Human Breast Milk miRNAs: Their Diversity and Potential for Preventive Strategies in Nutritional Therapy
Abstract
1. Introduction
2. RNA Content and Mature miRNA Diversity

3. Bioavailability of Mature miRNAs in the Digestive System
4. Potential of miRNAs for In Vivo Treatment of Offspring
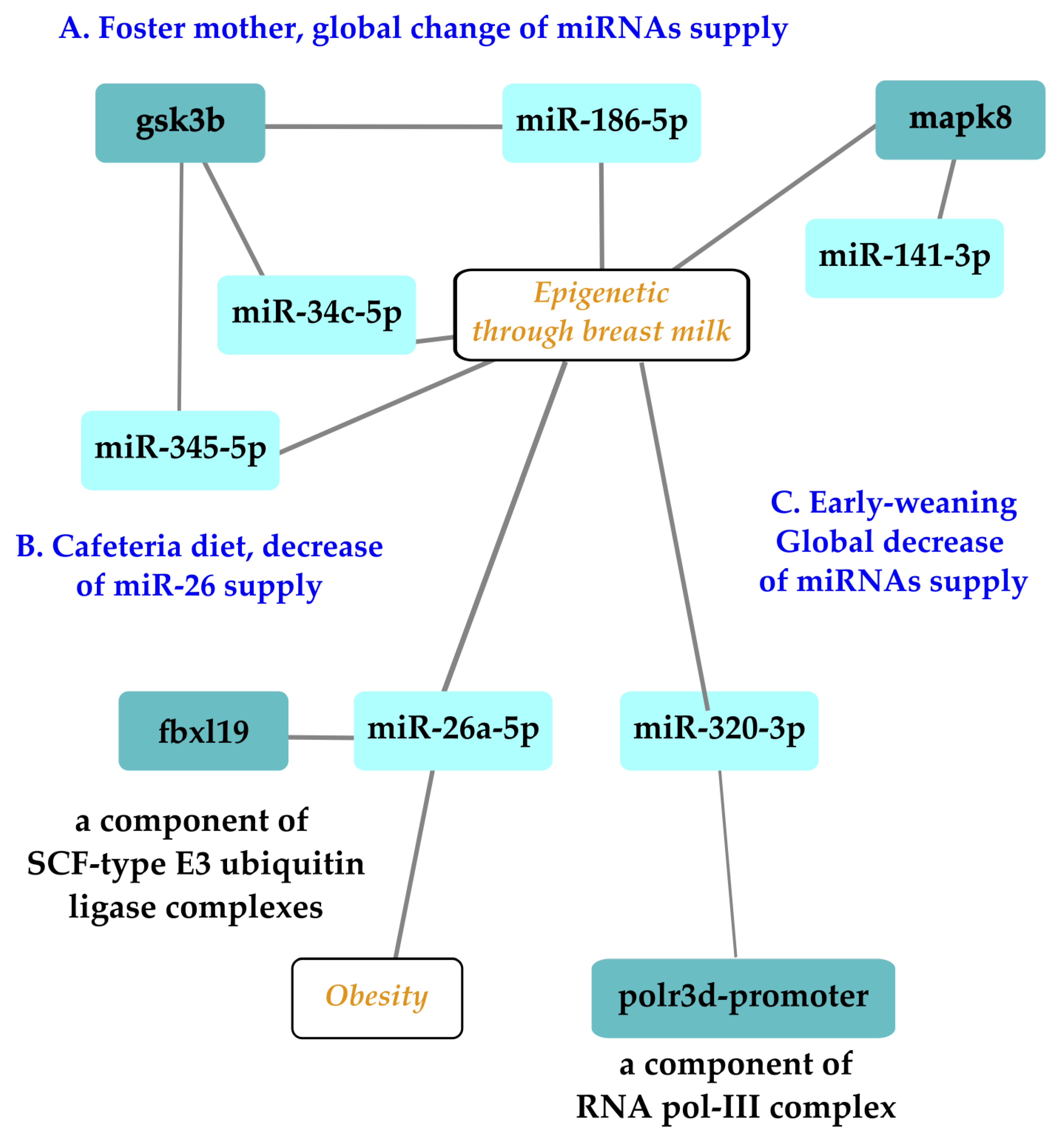
5. Conclusions and Perspectives
Funding
Acknowledgments
Conflicts of Interest
References
- Playford, R.J.; Macdonald, C.E.; Johnson, W.S. Colostrum and milk-derived peptide growth factors for the treatment of gastrointestinal disorders. Am. J. Clin. Nutr. 2000, 72, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Ambros, V.; Bartel, B.; Bartel, D.P.; Burge, C.B.; Carrington, J.C.; Chen, X.; Dreyfuss, G.; Eddy, S.R.; Griffiths-Jones, S.; Marshall, M.; et al. A uniform system for microRNA annotation. RNA 2003, 9, 277–279. [Google Scholar] [CrossRef] [PubMed]
- Fromm, B.; Billipp, T.; Peck, L.E.; Johansen, M.; Tarver, J.E.; King, B.L.; Newcomb, J.M.; Sempere, L.F.; Flatmark, K.; Hovig, E.; et al. A Uniform System for the Annotation of Vertebrate microRNA Genes and the Evolution of the Human microRNAome. Annu. Rev. Genet. 2015, 49, 213–242. [Google Scholar] [CrossRef] [PubMed]
- Alles, J.; Fehlmann, T.; Fischer, U.; Backes, C.; Galata, V.; Minet, M.; Hart, M.; Abu-Halima, M.; Grässer, F.A.; Lenhof, H.-P.; et al. An estimate of the total number of true human miRNAs. Nucleic Acids Res. 2019, 47, 3353–3364. [Google Scholar] [CrossRef]
- Bartel, D.P. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef]
- Lytle, J.R.; Yario, T.A.; Steitz, J.A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl. Acad. Sci. USA 2007, 104, 9667–9672. [Google Scholar] [CrossRef]
- Li, Z.F.; Liang, Y.M.; Lau, P.N.; Shen, W.; Wang, D.K.; Cheung, W.T.; Xue, C.J.; Poon, L.M.; Lam, Y.W. Dynamic Localisation of Mature MicroRNAs in Human Nucleoli is Influenced by Exogenous Genetic Materials. PLoS ONE 2013, 8, e70869. [Google Scholar] [CrossRef]
- Kren, B.T.; Wong, P.Y.-P.; Sarver, A.; Zhang, X.; Zeng, Y.; Steer, C.J. MicroRNAs identified in highly purified liver-derived mitochondria may play a role in apoptosis. RNA Biol. 2009, 6, 65–72. [Google Scholar] [CrossRef]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The MicroRNA Spectrum in 12 Body Fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- Melnik, B.C.; John, S.M.; Schmitz, G. Milk is not just food but most likely a genetic transfection system activating mTORC1 signaling for postnatal growth. Nutr. J. 2013, 12, 103. [Google Scholar] [CrossRef]
- Melnik, B.C.; Schmitz, G. DNA methyltransferase 1-targeting miRNA-148a of dairy milk: A potential bioactive modifier of the human epigenome. Funct. Foods Health Dis. 2017, 7, 671. [Google Scholar] [CrossRef][Green Version]
