Probing Liver Injuries Induced by Thioacetamide in Human In Vitro Pooled Hepatocyte Experiments
Abstract
1. Introduction
2. Results and Discussion
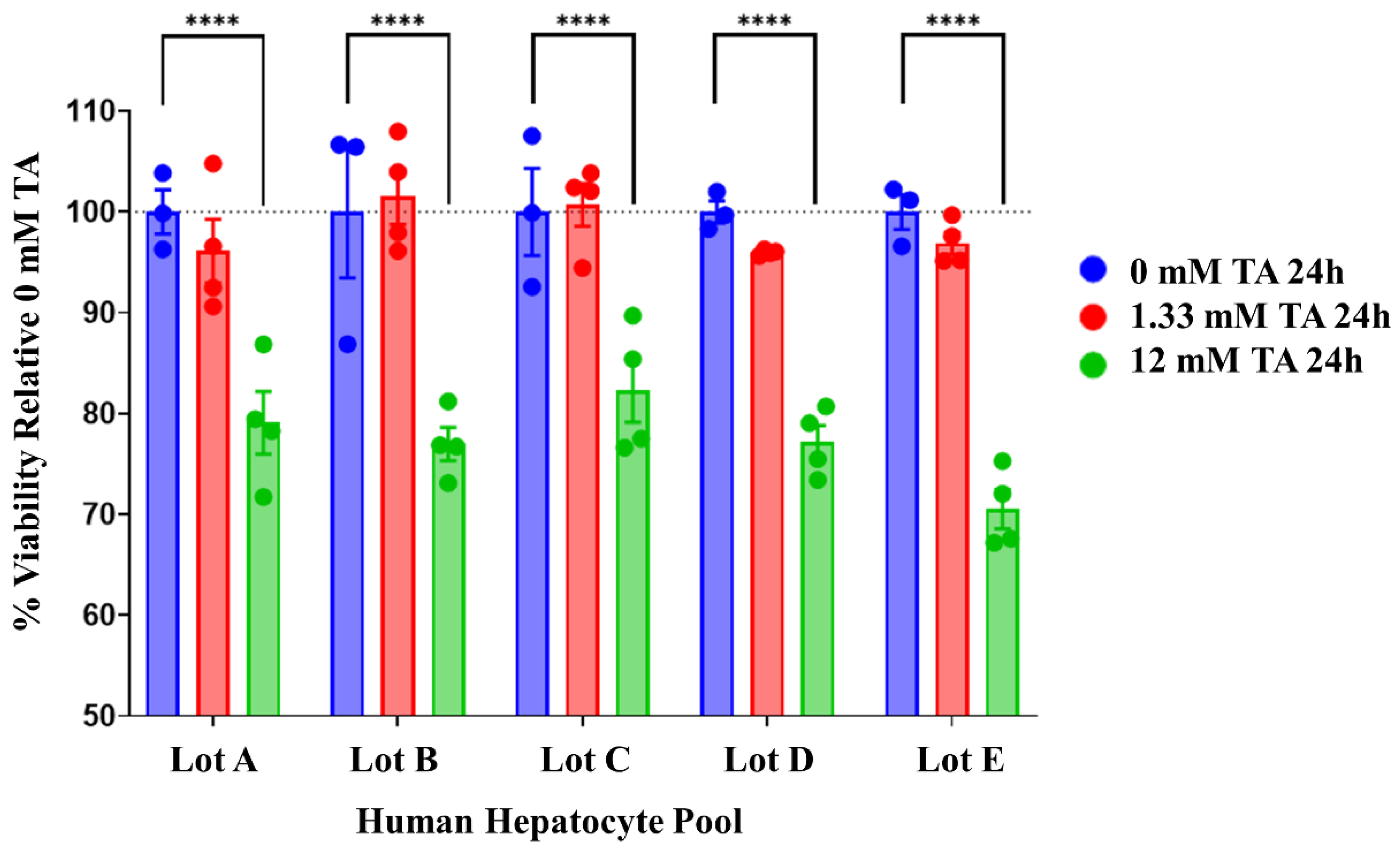
2.1. Dose Optimization for Thioacetamide Exposure
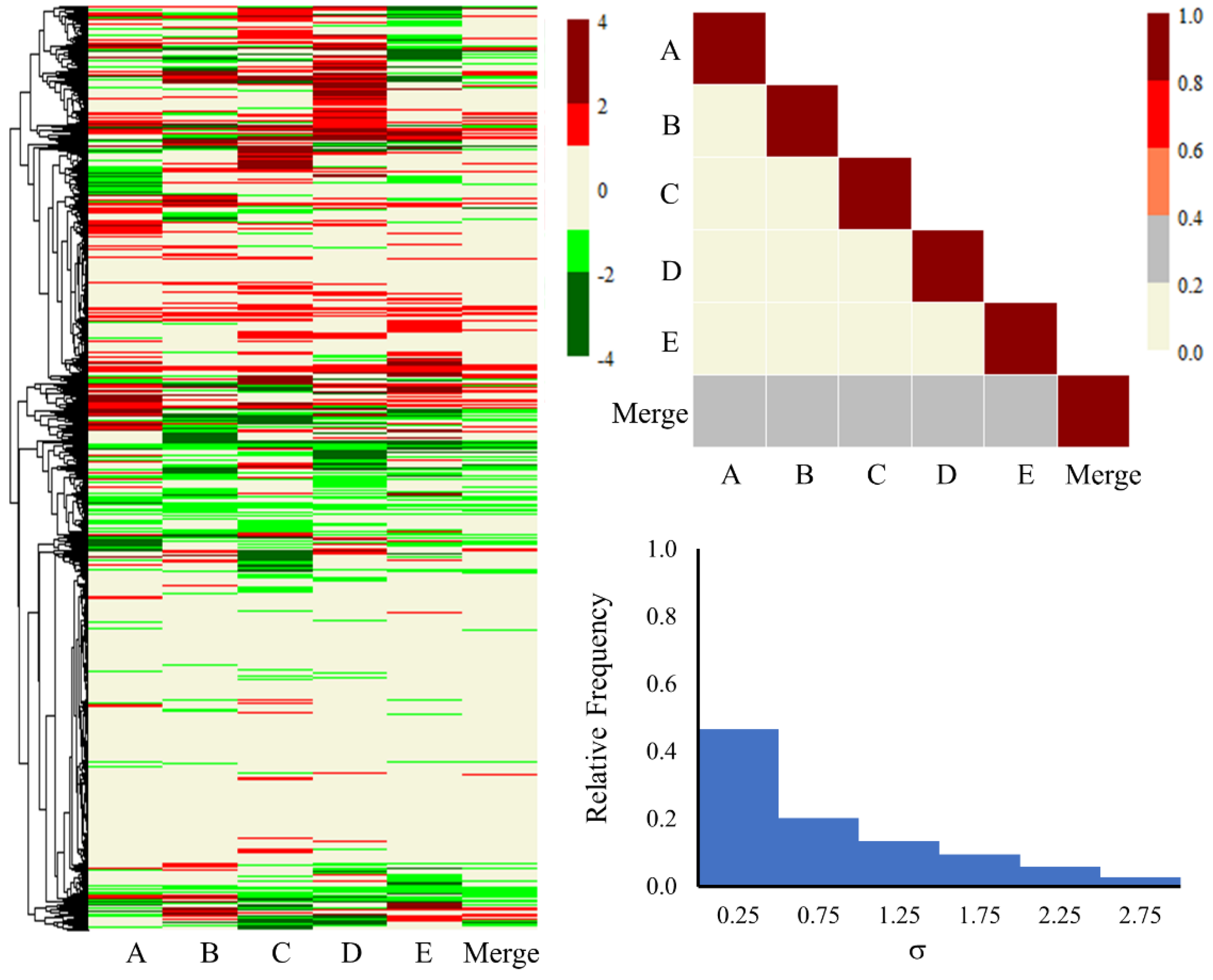
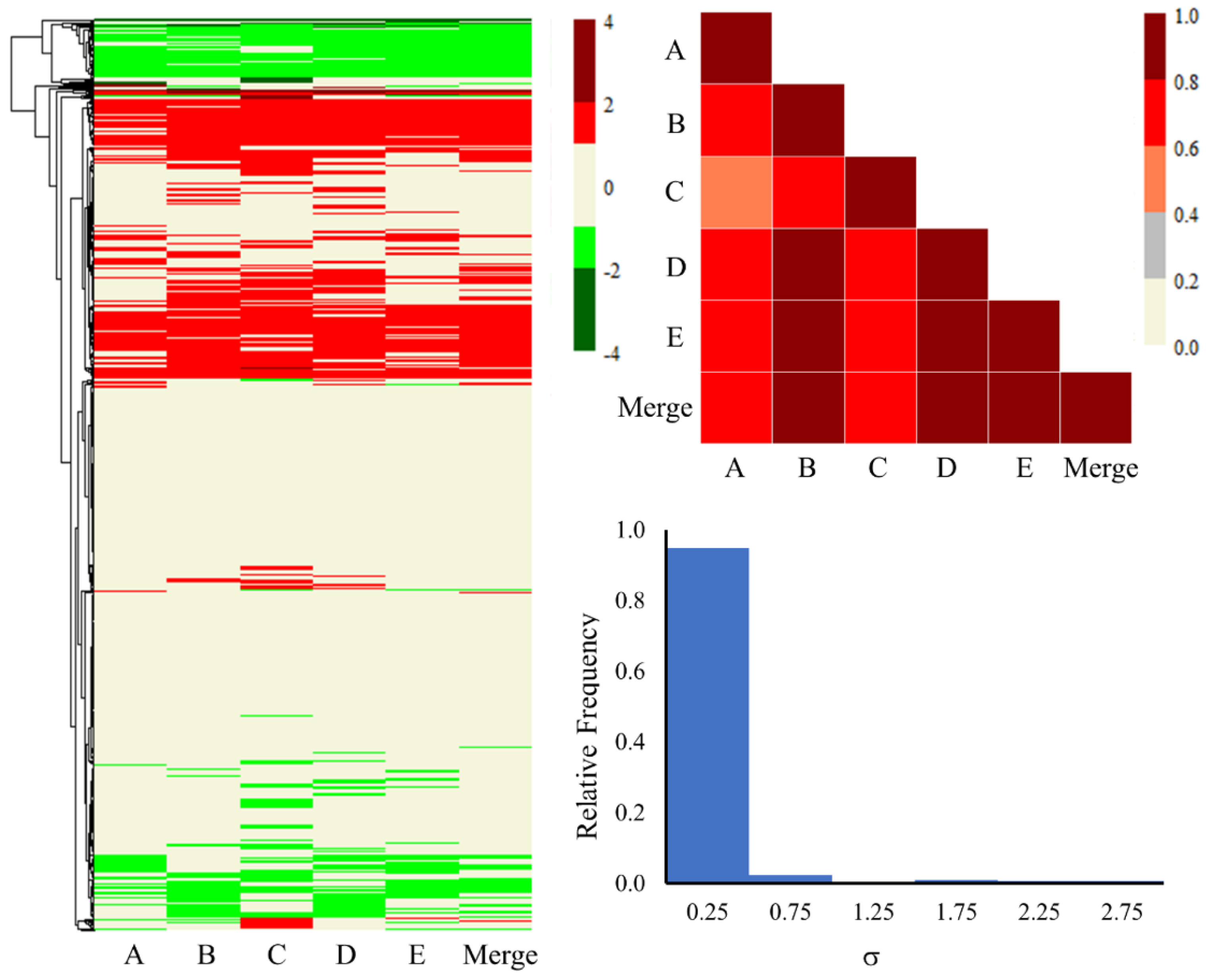
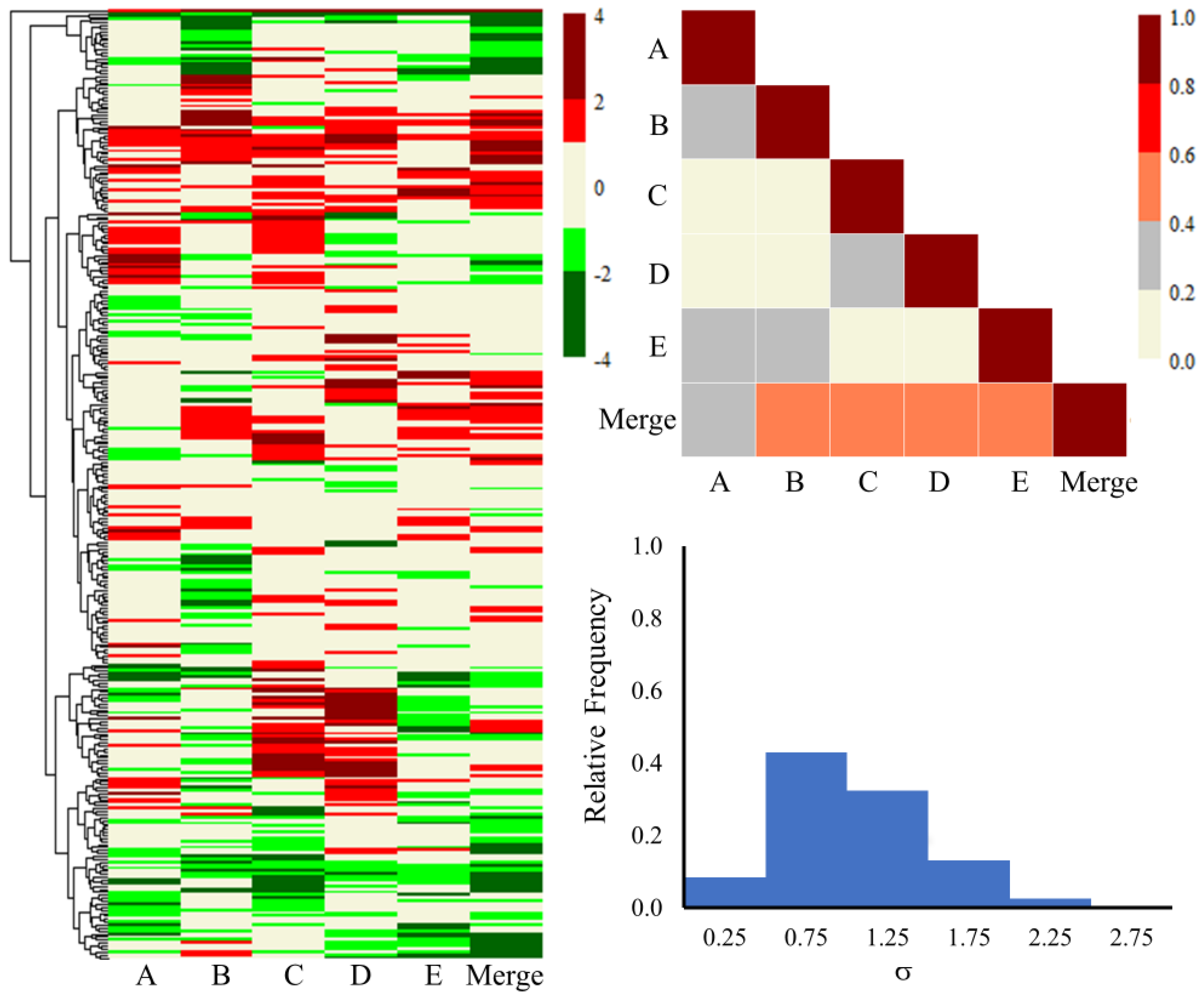
2.2. Gene-Level Analysis
2.3. KEGG Pathway Analysis
2.4. Liver Injury Module Analysis
2.5. Hepatocyte Transporters and Cytochrome P450 Enzymes
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. Cell Viability Assays
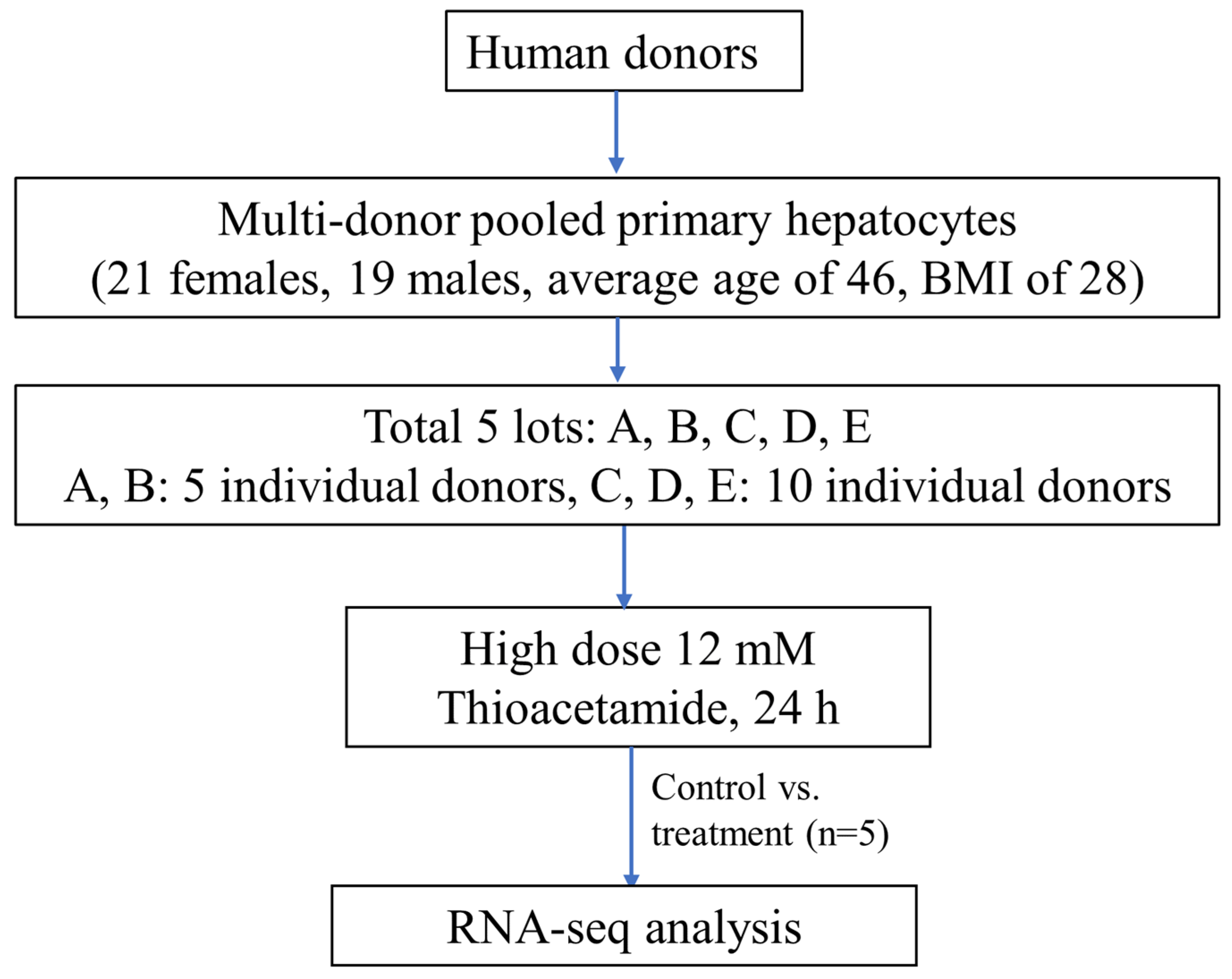
3.3. Experimental Procedures
3.4. Computational Details for RNA-Seq Data, KEGG Pathways, and Injury Module Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Saeidnia, S.; Manayi, A.; Abdollahi, M. From in vitro experiments to in vivo and clinical studies; pros and cons. Curr. Drug Discov. Technol. 2015, 12, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Vinken, M. Primary hepatocyte cultures for liver disease modeling. Curr. Opin. Toxicol. 2021, 25, 1–5. [Google Scholar] [CrossRef]
- Gomez-Lechon, M.; Donato, M.; Castell, J.; Jover, R. Human hepatocytes in primary culture: The choice to investigate drug metabolism in man. Curr. Drug Metab. 2004, 5, 443–462. [Google Scholar] [CrossRef] [PubMed]
- Kaur, I.; Vasudevan, A.; Rawal, P.; Tripathi, D.M.; Ramakrishna, S.; Kaur, S.; Sarin, S.K. Primary hepatocyte isolation and cultures: Technical aspects, challenges and advancements. Bioengineering 2023, 10, 131. [Google Scholar] [CrossRef] [PubMed]
- Su, S.; Di Poto, C.; Roy, R.; Liu, X.; Cui, W.; Kroemer, A.; Ressom, H.W. Long-term culture and characterization of patient-derived primary hepatocytes using conditional reprogramming. Exp. Biol. Med. 2019, 244, 857–864. [Google Scholar] [CrossRef] [PubMed]
- Alexander-Dann, B.; Pruteanu, L.L.; Oerton, E.; Sharma, N.; Berindan-Neagoe, I.; Módos, D.; Bender, A. Developments in toxicogenomics: Understanding and predicting compound-induced toxicity from gene expression data. Mol. Omics 2018, 14, 218–236. [Google Scholar] [CrossRef] [PubMed]
- Schyman, P.; Printz, R.L.; Estes, S.K.; Boyd, K.L.; Shiota, M.; Wallqvist, A. Identification of the toxicity pathways associated with thioacetamide-induced injuries in rat liver and kidney. Front. Pharmacol. 2018, 9, 1272. [Google Scholar] [CrossRef] [PubMed]
- Schyman, P.; Printz, R.L.; Estes, S.K.; O’Brien, T.P.; Shiota, M.; Wallqvist, A. Assessing chemical-induced liver injury in vivo from in vitro gene expression data in the rat: The case of thioacetamide toxicity. Front. Genet. 2019, 10, 1233. [Google Scholar] [CrossRef]
- Schyman, P.; Printz, R.L.; AbdulHameed, M.D.M.; Estes, S.K.; Shiota, C.; Shiota, M.; Wallqvist, A. A toxicogenomic approach to assess kidney injury induced by mercuric chloride in rats. Toxicology 2020, 442, 152530. [Google Scholar] [CrossRef]
