The RopGEF Gene Family and Their Potential Roles in Responses to Abiotic Stress in Brassica rapa
Abstract
1. Introduction
2. Results
2.1. Identification of RopGEF Genes and Analysis of the Physical–Chemical Properties and Subcellular Localization of Their Encoded Proteins
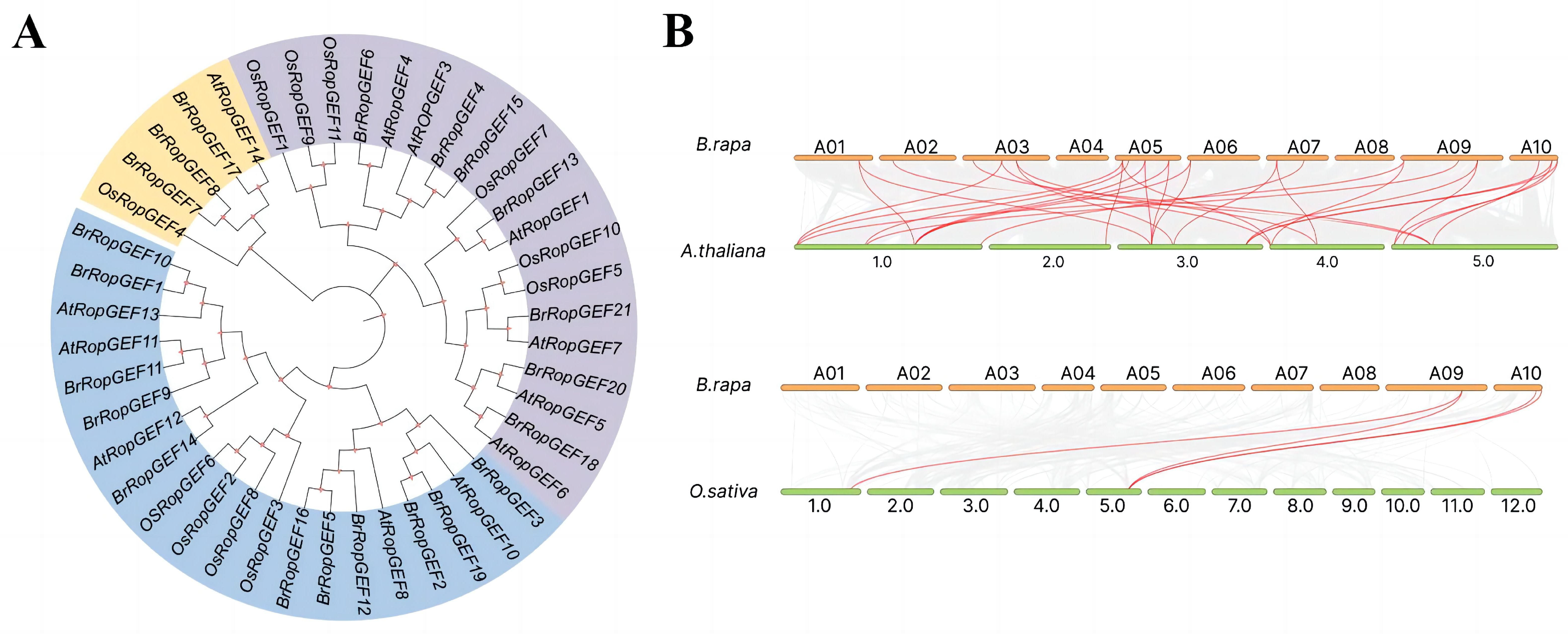
2.2. Chromosomal Mapping Analysis of BrRopGEF Genes in B. rapa
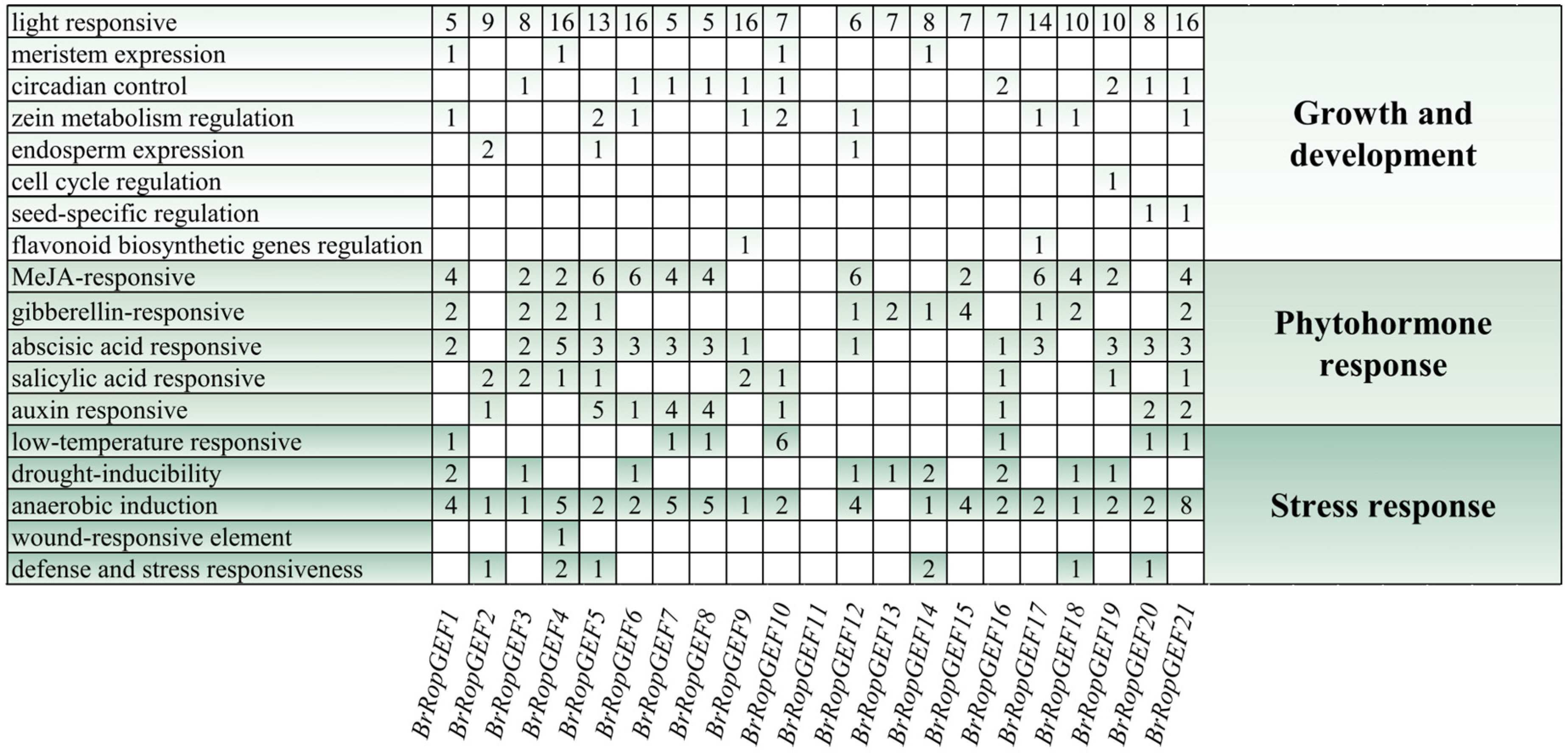
2.3. Analysis of Promoter Cis-Regulatory Elements of RopGEFs in B. rapa
2.4. Analysis of BrRopGEF Gene Structure and the Domain Distribution of Their Encoded Proteins
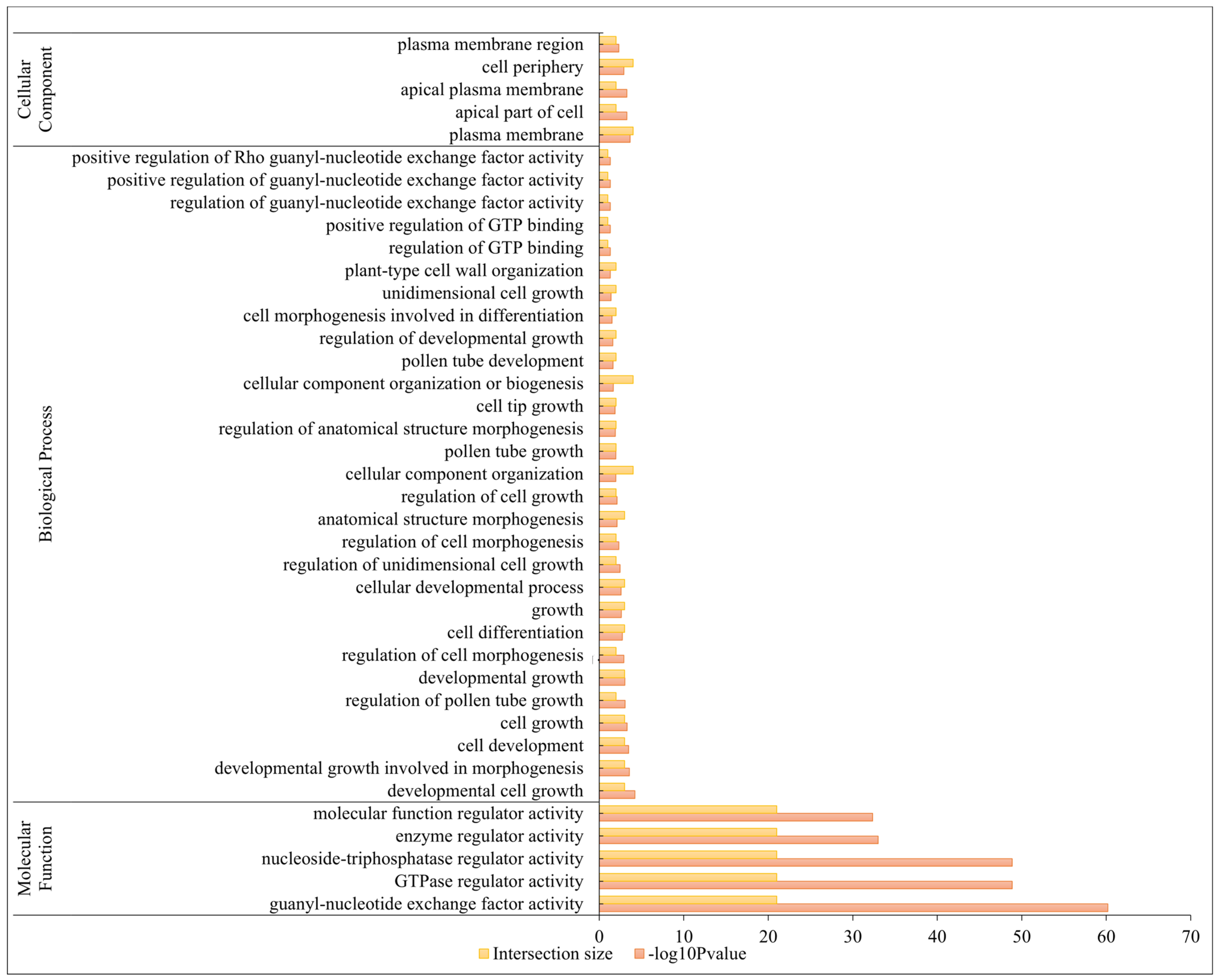
2.5. Gene Ontology (GO) Annotation Analysis of BrRopGEFs
2.6. Gene Expression Analysis of BrRopGEFs
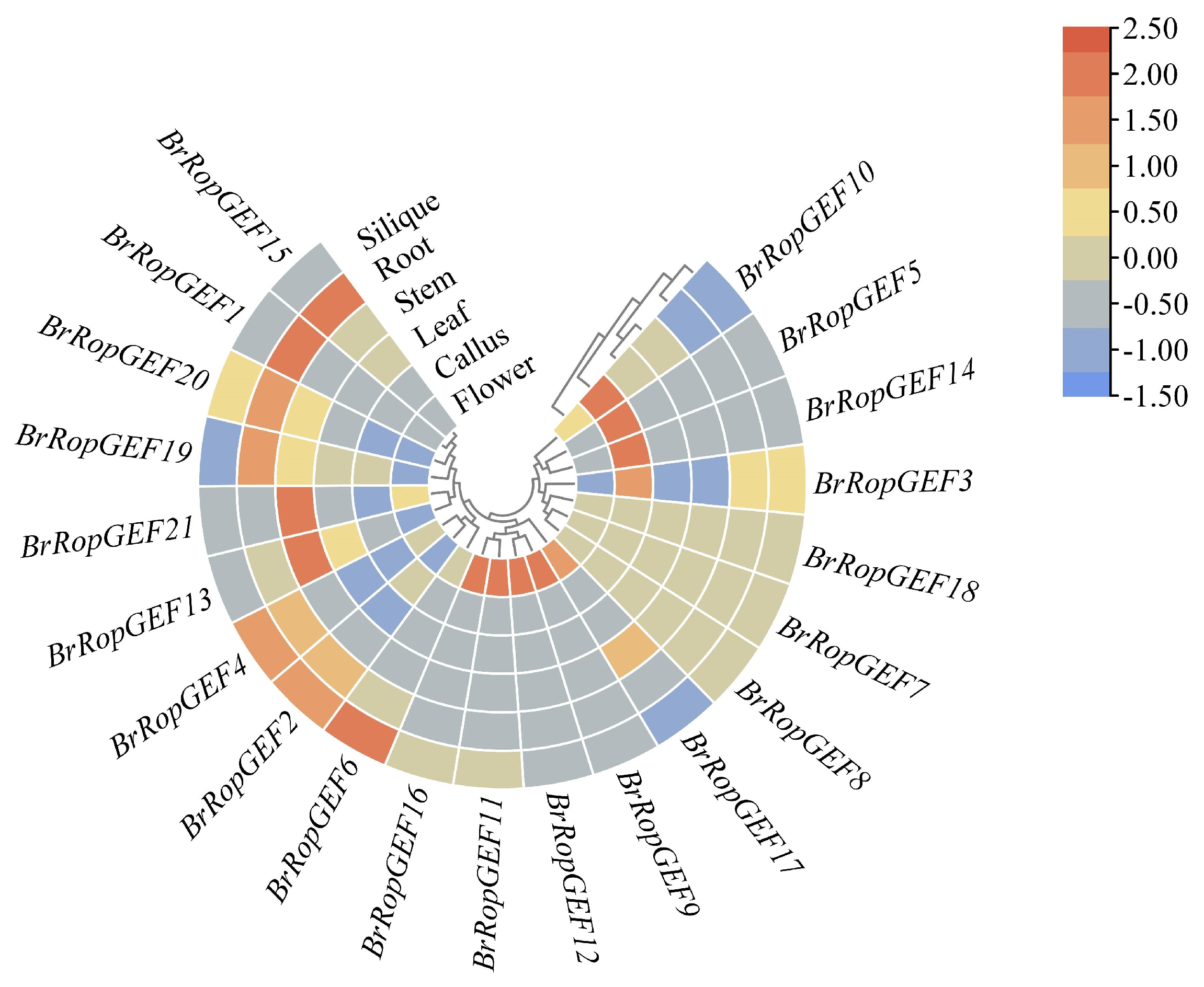
2.6.1. Analysis of the Organ-Specific Expression of BrRopGEF Genes in Organs
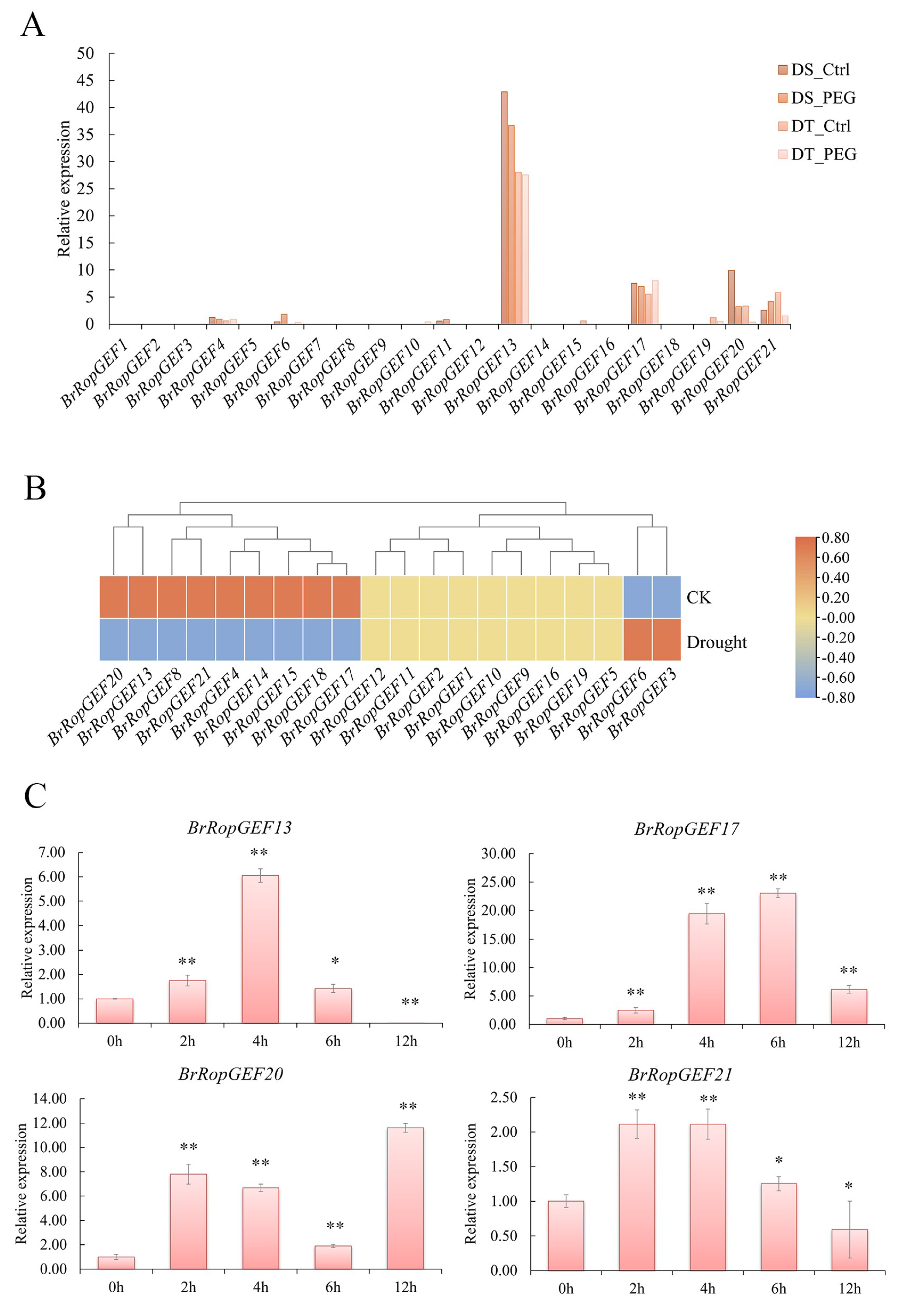
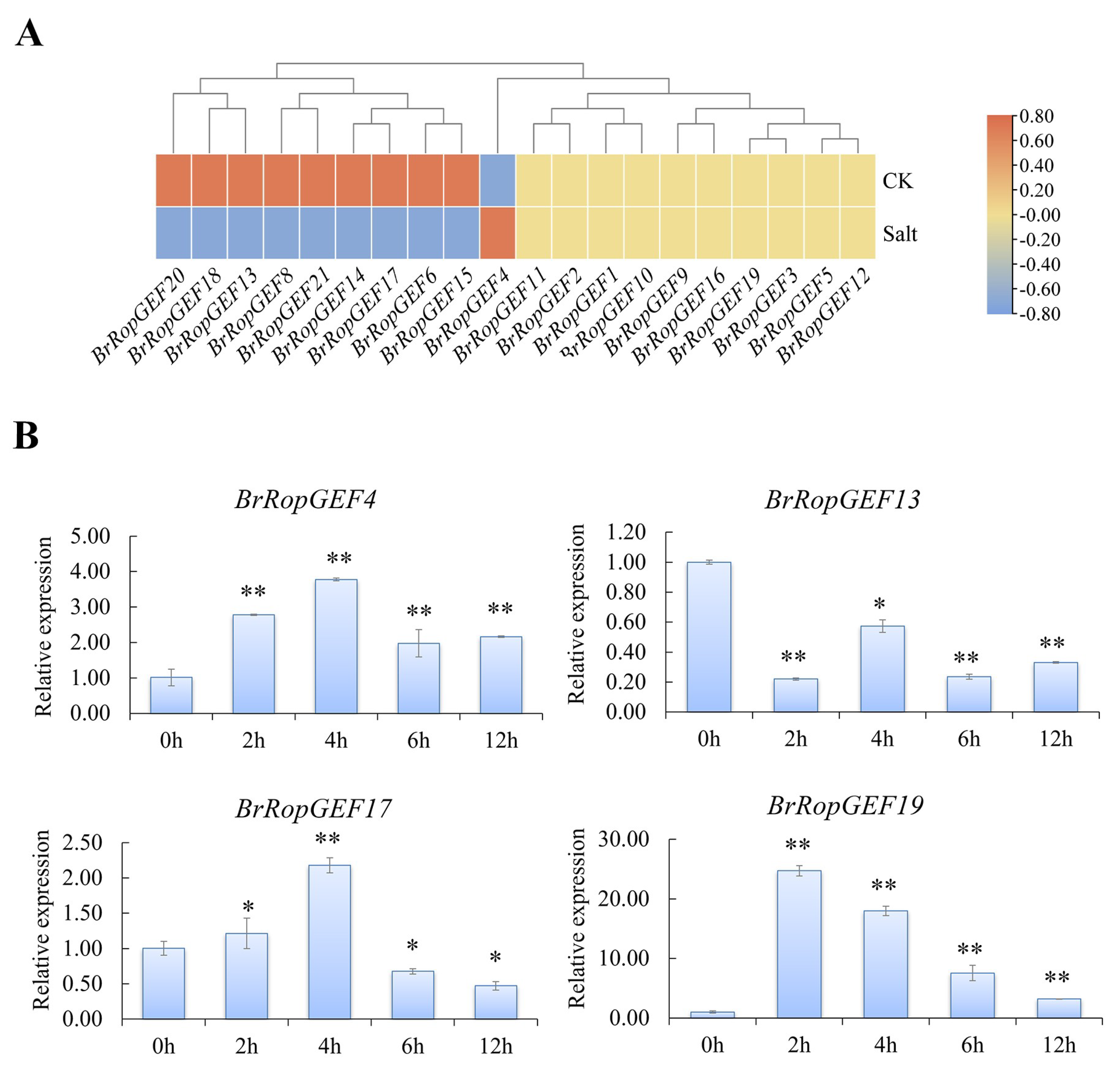
2.6.2. Analysis of BrRopGEFs Involved in Responses to Abiotic Stress
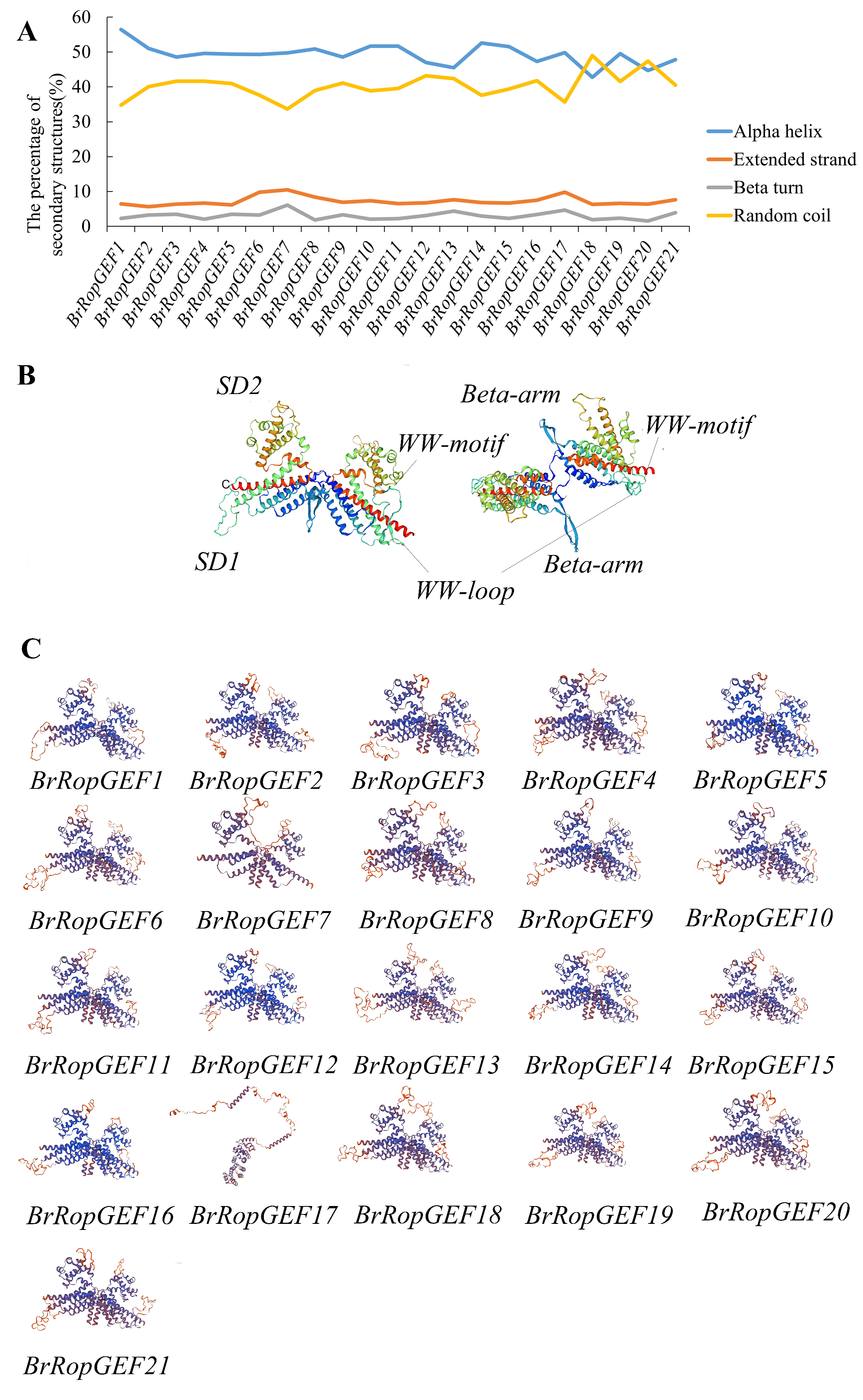
2.7. Analysis of the Protein Secondary and Tertiary Structures of BrRopGEFs
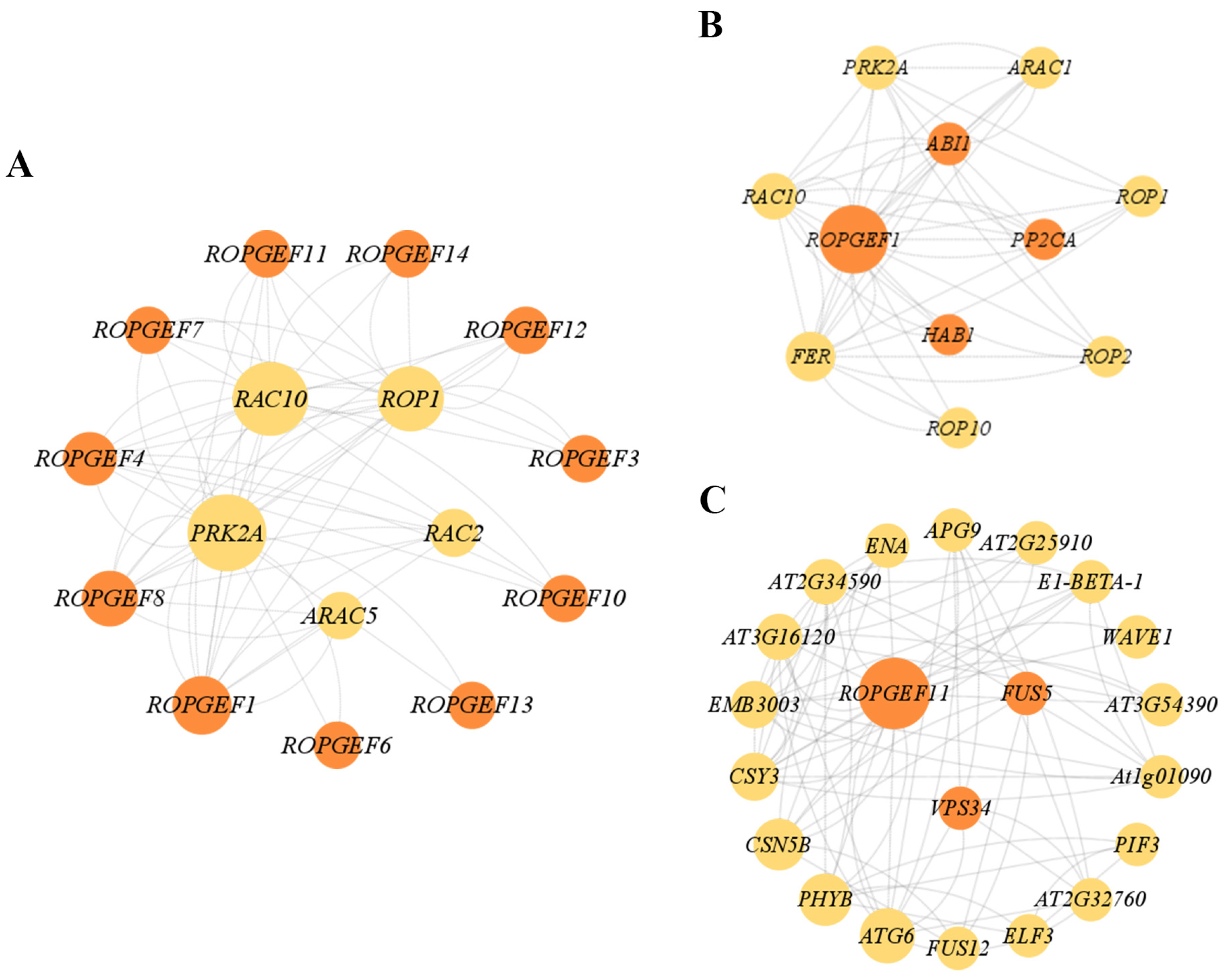
2.8. Protein–Protein Interaction (PPI) Network Analysis of BrRopGEFs
3. Discussion
4. Materials and Methods
4.1. Identification of RopGEF Genes and Analysis of the Physical-Chemical Properties and Subcellular Localization of Their Encoded Proteins
4.2. Chromosome Localization, Multiple Sequence Alignment, and Phylogenetic Analysis
4.3. Analysis of CAREs
4.4. Motif Analysis and GO
4.5. Expression Profiling of BrRopGEF Genes
4.6. Total RNA Extraction and RT-qPCR
4.7. Analysis of Protein Secondary Structure, Tertiary Structure, and PPI Networks
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Berken, A.; Thomas, C.; Wittinghofer, A. A new family of RhoGEFs activates the Rop molecular switch in plants. Nature 2005, 436, 1176–1180. [Google Scholar] [CrossRef]
- Berken, A.; Wittinghofer, A. Structure and function of Rho-type molecular switches in plants. Plant Physiol. Biochem. 2008, 46, 380–393. [Google Scholar] [CrossRef]
