Disease-Associated Shifts in Minor T Cell Subpopulations Define Distinct Immunopathology in HBV vs. HCV Infection
Abstract
1. Introduction
2. Results
3. Discussion
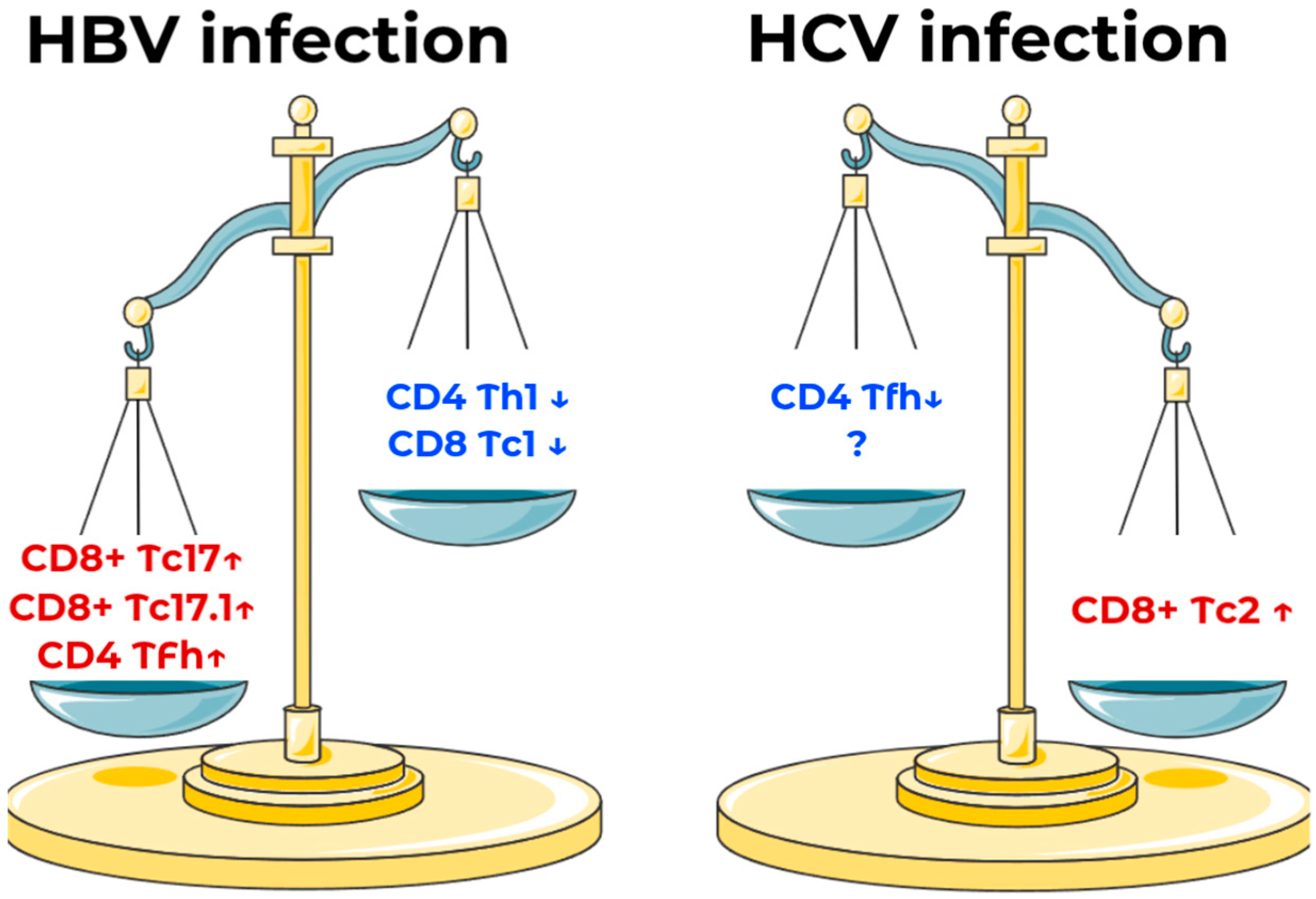
3.1. Changes in T Cellular Responses Associated with HCV Infection
3.2. Changes in T Cellular Responses Associated with HBV Infection
3.3. Disease-Specific Shifts in Immune Cell Populations: HBV vs. HCV
3.4. Study Limitations and Future Perspective
4. Materials and Methods
4.1. Studied Cohorts
4.2. Sample Collection
4.3. T Cell Immunophenotyping by Flow Cytometry
4.4. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ALT | Alanine transaminase |
| AST | Aspartate transaminase |
| CM | Central memory |
| CTL | Cytotoxic T cells |
| EM | Effector memory |
| HBV | Hepatitis B virus |
| HCV | Hepatitis C virus |
| IQR | Interquartile range |
| Mcmol/L | Micromole per liter |
| Me | Median |
| Tc | Cytokine-producing cytotoxic T cells |
| TEMRA | Terminally differentiated effector memory T cells |
| Tfh | T follicular helper cells |
| Th | Helper T cells |
| U/L | Units per liter |
| WHO | World Health Organization |
References
- World Health Organization (WHO). Chronic Viral Hepatitis. Available online: https://www.who.int/data/gho/data/themes/chronic-viral-hepatitis (accessed on 2 July 2025).
