Deep Paleoproteotyping and Microtomography Revealed No Heart Defect nor Traces of Embalming in the Cardiac Relics of Blessed Pauline Jaricot
Abstract
1. Introduction
2. Results
2.1. Macroscopic Examination
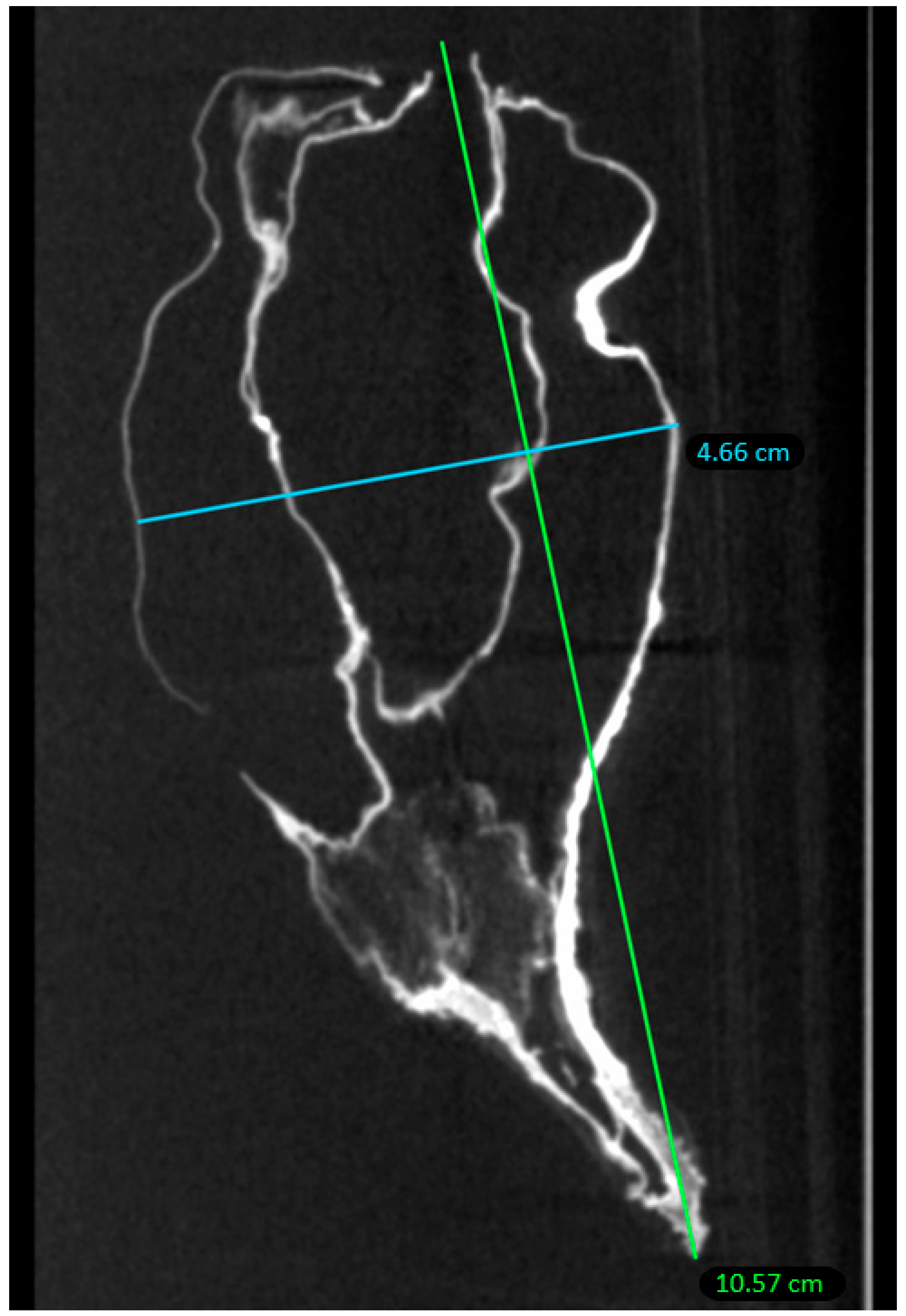
2.2. Micro-CT Scan
2.3. Tandem Mass Spectrometry Proteotyping
2.3.1. Identification of the Eukaryota Taxa Evidenced by Metaproteomics-Based Proteotyping
2.3.2. Characterization of Human Proteins
2.3.3. Identification of the Bacterial Components of the Sample
3. Discussion
4. Materials and Methods
4.1. Cardiotaph Description
4.2. Micro-CT Scan
4.3. Tandem Mass Spectrometry Proteotyping
4.4. Mass Spectrometry Data
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Charlier, P.; Augias, A.; Benmoussa, N.; Rainsard, P.; Froesch, P.; Richardin, P.; Froment, A.; Bianucci, R.; Appenzeller, O.; Perciaccante, A.; et al. The mandible of Saint-Louis (1270 AD): Retrospective diagnosis and circumstances of death. J. Stomatol. Oral Maxillofac. Surg. 2020, 121, 172–174. [Google Scholar] [CrossRef] [PubMed]
- Rubini, G.; Altini, C.; Iuele, F.; Nappi, A.G.; Sardaro, A.; Sablone, S.; Ferrari, C.; Introna, F. The paleoradiology importance in the study of relics: The unique densitometric analysis of a bone relic of Saint Nicholas. Hell J. Nucl. Med. 2019, 22 (Suppl. S2), 164–173. [Google Scholar] [PubMed]
- Charlier, P.; Poupon, J.; Eb, A.; De Mazancourt, P.; Gilbert, M.; Huynh-Charlier, I.; Loublier, Y.; Verhille, A.; Moulheirat, C.; Patou-Mathis, M.; et al. The ‘relics of Joan of Arc’: A forensic multidisciplinary analysis. Forensic Sci. Int. 2010, 194, e9–e15. [Google Scholar] [CrossRef] [PubMed]
- Molin, Y.A.; Agafonov, A.V.; Smolyanitskiy, A.G.; Kravtsov, A.I.; Rubin, A.A. The forensic medical expertise of the holy relics of the righteous martyr Anastasiya of Uglich. Sud. Med. Ekspert 2019, 62, 26–30. [Google Scholar] [CrossRef] [PubMed]
- Martinetto, P.; Riggi, G.; Valtz, A.; Cavallo, G.P. Microbiological research on specimens (dusts) drawn from the Holy Shroud. G. di Batteriol. Virol. Ed Immunol. 1981, 74, 202–208. [Google Scholar]
- Tamburelli, G. Some results in the processing of the holy shroud of turin. IEEE Trans. Pattern Anal. Mach. Intell. 1981, 3, 670–676. [Google Scholar] [CrossRef]
- Charlier, P.; Froesch, P.; Benmoussa, N.; Morin, S.; Augias, A.; Ubelmann, Y.; Weil, R.; Morin, S.; Straub, F.; Deo, S. Computer-Aided Facial Reconstruction of ‘Mary-Magdalene’ Relics Following Hair and Skull Analyses. Clin. Med. Insights Ear Nose Throat 2019, 12, 1179550618821933. [Google Scholar] [CrossRef]
