Kim, Y.S.; Pagire, H.S.; Ahn, J.H.; Ahn, Y.; et al. ENOblock inhibits the pathology of diet-induced obesity. Sci. Rep. 2019, 9, 493. [Google Scholar] [CrossRef]
- Pancholi, V. Multifunctional alpha-enolase: Its role in diseases. Cell. Mol. Life Sci. 2001, 58, 902–920. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Kim, H.; Kim, Y.; Jang, M.; Jeon, J.; Hwang, Y.I.; Shon, W.J.; Song, Y.W.; Kang, J.S.; Lee, W.J. The Anti-inflammatory Effect of GV1001 Mediated by the Downregulation of ENO1-induced Pro-inflammatory Cytokine Production. Immune Netw. 2015, 15, 291–303. [Google Scholar] [CrossRef]
- Ray, A.; Song, Y.; Du, T.; Chauhan, D.; Anderson, K.C. Preclinical validation of Alpha-Enolase (ENO1) as a novel immunometabolic target in multiple myeloma. Oncogene 2020, 39, 2786–2796. [Google Scholar] [CrossRef]
- Sedoris, K.C.; Thomas, S.D.; Miller, D.M. Hypoxia induces differential translation of enolase/MBP-1. BMC Cancer 2010, 10, 157. [Google Scholar] [CrossRef]
- Caramelo, I.; Coelho, M.; Rosado, M.; Cardoso, C.M.P.; Dinis, A.; Duarte, C.B.; Grãos, M.; Manadas, B. Biomarkers of hypoxic-ischemic encephalopathy: A systematic review. World J. Pediatr. 2023, 19, 505–548. [Google Scholar] [CrossRef]
- Shi, Y.; Liu, J.; Zhang, R.; Zhang, M.; Cui, H.; Wang, L.; Cui, Y.; Wang, W.; Sun, Y.; Wang, C. Targeting Endothelial ENO1 (Alpha-Enolase) -PI3K-Akt-mTOR Axis Alleviates Hypoxic Pulmonary Hypertension. Hypertension 2023, 80, 1035–1047. [Google Scholar] [CrossRef]
- Chen, L.J.; Li, J.Y.; Nguyen, P.; He, M.; Chen, Z.B.; Subramaniam, S.; Shyy, J.Y.; Chien, S. Single-cell RNA sequencing unveils unique transcriptomic signatures of endothelial cells and role of ENO1 in response to disturbed flow. Proc. Natl. Acad. Sci. USA 2024, 121, e2318904121. [Google Scholar] [CrossRef]
- Lin, J.; Feng, B.S.; Huang, N.; Ma, N.; Liu, Z.Q.; Shu, Q.; Zhong, N.; Liu, Z.G.; Yang, P.C. Enolase-specific cross antibodies induce neutrophilic inflammation in the intestine. J. Leukoc. Biol. 2021, 109, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.C.; Chuang, C.F.; Huang, Y.T.; Chung, I.C.; Chen, M.L.; Chuang, T.Y.; Yang, X.L.; Chou, Y.Y.; Liu, C.H.; Chen, N.Y.; et al. Monoclonal enolase-1 blocking antibody ameliorates pulmonary inflammation and fibrosis. Respir. Res. 2023, 24, 280. [Google Scholar] [CrossRef]
- Tan, C.; Ma, H.; Chen, J.; Ma, G.; Jha, A.; Tan, S.; Zhu, Y.; Liu, M.; Liu, K.; Xiao, X.; et al. Critical Role of IL1R2-ENO1 Interaction in Inhibiting Glycolysis-Mediated Pyroptosis for Protection Against Lethal Sepsis. Adv. Sci. 2025, e02297. [Google Scholar] [CrossRef]
- Jung, D.W.; Ha, H.H.; Zheng, X.; Chang, Y.T.; Williams, D.R. Novel use of fluorescent glucose analogues to identify a new class of triazine-based insulin mimetics possessing useful secondary effects. Mol. Biosyst. 2011, 7, 346–358. [Google Scholar] [CrossRef]
- Jung, D.W.; Kim, W.H.; Park, S.H.; Lee, J.; Kim, J.; Su, D.; Ha, H.H.; Chang, Y.T.; Williams, D.R. A unique small molecule inhibitor of enolase clarifies its role in fundamental biological processes. ACS Chem. Biol. J. 2013, 8, 1271–1282. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.; Um, J.; Lee, J.H.; Kim, W.H.; Kang, W.S.; Kim, S.H.; Ha, H.H.; Kim, Y.C.; Ahn, Y.K.; Jung, D.W.; et al. ENOblock, a unique small molecule inhibitor of the non-glycolytic functions of enolase, alleviates the symptoms of type 2 diabetes. Sci. Rep. 2017, 7, 44186. [Google Scholar] [CrossRef]
- Shimizu, J.; Murao, A.; Lee, Y.; Aziz, M.; Wang, P. Extracellular CIRP promotes Kupffer cell inflammatory polarization in sepsis. Front. Immunol. 2024, 15, 1411930. [Google Scholar] [CrossRef]
