Identification of miRNA Associated with Trichomonas gallinae Resistance in Pigeon (Columba livia)
Abstract
:1. Introduction
2. Results
2.1. Identification and Characterization of miRNAs
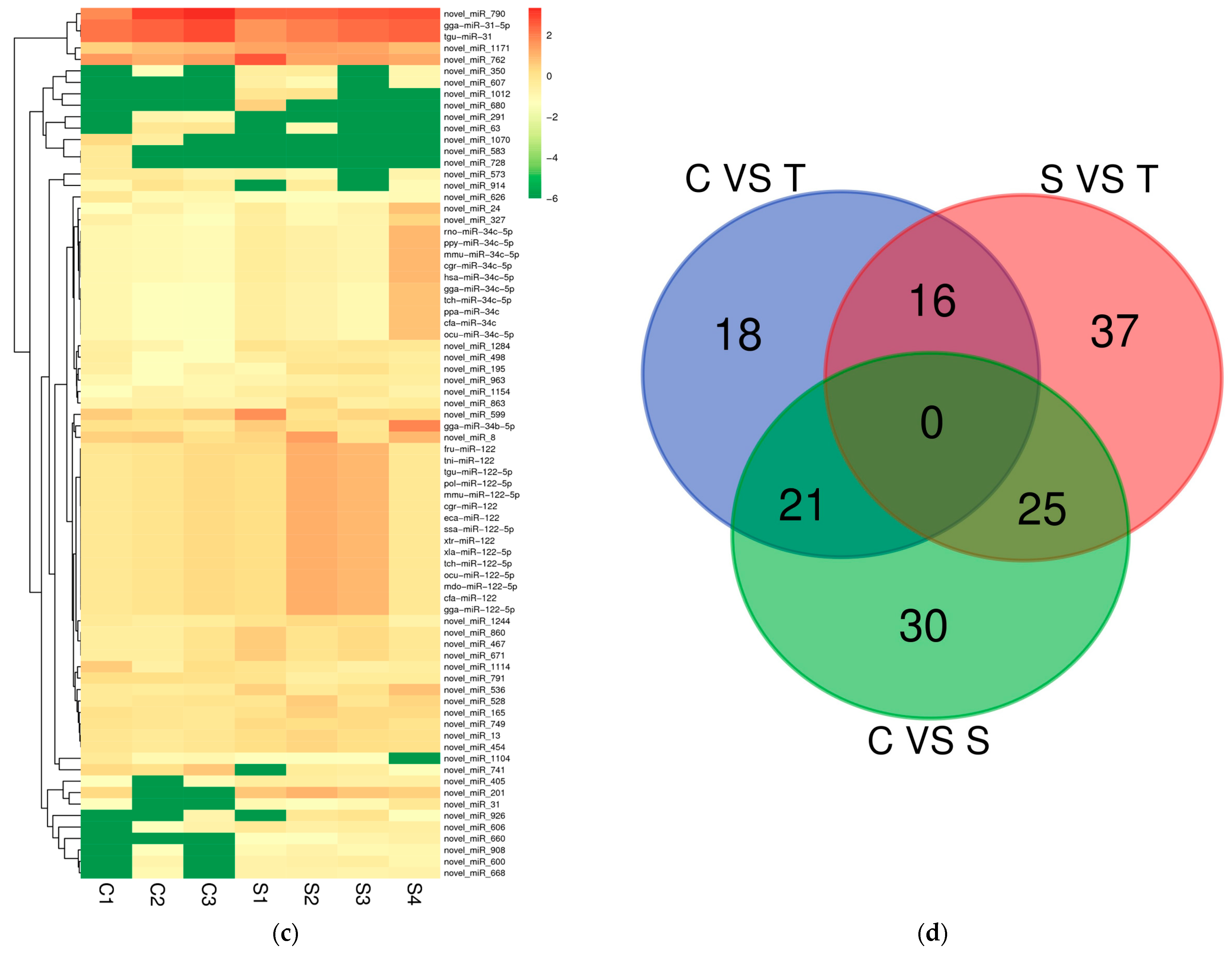
2.2. Differential Expression miRNAs
2.3. miRNA Target Prediction and Analysis
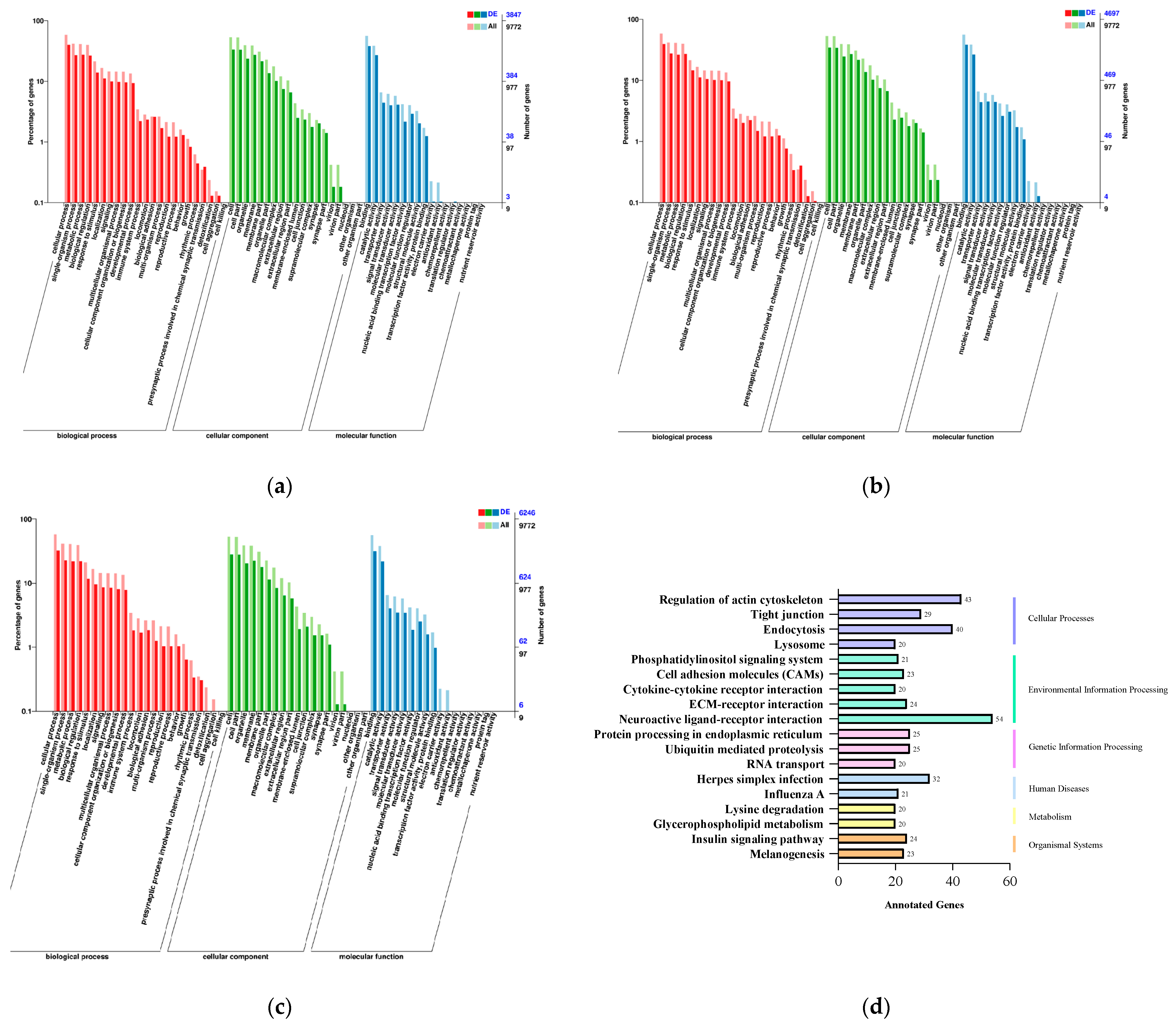
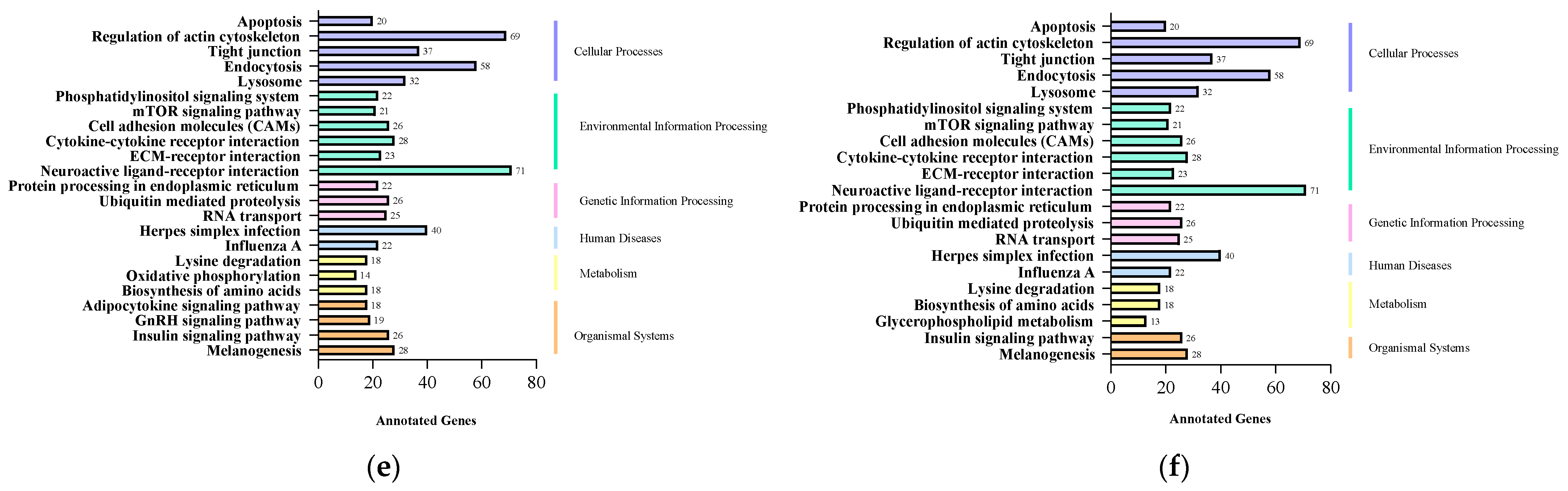
2.4. GO and KEGG Enrichment Analysis of Target Genes of Differentially Expressed miRNAs
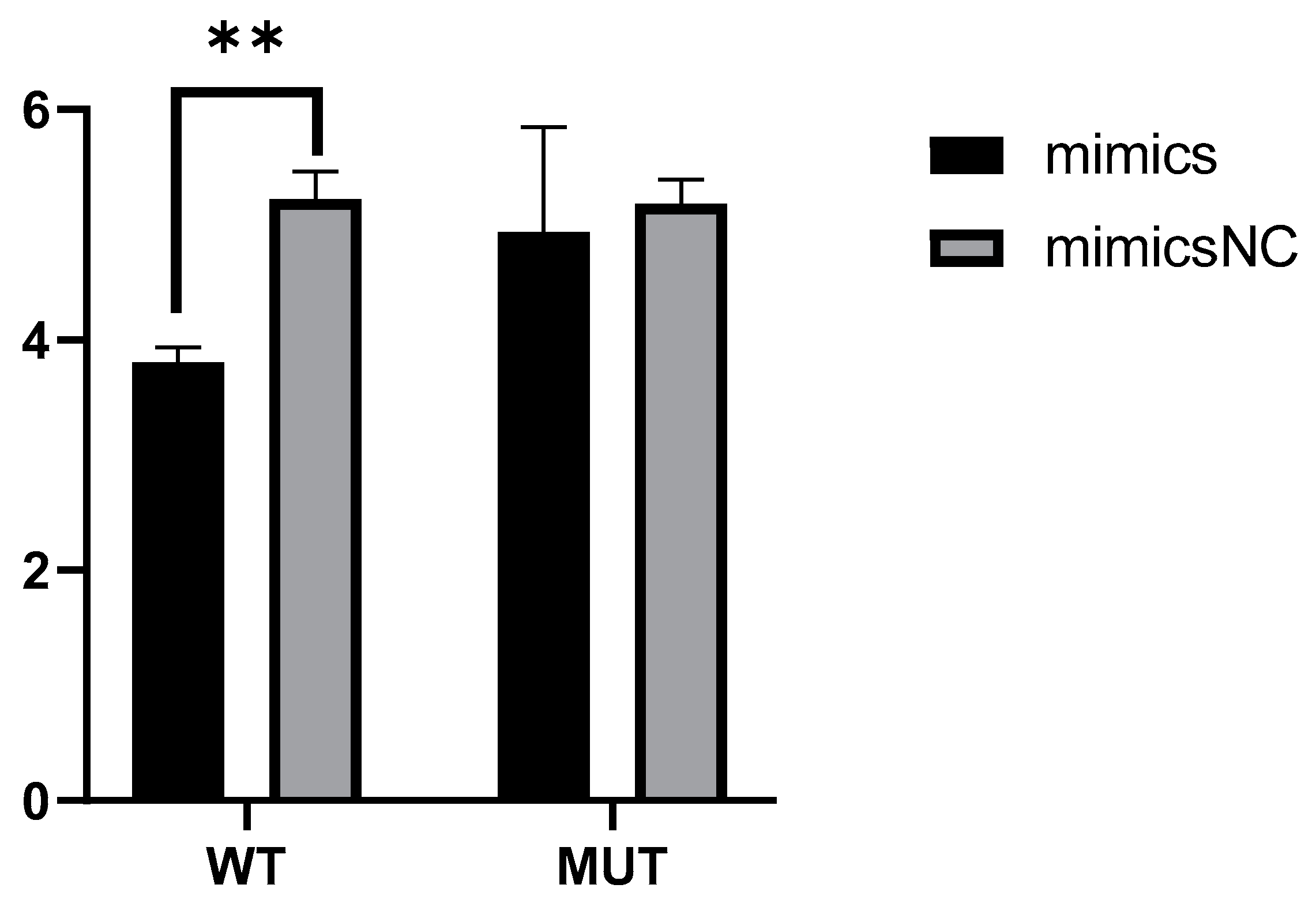
2.5. Novel_miR_741 Down-Regulate the Transcriptional Activity of PRKCQ
3. Discussion
4. Materials and Methods
4.1. T. gallinae Inoculation and Examination
4.2. RNA Isolation and Quality Assessment
4.3. RNA Sequencing and Quality Control
4.4. miRNA Identification
4.5. Differentially Expressed Analysis of miRNA
4.6. Luciferase Reporter Assays
4.7. Data Accessibility
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Anderson, C.; Parra, R.S.; Chapman, H.; Steinemer, A.; Porter, B.; Colombo, M. Pigeon nidopallium caudolaterale, entopallium, and mesopallium ventrolaterale neural responses during categorisation of Monet and Picasso paintings. Sci. Rep. 2020, 10, 15971. [Google Scholar] [CrossRef] [PubMed]
- Castro, L.; Remund Wiger, E.; Wasserman, E. Focusing and shifting attention in pigeon category learning. J. Exp. Psychol. Anim. Learn. Cogn. 2021, 47, 371–383. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Chen, C.; Han, S.; Chen, L.; Ding, H.; Lin, Y.; Zhang, G.; Xie, K.; Wang, J.; Dai, G. Integrated Analysis Reveals a lncRNA-miRNA-mRNA Network Associated with Pigeon Skeletal Muscle Development. Genes 2021, 12, 1787. [Google Scholar] [CrossRef]
- Ge, P.; Ma, H.; Li, Y.; Ni, A.; Isa, A.M.; Wang, P.; Bian, S.; Shi, L.; Zong, Y.; Wang, Y.; et al. Identification of microRNA-Associated-ceRNA Networks Regulating Crop Milk Production in Pigeon (Columba livia). Genes 2020, 12, 39. [Google Scholar] [CrossRef]