- Thomas, C.; Fricke, I.; Scrima, A.; Berken, A.; Wittinghofer, A. Structural Evidence for a Common Intermediate in Small G Protein-GEF Reactions. Mol. Cell 2007, 25, 141–149. [Google Scholar] [CrossRef]
- Chang, F.; Gu, Y.; Ma, H.; Yang, Z. AtPRK2 promotes ROP1 activation via RopGEFs in the control of polarized pollen tube growth. Mol. Plant 2013, 6, 1187–1201. [Google Scholar] [CrossRef]
- Zhang, Y.; McCormick, S. A distinct mechanism regulating a pollen-specific guanine nucleotide exchange factor for the small GTPase Rop in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2007, 104, 18830–18835. [Google Scholar] [CrossRef]
- Shin, D.H.; Kim, T.-L.; Kwon, Y.-K.; Cho, M.-H.; Yoo, J.; Jeon, J.-S.; Hahn, T.-R.; Bhoo, S.H. Characterization of Arabidopsis RopGEF family genes in response to abiotic stresses. Plant Biotechnol. Rep. 2009, 3, 183–190. [Google Scholar] [CrossRef]
- Gu, Y.; Li, S.; Lord, E.M.; Yang, Z. Members of a novel class of Arabidopsis Rho guanine nucleotide exchange factors control Rho GTPase-dependent polar growth. Plant Cell 2006, 18, 366–381. [Google Scholar] [CrossRef]
- Shin, D.H.; Cho, M.-H.; Kim, T.-L.; Yoo, J.; Kim, J.-I.; Han, Y.-J.; Song, P.-S.; Jeon, J.-S.; Bhoo, S.H.; Hahn, T.-R. A Small GTPase Activator Protein Interacts with Cytoplasmic Phytochromes in Regulating Root Development. J. Biol. Chem. 2010, 285, 32151–32159. [Google Scholar] [CrossRef]
- Li, Z.; Liu, D. ROPGEF1 and ROPGEF4 are functional regulators of ROP11 GTPase in ABA-mediated stomatal closure in Arabidopsis. FEBS Lett. 2012, 586, 1253–1258. [Google Scholar] [CrossRef]
- Chen, M.; Liu, H.; Kong, J.; Yang, Y.; Zhang, N.; Li, R.; Yue, J.; Huang, J.; Li, C.; Cheung, A.Y.; et al. RopGEF7 regulates PLETHORA-dependent maintenance of the root stem cell niche in Arabidopsis. Plant Cell 2011, 23, 2880–2894. [Google Scholar] [CrossRef]
- Liu, Y.; Dong, Q.; Kita, D.; Huang, J.B.; Liu, G.; Wu, X.; Zhu, X.; Cheung, A.Y.; Wu, H.M.; Tao, L.Z. RopGEF1 Plays a Critical Role in Polar Auxin Transport in Early Development. Plant Physiol. 2017, 175, 157–171. [Google Scholar] [CrossRef]