- Melnik, B.C.; Schmitz, G. MicroRNAs: Milk’s epigenetic regulators. Best Pract. Res. Clin. Endocrinol. Metab. 2017, 31, 427–442. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Milk disrupts p53 and DNMT1, the guardians of the genome: Implications for acne vulgaris and prostate cancer. Nutr. Metab. 2017, 14, 55. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Fung-Leung, W.P.; Bittner, A.; Ngo, K.; Liu, X. Comparison of RNA-Seq and microarray in transcriptome profiling of activated T cells [Comparative Study]. PLoS ONE. 2014, 9, e78644. [Google Scholar]
- Songia, P.; Chiesa, M.; Valerio, V.; Moschetta, D.; Myasoedova, V.A.; D’alessandra, Y.; Poggio, P. Direct screening of plasma circulating microRNAs. RNA Biol. 2018, 15, 1268–1272. [Google Scholar] [CrossRef] [PubMed]
- Raymond, F.; Lefebvre, G.; Texari, L.; Pruvost, S.; Metairon, S.; Cottenet, G.; Zollinger, A.; Mateescu, B.; Billeaud, C.; Picaud, J.-C.; et al. Longitudinal Human Milk miRNA Composition over the First 3 mo of Lactation in a Cohort of Healthy Mothers Delivering Term Infants. J. Nutr. 2022, 152, 94–106. [Google Scholar] [CrossRef]
- Hicks, S.D.; Carney, M.C.; Tarasiuk, A.; DiAngelo, S.L.; Birch, L.L.; Paul, I.M. Breastmilk microRNAs are stable throughout feeding and correlate with maternal weight. Transl. Genet. Genom. 2017, 5, 1–8. [Google Scholar]
- Alsaweed, M.; Lai, C.T.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human milk miRNAs primarily originate from the mammary gland resulting in unique miRNA profiles of fractionated milk. Sci. Rep. 2016, 6, 20680. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, M.; Wang, X.; Li, Q.; Wang, T.; Zhou, X.; Wang, X.; Gao, X.; Li, X. Immune-related MicroRNAs are Abundant in Breast Milk Exosomes. Int. J. Biol. Sci. 2012, 8, 118–123. [Google Scholar] [CrossRef]
- Floris, I.; Billard, H.; Boquien, C.-Y.; Joram-Gauvard, E.; Simon, L.; Legrand, A.; Boscher, C.; Rozé, J.-C.; Bolaños-Jiménez, F.; Kaeffer, B. MiRNA Analysis by Quantitative PCR in Preterm Human Breast Milk Reveals Daily Fluctuations of hsa-miR-16-5p. PLoS ONE 2015, 10, e0140488. [Google Scholar] [CrossRef]
- Holzhausen, E.A.; Kupsco, A.; Chalifour, B.N.; Patterson, W.B.; Schmidt, K.A.; Mokhtari, P.; Baccarelli, A.A.; Goran, M.I.; Alderete, T.L. Influence of technical and maternal-infant factors on the measurement and expression of extracellular miRNA in human milk. Front. Immunol. 2023, 14, 1151870. [Google Scholar] [CrossRef]
- Alexandre-Gouabau, M.-C.; Le Dréan, G.; Kaeffer, B.; Abderrahlan, A.; De Coppet, P.; Bobin, P.; Croyal, M.; De Luca, A.; Hankard, R.; Robitaille, J. Gestational Diabetes Meletis modifies human breast milk content in insulin sensitivity regulators. In Proceedings of the DOHAD, Vancouver, BC, Canada, 27–31 August 2022. [Google Scholar]
- Lukasik, A.; Brzozowska, I.; Zielenkiewicz, U.; Zielenkiewicz, P. Detection of Plant miRNAs Abundance in Human Breast Milk. Int. J. Mol. Sci. 2017, 19, 37. [Google Scholar] [CrossRef]