- Amin, A.; Bilic, I.; Liebhart, D.; Hess, M. Trichomonads in birds—A review. Parasitology 2014, 141, 733–747. [Google Scholar] [CrossRef]
- Borji, H.; Razmi, G.H.; Movassaghi, A.H.; Moghaddas, E.; Azad, M. Correction to: Prevalence and pathological lesion of Trichomonas gallinae in pigeons of Iran. J. Parasit. Dis. 2021, 45, 1172. [Google Scholar] [CrossRef]
- Chen, L.; Heikkinen, L.; Wang, C.; Yang, Y.; Sun, H.; Wong, G. Trends in the development of miRNA bioinformatics tools. Brief. Bioinform. 2019, 20, 1836–1852. [Google Scholar] [CrossRef]
- Ricafrente, A.; Cwiklinski, K.; Nguyen, H.; Dalton, J.P.; Tran, N.; Donnelly, S. Stage-specific miRNAs regulate gene expression associated with growth, development and parasite-host interaction during the intra-mammalian migration of the zoonotic helminth parasite Fasciola hepatica. BMC Genom. 2022, 23, 419. [Google Scholar] [CrossRef]
- Xu, Y.; Li, H.-Y.; Cai, L.; Xie, S.-C.; Zou, Y.; Zhu, X.-Q.; Zheng, W.-B. Differential Spleen miRNA Expression Profile of Beagle Dogs Infected with Toxocara canis. Animals 2022, 12, 2638. [Google Scholar] [CrossRef]
- Ming, Z.; Zhou, R.; Chen, X.-M. Regulation of Host Epithelial Responses to Cryptosporidium Infection by MicroRNAs. Parasite Immunol. 2017, 39, e12408. [Google Scholar] [CrossRef] [PubMed]
- Zeiner, G.M.; Norman, K.L.; Thomson, J.M.; Hammond, S.M.; Boothroyd, J.C. Toxoplasma gondii infection specifically increases the levels of key host microRNAs. PLoS ONE 2010, 5, e8742. [Google Scholar] [CrossRef] [PubMed]
- Heydarzadeh, S.; Ranjbar, M.; Karimi, F.; Seif, F.; Alivand, M.R. Overview of host miRNA properties and their association with epigenetics, long non-coding RNAs, and Xeno-infectious factors. Cell Biosci. 2021, 11, 43. [Google Scholar] [CrossRef]