- Schyman, P.; Printz, R.L.; Estes, S.K.; O’Brien, T.P.; Shiota, M.; Wallqvist, A. Concordance between thioacetamide-induced liver injury in rat and human in vitro gene expression data. Int. J. Mol. Sci. 2020, 21, 4017. [Google Scholar] [CrossRef]
- Schyman, P.; Printz, R.L.; Pannala, V.R.; AbdulHameed, M.D.M.; Estes, S.K.; Shiota, C.; Boyd, K.L.; Shiota, M.; Wallqvist, A. Genomics and metabolomics of early-stage thioacetamide-induced liver injury: An interspecies study between guinea pig and rat. Toxicol. Appl. Pharmacol. 2021, 430, 115713. [Google Scholar] [CrossRef]
- Goel, H.; Printz, R.L.; Shiota, C.; Estes, S.K.; Pannala, V.; AbdulHameed, M.D.M.; Shiota, M.; Wallqvist, A. Assessing kidney injury induced by mercuric chloride in guinea pigs with in vivo and in vitro experiments. Int. J. Mol. Sci. 2023, 24, 7434. [Google Scholar] [CrossRef] [PubMed]
- Melis, M.; Tang, X.-H.; Trasino, S.E.; Gudas, L.J. Retinoids in the pathogenesis and treatment of liver diseases. Nutrients 2022, 14, 1456. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, S.K.; Thomas, S.; Ramachandran, A.; Pulimood, A.B.; Balasubramanian, K.A. Retinoid metabolism during development of liver cirrhosis. Arch. Biochem. Biophys. 2005, 443, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Xi, J.; Yang, Z. Expression of RALDHs (ALDH1As) and CYP26s in human tissues and during the neural differentiation of P19 embryonal carcinoma stem cell. Gene Expr. Patterns 2008, 8, 438–442. [Google Scholar] [CrossRef] [PubMed]
- Tay, S.; Dickmann, L.; Dixit, V.; Isoherranen, N. A comparison of the roles of peroxisome proliferator-activated receptor and retinoic acid receptor on CYP26 regulation. Mol. Pharmacol. 2010, 77, 218–227. [Google Scholar] [CrossRef]
- Thatcher, J.E.; Zelter, A.; Isoherranen, N. The relative importance of CYP26A1 in hepatic clearance of all-trans retinoic acid. Biochem. Pharmacol. 2010, 80, 903–912. [Google Scholar] [CrossRef] [PubMed]
- Shmarakov, I.O.; Borschovetska, V.L.; Marchenko, M.M.; Blaner, W.S. Retinoids modulate thioacetamide-induced acute hepatotoxicity. Toxicol. Sci. 2014, 139, 284–292. [Google Scholar] [CrossRef]
- Kedishvili, N.Y. Enzymology of retinoic acid biosynthesis and degradation: Thematic review series: Fat-soluble vitamins: Vitamin A. J. Lipid Res. 2013, 54, 1744–1760. [Google Scholar] [CrossRef]
- Guo, L.; Dial, S.; Shi, L.; Branham, W.; Liu, J.; Fang, J.-L.; Green, B.; Deng, H.; Kaput, J.; Ning, B. Similarities and differences in the expression of drug-metabolizing enzymes between human hepatic cell lines and primary human hepatocytes. Drug Metab. Dispos. 2011, 39, 528–538. [Google Scholar] [CrossRef]
- Starlinger, P.; Brunnthaler, L.; McCabe, C.; Pereyra, D.; Santol, J.; Steadman, J.; Hackl, M.; Skalicky, S.; Hackl, H.; Gronauer, R. Transcriptomic landscapes of effective and failed liver regeneration in humans. JHEP Rep. 2023, 5, 100683. [Google Scholar] [CrossRef] [PubMed]