- Xu, Z.Y.; Gao, J.S.; He, Y.; Xiao, X.Q.; Gong, G.Z.; Zhang, M. Hepatitis B virus confers innate immunity evasion through hepatitis B virus-miR-3 down-regulation of cGAS-Sting-IFN signaling. World J. Hepatol. 2025, 17, 99292. [Google Scholar] [CrossRef]
- Moorman, J.P.; Joo, M.; Hahn, Y.S. Evasion of host immune surveillance by hepatitis C virus: Potential roles in viral persistence. Arch. Immunol. Ther. Exp. 2001, 49, 189–194. [Google Scholar]
- Wang, L.; Zeng, X.; Wang, Z.; Fang, L.; Liu, J. Recent advances in understanding T cell activation and exhaustion during HBV infection. Virol. Sin. 2023, 38, 851–859. [Google Scholar] [CrossRef]
- Andreata, F.; Iannacone, M. The hidden strength of CD8+ T cells in chronic hepatitis B. Nat. Immunol. 2024, 25, 1515–1516. [Google Scholar] [CrossRef]
- Heim, K.; Sagar; Sogukpinar, Ö.; Llewellyn-Lacey, S.; Price, D.A.; Emmerich, F.; Kraft, A.R.M.; Cornberg, M.; Kielbassa, S.; Knolle, P.; et al. Attenuated effector T cells are linked to control of chronic HBV infection. Nat. Immunol. 2024, 25, 1650–1662. [Google Scholar] [CrossRef]
- Shoukry, N.H.; Grakoui, A.; Houghton, M.; Chien, D.Y.; Ghrayeb, J.; Reimann, K.A.; Walker, C.M. Memory CD8+ T cells are required for protection from persistent hepatitis C virus infection. J. Exp. Med. 2003, 197, 1645–1655. [Google Scholar] [CrossRef] [PubMed]
- Lechner, F.; Wong, D.K.; Dunbar, P.R.; Chapman, R.; Chung, R.T.; Dohrenwend, P.; Robbins, G.; Phillips, R.; Klenerman, P.; Walker, B.D. Analysis of successful immune responses in persons infected with hepatitis C virus. J. Exp. Med. 2000, 191, 1499–1512. [Google Scholar] [CrossRef] [PubMed]
- Erickson, A.L.; Kimura, Y.; Igarashi, S.; Eichelberger, J.; Houghton, M.; Sidney, J.; McKinney, D.; Sette, A.; Hughes, A.L.; Walker, C.M. The outcome of hepatitis C virus infection is predicted by escape mutations in epitopes targeted by cytotoxic T lymphocytes. Immunity 2001, 15, 883–895. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Rehermann, B. Immune responses to HCV and other hepatitis viruses. Immunity 2014, 40, 13–24. [Google Scholar] [CrossRef]
- Osburn, W.O.; Fisher, B.E.; Dowd, K.A.; Urban, G.; Liu, L.; Ray, S.C.; Thomas, D.L.; Cox, A.L. Spontaneous control of primary hepatitis C virus infection and immunity against persistent reinfection. Gastroenterology 2010, 138, 315–324. [Google Scholar] [CrossRef]