- Lodde, V.; Floris, M.; Muroni, M.R.; Cucca, F.; Idda, M.L. Non-coding RNAs in malaria infection. Wiley Interdiscip Rev. RNA 2021, 13, e1697. [Google Scholar] [CrossRef]
- Yuan, D.; Luo, S.; Xu, L.; Zeng, X.; Wu, Z. Regulatory effect of host miR-101b-3p on parasitism of nematode Angiostrongylus cantonensis via superoxide dismutase 3. Biochim. Biophys Acta Gene Regul. Mech. 2019, 1862, 557–566. [Google Scholar] [CrossRef]
- Acuña, S.M.; Floeter-Winter, L.M.; Muxel, S.M. MicroRNAs: Biological Regulators in Pathogen-Host Interactions. Cells 2020, 9, 113. [Google Scholar] [CrossRef]
- He, X.; Sai, X.; Chen, C.; Zhang, Y.; Xu, X.; Zhang, D.; Pan, W. Host serum miR-223 is a potential new biomarker for Schistosoma japonicum infection and the response to chemotherapy. Parasit Vectors 2013, 6, 272. [Google Scholar] [CrossRef]
- Wang, X.; Yan, P.; Feng, S.; Luo, Y.; Liang, J.; Zhao, L.; Liu, H.; Tang, Q.; Long, K.; Jin, L.; et al. Identification and expression pattern analysis of miRNAs in pectoral muscle during pigeon (Columba livia) development. PeerJ 2021, 9, e11438. [Google Scholar] [CrossRef]
- Yuan, J.; Ni, A.; Li, Y.; Bian, S.; Liu, Y.; Wang, P.; Shi, L.; Isa, A.M.; Ge, P.; Sun, Y.; et al. Transcriptome Analysis Revealed Potential Mechanisms of Resistance to Trichomoniasis gallinae Infection in Pigeon (Columba livia). Front. Vet. Sci. 2021, 8, 672270. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Cai, X.; Bradley, J.E. microRNAs in parasites and parasite infection. RNA Biol. 2013, 10, 371–379. [Google Scholar] [CrossRef]
- Wang, Y.; Yang, H.-M.; Cao, W.; Li, Y.-B.; Wang, Z.-Y. Deep sequencing identification of miRNAs in pigeon ovaries illuminated with monochromatic light. BMC Genom. 2018, 19, 446. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Feng, S.; Wang, X.; Qazi, I.H.; Long, K.; Luo, Y.; Li, G.; Ning, C.; Wang, Y.; Hu, S.; et al. Exploration of exosomal microRNA expression profiles in pigeon “Milk” during the lactation period. BMC Genom. 2018, 19, 828. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Nicolas, E.; Marks, D.; Sander, C.; Lerro, A.; Buendia, M.A.; Xu, C.; Mason, W.S.; Moloshok, T.; Bort, R.; et al. miR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol. 2004, 1, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Elmén, J.; Lindow, M.; Silahtaroglu, A.; Bak, M.; Christensen, M.; Lind-Thomsen, A.; Hedtjärn, M.; Hansen, J.B.; Hansen, H.F.; Straarup, E.M. Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Res. 2008, 36, 1153–1162. [Google Scholar] [CrossRef]
- Long, J.-K.; Dai, W.; Zheng, Y.-W.; Zhao, S.-P. miR-122 promotes hepatic lipogenesis via inhibiting the LKB1/AMPK pathway by targeting Sirt1 in non-alcoholic fatty liver disease. Mol. Med. 2019, 25, 26. [Google Scholar] [CrossRef]
- Ghosh, J.; Bose, M.; Roy, S.; Bhattacharyya, S.N. Leishmania donovani targets Dicer1 to downregulate miR-122, lower serum cholesterol, and facilitate murine liver infection. Cell Host Microbe 2013, 13, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Giles, T.; van Limbergen, T.; Sakkas, P.; Quinn, L.; Belkhiri, A.; Maes, D.; Kyriazakis, I.; Barrow, P.; Foster, N. Diagnosis of sub-clinical coccidiosis in fast growing broiler chickens by MicroRNA profiling. Genomics 2020, 112, 3218–3225. [Google Scholar] [CrossRef]
- Guo, Y.-X.; Wang, N.; Wu, W.-C.; Li, C.-Q.; Chen, R.-H.; Zhang, Y.; Li, X. The Role of miR-23b in Cancer and Autoimmune Disease. J. Oncol. 2021, 2021, 6473038. [Google Scholar] [CrossRef]
- Yang, L.-Y.; Song, G.-L.; Zhai, X.-Q.; Wang, L.; Liu, Q.-L.; Zhou, M.-S. MicroRNA-331 inhibits development of gastric cancer through targeting musashi1. World J. Gastrointest. Oncol. 2019, 11, 705–716. [Google Scholar] [CrossRef]