- Warinner, C.; Richter, K.K.; Collins, M.J. Paleoproteomics. Chem. Rev. 2022, 122, 13401–13446. [Google Scholar] [CrossRef]
- Hardouin, P.; Pible, O.; Marchandin, H.; Culotta, K.; Armengaud, J.; Chiron, R.; Grenga, L. Quick and wide-range taxonomical repertoire establishment of the cystic fibrosis lung microbiota by tandem mass spectrometry on sputum samples. Front. Microbiol. 2022, 13, 975883. [Google Scholar] [CrossRef]
- Naïdenoff, G. Pauline Jaricot; Bibliothèque Diocésaine de Versailles, Médiaspaul & Editions Paulines: Versailles, France, 1986. [Google Scholar]
- Truchet, B.; Comby, J.; Paisant, C. Documents Episcopat-Pauline Marie Jaricot, une Œuvre D’amour-n°6/2013; Bibliothèque Diocésaine de Versailles, Publié par le Secrétariat général de la Conférence des Evèques de France: Versailles, France, 2013. [Google Scholar]
- Maurin, J. La Fondatrice de la Propagation de la Foi et du Rosaire-Vivant, Souvenirs D’une Amie sur La Vie, Les Œuvres Et les Epreuves de Pauline-Marie Jaricot; Palme, V., Albanel, J., Eds.; Bibliothèque Diocésaine de Versailles, Société Générale de Librairie Catholique: Versailles, France, 1879. [Google Scholar]
- Servel, J. Un Autre Visage-Textes Inédits de Pauline Jaricot; Bibliothèque Diocésaine de Versailles: Versailles, France, 1963. [Google Scholar]
- Tran, E. Sauvée Par un Miracle; Artège: Paris, France, 2022. [Google Scholar]
- Barabesi, C.; Galizzi, A.; Mastromei, G.; Rossi, M.; Tamburini, E.; Perito, B. Bacillus subtilis gene cluster involved in calcium carbonate biomineralization. J. Bacteriol. 2007, 189, 228–235. [Google Scholar] [CrossRef]
- Ortega-Villamagua, E.; Gudiño-Gomezjurado, M.; Palma-Cando, A. Microbiologically Induced Carbonate Precipitation in the Restoration and Conservation of Cultural Heritage Materials. Molecules 2020, 25, 5499. [Google Scholar] [CrossRef]
- Han, Z.; Zhao, Y.; Yan, H.; Zhao, H.; Han, M.; Sun, B.; Meng, R.; Zhuang, D.; Li, D.; Gao, W.; et al. The Characterization of Intracellular and Extracellular Biomineralization Induced by Synechocystis sp. PCC6803 Cultured under Low Mg/Ca Ratios Conditions. Geomicrobiol. J. 2017, 34, 362–373. [Google Scholar] [CrossRef]
- Rodriguez-Navarro, C.; Jroundi, F.; Gonzalez-Muñoz, M.T. Stone Consolidation by Bacterial Carbonatogenesis: Evaluation of in situ Applications. Restor. Build. Monum. 2015, 21, 9–20. [Google Scholar] [CrossRef]