- Duan, Q.; Kita, D.; Li, C.; Cheung, A.Y.; Wu, H.-M. FERONIA receptor-like kinase regulates RHO GTPase signaling of root hair development. Proc. Natl. Acad. Sci. USA 2010, 107, 17821–17826. [Google Scholar] [CrossRef]
- Yoshida, M.W.; Hakozaki, M.; Goshima, G. Armadillo repeat-containing kinesin represents the versatile plus-end-directed transporter in Physcomitrella. Nat. Plants 2023, 9, 733–748. [Google Scholar] [CrossRef]
- Kim, E.J.; Park, S.W.; Hong, W.J.; Silva, J.; Liang, W.; Zhang, D.; Jung, K.H.; Kim, Y.J. Genome-wide analysis of RopGEF gene family to identify genes contributing to pollen tube growth in rice (Oryza sativa). BMC Plant Biol. 2020, 20, 95. [Google Scholar] [CrossRef]
- Kim, E.J.; Hong, W.J.; Tun, W.; An, G.; Kim, S.T.; Kim, Y.J.; Jung, K.H. Interaction of OsRopGEF3 Protein With OsRac3 to Regulate Root Hair Elongation and Reactive Oxygen Species Formation in Rice (Oryza sativa). Front. Plant Sci. 2021, 12, 661352. [Google Scholar] [CrossRef]
- Ali Shad, M.; Wang, Y.; Zhang, H.; Zhai, S.; Shalmani, A.; Li, Y. Genetic analysis of GEFs and GDIs in rice reveals the roles of OsGEF5, OsGDI1, and OsGEF3 in the regulation of grain size and plant height. Crop J. 2022, 11, 345–360. [Google Scholar] [CrossRef]
- Riely, B.K.; He, H.; Venkateshwaran, M.; Sarma, B.; Schraiber, J.; Ane, J.M.; Cook, D.R. Identification of legume RopGEF gene families and characterization of a Medicago truncatula RopGEF mediating polar growth of root hairs. Plant J. 2011, 65, 230–243. [Google Scholar] [CrossRef]
- Ruan, J.; Lai, L.; Ou, H.; Yi, P. Two subtypes of GTPase-activating proteins coordinate tip growth and cell size regulation in Physcomitrium patens. Nat. Commun. 2023, 14, 7084. [Google Scholar] [CrossRef]
- Salunkhe, D.K.; Kadam, S. Handbook of Vegetable Science and Technology: Production, Compostion, Storage, and Processing; CRC Press: Boca Raton, FL, USA, 1998. [Google Scholar]
- Franks, S.J. Plasticity and evolution in drought avoidance and escape in the annual plant Brassica rapa. New Phytol. 2011, 190, 249–257. [Google Scholar] [CrossRef]