- Eisa, M.; Gomez-Escobar, E.; Bédard, N.; Abdeltawab, N.F.; Flores, N.; Mazouz, S.; Fieffé-Bédard, A.; Sakayan, P.; Gridley, J.; Abdel-Hakeem, M.S.; et al. Coordinated expansion of memory T follicular helper and B cells mediates spontaneous clearance of HCV reinfection. Front. Immunol. 2024, 15, 1403769. [Google Scholar] [CrossRef] [PubMed]
- Salinas, E.; Boisvert, M.; Upadhyay, A.A.; Bédard, N.; Nelson, S.A.; Bruneau, J.; Derdeyn, C.A.; Marcotrigiano, J.; Evans, M.J.; Bosinger, S.E.; et al. Early T follicular helper cell activity accelerates hepatitis C virus-specific B cell expansion. J. Clin. Investig. 2021, 131, e140590. [Google Scholar] [CrossRef] [PubMed]
- Goronzy, J.J.; Weyand, C.M. Successful and maladaptive T cell aging. Immunity 2017, 46, 364–378. [Google Scholar] [CrossRef]
- Osuch, S.; Metzner, K.J.; Cortés, K.C. Reversal of T cell exhaustion in chronic HCV infection. Viruses 2020, 12, 799. [Google Scholar] [CrossRef] [PubMed]
- Hensel, N.; Gu, Z.; Sagar; Wieland, D.; Jechow, K.; Kemming, J.; Llewellyn-Lacey, S.; Gostick, E.; Sogukpinar, O.; Emmerich, F.; et al. Memory-like HCV-specific CD8+ T cells retain a molecular scar after cure of chronic HCV infection. Nat. Immunol. 2021, 22, 229–239. [Google Scholar] [CrossRef]
- Hofmann, M.; Wieland, D.; Pircher, H.; Thimme, R. Memory vs memory-like: The different facets of CD8+ T-cell memory in HCV infection. Immunol. Rev. 2018, 283, 232–237. [Google Scholar] [CrossRef]
- Sobue, S.; Nomura, T.; Ishikawa, T.; Ito, S.; Saso, K.; Ohara, H.; Joh, T.; Itoh, M.; Kakumu, S. Th1/Th2 cytokine profiles and their relationship to clinical features in patients with chronic hepatitis C virus infection. J. Gastroenterol. 2001, 36, 544–551. [Google Scholar] [CrossRef]
- Yue, M.; Deng, X.; Zhai, X.; Xu, K.; Kong, J.; Zhang, J.; Zhou, Z.; Yu, X.; Xu, X.; Liu, Y.; et al. Th1 and Th2 cytokine profiles induced by hepatitis C virus F protein in peripheral blood mononuclear cells from chronic hepatitis C patients. Immunol. Lett. 2013, 152, 89–95. [Google Scholar] [CrossRef]
- Essa, S.; Siddiqque, I.; Al-Nakib, W.; Raghupathy, R. Relevance of T helper Type 1, 2 and 17 cytokine profiles to the outcome of therapy in hepatitis C patients. J. Antivir. Antiretrovir. 2020, 12, 193. [Google Scholar] [CrossRef]
- Rowan, A.G.; Fletcher, J.M.; Ryan, E.J.; Moran, B.; Hegarty, J.E.; O’Farrelly, C.; Mills, K.H.G. Hepatitis C virus-specific Th17 cells are suppressed by virus-induced TGF-β1. J. Immunol. 2008, 181, 4485–4494. [Google Scholar] [CrossRef]