- Petrenko, O.; Königshofer, P.; Brusilovskaya, K.; Hofer, B.S.; Bareiner, K.; Simbrunner, B.; Jühling, F.; Baumert, T.F.; Lupberger, J.; Trauner, M. Transcriptomic signatures of progressive and regressive liver fibrosis and portal hypertension. iScience 2024, 27, 109301. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Wang, L.; Wang, M.; Zhou, S.; Lu, Y.; Cui, H.; Racanelli, A.C.; Zhang, L.; Ye, T.; Ding, B. Targeting fibrosis: Mechanisms and clinical trials. Signal Transduct. Target. Ther. 2022, 7, 206. [Google Scholar] [CrossRef] [PubMed]
- Chang, G.-R.; Lin, W.-L.; Lin, T.-C.; Liao, H.-J.; Lu, Y.-W. The ameliorative effects of saikosaponin in thioacetamide-induced liver injury and non-alcoholic fatty liver disease in mice. Int. J. Mol. Sci. 2021, 22, 11383. [Google Scholar] [CrossRef]
- Kang, H.; Seo, E.; Park, J.M.; Han, N.Y.; Lee, H.; Jun, H.S. Effects of FGF 21-secreting adipose-derived stem cells in thioacetamide-induced hepatic fibrosis. J. Cell. Mol. Med. 2018, 22, 5165–5169. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.-Y.; Yang, W.-C.; Lin, C.-F.; Wang, C.-M.; Liu, H.-Y.; Lin, C.-S.; Lin, J.-W.; Lin, W.-L.; Lin, T.-C.; Fan, P.-S. The ameliorative effects of fucoidan in thioacetaide-induced liver injury in mice. Molecules 2021, 26, 1937. [Google Scholar] [CrossRef] [PubMed]
- Tillman, E.J.; Rolph, T. FGF21: An emerging therapeutic target for non-alcoholic steatohepatitis and related metabolic diseases. Front. Endocrinol. 2020, 11, 601290. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Chen, W.; Athavale, D.; Ge, X.; Desert, R.; Das, S.; Han, H.; Nieto, N. Osteopontin takes center stage in chronic liver disease. Hepatology 2021, 73, 1594–1608. [Google Scholar] [CrossRef]
- Ferrigno, A.; Di Pasqua, L.G.; Berardo, C.; Richelmi, P.; Vairetti, M. Liver plays a central role in asymmetric dimethylarginine-mediated organ injury. World J. Gastroenterol. 2015, 21, 5131. [Google Scholar] [CrossRef]
- Ritter, M.J.; Amano, I.; Hollenberg, A.N. Thyroid hormone signaling and the liver. Hepatology 2020, 72, 742–752. [Google Scholar] [CrossRef]
- Weber, S.N.; Lammert, F. Genetics in liver diseases: From diagnostics to precise therapy. Clin. Liver Dis. 2017, 9, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Schumann, T.; König, J.; Henke, C.; Willmes, D.M.; Bornstein, S.R.; Jordan, J.; Fromm, M.F.; Birkenfeld, A.L. Solute carrier transporters as potential targets for the treatment of metabolic disease. Pharmacol. Rev. 2020, 72, 343–379. [Google Scholar] [CrossRef] [PubMed]
- Cooper, G.M.; Adams, K. The Cell: A Molecular Approach; Oxford University Press: Oxford, UK, 2023. [Google Scholar]
- Ferdinandusse, S.; Houten, S.M. Peroxisomes and bile acid biosynthesis. Biochim. Biophys. Acta Mol. Cell Res. 2006, 1763, 1427–1440. [Google Scholar] [CrossRef] [PubMed]
- Ferdinandusse, S.; Denis, S.; Faust, P.L.; Wanders, R.J. Bile acids: The role of peroxisomes. J. Lipid Res. 2009, 50, 2139–2147. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Ji, G.; Zhang, L. The role of p53 in liver fibrosis. Front. Pharmacol. 2022, 13, 1057829. [Google Scholar] [CrossRef] [PubMed]