- González-Muñoz, M.T.; Rodriguez-Navarro, C.; Martínez-Ruiz, F.; Arias, J.M.; Merroun, M.L.; Rodriguez-Gallego, M. Bacterial biomineralization: New insights from Myxococcus-induced mineral precipitation. Geol. Soc. Lond. Spec. Publ. 2010, 336, 31–50. [Google Scholar] [CrossRef]
- Charlier, P.; Perciaccante, A.; Herbin, M.; Appenzeller, O.; Bianucci, R. The Heart of Frederic Chopin (1810–1849). Am. J. Med. 2018, 131, e173–e174. [Google Scholar] [CrossRef]
- Charlier, P.; Huynh-Charlier, I.; Poupon, J.; Fox, C.L.; Keyser, C.; Mougniot, C.; Popescu, S.-M.; Brun, L.; Pietri, S.; Thévenard, F.; et al. The heart of Blessed Anne-Madeleine Remuzat: A biomedical approach of ‘miraculous’ heart conservation. Cardiovasc. Pathol. 2014, 23, 344–350. [Google Scholar] [CrossRef]
- Armengaud, J. Metaproteomics to understand how microbiota function: The crystal ball predicts a promising future. Environ. Microbiol. 2023, 25, 115–125. [Google Scholar] [CrossRef]
- Yimagou, E.K.; Tall, M.; Baudoin, J.-P.; Raoult, D.; Khalil, J.B. Clostridium transplantifaecale sp. nov., a new bacterium isolated from patient with recurrent Clostridium difficile infection. New Microbe. New Infect. 2019, 32, 100598. [Google Scholar] [CrossRef]
- Sen, B.; Asan, A. Fungal flora in indoor and outdoor air of different residential houses in Tekirdag City (Turkey): Seasonal distribution and relationship with climatic factors. Environ. Monit. Assess. 2009, 151, 209–219. [Google Scholar] [CrossRef]
- Fouladi, B.; Tadjrobehkar, O.; Zamanpour, S.; Nooshiravani, Y. Comparison of airborne fungal flora in indoors and outdoors of Zabol city in spring and summer. Arch. Pharm. Pract. 2020, 11, 8. [Google Scholar]
- Šimonovičová, A.; Kraková, L.; Pangallo, D.; Majorošová, M.; Piecková, E.; Bodoriková, S.; Dörnhoferová, M. Fungi on mummified human remains and in the indoor air in the Kuffner family crypt in Sládkovičovo (Slovakia). Int. Biodeterior. Biodegrad. 2015, 99, 157–164. [Google Scholar] [CrossRef]
- Burkowska-But, A.; Drążkowska, A.; Swiontek Brzezinska, M.; Deja-Sikora, E.; Walczak, M. Microbiological hazards associated with archaeological works, illustrated with an example of Fredro Crypt (Przemyśl, Poland). Collegium Antropologicum 2019, 43, 61–68. Available online: https://omega.umk.pl/info/article/UMKeba224c361f6437f9087ae82bc4bf1c2/Publikacja%2B%25E2%2580%2593%2BMicrobiological%2Bhazards%2Bassociated%2Bwith%2Barchaeological%2Bworks%252C%2Billustrated%2Bwith%2Ban%2Bexample%2Bof%2BFredro%2BCrypt%2B%2528Przemy%25C5%259Bl%252C%2BPoland%2529%2B%25E2%2580%2593%2BUniwersytet%2BMiko%25C5%2582aja%2BKopernika%2Bw%2BToruniu?r=publication&ps=20&tab=&lang=pl (accessed on 29 September 2022).