- Chen, J.; Li, T.C.; Pang, R.; Yue, X.Z.; Hu, J.; Zhang, W.Q. Genome-Wide Screening and Functional Analysis Reveal That the Specific microRNA nlu-miR-173 Regulates Molting by Targeting Ftz-F1 in Nilaparvata lugens. Front. Physiol. 2018, 9, 1854. [Google Scholar] [CrossRef]
- Li, H.; Zheng, D.; Zhang, B.; Liu, L.; Ou, J.; Chen, W.; Xiong, S.; Gu, Y.; Yang, J. Mir-208 promotes cell proliferation by repressing SOX6 expression in human esophageal squamous cell carcinoma. J. Transl. Med. 2014, 12, 196. [Google Scholar] [CrossRef]
- Wang, Q.; Zhou, H.; Zhu, X.; Jiang, F.; Yu, Q.; Zhang, J.; Ji, Y. miR-208 inhibits myocardial tissues apoptosis in mice with acute myocardial infarction by targeting inhibition of PDCD4. J. Biochem. Mol. Toxicol. 2022, 36, e23202. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.-J.; Liu, H.; Li, P.; Wang, H.-F.; Yang, A.-K.; Di, J.-M.; Jiang, Q.-W.; Yang, Y.; Huang, J.-R.; Yuan, M.-L.; et al. miR-936 Suppresses Cell Proliferation, Invasion, and Drug Resistance of Laryngeal Squamous Cell Carcinoma and Targets GPR78. Front. Oncol. 2020, 10, 60. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Chen, J. miR-583 inhibits the proliferation and invasion of prostate cancer cells by targeting JAK1. Mol. Med. Rep. 2021, 23, 199. [Google Scholar] [CrossRef] [PubMed]
- Zhang, E.Y.; Kong, K.-F.; Altman, A. The yin and yang of protein kinase C-theta (PKCθ): A novel drug target for selective immunosuppression. Adv. Pharmacol. 2013, 66, 267–312. [Google Scholar] [PubMed]
- Guéry, L.; Hugues, S. Th17 Cell Plasticity and Functions in Cancer Immunity. Biomed Res. Int. 2015, 2015, 314620. [Google Scholar] [CrossRef]
- Tan, S.-L.; Zhao, J.; Bi, C.; Chen, X.C.; Hepburn, D.L.; Wang, J.; Sedgwick, J.D.; Chintalacharuvu, S.R.; Na, S. Resistance to experimental autoimmune encephalomyelitis and impaired IL-17 production in protein kinase C theta-deficient mice. J. Immunol. 2006, 176, 2872–2879. [Google Scholar] [CrossRef]