- Benmoussa, A.; Laugier, J.; Beauparlant, C.J.; Lambert, M.; Droit, A.; Provost, P. Complexity of the microRNA transcriptome of cow milk and milk-derived extracellular vesicles isolated via differential ultracentrifugation. J. Dairy Sci. 2020, 103, 16–29. [Google Scholar] [CrossRef]
- Karlsson, O.; Rodosthenous, R.S.; Jara, C.; Brennan, K.J.; Wright, R.O.; Baccarelli, A.A.; Wright, R.J. Detection of long non-coding RNAs in human breastmilk extracellular vesicles: Implications for early child development. Epigenetics 2016, 11, 721–729. [Google Scholar] [CrossRef]
- Mourtzi, N.; Siahanidou, T.; Tsifintaris, M.; Karamichali, E.; Tasiopoulou, A.; Sertedaki, A.; Pesmatzoglou, M.; Kapetanaki, A.; Liosis, G.; Baltatzis, G.; et al. lncRNA NORAD is consistently detected in breastmilk exosomes and its expression is downregulated in mothers of preterm infants. Int. J. Mol. Med. 2021, 48, 216. [Google Scholar] [CrossRef]
- Leiferman, A.; Shu, J.; Upadhyaya, B.; Cui, J.; Zempleni, J. Storage of Extracellular Vesicles in Human Milk, and MicroRNA Profiles in Human Milk Exosomes and Infant Formulas. J. Pediatr. Gastroenterol. Nutr. 2019, 69, 235–238. [Google Scholar] [CrossRef]
- Alsaweed, M.; Lai, C.T.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human milk cells and lipids conserve numerous known and novel miRNAs, some of which are differentially expressed during lactation. PLoS ONE 2016, 11, e0152610. [Google Scholar] [CrossRef]
- Alsaweed, M.; Lai, C.T.; Hartmann, P.E.; Geddes, D.T.; Kakulas, F. Human Milk Cells Contain Numerous miRNAs that May Change with Milk Removal and Regulate Multiple Physiological Processes. Int. J. Mol. Sci. 2016, 17, 956. [Google Scholar] [CrossRef]
- Munch, E.M.; Harris, R.A.; Mohammad, M.; Benham, A.L.; Pejerrey, S.M.; Showalter, L.; Hu, M.; Shope, C.D.; Maningat, P.D.; Gunaratne, P.H.; et al. Transcriptome Profiling of microRNA by Next-Gen Deep Sequencing Reveals Known and Novel miRNA Species in the Lipid Fraction of Human Breast Milk. PLoS ONE 2013, 8, e50564. [Google Scholar] [CrossRef]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef]
- Desvignes, T.; Batzel, P.; Berezikov, E.; Eilbeck, K.; Eppig, J.T.; McAndrews, M.S.; Singer, A.; Postlethwait, J.H. miRNA No-menclature: A View Incorporating Genetic Origins, Biosynthetic Pathways, and Sequence Variants. Trends Genet. 2015, 31, 613–626. [Google Scholar] [CrossRef] [PubMed]
- Zonneveld, M.I.; Brisson, A.R.; van Herwijnen, M.J.C.; Tan, S.; van de Lest, C.H.A.; Redegeld, F.A.; Garssen, J.; Wauben, M.H.M.; Nolte-’t Hoen, E.N. Recovery of extracellular vesicles from human breast milk is influenced by sample collection and vesicle isolation procedures. J. Extracell. Vesicles 2014, 3, 24215. [Google Scholar] [CrossRef]
- Zheng, Z.; Mo, J.; Lin, F.; Wang, J.; Chen, J.; Luo, H.; Liu, Y.; Su, C.; Gu, X.; Xiong, F.; et al. Milk Exosomes from Gestational Diabetes Mellitus (GDM) and Healthy Parturient Exhibit Differential miRNAs Profiles and Distinct Regulatory Bioactivity on Hepatocyte Proliferation. Mol. Nutr. Food Res. 2023, 67, e2300005. [Google Scholar] [CrossRef]