- Naji, K.M.; Abdullah, Q.Y.M.; Al-Zaqri, A.Q.M.; Alghalibi, S.M. Evaluating the Biodeterioration Enzymatic Activities of Fungal Contamination Isolated from Some Ancient Yemeni Mummies Preserved in the National Museum. Biochem. Res. Int. 2014, 2014, 481508. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.F. Mycotoxin production by indoor molds. Fungal Genet. Biol. 2003, 39, 103–117. [Google Scholar] [CrossRef] [PubMed]
- Oyeka, C. Trichophyton mentagrophytes a keratinophilic fungus. Rev. Iberoam. de Micol. 2000, 17, 60–65. [Google Scholar]
- Anbu, P.; Gopinath, S.C.; Hilda, A.; Mathivanan, N.; Annadurai, G. Secretion of keratinolytic enzymes and keratinolysis by Scopulariopsis brevicaulis and Trichophyton mentagrophytes: Regression analysis. Can. J. Microbiol. 2006, 52, 1060–1069. [Google Scholar] [CrossRef]
- Moreno, G.; Arenas, R. Other fungi causing onychomycosis. Clin. Dermatol. 2010, 28, 160–163. [Google Scholar] [CrossRef]
- Rotoli, M.; Sascaro, G.; Cavalieri, S. Aspergillus versicolor infection of the external auditory canal successfully treated with terbinafine. Dermatology 2001, 202, 143. [Google Scholar] [CrossRef]
- Kane, S.; Pinto, J.M.; Dadzie, C.K.; Dawis, M.A.C. Aspergilloma caused by Aspergillus versicolor. Pediatr. Infect. Dis. J. 2014, 33, 891. [Google Scholar] [CrossRef]
- Veraldi, S.; Chiaratti, A.; Harak, H. Onychomycosis caused by Aspergillus versicolor. Mycoses 2010, 53, 363–365. [Google Scholar] [CrossRef]
- Bifrare, Y.-D.; Wolfensberger, T.J. Protracted Aspergillus versicolor endophthalmitis caused by corneal microperforation. Klin. Monbl. Augenheilkd. 2007, 224, 314–316. [Google Scholar] [CrossRef]
- Hodgson, M.J.; Morey, P.; Leung, W.-Y.; Morrow, L.; Miller, D.; Jarvis, B.B.; Robbins, H.; Halsey, J.F.; Storey, E. Building-associated pulmonary disease from exposure to Stachybotrys chartarum and Aspergillus versicolor. J. Occup. Environ. Med. 1998, 40, 241–249. [Google Scholar] [CrossRef]
- Traboulsi, R.S.; Kattar, M.M.; Dbouni, O.; Araj, G.F.; Kanj, S.S. Fatal brain infection caused by Aspergillus glaucus in an immunocompetent patient identified by sequencing of the ribosomal 18S-28S internal transcribed spacer. Eur. J. Clin. Microbiol. Infect. Dis. 2007, 26, 747–750. [Google Scholar] [CrossRef]
- Venugopal, T.V.; Venugopal, P.V. Primary cutaneous aspergillosis from Tamilnadu diagnosed by fine needle aspiration cytology. Med. Mycol. Case Rep. 2012, 1, 103–106. [Google Scholar] [CrossRef]
- Araujo, R.; Pina-Vaz, C.; Rodrigues, A.G. Rodrigues, Susceptibility of environmental versus clinical strains of pathogenic Aspergillus. Int. J. Antimicrob. Agents 2007, 29, 108–111. [Google Scholar] [CrossRef]
- Dagenais, T.R.T.; Keller, N.P. Pathogenesis of Aspergillus fumigatus in Invasive Aspergillosis. Clin. Microbiol. Rev. 2009, 22, 447–465. [Google Scholar] [CrossRef]
- Seif, M.; Kakoschke, T.K.; Ebel, F.; Bellet, M.M.; Trinks, N.; Renga, G.; Pariano, M.; Romani, L.; Tappe, B.; Espie, D.; et al. CAR T cells targeting Aspergillus fumigatus are effective at treating invasive pulmonary aspergillosis in preclinical models. Sci. Transl. Med. 2022, 14, eabh1209. [Google Scholar] [CrossRef]