- Oo, Y.H.; Banz, V.; Kavanagh, D.; Liaskou, E.; Withers, D.R.; Humphreys, E.; Reynolds, G.M.; Lee-Turner, L.; Kalia, N.; Hubscher, S.G.; et al. CXCR3-dependent recruitment and CCR6-mediated positioning of Th-17 cells in the inflamed liver. J. Hepatol. 2012, 57, 1044–1051. [Google Scholar] [CrossRef]
- Semmo, N.; Klenerman, P. CD4+ T cell responses in hepatitis C virus infection. World J. Gastroenterol. 2007, 13, 4831–4838. [Google Scholar] [CrossRef]
- Sallusto, F.; Geginat, J.; Lanzavecchia, A. Central memory and effector memory T cell subsets: Function, generation, and maintenance. Annu. Rev. Immunol. 2004, 22, 745–763. [Google Scholar] [CrossRef]
- Arsentieva, N.A.; Batsunov, O.K.; Lyubimova, N.E.; Basina, V.V.; Esaulenko, E.V.; Totolian, A.A. Cytokine profiling of plasma in patients with viral hepatitis C. Med. Immunol. 2024, 26, 1235–1248. (In Russian) [Google Scholar] [CrossRef]
- Stelma, F.; Willemse, S.B.; Erken, R.; de Niet, A.; Sinnige, M.J.; van Dort, K.; Zaaijer, H.L.; van Leeuwen, E.M.M.; A Kootstra, N.; Reesink, H.W. Dynamics of the Immune Response in Acute Hepatitis B Infection. Open Forum Infect. Dis. 2017, 4, ofx231. [Google Scholar] [CrossRef] [PubMed]
- Iannacone, M.; Guidotti, L.G. Immunobiology and pathogenesis of hepatitis B virus infection. Nat. Rev. Immunol. 2022, 22, 19–32. [Google Scholar] [CrossRef] [PubMed]
- Thimme, R.; Bertoletti, A.; Iannacone, M. Beyond exhaustion: The unique characteristics of CD8+ T cell dysfunction in chronic HBV infection. Nat. Rev. Immunol. 2024, 24, 775–776. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.; Vecchi, A.; Tiezzi, C.; Barili, V.; Fisicaro, P.; Penna, A.; Montali, I.; Daffis, S.; Fletcher, S.P.; Gaggar, A.; et al. Phenotypic CD8 T cell profiling in chronic hepatitis B to predict HBV-specific CD8 T cell susceptibility to functional restoration in vitro. Gut 2023, 72, 2123–2137. [Google Scholar] [CrossRef]
- Chen, X.; Liu, X.; Jiang, Y.; Xia, N.; Liu, C.; Luo, W. Abnormally primed CD8 T cells: The Achilles’ heel of CHB. Front. Immunol. 2023, 14, 1106700. [Google Scholar] [CrossRef]
- Huang, Y.; Chen, Z.; Jia, H.; Wu, W.; Zhong, S.; Zhou, C. Induction of Tc1 response and enhanced cytotoxic T lymphocyte activity in mice by dendritic cells transduced with adenovirus expressing HBsAg. Clin. Immunol. 2006, 119, 280–290. [Google Scholar] [CrossRef]