- Roehlen, N.; Roca Suarez, A.A.; El Saghire, H.; Saviano, A.; Schuster, C.; Lupberger, J.; Baumert, T.F. Tight junction proteins and the biology of hepatobiliary disease. Int. J. Mol. Sci. 2020, 21, 825. [Google Scholar] [CrossRef]
- Zeisel, M.B.; Dhawan, P.; Baumert, T.F. Tight junction proteins in gastrointestinal and liver disease. Gut 2019, 68, 547–561. [Google Scholar] [CrossRef]
- Farley-Barnes, K.I.; Baserga, S.J. Ribosome biogenesis and its role in cell growth and proliferation in the liver. In The Liver: Biology and Pathobiology; John Wiley & Sons: Hoboken, NJ, USA, 2020; Chapter 15; pp. 174–182. [Google Scholar]
- Schyman, P.; Xu, Z.; Desai, V.; Wallqvist, A. TOXPANEL: A gene-set analysis tool to assess liver and kidney injuries. Front. Pharmacol. 2021, 12, 601511. [Google Scholar] [CrossRef]
- Zheng, L.; Yin, L.; Xu, L.; Qi, Y.; Li, H.; Xu, Y.; Han, X.; Liu, K.; Peng, J. Protective effect of dioscin against thioacetamide-induced acute liver injury via FXR/AMPK signaling pathway in vivo. Biomed. Pharmacother. 2018, 97, 481–488. [Google Scholar] [CrossRef]
- Ezhilarasan, D. Molecular mechanisms in thioacetamide-induced acute and chronic liver injury models. Environ. Toxicol. Pharmacol. 2023, 99, 104093. [Google Scholar] [CrossRef]
- Roehlen, N.; Crouchet, E.; Baumert, T.F. Liver fibrosis: Mechanistic concepts and therapeutic perspectives. Cells 2020, 9, 875. [Google Scholar] [CrossRef] [PubMed]
- Omar, J.M.; Hai, Y.; Jin, S. Hypoxia-induced factor and its role in liver fibrosis. PeerJ 2022, 10, e14299. [Google Scholar] [CrossRef]
- Lv, X.; Kong, J.; Chen, W.-D.; Wang, Y.-D. The role of the apelin/APJ system in the regulation of liver disease. Front. Pharmacol. 2017, 8, 221. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Liu, X.; Ding, C.; Gu, Y.; Liu, W. Dihydromyricetin reverses thioacetamide-induced liver fibrosis through inhibiting NF-κB-mediated inflammation and TGF-β1-regulated of PI3K/Akt signaling pathway. Front. Pharmacol. 2021, 12, 783886. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Marzioni, M.; Meng, F.; Francis, H.; Glaser, S.; Alpini, G. Ductular reaction in liver diseases: Pathological mechanisms and translational significances. Hepatology 2019, 69, 420–430. [Google Scholar] [CrossRef] [PubMed]
- Gadd, V.L.; Skoien, R.; Powell, E.E.; Fagan, K.J.; Winterford, C.; Horsfall, L.; Irvine, K.; Clouston, A.D. The portal inflammatory infiltrate and ductular reaction in human nonalcoholic fatty liver disease. Hepatology 2014, 59, 1393–1405. [Google Scholar] [CrossRef] [PubMed]
- Desmet, V.; Roskams, T.; Van Eyken, P. Ductular reaction in the livers. Pathol.-Res. Pract. 1995, 191, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Gouw, A.S.; Clouston, A.D.; Theise, N.D. Ductular reactions in human liver: Diversity at the interface. Hepatology 2011, 54, 1853–1863. [Google Scholar] [CrossRef]