- Klinger, M.; Theiler, M.; Bosshard, P. Epidemiological and clinical aspects of Trichophyton mentagrophytes/Trichophyton interdigitale infections in the Zurich area: A retrospective study using genotyping. J. Eur. Acad. Dermatol. Venereol. 2021, 35, 1017–1025. [Google Scholar] [CrossRef]
- Ninet, B.; Jan, I.; Bontems, O.; Léchenne, B.; Jousson, O.; Panizzon, R.; Lew, D.; Monod, M. Identification of dermatophyte species by 28S ribosomal DNA sequencing with a commercial kit. J. Clin. Microbiol. 2003, 41, 826–830. [Google Scholar] [CrossRef]
- Heidemann, S.; Monod, M.; Gräser, Y. Signature polymorphisms in the internal transcribed spacer region relevant for the differentiation of zoophilic and anthropophilic strains of Trichophyton interdigitale and other species of T. mentagrophytes sensu lato. Br. J. Dermatol. 2010, 162, 282–295. [Google Scholar] [CrossRef]
- Nenoff, P.; Verma, S.B.; Uhrlaß, S.; Burmester, A.; Gräser, Y. A clarion call for preventing taxonomical errors of dermatophytes using the example of the novel Trichophyton mentagrophytes genotype VIII uniformly isolated in the Indian epidemic of superficial dermatophytosis. Mycoses 2019, 62, 6–10. [Google Scholar] [CrossRef] [PubMed]
- Taghipour, S.; Pchelin, I.M.; Mahmoudabadi, A.Z.; Ansari, S.; Katiraee, F.; Rafiei, A.; Shokohi, T.; Abastabar, M.; Taraskina, A.E.; Kermani, F.; et al. Trichophyton mentagrophytes and T. interdigitale genotypes are associated with particular geographic areas and clinical manifestations. Mycoses 2019, 62, 1084–1091. [Google Scholar] [CrossRef] [PubMed]
- Kosmidis, C.; Denning, D.W. The clinical spectrum of pulmonary aspergillosis. Thorax 2015, 70, 270–277. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.; Heaf, P. Aspergillus in persistent lung cavities after tuberculosis. A report from the Research Committee of the British Tuberculosis Association. Tubercle 1968, 49, 1–11. [Google Scholar]
- Hayoun, K.; Gouveia, D.; Grenga, L.; Pible, O.; Armengaud, J.; Alpha-Bazin, B. Evaluation of Sample Preparation Methods for Fast Proteotyping of Microorganisms by Tandem Mass Spectrometry. Front. Microbiol. 2019, 10, 1985. [Google Scholar] [CrossRef]
- Rubiano-Labrador, C.; Bland, C.; Miotello, G.; Guérin, P.; Pible, O.; Baena, S.; Armengaud, J. Proteogenomic insights into salt tolerance by a halotolerant alpha-proteobacterium isolated from an Andean saline spring. J. Proteom. 2014, 97, 36–47. [Google Scholar] [CrossRef]
- Lozano, C.; Grenga, L.; Gallais, F.; Miotello, G.; Bellanger, L.; Armengaud, J. Mass spectrometry detection of monkeypox virus: Comprehensive coverage for ranking the most responsive peptide markers. Proteomics 2022, 23, e2200253. [Google Scholar] [CrossRef]