- Wang, B.; Zhang, H.; Huai, J.; Peng, F.; Wu, J.; Lin, R.; Fang, X. Condensation of SEUSS promotes hyperosmotic stress tolerance in Arabidopsis. Nat. Chem. Biol. 2022, 18, 1361–1369. [Google Scholar] [CrossRef]
- Guo, Y.M.; Samans, B.; Chen, S.; Kibret, K.B.; Hatzig, S.; Turner, N.C.; Nelson, M.N.; Cowling, W.A.; Snowdon, R.J. Drought-Tolerant Brassica rapa Shows Rapid Expression of Gene Networks for General Stress Responses and Programmed Cell Death Under Simulated Drought Stress. Plant Mol. Biol. Rep. 2017, 35, 416–430. [Google Scholar] [CrossRef]
- Thomas, C.; Fricke, I.; Weyand, M.; Berken, A. 3D structure of a binary ROP-PRONE complex: The final intermediate for a complete set of molecular snapshots of the RopGEF reaction. Biol. Chem. 2009, 390, 427–435. [Google Scholar] [CrossRef]
- Liu, C.; Liu, L.; Zhou, C.; Zhuang, J.; Wang, L.; Sun, Y.; Sun, C. Protein-protein interaction networks and different clustering analysis in Burkitt’s lymphoma. Hematology 2018, 23, 391–398. [Google Scholar] [CrossRef] [PubMed]
- Bader, G.D.; Hogue, C.W.V. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Movahedi, A.; Liu, G.; Li, Y.; Liu, S.; Yu, C.; Chen, Y.; Zhong, F.; Zhang, J. Comprehensive Analysis of Carotenoid Cleavage Dioxygenases Gene Family and Its Expression in Response to Abiotic Stress in Poplar. Int. J. Mol. Sci. 2022, 23, 1418. [Google Scholar] [CrossRef]
- Alberts, B. The cell as a collection of protein machines: Preparing the next generation of molecular biologists. Cell 1998, 92, 291–294. [Google Scholar] [CrossRef]
- Liu, Z.; Li, Y.; Zhu, J.; Ma, W.; Li, Z.; Bi, Z.; Sun, C.; Bai, J.; Zhang, J.; Liu, Y. Genome-Wide Identification and Analysis of the NF-Y Gene Family in Potato (Solanum tuberosum L.). Front. Genet. 2021, 12, 739989. [Google Scholar] [CrossRef]
- Li, F.; Guo, X.; Liu, J.; Zhou, F.; Liu, W.; Wu, J.; Zhang, H.; Cao, H.; Su, H.; Wen, R. Genome-Wide Identification, Characterization, and Expression Analysis of the NAC Transcription Factor in Chenopodium quinoa. Genes 2019, 10, 500. [Google Scholar] [CrossRef]
- Li, Z.; Waadt, R.; Schroeder, J.I. Release of GTP Exchange Factor Mediated Down-Regulation of Abscisic Acid Signal Transduction through ABA-Induced Rapid Degradation of RopGEFs. PLoS Biol. 2016, 14, e1002461. [Google Scholar] [CrossRef]