- Kohonen, P.; Parkkinen, J.A.; Willighagen, E.L.; Ceder, R.; Wennerberg, K.; Kaski, S.; Grafström, R.C. A transcriptomics data-driven gene space accurately predicts liver cytopathology and drug-induced liver injury. Nat. Commun. 2017, 8, 15932. [Google Scholar] [CrossRef]
- Tawa, G.J.; AbdulHameed, M.D.M.; Yu, X.; Kumar, K.; Ippolito, D.L.; Lewis, J.A.; Stallings, J.D.; Wallqvist, A. Characterization of chemically induced liver injuries using gene co-expression modules. PLoS ONE 2014, 9, e107230. [Google Scholar] [CrossRef]
- Te, J.A.; AbdulHameed, M.D.M.; Wallqvist, A. Systems toxicology of chemically induced liver and kidney injuries: Histopathology-associated gene co-expression modules. J. Appl. Toxicol. 2016, 36, 1137–1149. [Google Scholar] [CrossRef] [PubMed]
- Jetter, A.; Kullak-Ublick, G.A. Drugs and hepatic transporters: A review. Pharmacol. Res. 2020, 154, 104234. [Google Scholar] [CrossRef] [PubMed]
- Tátrai, P.; Erdő, F.; Krajcsi, P. Role of hepatocyte transporters in drug-induced liver injury (DILI)—In Vitro testing. Pharmaceutics 2022, 15, 29. [Google Scholar] [CrossRef] [PubMed]
- Davit-Spraul, A.; Gonzales, E.; Baussan, C.; Jacquemin, E. Progressive familial intrahepatic cholestasis. Orphanet J. Rare Dis. 2009, 4, 1. [Google Scholar] [CrossRef] [PubMed]
- Nicolaou, M.; Andress, E.J.; Zolnerciks, J.K.; Dixon, P.H.; Williamson, C.; Linton, K.J. Canalicular ABC transporters and liver disease. J. Pathol. 2012, 226, 300–315. [Google Scholar] [CrossRef] [PubMed]
- Adam, M.P.; Mirzaa, G.M.; Pagon, R.A.; Wallace, S.E.; Bean, L.J.; Gripp, K.W.; Amemiya, A. GeneReviews® [Internet]; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Paulusma, C.C.; de Waart, D.R.; Kunne, C.; Mok, K.S.; Elferink, R.P.O. Activity of the bile salt export pump (ABCB11) is critically dependent on canalicular membrane cholesterol content. J. Biol. Chem. 2009, 284, 9947–9954. [Google Scholar] [CrossRef] [PubMed]
- Andersen, J.P.; Vestergaard, A.L.; Mikkelsen, S.A.; Mogensen, L.S.; Chalat, M.; Molday, R.S. P4-ATPases as phospholipid flippases—Structure, function, and enigmas. Front. Physiol. 2016, 7, 275. [Google Scholar] [CrossRef]
- Waring, R.H. Cytochrome P450: Genotype to phenotype. Xenobiotica 2020, 50, 9–18. [Google Scholar] [CrossRef]
- Zhao, M.; Ma, J.; Li, M.; Zhang, Y.; Jiang, B.; Zhao, X.; Huai, C.; Shen, L.; Zhang, N.; He, L. Cytochrome P450 enzymes and drug metabolism in humans. Int. J. Mol. Sci. 2021, 22, 12808. [Google Scholar] [CrossRef]
- Hewitt, N.J.; Gómez Lechón, M.J.; Houston, J.B.; Hallifax, D.; Brown, H.S.; Maurel, P.; Kenna, J.G.; Gustavsson, L.; Lohmann, C.; Skonberg, C. Primary hepatocytes: Current understanding of the regulation of metabolic enzymes and transporter proteins, and pharmaceutical practice for the use of hepatocytes in metabolism, enzyme induction, transporter, clearance, and hepatotoxicity studies. Drug Metab. Rev. 2007, 39, 159–234. [Google Scholar] [CrossRef]