- Grenga, L.; Pible, O.; Miotello, G.; Culotta, K.; Ruat, S.; Roncato, M.; Gas, F.; Bellanger, L.; Claret, P.; Dunyach-Remy, C.; et al. Taxonomical and functional changes in COVID-19 faecal microbiome could be related to SARS-CoV-2 faecal load. Environ. Microbiol. 2022, 24, 4299–4316. [Google Scholar] [CrossRef]
- Pible, O.; Allain, F.; Jouffret, V.; Culotta, K.; Miotello, G.; Armengaud, J. Estimating relative biomasses of organisms in microbiota using ‘phylopeptidomics’. Microbiome 2020, 8, 30. [Google Scholar] [CrossRef]
- Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Gonzales, N.R.; Gwadz, M.; Lu, S.; Marchler, G.H.; Song, J.S.; Thanki, N.; Yamashita, R.A.; et al. The conserved domain database in 2023. Nucleic Acids Res. 2022, 51, gkac1096. [Google Scholar] [CrossRef]





| SK | TSMs | spePEPs | PHYLUM | TSMs | spePEPs | CLASS | TSMs | spePEPs | ORDER | TSMs | spePEPs |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Eukaryota | 910 | 301 | Chordata | 367 | 88 | Mammalia | 370 | 88 | Primates | 370 | 88 |
| Ascomycota | 487 | 128 | Eurotiomycetes | 487 | 128 | Eurotiales | 429 | 55 | |||
| Onygenales | 56 | 24 | |||||||||
| Streptophyta | 29 | 14 | Magnoliopsida | 29 | 14 | Brassicales | 31 | 14 | |||
| Arthropoda | 0 | 6 | Insecta | 24 | 6 | Coleoptera | 24 | 6 | |||
| Bacteria | 58 | 25 | Actinobacteria | 26 | 14 | Actinobacteria | 23 | 14 | Pseudonocardiales | 11 | 3 |
| Micrococcales | 8 | 5 | |||||||||
| Streptomycetales | 4 | 4 | |||||||||
| Firmicutes | 0 | 6 | Bacilli | 0 | 3 | Bacillales | 6 | 3 | |||
| Clostridia | 7 | 3 | Clostridiales | 7 | 3 | ||||||
| Proteobacteria | 0 | 1 | Alphaproteobacteria | 15 | 1 | Rhodobacterales | 15 | 1 | |||
| Bacteroidetes | 0 | 4 | Flavobacteriia | 7 | 4 | Flavobacteriales | 7 | 4 | |||
| FAMILY | TSMs | spePEPs | GENUS | TSMs | spePEPs | SPECIES | TSMs | spePEPs | |||
| Hominidae | 370 | 88 | Homo | 370 | 88 | H. sapiens | 374 | 88 | |||
| Aspergillaceae | 429 | 55 | Aspergillus | 429 | 55 | A. versicolor | 378 | 28 | |||
| A. glaucus | 37 | 13 | |||||||||
| Arthrodermataceae | 58 | 24 | Trichophyton | 58 | 24 | T. mentagrophytes | 68 | 24 | |||
| Brassicaceae | 29 | 14 | Brassica | 29 | 14 | B. napus | 29 | 14 | |||
| Chrysomelidae | 24 | 6 | Leptinotarsa | 24 | 6 | L. decemlineata | 24 | 6 | |||
| Pseudonocardiaceae | 12 | 3 | Pseudonocardia | 12 | 3 | P. thermophila | 12 | 3 | |||
| Microbacteriaceae | 7 | 5 | Microbacterium | 7 | 5 | M. hydrocarbonoxydans | 7 | 5 | |||
| Streptomycetaceae | 4 | 4 | Streptomyces | 4 | 4 | S. bungoensis | 4 | 4 | |||
| Bacillaceae | 2 | 2 | Bacillus | 2 | 2 | B. caseinilyticus | 2 | 2 | |||
| Paenibacillaceae | 0 | 1 | Paenibacillus | 4 | 1 | P. albidus | 4 | 1 | |||
| Clostridiaceae | 7 | 3 | Clostridium | 7 | 3 | C. transplantifaecale | 7 | 3 | |||
| Rhodobacteraceae | 15 | 1 | Rhodobacter | 15 | 1 | R. ovatus | 15 | 1 | |||
| Flavobacteriaceae | 7 | 4 | Flavobacterium | 7 | 4 | F. cucumis | 4 | 1 | |||
| Protein Accession | Functional Description | Assigned Peptides | MS/MS Spectra | Protein Group |
|---|---|---|---|---|
| EAW81588.1 | Serpin peptidase inhibitor | 11 | 85 | A (8 shared peptides) |