- Shohat, H.; Cheriker, H.; Kilambi, H.V.; Illouz Eliaz, N.; Blum, S.; Amsellem, Z.; Tarkowská, D.; Aharoni, A.; Eshed, Y.; Weiss, D. Inhibition of gibberellin accumulation by water deficiency promotes fast and long-term ‘drought avoidance’ responses in tomato. New Phytol. 2021, 232, 1985–1998. [Google Scholar] [CrossRef]
- Rehman, A.; Azhar, M.T.; Hinze, L.; Qayyum, A.; Li, H.; Peng, Z.; Qin, G.; Jia, Y.; Pan, Z.; He, S.; et al. Insight into abscisic acid perception and signaling to increase plant tolerance to abiotic stress. J. Plant Interact. 2021, 16, 222–237. [Google Scholar] [CrossRef]
- Li, Z.; Takahashi, Y.; Scavo, A.; Brandt, B.; Nguyen, D.; Rieu, P.; Schroeder, J.I. Abscisic acid-induced degradation of Arabidopsis guanine nucleotide exchange factor requires calcium-dependent protein kinases. Proc. Natl. Acad. Sci. USA 2018, 115, E4522–E4531. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, Z.; Gao, X.; Chinnusamy, V.; Bressan, R.; Wang, Z.X.; Zhu, J.K.; Wu, J.W.; Liu, D. ROP11 GTPase negatively regulates ABA signaling by protecting ABI1 phosphatase activity from inhibition by the ABA receptor RCAR1/PYL9 in Arabidopsis. J. Integr. Plant Biol. 2012, 54, 180–188. [Google Scholar] [CrossRef]
- Yu, F.; Qian, L.; Nibau, C.; Duan, Q.; Kita, D.; Levasseur, K.; Li, X.; Lu, C.; Li, H.; Hou, C.; et al. FERONIA receptor kinase pathway suppresses abscisic acid signaling in Arabidopsis by activating ABI2 phosphatase. Proc. Natl. Acad. Sci. USA 2012, 109, 14693–14698. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.L.; Nafisi, M.; Tam, A.; Li, H.; Crowell, D.N.; Chary, S.N.; Schroeder, J.I.; Shen, J.; Yang, Z. Plasma membrane-associated ROP10 small GTPase is a specific negative regulator of abscisic acid responses in Arabidopsis. Plant Cell 2002, 14, 2787–2797. [Google Scholar] [CrossRef]
- Lim, C.W.; Baek, W.; Jung, J.; Kim, J.H.; Lee, S.C. Function of ABA in Stomatal Defense against Biotic and Drought Stresses. Int. J. Mol. Sci. 2015, 16, 15251–15270. [Google Scholar] [CrossRef]