- Tan, Z.; Xie, N.; Banerjee, S.; Cui, H.; Fu, M.; Thannickal, V.J.; Liu, G. The monocarboxylate transporter 4 is required for glycolytic reprogramming and inflammatory response in macrophages. J. Biol. Chem. 2015, 290, 46–55. [Google Scholar] [CrossRef]
- Luo, Y.; Pang, B.; Hao, J.; Li, Q.; Qiao, P.; Zhang, C.; Bai, Y.; Xiao, C.; Chen, J.; Zhi, D.; et al. Keratin 17 covalently binds to alpha-enolase and exacerbates proliferation of keratinocytes in psoriasis. Int. J. Biol. Sci. 2023, 19, 3395–3411. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Shu, X.; Zhang, H.W.; Sun, L.X.; Yu, L.; Liu, J.; Sun, L.C.; Yang, Z.H.; Ran, Y.L. Enolase 1 regulates stem cell-like properties in gastric cancer cells by stimulating glycolysis. Cell Death Dis. 2020, 11, 870. [Google Scholar] [CrossRef]
- Kholmukhamedov, A.; Czerny, C.; Hu, J.; Schwartz, J.; Zhong, Z.; Lemasters, J.J. Minocycline and doxycycline, but not tetracycline, mitigate liver and kidney injury after hemorrhagic shock/resuscitation. Shock 2014, 42, 256–263. [Google Scholar] [CrossRef] [PubMed]
- Petrak, J.; Ivanek, R.; Toman, O.; Cmejla, R.; Cmejlova, J.; Vyoral, D.; Zivny, J.; Vulpe, C.D. Deja vu in proteomics. A hit parade of repeatedly identified differentially expressed proteins. Proteomics 2008, 8, 1744–1749. [Google Scholar] [CrossRef]
- Jung, D.W.; Kim, W.H.; Williams, D.R. Chemical genetics and its application to moonlighting in glycolytic enzymes. Biochem. Soc. Trans. 2014, 42, 1756–1761. [Google Scholar] [CrossRef] [PubMed]
- Baik, S.H.; Kang, S.; Lee, W.; Choi, H.; Chung, S.; Kim, J.I.; Mook-Jung, I. A Breakdown in Metabolic Reprogramming Causes Microglia Dysfunction in Alzheimer’s Disease. Cell Metab. 2019, 30, 493–507.e496. [Google Scholar] [CrossRef]
- Guo, C.; Islam, R.; Zhang, S.; Fang, J. Metabolic reprogramming of macrophages and its involvement in inflammatory diseases. EXCLI J. 2021, 20, 628–641. [Google Scholar]
- Orihuela, R.; McPherson, C.A.; Harry, G.J. Microglial M1/M2 polarization and metabolic states. Br. J. Pharmacol. 2016, 173, 649–665. [Google Scholar] [CrossRef]
- Fan, N.; Zhang, X.; Zhao, W.; Zhao, J.; Luo, D.; Sun, Y.; Li, D.; Zhao, C.; Wang, Y.; Zhang, H.; et al. Covalent Inhibition of Pyruvate Kinase M2 Reprograms Metabolic and Inflammatory Pathways in Hepatic Macrophages against Non-alcoholic Fatty Liver Disease. Int. J. Biol. Sci. 2022, 18, 5260–5275. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Bai, J.; Lu, P.; Jing, Y.M.; Zheng, W.C.; Wang, L.Y.; Wang, J.H.; Wang, F. Hirudin ameliorates myocardial ischemia-reperfusion injury in a rat model of hemorrhagic shock and resuscitation: Roles of NLRP3-signaling pathway. Mol. Cell. Biochem. 2024, 479, 63–72. [Google Scholar] [CrossRef]
- Sanman, L.E.; Qian, Y.; Eisele, N.A.; Ng, T.M.; van der Linden, W.A.; Monack, D.M.; Weerapana, E.; Bogyo, M. Disruption of glycolytic flux is a signal for inflammasome signaling and pyroptotic cell death. eLife 2016, 5, e13663. [Google Scholar] [CrossRef]
- Xie, M.; Yu, Y.; Kang, R.; Zhu, S.; Yang, L.; Zeng, L.; Sun, X.; Yang, M.; Billiar, T.R.; Wang, H.; et al. PKM2-dependent glycolysis promotes NLRP3 and AIM2 inflammasome activation. Nat. Commun. 2016, 7, 13280, Erratum in Nat. Commun. 2025, 16, 6158. [Google Scholar] [CrossRef]
- Yu, Q.; Guo, M.; Zeng, W.; Zeng, M.; Zhang, X.; Zhang, Y.; Zhang, W.; Jiang, X.; Yu, B. Interactions between NLRP3 inflammasome and glycolysis in macrophages: New insights into chronic inflammation pathogenesis. Immun. Inflamm. Dis. 2022, 10, e581. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.; Yu, H.; Peng, T.; Yang, K.; Xu, X.; Gu, W. Glycolytic enzyme PGK1 promotes M1 macrophage polarization and induces pyroptosis of acute lung injury via regulation of NLRP3. Respir. Res. 2024, 25, 291. [Google Scholar] [CrossRef]