- Pfeifhofer, C.; Kofler, K.; Gruber, T.; Tabrizi, N.G.; Lutz, C.; Maly, K.; Leitges, M.; Baier, G. Protein kinase C theta affects Ca2+ mobilization and NFAT cell activation in primary mouse T cells. J. Exp. Med. 2003, 197, 1525–1535. [Google Scholar] [CrossRef]
- Altman, A.; Kaminski, S.; Busuttil, V.; Droin, N.; Hu, J.; Tadevosyan, Y.; Hipskind, R.A.; Villalba, M. Positive feedback regulation of PLCgamma1/Ca(2+) signaling by PKCtheta in restimulated T cells via a Tec kinase-dependent pathway. Eur. J. Immunol. 2004, 34, 2001–2011. [Google Scholar] [CrossRef]
- Valenzuela, J.O.; Iclozan, C.; Hossain, M.S.; Prlic, M.; Hopewell, E.; Bronk, C.C.; Wang, J.; Celis, E.; Engelman, R.W.; Blazar, B.R.; et al. PKCtheta is required for alloreactivity and GVHD but not for immune responses toward leukemia and infection in mice. J. Clin. Investig. 2009, 119, 3774–3786. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Friedländer, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformaticsv 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Betel, D.; Wilson, M.; Gabow, A.; Marks, D.S.; Sander, C. The microRNA.org resource: Targets and expression. Nucleic Acids Res. 2008, 36, D149–D153. [Google Scholar] [CrossRef] [PubMed]
- Lewis, B.P.; Shih, I.-h.; Jones-Rhoades, M.W.; Bartel, D.P.; Burge, C.B. Prediction of mammalian microRNA targets. Cell 2003, 115, 787–798. [Google Scholar] [CrossRef] [PubMed]
- UniProt Consortium, T. UniProt: The universal protein knowledgebase. Nucleic Acids Res. 2018, 46, 2699. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Tatusov, R.L.; Galperin, M.Y.; Natale, D.A.; Koonin, E.V. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28, 33–36. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Kawashima, S.; Okuno, Y.; Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004, 32, D277–D280. [Google Scholar] [CrossRef]