- Wu, F.; Zhi, X.; Xu, R.; Liang, Z.; Wang, F.; Li, X.; Li, Y.; Sun, B. Exploration of microRNA profiles in human colostrum. Ann. Transl. Med. 2020, 8, 1170. [Google Scholar] [CrossRef] [PubMed]
- Ishibashi, O.; Ohkuchi, A.; Ali, M.; Kurashina, R.; Luo, S.-S.; Ishikawa, T.; Takizawa, T.; Hirashima, C.; Takahashi, K.; Migita, M.; et al. Hydroxysteroid (17-β) Dehydrogenase 1 Is Dysregulated by Mir-210 and Mir-518c That Are Aberrantly Expressed in Preeclamptic Placentas. Hypertension 2012, 59, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Patuleia, S.I.S.; van Gils, C.H.; Cao, A.M.O.; Bakker, M.F.; van Diest, P.J.; van der Wall, E.; Moelans, C.B. The Physiological MicroRNA Landscape in Nipple Aspirate Fluid: Differences and Similarities with Breast Tissue, Breast Milk, Plasma and Serum. Int. J. Mol. Sci. 2020, 21, 8466. [Google Scholar] [CrossRef] [PubMed]
- Layne, T.R.; Green, R.A.; Lewis, C.A.; Nogales, F.; Cruz, T.C.D.; Zehner, Z.E.; Seashols-Williams, S.J. micro RNA Detection in Blood, Urine, Semen, and Saliva Stains After Compromising Treatments. J. Forensic Sci. 2019, 64, 1831–1837. [Google Scholar] [CrossRef]
- Mirza, A.H.; Kaur, S.; Nielsen, L.B.; Størling, J.; Yarani, R.; Roursgaard, M.; Mathiesen, E.R.; Damm, P.; Svare, J.; Mortensen, H.B.; et al. Breast Milk-Derived Extracellular Vesicles Enriched in Exosomes From Mothers With Type 1 Diabetes Contain Aberrant Levels of microRNAs. Front. Immunol. 2019, 10, 2543. [Google Scholar] [CrossRef]
- Ahlberg, E.; Al-Kaabawi, A.; Thune, R.; Simpson, M.R.; Pedersen, S.A.; Cione, E.; Jenmalm, M.C.; Tingö, L. Breast milk microRNAs: Potential players in oral tolerance development. Front. Immunol. 2023, 14, 1154211. [Google Scholar] [CrossRef]
- Chiba, T.; Kooka, A.; Kowatari, K.; Yoshizawa, M.; Chiba, N.; Takaguri, A.; Fukushi, Y.; Hongo, F.; Sato, H.; Wada, S. Expression profiles of hsa-miR-148a-3p and hsa-miR-125b-5p in human breast milk and infant formulae. Int. Breastfeed. J. 2022, 17, 1. [Google Scholar] [CrossRef]
- Li, A.; Li, Y.; Zhang, X.; Zhang, C.; Li, T.; Zhang, J.; Li, C. The human milk oligosaccharide 2′-fucosyllactose attenuates β-lactoglobulin–induced food allergy through the miR-146a–mediated toll-like receptor 4/nuclear factor-κB signaling pathway. J. Dairy Sci. 2021, 104, 10473–10484. [Google Scholar] [CrossRef] [PubMed]
- Kosaka, N.; Izumi, H.; Sekine, K.; Ochiya, T. microRNA as a new immune-regulatory agent in breast milk. Silence 2010, 1, 7. [Google Scholar] [CrossRef]
- Elsarraj, H.S.; Hong, Y.; Valdez, K.; Carletti, M.; Salah, S.M.; Raimo, M.; Taverna, D.; Prochasson, P.; Bharadwaj, U.; Tweardy, D.J.; et al. Novel role of microRNA146b in promoting mammary alveolar progenitor cell maintenance. J. Cell Sci. 2013, 126, 2446–2458. [Google Scholar] [CrossRef] [PubMed]