- Zolfaghari, R.; Cifelli, C.J.; Lieu, S.O.; Chen, Q.; Li, N.-q.; Ross, A.C. Lipopolysaccharide opposes the induction of CYP26A1 and CYP26B1 gene expression by retinoic acid in the rat liver in vivo. Am. J. Physiol.-Gastrointest. Liver Physiol. 2007, 292, G1029–G1036. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bawa, F.N.C.; Zhang, Y. Retinoic acid signaling in fatty liver disease. Liver Res. 2023, 7, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Czuba, L.C.; Wu, X.; Huang, W.; Hollingshead, N.; Roberto, J.B.; Kenerson, H.L.; Yeung, R.S.; Crispe, I.N.; Isoherranen, N. Altered vitamin A metabolism in human liver slices corresponds to fibrogenesis. Clin. Transl. Sci. 2021, 14, 976–989. [Google Scholar] [CrossRef] [PubMed]
- Shirakami, Y.; Lee, S.-A.; Clugston, R.D.; Blaner, W.S. Hepatic metabolism of retinoids and disease associations. Biochim. Et Biophys. Acta Mol. Cell Biol. Lipids 2012, 1821, 124–136. [Google Scholar] [CrossRef] [PubMed]
- Pettinelli, P.; Arendt, B.M.; Teterina, A.; McGilvray, I.; Comelli, E.M.; Fung, S.K.; Fischer, S.E.; Allard, J.P. Altered hepatic genes related to retinol metabolism and plasma retinol in patients with non-alcoholic fatty liver disease. PLoS ONE 2018, 13, e0205747. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Gong, L.; Fang, Y.; Zhan, Q.; Liu, H.-X.; Lu, Y.; Guo, G.L.; Lehman-McKeeman, L.; Fang, J.; Wan, Y.-J.Y. The role of retinoic acid in hepatic lipid homeostasis defined by genomic binding and transcriptome profiling. BMC Genom. 2013, 14, 575. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.S.; Wanibuchi, H.; Morimura, K.; Wongpoomchai, R.; Chusiri, Y.; Gonzalez, F.J.; Fukushima, S. Role of CYP2E1 in thioacetamide-induced mouse hepatotoxicity. Toxicol. Appl. Pharmacol. 2008, 228, 295–300. [Google Scholar] [CrossRef]
- Leung, T.-M.; Nieto, N. CYP2E1 and oxidant stress in alcoholic and non-alcoholic fatty liver disease. J. Hepatol. 2013, 58, 395–398. [Google Scholar] [CrossRef]
- Cichoż-Lach, H.; Michalak, A. Oxidative stress as a crucial factor in liver diseases. World J. Gastroenterol. 2014, 20, 8082. [Google Scholar] [CrossRef]
- Marciniak, S.; Wnorowski, A.; Smolińska, K.; Walczyna, B.; Turski, W.; Kocki, T.; Paluszkiewicz, P.; Parada-Turska, J. Kynurenic acid protects against thioacetamide-induced liver injury in rats. Anal. Cell. Pathol. 2018, 2018, 1270483. [Google Scholar] [CrossRef]
- Tracy, T.S.; Chaudhry, A.S.; Prasad, B.; Thummel, K.E.; Schuetz, E.G.; Zhong, X.-B.; Tien, Y.-C.; Jeong, H.; Pan, X.; Shireman, L.M. Interindividual variability in cytochrome P450-mediated drug metabolism. Drug Metab. Dispos. 2016, 44, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Shirasaka, Y.; Chaudhry, A.S.; McDonald, M.; Prasad, B.; Wong, T.; Calamia, J.C.; Fohner, A.; Thornton, T.A.; Isoherranen, N.; Unadkat, J.D. Interindividual variability of CYP2C19-catalyzed drug metabolism due to differences in gene diplotypes and cytochrome P450 oxidoreductase content. Pharmacogenomics J. 2016, 16, 375–387. [Google Scholar] [CrossRef]