- Koonin, E.V.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Krylov, D.M.; Makarova, K.S.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N.; Rao, B.S.; et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 2004, 5, R7. [Google Scholar] [CrossRef]
- Eddy, S.R. Profile hidden Markov models. Bioinformatics 1998, 14, 755–763. [Google Scholar] [CrossRef]


| Samples 1 | Raw Reads | Low Quality | Containing ‘N’ Reads | Length < 18 | Length > 30 | Clean Reads | Q30 (%) |
|---|---|---|---|---|---|---|---|
| C1 | 26,719,180 | 0 | 0 | 140,044 | 0 | 26,579,136 | 95.47 |
| C2 | 27,186,806 | 0 | 0 | 131,113 | 0 | 27,055,693 | 94.80 |
| C3 | 32,297,099 | 0 | 0 | 201,598 | 0 | 32,095,501 | 95.84 |
| S1 | 29,038,069 | 0 | 0 | 122,007 | 0 | 28,916,062 | 96.75 |
| S2 | 33,717,404 | 0 | 0 | 250,965 | 0 | 33,466,439 | 97.46 |
| S3 | 30,511,399 | 0 | 0 | 219,741 | 0 | 30,291,658 | 96.65 |
| S4 | 30,764,981 | 0 | 0 | 49,885 | 0 | 30,715,096 | 96.33 |
| T1 | 31,278,720 | 0 | 0 | 184,431 | 0 | 31,094,289 | 96.95 |
| T2 | 33,971,942 | 0 | 0 | 237,057 | 0 | 33,734,885 | 97.19 |
| T3 | 31,791,900 | 0 | 0 | 125,872 | 0 | 31,666,028 | 96.75 |
| T4 | 34,630,509 | 0 | 0 | 106,722 | 0 | 34,523,787 | 96.78 |
| Samples 1 | Total_Reads | Mapped_Reads | Mapped_Reads (+) | Mapped_Reads (−) |
|---|---|---|---|---|
| C1 | 22,132,444 | 15,780,016 (71.30%) | 4,616,447 (20.86%) | 11,163,569 (50.44%) |
| C2 | 24,296,565 | 17,895,382 (73.65%) | 4,593,190 (18.90%) | 13,302,192 (54.75%) |
| C3 | 29,065,079 | 21,596,946 (74.31%) | 6,078,851 (20.91%) | 15,518,095 (53.39%) |
| S1 | 24,687,783 | 18,275,595 (74.03%) | 4,900,642 (19.85%) | 13,374,953 (54.18%) |
| S2 | 27,049,727 | 19,922,935 (73.65%) | 6,423,420 (23.75%) | 13,499,515 (49.91%) |