- Benmoussa, A.; Provost, P. Milk microRNAs in health and disease. Compr. Rev. Food Sci. Food Saf. 2019, 18, 703–722. [Google Scholar] [CrossRef] [PubMed]
- Shah, K.B.; Fields, D.A.; Pezant, N.P.; Kharoud, H.K.; Gulati, S.; Jacobs, K.; Gale, C.A.; Kharbanda, E.O.; Nagel, E.M.; Demerath, E.W.; et al. Gestational Diabetes Mellitus Is Associated with Altered Abundance of Exosomal MicroRNAs in Human Milk. Clin. Ther. 2022, 44, 172–185.e1, Erratum in: Clin Ther. 2022, 44, 1034. [Google Scholar] [CrossRef]
- Golan-Gerstl, R.; Shiff, Y.E.; Moshayoff, V.; Schecter, D.; Leshkowitz, D.; Reif, S. Characterization and biological function of milk-derived miRNAs. Mol. Nutr. Food Res. 2017, 61, 1700009. [Google Scholar] [CrossRef]
- Xi, Y.; Jiang, X.; Li, R.; Chen, M.; Song, W.; Li, X. The levels of human milk microRNAs and their association with maternal weight characteristics. Eur. J. Clin. Nutr. 2015, 70, 445–449. [Google Scholar] [CrossRef]
- Zamanillo, R.; Sánchez, J.; Serra, F.; Palou, A. Breast Milk Supply of MicroRNA Associated with Leptin and Adiponectin Is Affected by Maternal Overweight/Obesity and Influences Infancy BMI. Nutrients 2019, 11, 2589. [Google Scholar] [CrossRef]
- Karbiener, M.; Pisani, D.F.; Frontini, A.; Oberreiter, L.M.; Lang, E.; Vegiopoulos, A.; Mössenböck, K.; Bernhardt, G.A.; Mayr, T.; Hildner, F.; et al. MicroRNA-26 Family Is Required for Human Adipogenesis and Drives Characteristics of Brown Adipocytes. Stem Cells 2013, 32, 1578–1590. [Google Scholar] [CrossRef]
- Kupsco, A.; Prada, D.; Valvi, D.; Hu, L.; Petersen, M.S.; Coull, B.; Grandjean, P.; Weihe, P.; Baccarelli, A.A. Human milk extracellular vesicle miRNA expression and associations with maternal characteristics in a population-based cohort from the Faroe Islands. Sci. Rep. 2021, 11, 5840. [Google Scholar] [CrossRef]
- Shiff, Y.E.; Reif, S.; Marom, R.; Shiff, K.; Reifen, R.; Golan-Gerstl, R. MiRNA-320a is less expressed and miRNA-148a more expressed in preterm human milk compared to term human milk. J. Funct. Foods 2019, 57, 68–74. [Google Scholar] [CrossRef]
- Chen, Q.; Zhang, F.; Dong, L.; Wu, H.; Xu, J.; Li, H.; Wang, J.; Zhou, Z.; Liu, C.; Wang, Y.; et al. SIDT1-dependent absorption in the stomach mediates host uptake of dietary and orally administered microRNAs. Cell Res. 2020, 31, 247–258. [Google Scholar] [CrossRef] [PubMed]
- Title, A.C.; Denzler, R.; Stoffel, M. Uptake and Function Studies of Maternal Milk-derived MicroRNAs. J. Biol. Chem. 2015, 290, 23680–23691. [Google Scholar] [CrossRef]
- Laubier, J.; Castille, J.; Le Guillou, S.; Le Provost, F. No effect of an elevated miR-30b level in mouse milk on its level in pup tissues. RNA Biol. 2015, 12, 26–29. [Google Scholar] [CrossRef] [PubMed]
- Mullokandov, G.; Baccarini, A.; Ruzo, A.; Jayaprakash, A.D.; Tung, N.; Israelow, B.; Evans, M.J.; Sachidanandam, R.; Brown, B.D. High-throughput assessment of microRNA activity and function using microRNA sensor and decoy libraries. Nat. Methods 2012, 9, 840–846. [Google Scholar] [CrossRef] [PubMed]