- Li, A.P.; Ho, M.C.D.; Alam, N.; Mitchell, W.; Wong, S.; Yan, Z.; Kenny, J.R.; Hop, C.E.C.A. Inter-individual and inter-regional variations in enteric drug metabolizing enzyme activities: Results with cryopreserved human intestinal mucosal epithelia (CHIM) from the small intestines of 14 donors. Pharmacol. Res. Perspect. 2020, 8, e00645. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhang, B.; Lin, L.-L.; Zhao, S. Evaluation and comparison of computational tools for RNA-seq isoform quantification. BMC Genom. 2017, 18, 583. [Google Scholar] [CrossRef]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 2016, 34, 525. [Google Scholar]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef] [PubMed]
- Pimentel, H.; Bray, N.L.; Puente, S.; Melsted, P.; Pachter, L. Differential analysis of RNA-seq incorporating quantification uncertainty. Nat. Methods 2017, 14, 687. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Tan, Q.; Collins, J.R.; Alvord, W.G.; Roayaei, J.; Stephens, R.; Baseler, M.W.; Lane, H.C.; Lempicki, R.A. The DAVID Gene Functional Classification Tool: A novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 2007, 8, R183. [Google Scholar] [CrossRef]




| Overlap | 12 mM Thioacetamide | ||||
|---|---|---|---|---|---|
| Lots | A | B | C | D | E |
| A | 2821 | 1631 | 1542 | 1302 | 1609 |
| B | 3128 | 1970 | 1600 | 1567 | |
| C | 3521 | 1548 | 1444 | ||
| D | 2241 | 1275 | |||
| E | 2347 | ||||
| Gene (Down) | log2(FC) | SD | Gene (Up) | log2(FC) | SD |
|---|---|---|---|---|---|
| CYP26B1 | −4.50 | 1.66 | C2CD4B | 2.42 | 0.39 |
| CYP26A1 | −3.75 | 2.00 | C2CD4A | 2.40 | 0.27 |
| IFNGR1 | −2.53 | 3.03 | SERPINA1 | 2.32 | 4.92 |
| MCM10 | −2.15 | 0.78 | DDAH2 | 2.08 | 2.17 |
| STEAP4 | −1.95 | 0.62 | THRSP | 1.85 | 0.52 |
| GNAO1 | −1.75 | 1.74 | FGF21 | 1.81 | 0.22 |
| SPP1 | −1.44 | 0.28 | PIWIL2 | 1.35 | 0.37 |
| LPCAT1 | −1.44 | 1.12 | FUT1 | 1.32 | 0.23 |
| DTL | −1.41 | 0.18 | SNORC | 1.26 | 0.08 |
| WDR76 | −1.33 | 0.23 | SLCO4C1 | 1.21 | 0.35 |
| KEGG Pathway (HD, 24 h) | Benjamini p-Value |
|---|---|
| Steroid biosynthesis | 9.4 × 10−6 |
| Peroxisome | 6.5 × 10−5 |
| Metabolic pathways | 1.3 × 10−3 |
| Carbon metabolism | 4.8 × 10−3 |
| Steroid hormone biosynthesis | 8.4 × 10−3 |
| Glyoxylate and dicarboxylate metabolism | 9.2 × 10−3 |
| Glycine, serine, and threonine metabolism | 1.2 × 10−2 |
| Biosynthesis of cofactors | 1.2 × 10−2 |
| Tight junction | 1.6 × 10−2 |
| p53 signaling pathway | 2.0 × 10−2 |
| Biosynthesis of amino acids | 2.4 × 10−2 |
| Terpenoid backbone biosynthesis | 3.5 × 10−2 |
| Ribosome biogenesis in eukaryotes | 5.0 × 10−2 |
| Biosynthesis of unsaturated fatty acids | 7.0 × 10−2 |
| DNA replication | 7.0 × 10−2 |
| Lots | Gene Level Feature Sets | ||
|---|---|---|---|
| PTGS (All) | PTGS (Core) | TMG | |
| A | 0.75 | 0.71 | 0.68 |
| B | 0.80 | 0.78 | 0.70 |
| C | 0.94 | 0.89 | 0.83 |
| D | 0.88 | 0.80 | 0.79 |
| E | 0.85 | 0.81 | 0.74 |
| Drug Transporters | Names | Gene Symbol | A | B | C | D | E | Avg. | SD |
|---|---|---|---|---|---|---|---|---|---|