- Okuma, E.; Nozawa, R.; Murata, Y.; Miura, K. Accumulation of endogenous salicylic acid confers drought tolerance to Arabidopsis. Plant Signal. Behav. 2014, 9, e28085. [Google Scholar] [CrossRef]
- Shi, H.; Chen, L.; Ye, T.; Liu, X.; Ding, K.; Chan, Z. Modulation of auxin content in Arabidopsis confers improved drought stress resistance. Plant Physiol. Biochem. 2014, 82, 209–217. [Google Scholar] [CrossRef]
- Creelman, R.A.; Mullet, J.E. Jasmonic acid distribution and action in plants: Regulation during development and response to biotic and abiotic stress. Proc. Natl. Acad. Sci. USA 1995, 92, 4114–4119. [Google Scholar] [CrossRef]
- Javid, M.G.; Sorooshzadeh, A.; Moradi, F.; Sanavy, S.A.M.M.; Allahdadi, I. The Role of Phytohormones in Alleviating Salt Stress in Crop Plants. Aust. J. Crop Sci. 2011, 5, 726–734. [Google Scholar]
- Chen, H.; Wang, T.; He, X.; Cai, X.; Lin, R.; Liang, J.; Wu, J.; King, G.; Wang, X. BRAD V3.0: An upgraded Brassicaceae database. Nucleic Acids Res. 2022, 50, D1432–D1441. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef]
- Gasteiger, E.; Gattiker, A.; Hoogland, C.; Ivanyi, I.; Appel, R.D.; Bairoch, A. ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res. 2003, 31, 3784–3788. [Google Scholar] [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2011, 40, D1178–D1186. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019, 47, W191–W198. [Google Scholar] [CrossRef]
- Barrett, T.; Edgar, R. [19] Gene Expression Omnibus: Microarray Data Storage, Submission, Retrieval, and Analysis. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2006; Volume 411, pp. 352–369. [Google Scholar]
- Tong, C.; Wang, X.; Yu, J.; Wu, J.; Li, W.; Huang, J.; Dong, C.; Hua, W.; Liu, S. Comprehensive analysis of RNA-seq data reveals the complexity of the transcriptome in Brassica rapa. BMC Genom. 2013, 14, 689. [Google Scholar] [CrossRef]
- Lu, K.; Li, T.; He, J.; Chang, W.; Zhang, R.; Liu, M.; Yu, M.; Fan, Y.; Ma, J.; Sun, W.; et al. qPrimerDB: A thermodynamics-based gene-specific qPCR primer database for 147 organisms. Nucleic Acids Res. 2017, 46, D1229–D1236. [Google Scholar] [CrossRef]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]