- Beuzelin, D.; Pitard, B.; Kaeffer, B. Oral Delivery of miRNA With Lipidic Aminoglycoside Derivatives in the Breastfed Rat. Front. Physiol. 2019, 10, 1037. [Google Scholar] [CrossRef]
- Tavares, G.A.; Torres, A.; Le Drean, G.; Queignec, M.; Castellano, B.; Tesson, L.; Remy, S.; Anegon, I.; Pitard, B.; Kaeffer, B. Oral Delivery of miR-320-3p with Lipidic Aminoglycoside Derivatives at Mid-Lactation Alters miR-320-3p Endogenous Levels in the Gut and Brain of Adult Rats According to Early or Regular Weaning. Int. J. Mol. Sci. 2022, 24, 191. [Google Scholar] [CrossRef]
- Liao, Y.; Du, X.; Yalin, L.; Lönnerdal, B. Human milk exosomes and their microRNAs survive digestion in vitro and are taken up by human intestinal cells. Mol. Nutr. Food Res. 2017, 61, 1700082. [Google Scholar] [CrossRef]
- Manca, S.; Upadhyaya, B.; Mutai, E.; Desaulniers, A.T.; Cederberg, R.A.; White, B.R.; Zempleni, J. Milk exosomes are bioavailable and distinct microRNA cargos have unique tissue distribution patterns. Sci. Rep. 2018, 8, 11321. [Google Scholar] [CrossRef]
- Riquelme, I.; Tapia, O.; Leal, P.; Sandoval, A.; Varga, M.G.; Letelier, P.; Buchegger, K.; Bizama, C.; Espinoza, J.A.; Peek, R.M.; et al. miR-101-2, miR-125b-2 and miR-451a act as potential tumor suppressors in gastric cancer through regulation of the PI3K/AKT/mTOR pathway. Cell Oncol. 2015, 39, 23–33. [Google Scholar] [CrossRef]
- Horns, F.; Martinez, J.A.; Fan, C.; Haque, M.; Linton, J.M.; Tobin, V.; Santat, L.; Maggiolo, A.O.; Bjorkman, P.J.; Lois, C.; et al. Engineering RNA export for measurement and manipulation of living cells. Cell 2023, 186, 3642–3658.e32. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Martin, R.; Wang, G.; Brandão, B.B.; Zanotto, T.M.; Shah, S.; Patel, S.K.; Schilling, B.; Kahn, C.R. MicroRNA sequence codes for small extracellular vesicle release and cellular retention. Nature 2021, 601, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Ghoshal, B.; Bertrand, E.; Bhattacharyya, S.N. Non-canonical argonaute loading of extracellular vesicle-derived exogenous single-stranded miRNA in recipient cells. J. Cell Sci. 2021, 134, jcs253914. [Google Scholar] [CrossRef] [PubMed]
- Beuzelin, D.; Kaeffer, B. Exosomes and miRNA-Loaded Biomimetic Nanovehicles, a Focus on Their Potentials Preventing Type-2 Diabetes Linked to Metabolic Syndrome. Front. Immunol. 2018, 9, 2711. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Wang, P.; Zhang, P.; Zhang, X.; Du, H.; Liu, Q.; Huang, B.; Qian, C.; Zhang, S.; Zhu, W.; et al. An integrative bioinformatics analysis identified miR-375 as a candidate key regulator of malignant breast cancer. J. Appl. Genet. 2019, 60, 335–346. [Google Scholar] [CrossRef] [PubMed]