- Vanwolleghem, T.; Adomati, T.; Van Hees, S.; Janssen, H.L.A. Humoral immunity in hepatitis B virus infection: Rehabilitating the B in HBV. JHEP Rep. 2021, 4, 100398. [Google Scholar] [CrossRef]
- Svicher, V.; Salpini, R.; D’aNna, S.; Piermatteo, L.; Iannetta, M.; Malagnino, V.; Sarmati, L. New insights into hepatitis B virus lymphotropism: Implications for HBV-related lymphomagenesis. Front Oncol. 2023, 13, 1143258. [Google Scholar] [CrossRef]
- Batsunov, O.K.; Arsentieva, N.A.; Lyubimova, N.E.; Basina, V.V.; Esaulenko, E.V.; Totolian, A.A. Cytokine Levels in Blood Plasma of Patients with Chronic Viral Hepatitis B at Various Stages of Liver Fibrosis. Russ. J. Infect. Immun. 2025, 15, 3. (In Russian) [Google Scholar]
- Liang, Y.; Pan, H.F.; Ye, D.Q. Tc17 Cells in Immunity and Systemic Autoimmunity. Int. Rev. Immunol. 2015, 34, 318–331. [Google Scholar] [CrossRef] [PubMed]
- Koh, C.H.; Lee, S.; Kwak, M.; Kim, B.S.; Chung, Y. CD8 T-cell subsets: Heterogeneity, functions, and therapeutic potential. Exp. Mol. Med. 2023, 55, 2287–2299. [Google Scholar] [CrossRef]
- Wang, X.; Dong, Q.; Li, Q.; Li, Y.; Zhao, D.; Sun, J.; Fu, J.; Meng, F.; Lin, H.; Tian, Z. Dysregulated Response of Follicular Helper T Cells to Hepatitis B Surface Antigen Promotes HBV Persistence in Mice and Associates With Outcomes of Patients. Gastroenterology 2018, 154, 2222–2236. [Google Scholar] [CrossRef] [PubMed]
- Golovkin, A.; Kalinina, O.; Bezrukikh, V.; Aquino, A.; Zaikova, E.; Karonova, T.; Melnik, O.; Vasilieva, E.; Kudryavtsev, I. Imbalanced immune response of T-cell and B-cell subsets in patients with moderate and severe COVID-19. Viruses 2021, 13, 1966. [Google Scholar] [CrossRef]
- Kudryavtsev, I.V.; Arsentieva, N.A.; Korobova, Z.R.; Isakov, D.V.; Rubinstein, A.A.; Batsunov, O.K.; Khamitova, I.V.; Kuznetsova, R.N.; Savin, T.V.; Akisheva, T.V.; et al. Heterogenous CD8+ T cell maturation and ‘polarization’ in acute and convalescent COVID-19 patients. Viruses 2022, 14, 1906. [Google Scholar] [CrossRef]
| HCV (1) | HBV (2) | HD (3) | p-Value (Kruskal–Wallis) | ||
|---|---|---|---|---|---|
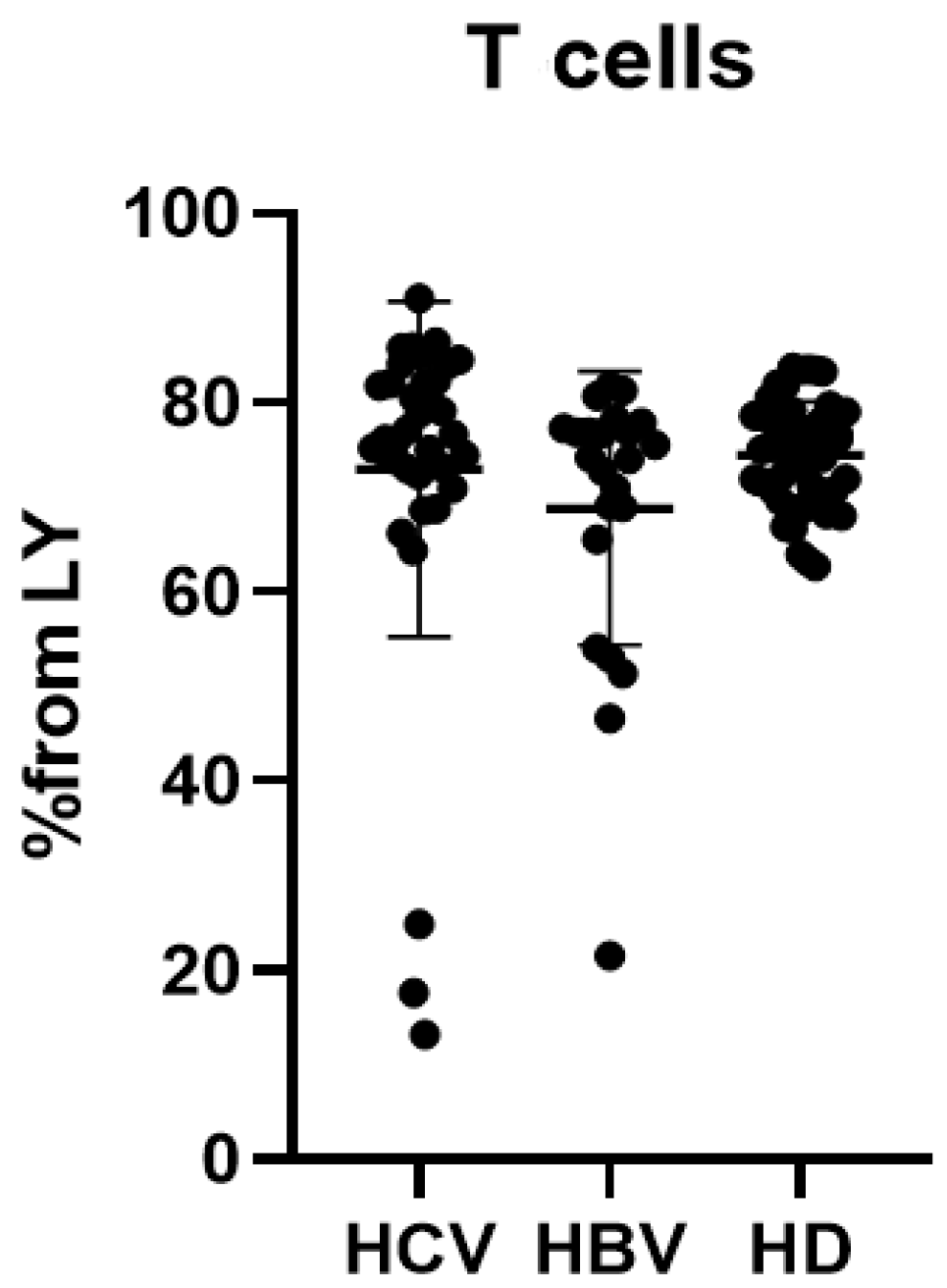
| T cell | % | 76.4 (72.54; 82.23) | 74.22 (65.58; 77.3) | 75.09 (70.42; 78.71) | ns |
| abs | 1.64 (1.25; 1.86) | 0.98 (0.69; 1.61) | 1.25 (1.06; 1.60) | p (1–2) = 0.0103 |