| Gene Name | Gene ID | Chromosome | pI | MW (Da) | Protein Length (aa) | Subcellular Location | A. thaliana ID | A. thaliana Name |
|---|---|---|---|---|---|---|---|---|
| BrRopGEF1 | Bra021162 | A01:23638077-23639663 | 5.82 | 49,216.11 | 434 | nucleus | AT3G16130 | AtRopGEF13 |
| BrRopGEF2 | Bra020048 | A02:4812438-4815205 | 5.56 | 565,501 | 494 | nucleus | AT5G19560 | AtRopGEF10 |
| BrRopGEF3 | Bra006510 | A03:3845994-3848462 | 7.75 | 56,166.59 | 488 | nucleus cytosol | AT5G19560 | AtRopGEF10 |
| BrRopGEF4 | Bra000971 | A03:14185129-14186999 | 4.98 | 53,899.85 | 478 | nucleus | AT4G00460 | AtRopGEF3 |
| BrRopGEF5 | Bra013249 | A03:19623928-19627223 | 5.66 | 58,016.66 | 518 | nucleus | AT3G24620 | AtRopGEF8 |
| BrRopGEF6 | Bra004945 | A05:2591812-2594623 | 4.95 | 51,723.86 | 460 | nucleus | AT2G45890 | AtRopGEF4 |
| BrRopGEF7 | Bra005322 | A05:4825068-4826940 | 6.06 | 44,107.32 | 391 | nucleus | AT1G31650 | AtRopGEF14 |
| BrRopGEF8 | Bra005323 | A05:4825068-4826943 | 6.6 | 56,592.05 | 501 | nucleus | AT1G31650 | AtRopGEF14 |
| BrRopGEF9 | Bra030396 | A05:10855878-10857882 | 5.61 | 60,575.65 | 536 | nucleus | AT1G52240 | AtRopGEF11 |
| BrRopGEF10 | Bra027189 | A05:19599734-19601858 | 5.68 | 60,452.88 | 530 | nucleus | AT3G16130 | AtRopGEF13 |
| BrRopGEF11 | Bra018956 | A06:1021072-1023077 | 5.64 | 60,598.73 | 536 | nucleus | AT1G52240 | AtRopGEF11 |
| BrRopGEF12 | Bra015068 | A07:3804713-3807050 | 5.65 | 58,233.8 | 521 | nucleus | AT3G24620 | AtRopGEF8 |
| BrRopGEF13 | Bra015010 | A07:4301480-4303430 | 5.84 | 60,516.64 | 547 | nucleus | AT4G38430 | AtRopGEF1 |
| BrRopGEF14 | Bra003536 | A07:13678585-13681092 | 5.5 | 57,786.29 | 513 | nucleus cytosol | AT1G79860 | AtRopGEF12 |
| BrRopGEF15 | Bra037342 | A09:971667-973569 | 5.03 | 53,733.8 | 477 | nucleus cytosol | AT4G00460 | AtRopGEF3 |
| BrRopGEF16 | Bra036671 | A09:5723788-5725881 | 5.86 | 56,589.91 | 505 | chloroplast | AT3G24620 | AtRopGEF8 |
| BrRopGEF17 | Bra023198 | A09:20826415-20833784 | 6.41 | 106,720.03 | 941 | cytosol | AT1G31650 | AtRopGEF14 |