- Hicks, S.D.; Beheshti, R.; Chandran, D.; Warren, K.; Confair, A. Infant consumption of microRNA miR-375 in human milk lipids is associated with protection from atopy. Am. J. Clin. Nutr. 2022, 116, 1654–1662. [Google Scholar] [CrossRef]
- Simpson, M.R.; Brede, G.; Johansen, J.; Johnsen, R.; Storrø, O.; Sætrom, P.; Øien, T. Human breast milk miRNA, maternal probiotic supplementation and atopic dermatitis in offspring. PLoS ONE 2015, 10, e0143496. [Google Scholar] [CrossRef]
- Acevedo, N.; Alashkar Alhamwe, B.; Caraballo, L.; Ding, M.; Ferrante, A.; Garn, H.; Garssen, J.; Hii, C.S.; Irvine, J.; Llinás-Caballero, K.; et al. Perinatal and Early-Life Nutrition, Epigenetics, and Allergy. Nutrients 2021, 13, 724. [Google Scholar] [CrossRef]
- Liu, H.; Geng, Z.; Su, J. Engineered mammalian and bacterial extracellular vesicles as promising nanocarriers for targeted therapy. Extracell. Vesicles Circ. Nucleic Acids 2022, 3, 63–86. [Google Scholar] [CrossRef]
- Afrin, H.; Bai, R.G.; Kumar, R.; Ahmad, S.S.; Agarwal, S.K. Nurunnabi Oral delivery of RNAi for cancer therapy. Cancer Metastasis Rev. 2023, 42, 699–724. [Google Scholar] [CrossRef]
- Gao, F.; Wang, F.; Cao, H.; Chen, Y.; Diao, Y.; Kapranov, P. Evidence for Existence of Multiple Functional Human Small RNAs Derived from Transcripts of Protein-Coding Genes. Int. J. Mol. Sci. 2023, 24, 4163. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; Schmitz, G. Milk Exosomal microRNAs: Postnatal Promoters of β Cell Proliferation but Potential Inducers of β Cell De-Differentiation in Adult Life. Int. J. Mol. Sci. 2022, 23, 11503. [Google Scholar] [CrossRef] [PubMed]
- Ma, R.C.W.; Popkin, B.M. Intergenerational diabetes and obesity—A cycle to break? PLoS Med. 2017, 14, e1002415. [Google Scholar] [CrossRef]
- Armstrong, J.; Reilly, J.J. Breastfeeding and lowering the risk of childhood obesity. Lancet 2002, 359, 2003–2004. [Google Scholar] [CrossRef] [PubMed]
- Ozkan, H.; Tuzun, F.; Taheri, S.; Korhan, P.; Akokay, P.; Yılmaz, O.; Duman, N.; Özer, E.; Tufan, E.; Kumral, A.; et al. Epigenetic Programming Through Breast Milk and Its Impact on Milk-Siblings Mating. Front. Genet. 2020, 11, 569232. [Google Scholar] [CrossRef]
- Pomar, C.A.; Serra, F.; Palou, A.; Sánchez, J. Lower miR-26a levels in breastmilk affect gene expression in adipose tissue of offspring. FASEB J. 2021, 35, e21924. [Google Scholar] [CrossRef]
- Arslanoglu, S.; Ziegler, E.E.; Moro, G.E. the WAPM Working Group on Nutrition Donor human milk in preterm infant feeding: Evidence and recommendations. J. Perinat. Med. 2010, 38, 347–351. [Google Scholar] [CrossRef]
- Kakimoto, Y.; Matsushima, Y.; Tanaka, M.; Hayashi, H.; Wang, T.; Yokoyama, K.; Ochiai, E.; Osawa, M. MicroRNA profiling of gastric content from breast-fed and formula-fed infants to estimate last feeding: A pilot study. Int. J. Leg. Med. 2019, 134, 903–909. [Google Scholar] [CrossRef]
- Carrillo-Lozano, E.; Sebastián-Valles, F.; Knott-Torcal, C. Circulating microRNAs in Breast Milk and Their Potential Impact on the Infant. Nutrients 2020, 12, 3066. [Google Scholar] [CrossRef]