| CAA25459.1 | Serpin-like protein | 9 | 5 | A (8 shared peptides) |
| AQN67653.1 | Hemoglobin subunit alpha | 2 | 34 | |
| AAY46275.1 | Beta globin chain, partial | 3 | 22 | |
| NP_006112.3 | Keratin, type II cytoskeletal 1 | 9 | 19 | |
| EAX11019.1 | Titin, isoform CRA_e | 8 | 13 | |
| NP_000412.4 | Keratin, type I cytoskeletal 10 isoform 1 | 5 | 11 | B (3 shared peptides) |
| CAA60378.1 | Keratin, type I | 2 | 1 | B (3 shared peptides) |
| XP_005257116.2 | Collagen alpha-1(I) chain isoform X3 | 2 | 9 | |
| NP_005208.1 | Neutrophil defensin 3 | 3 | 9 | |
| NP_000217.2 | Keratin, type I cytoskeletal 9 | 4 | 8 | |
| AAC15854.1 | Ribosomal protein S13 | 1 | 7 | |
| CAB56534.1 | Fibrillin 15 | 1 | 7 | |
| AAI28106.1 | Histone H4B | 2 | 6 | |
| BAG60658.1 | Albumin domain-containing protein | 3 | 5 | |
| AAB59495.1 | Alpha-1-antitrypsin | 1 | 4 | |
| EAW56540.1 | Rho family protein | 1 | 4 | |
| BAB79461.1 | Ribosomal protein L15 | 1 | 3 | |
| NP_001952.1 | Elongation factor 2 | 1 | 3 | |
| AAH42586.1 | Collagen, type I, alpha 2 | 2 | 3 | |
| ACE96343.1 | 5-hydroxytryptamine (serotonin) receptor 2B | 1 | 2 | |
| BAD92412.1 | Collagen, type V, alpha 3 | 1 | 2 | |
| BAG61384.1 | Keratin type II | 2 | 1 | |
| NP_001373235.1 | Antithrombin-III isoform 7 | 1 | 1 | |
| XP_011533341.1 | Sacsin isoform X2 | 1 | 1 | |
| EAW90166.1 | Translation initiation factor 4A, isoform 1 | 1 | 1 | |
| NP_001837.2 | Collagen alpha-2(IV) chain | 1 | 1 | |
| EAW94269.1 | Proteasome 26S subunit, ATPase 5 | 1 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bourdin, V.; Charlier, P.; Crevat, S.; Slimani, L.; Chaussain, C.; Kielbasa, M.; Pible, O.; Armengaud, J. Deep Paleoproteotyping and Microtomography Revealed No Heart Defect nor Traces of Embalming in the Cardiac Relics of Blessed Pauline Jaricot. Int. J. Mol. Sci. 2023, 24, 3011. https://doi.org/10.3390/ijms24033011
Bourdin V, Charlier P, Crevat S, Slimani L, Chaussain C, Kielbasa M, Pible O, Armengaud J. Deep Paleoproteotyping and Microtomography Revealed No Heart Defect nor Traces of Embalming in the Cardiac Relics of Blessed Pauline Jaricot. International Journal of Molecular Sciences. 2023; 24(3):3011. https://doi.org/10.3390/ijms24033011
Chicago/Turabian StyleBourdin, Virginie, Philippe Charlier, Stéphane Crevat, Lotfi Slimani, Catherine Chaussain, Mélodie Kielbasa, Olivier Pible, and Jean Armengaud. 2023. "Deep Paleoproteotyping and Microtomography Revealed No Heart Defect nor Traces of Embalming in the Cardiac Relics of Blessed Pauline Jaricot" International Journal of Molecular Sciences 24, no. 3: 3011. https://doi.org/10.3390/ijms24033011
APA StyleBourdin, V., Charlier, P., Crevat, S., Slimani, L., Chaussain, C., Kielbasa, M., Pible, O., & Armengaud, J. (2023). Deep Paleoproteotyping and Microtomography Revealed No Heart Defect nor Traces of Embalming in the Cardiac Relics of Blessed Pauline Jaricot. International Journal of Molecular Sciences, 24(3), 3011. https://doi.org/10.3390/ijms24033011