| BrRopGEF18 | Bra007183 | A09:27809243-27812082 | 8.86 | 63,303.85 | 571 | cytosol | AT3G55660 | AtRopGEF6 |
| BrRopGEF19 | Bra002246 | A10:10684361-10686961 | 6.65 | 59,090.48 | 517 | chloroplast | AT5G19560 | AtRopGEF10 |
| BrRopGEF20 | Bra009152 | A10:15438568-15440923 | 9.12 | 66,383.94 | 595 | nucleus | AT5G05940 | AtRopGEF5 |
| BrRopGEF21 | Bra009621 | A10:17469516-17471749 | 5.61 | 63,438.88 | 521 | nucleus | AT5G02010 | AtRopGEF7 |
| Motif | Motif Consensus |
|---|---|
| Motif1 | EMMKERFAKLLLGEDMSGGGKGVCSALALSNAITNLAASVFGEQWRLZPL |
| Motif2 | FPGJPQSSLDISKIQYNKDVGKAILESYSRVLESLAYTILSRIEDVLYAD |
| Motif3 | QKDSVNQVLKAAMAINAQVLSEMEIPESYJDSLPKNGKASLGDSIYKMJT |
| Motif4 | QQTNKBGTSTEIMTTRQRSDLLMNJPALRKLDSMLJDTLDS |
| Motif5 | EMFDPDQFLSSLDLSSEHKALDLKNRIEASIVIWKRKMVZK |
| Motif6 | QRNDEKWWLPVVKVPPNGLSEESRKFLQS |
| Motif7 | RWRREMDWLLSVTDHIVEFVP |
| Motif8 | SPWGSAVSLEKRELFEERAETJLVLLKQR |
| Motif9 | KDQTEFWYVERDSEE |
| Motif10 | PTKSPRVTPKKLSYLEKLENMRSPTARH |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, M.; Wu, X.; Chen, L.; Yang, L.; Cui, X.; Cao, Y. The RopGEF Gene Family and Their Potential Roles in Responses to Abiotic Stress in Brassica rapa. Int. J. Mol. Sci. 2024, 25, 3541. https://doi.org/10.3390/ijms25063541
Zhang M, Wu X, Chen L, Yang L, Cui X, Cao Y. The RopGEF Gene Family and Their Potential Roles in Responses to Abiotic Stress in Brassica rapa. International Journal of Molecular Sciences. 2024; 25(6):3541. https://doi.org/10.3390/ijms25063541
Chicago/Turabian StyleZhang, Meiqi, Xiaoyu Wu, Luhan Chen, Lin Yang, Xiaoshuang Cui, and Yunyun Cao. 2024. "The RopGEF Gene Family and Their Potential Roles in Responses to Abiotic Stress in Brassica rapa" International Journal of Molecular Sciences 25, no. 6: 3541. https://doi.org/10.3390/ijms25063541
APA StyleZhang, M., Wu, X., Chen, L., Yang, L., Cui, X., & Cao, Y. (2024). The RopGEF Gene Family and Their Potential Roles in Responses to Abiotic Stress in Brassica rapa. International Journal of Molecular Sciences, 25(6), 3541. https://doi.org/10.3390/ijms25063541