| HCV (1) | HBV (2) | HD (3) | p-Value (Kruskal–Wallis) | ||
|---|---|---|---|---|---|
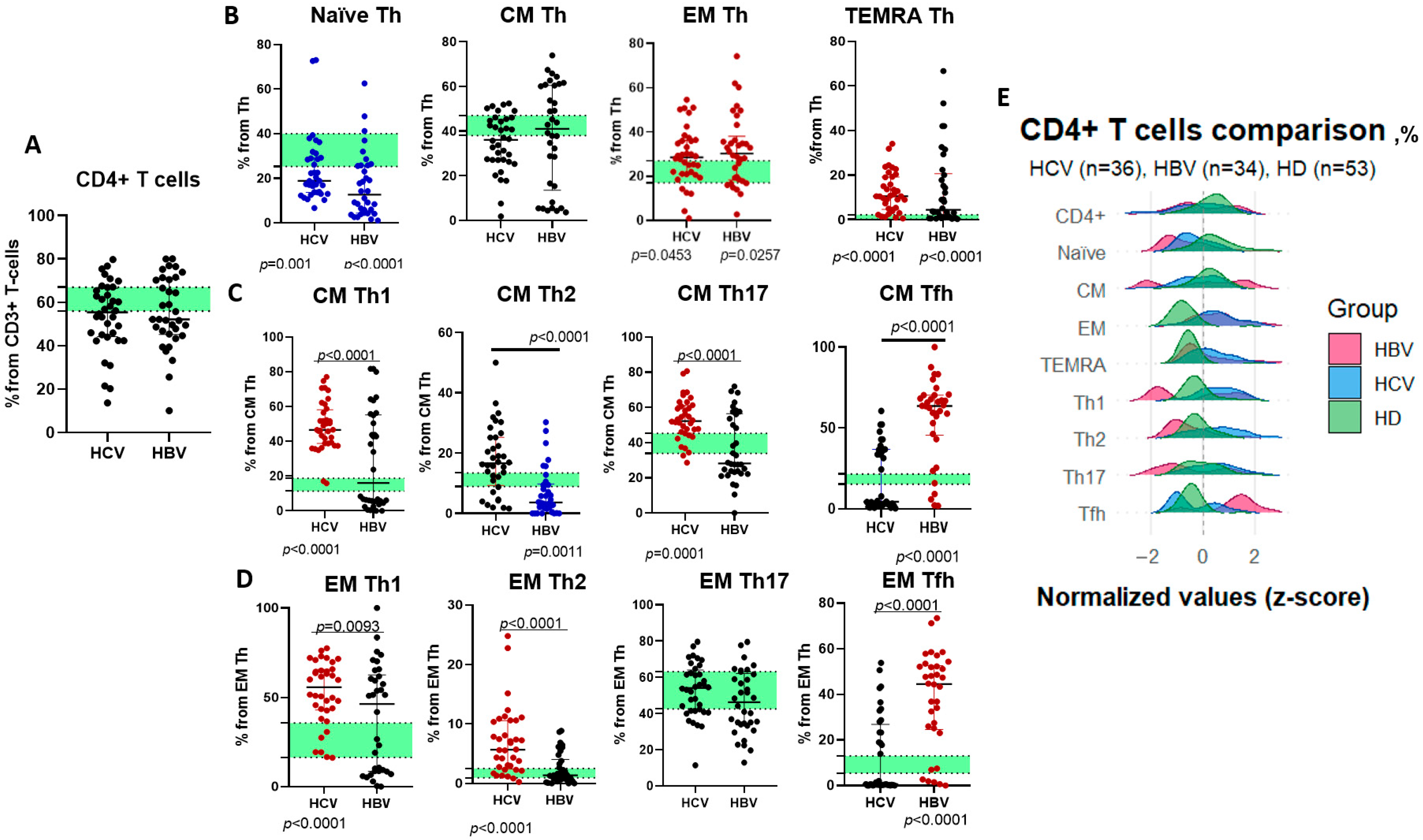
| CD4+ T cell | % | 55.46 (44.27; 66.55) | 52.23 (45.28; 70.79) | 62.94 (56.2; 66.82) | ns |
| abs | 0.60 (0.25; 0.99) | 0.31 (0.10; 0.59) | 0.77 (0.67; 1.07) | p (1–3) = 0.0338 p (2–3) < 0.0001 | |
| Naïve Th | % | 18.86 (13.71; 29.18) | 12.68 (4.465; 25.83) | 31.3 (25.3; 39.8) | p (1–3) = 0.001 p (2–3) < 0.0001 |
| abs | 0.12 (0.04; 0.20) | 0.0334 (0.01; 0.08) | 0.22 (0.19; 0.33) | p (1–2) = 0.0327 p (1–3) = 0.0016 p (2–3) < 0.0001 | |
| CM Th | % | 36.13 (27.1; 45.24) | 41.06 (13.6; 60.52) | 41.55 (37.66; 47.33) | ns |
| abs | 0.17 (0.08; 0.33) | 0.12 (0.06; 0.37) | 0.33 (0.27; 0.47) | p (1–3) = 0.0016 p (2–3) = 0.0012 | |
| EM Th | % | 28.59 (21.0; 36.32) | 30.33 (18.42; 38.06) | 22.94 (17.28; 27.45) | p (1–3) = 0.0453 p (2–3) = 0.0257 |
| abs | 0.19 (0.05; 0.20) | 0.07 (0.05; 0.17) | 0.18 (0.14; 0.23) | p (1–3) = 0.0212 p (2–3) = 0.0003 | |
| TEMRA Th | % | 10.57 (4.67; 19.23) | 4.505 (1.09; 20.68) | 0.68 (0.27; 2.58) | p (1–3) < 0.0001 p (2–3) < 0.0001 |
| abs | 0.03 (0.01; 0.09) | 0.01 (0.01; 0.03) | 0.001 (0.02; 0.02) | p (1–2) = 0.0151 p (1–3) < 0.0001 | |
| CM Th1 | % | 46.67 (38.79; 58.21) | 14.81 (11.4; 18.8) | 14.81 (11.4; 18.8) | p (1–2) < 0.0001 p (1–3) < 0.0001 |
| abs | 0.07 (0.03; 0.15) | 0.01 (0.01; 0.02) | 0.05 (0.03; 0.06) | p (1–2) < 0.0001 p (2–3) < 0.0001 | |
| CM Th2 | % | 16.61 (9.28; 25.22) | 10.72 (8.92; 13.4) | 10.72 (8.92; 13.4) | p (1–2) < 0.0001 p (2–3) = 0.0011 |
| abs | 0.02 (0.01; 0.07) | 0.02 (0.01; 0.02) | 0.04 (0.03; 0.05) | p (1–2) = 0.0005 p (2–3) < 0.0001 | |
| CM Th17 | % | 52.32 (45.42; 61.1) | 39.23 (33.93; 45.26) | 39.23 (33.93; 45.26) | p (1–2) < 0.0001 p (1–3) = 0.0001 |
| abs | 0.08 (0.04; 0.18) | 0.03 (0.01; 0.09) | 0.12 (0.09; 0.17) | p (1–2) = 0.0343 p (2–3) < 0.0001 | |
| CM Tfh | % | 4.38 (2.63; 36.77) | 17.87 (15.23; 21.63) | 17.87 (15.23; 21.63) | p (1–2) < 0.0001 p (2–3) < 0.0001 |
| abs | 0.02 (0.01; 0.07) | 0.07 (0.01; 0.19) | 0.06 (0.05; 0.08) | p (1–2) = 0.0073 p (1–3) = 0.0044 | |
| EM Th1 | % | 55.77 (42.93; 65.67) | 21.7 (16.41; 35.74) | 21.7 (16.41; 35.74) | p (1–2) = 0.0093 p (1–3) < 0.0001 |
| abs | 0.04 (0.02; 0.13) | 0.01 (0.01; 0.03) | 0.04 (0.02; 0.06) | p (1–2) = 0.0002 p (2–3) = 0.0001 | |