| Basolateral or sinusoidal uptake transporters | OATP1B3 | SLCO1B3 | 0.37 | −0.10 | −0.33 | −0.34 | 1.27 | 0.17 | 0.68 |
| OATP2B1 | SLCO2B1 | 0.76 | 0.29 | 1.19 | 0.58 | −2.49 | 0.07 | 1.46 | |
| OAT2 | SLC22A7 | 2.56 | −1.88 | −3.97 | −1.59 | 4.25 | −0.13 | 3.40 | |
| OAT7 | SLC22A9 | 0.08 | 0.67 | 0.86 | 0.45 | 0.14 | 0.44 | 0.33 | |
| OCT1 | SLC22A1 | 0.22 | −1.52 | 0.23 | 3.32 | 0.21 | 0.49 | 1.75 | |
| Basolateral efflux transporters | MRP3 | ABCC3 | −0.24 | 0.63 | 1.45 | −0.65 | 0.97 | 0.43 | 0.86 |
| MRP4 | ABCC4 | 0.24 | −1.81 | −1.57 | 2.19 | 0.37 | −0.11 | 1.63 | |
| Canalicular efflux transporters | MRP2 * | ABCC2 | 0.24 | 0.37 | 0.25 | 0.32 | 0.30 | 0.30 | 0.06 |
| FIC1 * | ATP8B1 | −0.22 | −0.36 | −0.55 | −0.49 | −0.16 | −0.36 | 0.17 | |
| BCRP * | ABCG2 | 0.25 | 0.44 | 0.14 | 0.18 | 0.27 | 0.26 | 0.12 | |
| Pgp (MDR1) | ABCB1 | 0.20 | −0.63 | 0.90 | 0.14 | 5.27 | 1.18 | 2.35 | |
| MDR3 | ABCB4 | −0.41 | 0.37 | 0.85 | −2.17 | −0.48 | −0.37 | 1.15 | |
| SLC transporters | SLC16A11 | 3.94 | 0.64 | 3.61 | 0.66 | 1.85 | 2.14 | 1.58 | |
| SLC16A13 | 0.44 | 0.78 | 0.93 | 1.08 | 0.52 | 0.75 | 0.27 |
| Gene Symbol | A | B | C | D | E | Avg. | SD |
|---|---|---|---|---|---|---|---|
| CYP2C9 | 0.33 | 0.36 | 1.16 | 0.90 | 0.14 | 0.58 | 0.43 |
| CYP3A4 | 0.51 | 2.11 | 1.22 | 0.86 | 0.35 | 1.01 | 0.70 |
| CYP3A5 | 1.05 | 0.87 | 0.87 | 0.91 | 0.90 | 0.92 | 0.07 |
| CYP26A1 | −2.20 | −7.22 | −2.66 | −3.33 | −3.37 | −3.75 | 2.00 |
| CYP26B1 | −2.98 | −5.48 | −4.07 | −3.10 | −6.86 | −4.50 | 1.66 |
| CYP1A1 | 1.02 | 2.02 | 2.86 | 2.89 | −1.46 | 1.46 | 1.81 |
| CYP1A2 | −0.20 | 0.68 | 0.14 | 0.60 | −0.12 | 0.22 | 0.40 |
| CYP2C19 | 0.27 | 0.33 | 0.85 | 0.69 | 0.82 | 0.59 | 0.27 |
| CYP2C8 | −0.63 | −0.29 | −0.28 | 3.45 | −0.38 | 0.37 | 1.73 |
| CYP2E1 | 0.31 | 0.23 | 0.28 | 0.24 | 0.22 | 0.26 | 0.04 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goel, H.; Printz, R.L.; Pannala, V.R.; AbdulHameed, M.D.M.; Wallqvist, A. Probing Liver Injuries Induced by Thioacetamide in Human In Vitro Pooled Hepatocyte Experiments. Int. J. Mol. Sci. 2024, 25, 3265. https://doi.org/10.3390/ijms25063265
Goel H, Printz RL, Pannala VR, AbdulHameed MDM, Wallqvist A. Probing Liver Injuries Induced by Thioacetamide in Human In Vitro Pooled Hepatocyte Experiments. International Journal of Molecular Sciences. 2024; 25(6):3265. https://doi.org/10.3390/ijms25063265
Chicago/Turabian StyleGoel, Himanshu, Richard L. Printz, Venkat R. Pannala, Mohamed Diwan M. AbdulHameed, and Anders Wallqvist. 2024. "Probing Liver Injuries Induced by Thioacetamide in Human In Vitro Pooled Hepatocyte Experiments" International Journal of Molecular Sciences 25, no. 6: 3265. https://doi.org/10.3390/ijms25063265
APA StyleGoel, H., Printz, R. L., Pannala, V. R., AbdulHameed, M. D. M., & Wallqvist, A. (2024). Probing Liver Injuries Induced by Thioacetamide in Human In Vitro Pooled Hepatocyte Experiments. International Journal of Molecular Sciences, 25(6), 3265. https://doi.org/10.3390/ijms25063265