- Rao, T.; Yang, W.; Ma, X.; Jiang, X.; Jiang, S.; Xu, S. Bergapten attenuates hemorrhagic shock induced multi-organ injury by inhibiting NLRP3 inflammasome activation and pyroptosis. Int. Immunopharmacol. 2024, 140, 112839. [Google Scholar] [CrossRef]
- Du, S.; Zhang, X.; Jia, Y.; Peng, P.; Kong, Q.; Jiang, S.; Li, Y.; Li, C.; Ding, Z.; Liu, L. Hepatocyte HSPA12A inhibits macrophage chemotaxis and activation to attenuate liver ischemia/reperfusion injury via suppressing glycolysis-mediated HMGB1 lactylation and secretion of hepatocytes. Theranostics 2023, 13, 3856–3871. [Google Scholar] [CrossRef] [PubMed]
- Tong, G.; Chen, Y.; Chen, X.; Fan, J.; Zhu, K.; Hu, Z.; Li, S.; Zhu, J.; Feng, J.; Wu, Z.; et al. FGF18 alleviates hepatic ischemia-reperfusion injury via the USP16-mediated KEAP1/Nrf2 signaling pathway in male mice. Nat. Commun. 2023, 14, 6107. [Google Scholar] [CrossRef]
- Zhou, H.; Li, L.; Sun, H.; Li, H.; Wu, Y.; Zhang, X.; Zhang, J. Remote Ischemic Preconditioning Attenuates Hepatic Ischemia/Reperfusion Injury after Hemorrhagic Shock by Increasing Autophagy. Int. J. Med. Sci. 2021, 18, 873–882. [Google Scholar] [CrossRef]
- Wen, Y.; Ju, C. New insights into liver injury and regeneration from single-cell transcriptomics. eGastroenterology 2025, 3, e100202. [Google Scholar] [CrossRef] [PubMed]
- Ryans, K.; Omosun, Y.; McKeithen, D.N.; Simoneaux, T.; Mills, C.C.; Bowen, N.; Eko, F.O.; Black, C.M.; Igietseme, J.U.; He, Q. The immunoregulatory role of alpha enolase in dendritic cell function during Chlamydia infection. BMC Immunol. 2017, 18, 27. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Cagliani, J.; Aziz, M.; Tan, C.; Brenner, M.; Wang, P. Extracellular CIRP activates STING to exacerbate hemorrhagic shock. JCI Insight. 2021, 6, e143715. [Google Scholar] [CrossRef]
- Hu, Z.; Li, J.; Zhang, F.; Jacob, A.; Wang, P. A novel oligonucleotide mRNA mimic attenuates hemorrhage-induced acute lung injury. Shock 2024, 61, 630–637. [Google Scholar] [CrossRef] [PubMed]
- Li, P.Z.; Li, J.Z.; Li, M.; Gong, J.P.; He, K. An efficient method to isolate and culture mouse Kupffer cells. Immunol. Lett. 2014, 158, 52–56. [Google Scholar] [CrossRef]
- Borjas, T.; Jacob, A.; Yen, H.; Patel, V.; Coppa, G.F.; Aziz, M.; Wang, P. Inhibition of the Interaction of TREM-1 and eCIRP Attenuates Inflammation and Improves Survival in Hepatic Ischemia/Reperfusion. Shock 2022, 57, 246–255. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hu, Z.; Li, J.; Rashid, N.; Jacob, A.; Wang, P. Targeting Kupffer Cell Enolase 1 Attenuates Liver Inflammation and Injury in Hemorrhagic Shock. Int. J. Mol. Sci. 2025, 26, 8340. https://doi.org/10.3390/ijms26178340
Hu Z, Li J, Rashid N, Jacob A, Wang P. Targeting Kupffer Cell Enolase 1 Attenuates Liver Inflammation and Injury in Hemorrhagic Shock. International Journal of Molecular Sciences. 2025; 26(17):8340. https://doi.org/10.3390/ijms26178340
Chicago/Turabian StyleHu, Zhijian, Jingsong Li, Naureen Rashid, Asha Jacob, and Ping Wang. 2025. "Targeting Kupffer Cell Enolase 1 Attenuates Liver Inflammation and Injury in Hemorrhagic Shock" International Journal of Molecular Sciences 26, no. 17: 8340. https://doi.org/10.3390/ijms26178340
APA StyleHu, Z., Li, J., Rashid, N., Jacob, A., & Wang, P. (2025). Targeting Kupffer Cell Enolase 1 Attenuates Liver Inflammation and Injury in Hemorrhagic Shock. International Journal of Molecular Sciences, 26(17), 8340. https://doi.org/10.3390/ijms26178340