| EM Th2 | % | 5.67 (2.43; 10.56) | 1.45 (0.93; 2.47) | 1.45 (0.93; 2.47) | p (1–2) < 0.0001 p (1–3) < 0.0001 |
| abs | 0.01 (0.01; 0.02) | 0.01 (0.002; 0.03) | 0.01 (0.0016; 0.03) | p (1–2) < 0.0001 p (1–3) = 0.0098 p (2–3) = 0.0483 | |
| EM Th17 | % | 54.17 (41.47; 64.16) | 55.45 (42.56; 63.19) | 55.45 (42.56; 63.19) | ns |
| abs | 0.05 (0.02; 0.14) | 0.03 (0.01; 0.07) | 0.10 (0.07; 0.13) | p (2–3) = 0.0001 | |
| EM Tfh | % | 0.65 (0.30; 26.88) | 8.29 (5.39; 13.14) | 8.29 (5.39; 13.14) | p (1–2) < 0.0001 p (2–3) < 0.0001 |
| abs | 0.02 (0.0001; 0.03) | 0.03 (0.02; 0.04) | 0.02 (0.01; 0.03) | p (1–2) < 0.0001 p (1–3) = 0.0478 p (2–3) = 0.0495 |
| HCV (1) | HBV (2) | HD (3) | p-Value (Kruskal–Wallis) | ||
|---|---|---|---|---|---|
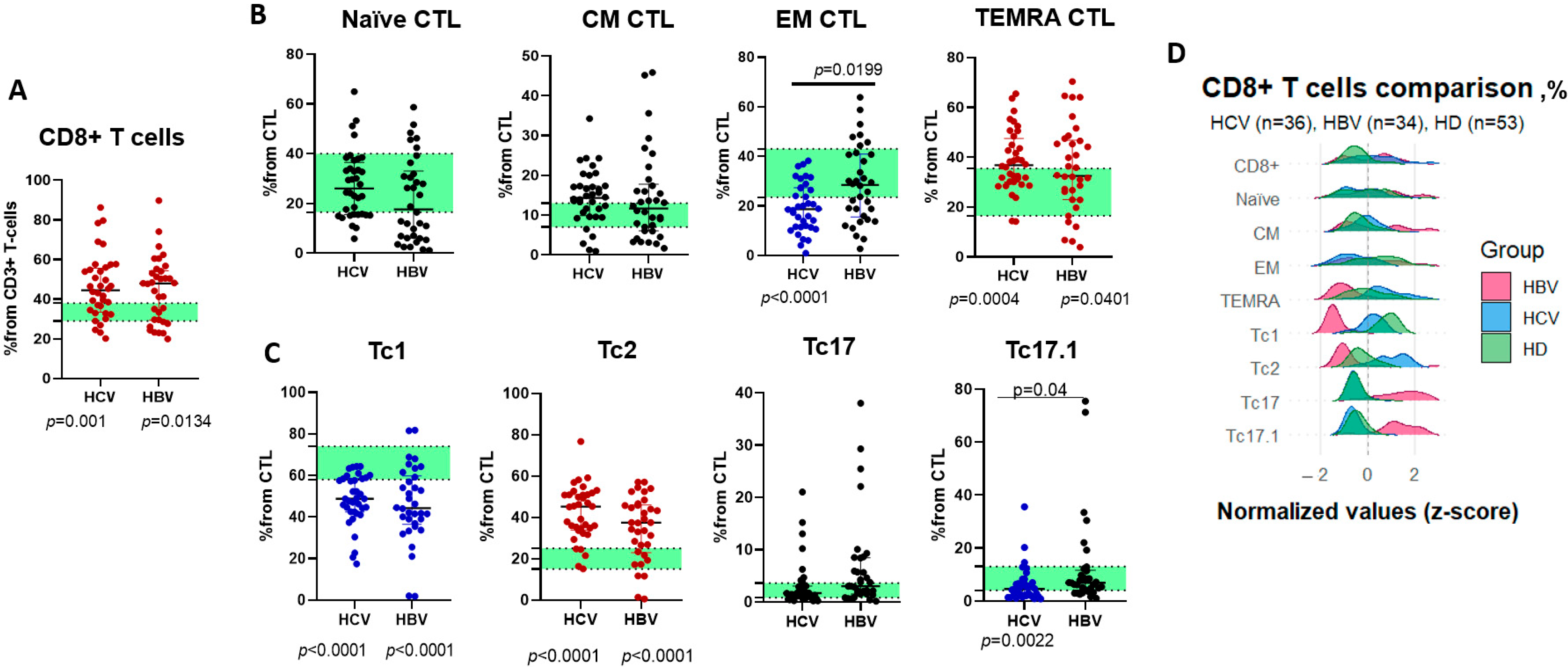
| CD8+ T cell | % | 44.55 (33.45; 55.73) | 47.93 (29.38; 54.4) | 33.34 (28.94; 38.26) | p (1–3) = 0.001 p (2–3) = 0.0134 |
| abs | 0.50 (0.27; 0.72) | 0.18 (0.09; 0.34) | 0.44 (0.35; 0.53) | p (1–2) = 0.0026 p (2–3) = 0.0012 | |
| Naïve CTL | % | 26.06 (15.55; 36.53) | 17.61 (6.093; 33.03) | 29.43 (16.6; 39.61) | ns |
| abs | 0.10 (0.06; 0.21) | 0.02 (0.01; 0.08) | 0.12 (0.07; 0.15) | p (1–2) = 0.0011 p (2–3) = 0.0008 | |
| CM CTL | % | 14.3 (9.88; 17.36) | 11.71 (6.05; 17.79) | 11.0 (6.99; 13.45) | ns |
| abs | 0.05 (0.02; 0.10) | 0.05 (0.01; 0.06) | 0.04 (0.03; 0.07) | ns | |
| EM CTL | % | 18.73 (11.56; 27.38) | 28.52 (15.59; 40.92) | 34.13 (23.53; 42.8) | p (1–2) = 0.0199 p (1–3) < 0.0001 |
| abs | 0.06 (0.03; 0.11) | 0.04 (0.03; 0.10) | 0.15 (0.10; 0.20) | p (1–3) = 0.0002 p (2–3) = 0.0001 | |
| TEMRA CTL | % | 36.8 (30.13; 47.58) | 32.51 (22.95; 46.81) | 25.18 (16.48; 35.55) | p (1–3) = 0.0004 p (2–3) = 0.0401 |
| abs | 0.19 (0.10; 0.31) | 0.05 (0.02; 0.14) | 0.11 (0.06; 0.17) | p (1–2) = 0.0002 p (2–3) = 0.0279 | |
| Tc1 | % | 48.76 (42.49; 58.44) | 44.29 (36.51; 59.79) | 68.62 (57.78; 74.34) | p (1–3) < 0.0001 p (2–3) < 0.0001 |
| abs | 0.24 (0.13; 0.35) | 0.10 (0.04; 0.21) | 0.30 (0.24; 0.37) | p (2–3) < 0.0001 | |
| Tc2 | % | 45.28 (33.94; 51.05) | 37.45 (22.98; 46.06) | 18.5 (14.78; 25.53) | p (1–3) < 0.0001 p (2–3) < 0.0001 |
| abs | 0.22 (0.11; 0.35) | 0.06 (0.03; 0.14) | 0.09 (0.06; 0.12) | p (1–2) < 0.0001 p (1–3) = 0.0003 | |
| Tc17 | % | 1.685 (0.83; 3.05) | 3.03 (1.523; 8.448) | 1.8 (0.845; 3.665) | ns |
| abs | 0.01 (0.01; 0.02) | 0.01 (0.01; 0.02) | 0.01 (0.01; 0.02) | ns | |