| S3 | 25,404,509 | 18,372,520 (72.32%) | 5,664,517 (22.30%) | 12,708,003 (50.02%) |
| S4 | 29,284,210 | 21,069,624 (71.95%) | 4,626,527 (15.80%) | 16,443,097 (56.15%) |
| T1 | 22,943,862 | 16,663,501 (72.63%) | 4,499,778 (19.61%) | 12,163,723 (53.02%) |
| T2 | 26,130,297 | 16,922,718 (64.76%) | 6,781,695 (25.95%) | 10,141,023 (38.81%) |
| T3 | 26,485,112 | 19,276,440 (72.78%) | 4,619,085 (17.44%) | 14,657,355 (55.34%) |
| T4 | 30,523,003 | 21,635,872 (70.88%) | 6,087,472 (19.94%) | 15,548,400 (50.94%) |
| Types | All_miRNA 1 | miRNA_with_Target 2 | Target_Gene 3 |
|---|---|---|---|
| Known_miRNA | 1162 | 998 | 9855 |
| Novel_miRNA | 1267 | 1123 | 21,920 |
| Total | 2429 | 2121 | 22,054 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Ni, A.; Zhang, R.; Li, Y.; Yuan, J.; Sun, Y.; Chen, J.; Ma, H. Identification of miRNA Associated with Trichomonas gallinae Resistance in Pigeon (Columba livia). Int. J. Mol. Sci. 2023, 24, 16453. https://doi.org/10.3390/ijms242216453
Li X, Ni A, Zhang R, Li Y, Yuan J, Sun Y, Chen J, Ma H. Identification of miRNA Associated with Trichomonas gallinae Resistance in Pigeon (Columba livia). International Journal of Molecular Sciences. 2023; 24(22):16453. https://doi.org/10.3390/ijms242216453
Chicago/Turabian StyleLi, Xinyi, Aixin Ni, Ran Zhang, Yunlei Li, Jingwei Yuan, Yanyan Sun, Jilan Chen, and Hui Ma. 2023. "Identification of miRNA Associated with Trichomonas gallinae Resistance in Pigeon (Columba livia)" International Journal of Molecular Sciences 24, no. 22: 16453. https://doi.org/10.3390/ijms242216453
APA StyleLi, X., Ni, A., Zhang, R., Li, Y., Yuan, J., Sun, Y., Chen, J., & Ma, H. (2023). Identification of miRNA Associated with Trichomonas gallinae Resistance in Pigeon (Columba livia). International Journal of Molecular Sciences, 24(22), 16453. https://doi.org/10.3390/ijms242216453