| Tc17.1 | % | 4.60 (2.04; 6.88) | 6.89 (4.10; 11.64) | 8.41 (4.00; 12.94) | p (1–2) = 0.0406 p (1–3) = 0.0022 |
| abs | 0.01 (0.01; 0.04) | 0.01 (0.01; 0.03) | 0.03 (0.02; 0.05) | p (1–3) = 0.0104 p (2–3) = 0.0022 |
| Group | Age (Me, Q25–Q75) | Sex (%) | Fibrosis Stage (%) | ALT Levels (U/L, Me, Q25–Q75) | AST Levels (U/L, Me, Q25–Q75) | Bilirubin (mcmol/L, Me, Q25–Q75) |
|---|---|---|---|---|---|---|
| Chronic HBV | 48, 36–66 | 29.4% male vs. 70.6% female | F0–F1—32.3% F2–F3—5% F4—55.8% | 24, 18.5–37.6 | 29, 20.3–44.9 | 16.5 (12.2–29) |
| Chronic HCV | 52, 39–67 | 41.6% male vs. 58.4% female | F0–F1—28% F2–F3—17% F4—11% 44% unspecified | 57, 29.3–95.5 | 45, 30.3–93.3 | 32.5, 10–92.8 |
| Healthy donors | 40, 34–49 | 43% male vs. 57% female | - | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Korobova, Z.R.; Arsentieva, N.A.; Butenko, A.A.; Batsunov, O.K.; Lyubimova, N.E.; Ostankova, Y.V.; Anufrieva, E.V.; Maslov, S.A.; Kozlov, K.V.; Sulima, D.L.; et al. Disease-Associated Shifts in Minor T Cell Subpopulations Define Distinct Immunopathology in HBV vs. HCV Infection. Int. J. Mol. Sci. 2025, 26, 7761. https://doi.org/10.3390/ijms26167761
Korobova ZR, Arsentieva NA, Butenko AA, Batsunov OK, Lyubimova NE, Ostankova YV, Anufrieva EV, Maslov SA, Kozlov KV, Sulima DL, et al. Disease-Associated Shifts in Minor T Cell Subpopulations Define Distinct Immunopathology in HBV vs. HCV Infection. International Journal of Molecular Sciences. 2025; 26(16):7761. https://doi.org/10.3390/ijms26167761
Chicago/Turabian StyleKorobova, Zoia R., Natalia A. Arsentieva, Anastasia A. Butenko, Oleg K. Batsunov, Natalia E. Lyubimova, Yulia V. Ostankova, Ekaterina V. Anufrieva, Sergey A. Maslov, Konstantin V. Kozlov, Dmitrii L. Sulima, and et al. 2025. "Disease-Associated Shifts in Minor T Cell Subpopulations Define Distinct Immunopathology in HBV vs. HCV Infection" International Journal of Molecular Sciences 26, no. 16: 7761. https://doi.org/10.3390/ijms26167761
APA StyleKorobova, Z. R., Arsentieva, N. A., Butenko, A. A., Batsunov, O. K., Lyubimova, N. E., Ostankova, Y. V., Anufrieva, E. V., Maslov, S. A., Kozlov, K. V., Sulima, D. L., Rishnyak, O. Y., & Totolian, A. A. (2025). Disease-Associated Shifts in Minor T Cell Subpopulations Define Distinct Immunopathology in HBV vs. HCV Infection. International Journal of Molecular Sciences, 26(16), 7761. https://doi.org/10.3390/ijms26167761