- Acharya, A.; Berry, D.C.; Zhang, H.; Jiang, Y.; Jones, B.T.; Hammer, R.E.; Graff, J.M.; Mendell, J.T. miR-26 suppresses adipocyte progenitor differentiation and fat production by targeting Fbxl. Genes Dev. 2019, 33, 1367–11380. [Google Scholar] [CrossRef]
- Babiarz, J.E.; Ruby, J.G.; Wang, Y.; Bartel, D.P.; Blelloch, R. Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev. 2008, 22, 2773–2785. [Google Scholar] [CrossRef]
- Cline, M.S.; Smoot, M.; Cerami, E.; Bartel, A.; Landys, N.; Workman, C.; Christmas, R.; Avila-Campilo, I.; Creech, M.; Gross, B.; et al. Integration of biological networks and gene expression data using Cytoscape. Nat. Protoc. 2007, 2, 2366–2382. [Google Scholar] [CrossRef]
- Kim, D.H.; Sætrom, P.; Snøve, O., Jr.; Rossi, J.J. MicroRNA-directed transcriptional gene silencing in mammalian cells. Proc. Natl. Acad. Sci. USA 2008, 105, 16230–16235. [Google Scholar] [CrossRef] [PubMed]
- Yeganeh, M.; Hernandez, N. RNA polymerase III transcription as a disease factor. Genes Dev. 2020, 34, 865–882. [Google Scholar] [CrossRef] [PubMed]
- Kulaberoglu, Y.; Malik, Y.; Borland, G.; Selman, C.; Alic, N.; Tullet, J.M.A. Corrigendum: RNA Polymerase III, Ageing and Longevity. Front. Genet. 2021, 12, 758135, Erratum for: Front. Genet. 2021, 12, 705122. [Google Scholar] [CrossRef] [PubMed]
- Shenderov, B.A.; Midtvedt, T. Epigenomic programing: A future way to health? Microb. Ecol. Health Dis. 2014, 25, 24145. [Google Scholar] [CrossRef]
- Yu, A.-M.; Choi, Y.H.; Tu, M.-J. RNA Drugs and RNA Targets for Small Molecules: Principles, Progress, and Challenges. Pharmacol. Rev. 2020, 72, 862–898. [Google Scholar] [CrossRef]
- Torri, A.; Jaeger, J.; Pradeu, T.; Saleh, M.-C. The origin of RNA interference: Adaptive or neutral evolution? PLoS Biol. 2022, 20, e3001715. [Google Scholar] [CrossRef]
- Denzler, R.; Agarwal, V.; Stefano, J.; Bartel, D.P.; Stoffel, M. Assessing the ceRNA Hypothesis with Quantitative Measurements of miRNA and Target Abundance. Mol. Cell 2014, 54, 766–776. [Google Scholar] [CrossRef]
- Harman, J.C.; Guidry, J.J.; Gidday, J.M. Intermittent Hypoxia Promotes Functional Neuroprotection from Retinal Ischemia in Untreated First-Generation Offspring: Proteomic Mechanistic Insights. Investig. Opthalmology Vis. Sci. 2020, 61, 15. [Google Scholar] [CrossRef]
- Ip, S.; Chung, M.; Raman, G.; Chew, P.; Magula, N.; DeVine, D.; Trikalinos, T.; Lau, J. Breastfeeding and maternal and infant health outcomes in developed countries. Evid. Rep. Technol. Assess. 2007, 153, 1–186. [Google Scholar]
- Huang, S. Towards a unification of the 2 meanings of “epigenetics”. PLOS Biol. 2022, 20, e3001944. [Google Scholar] [CrossRef] [PubMed]
- Smythies, J.; Edelstein, L.; Ramachandran, V. Molecular mechanisms for the inheritance of acquired characteristics-exosomes, microRNA shuttling, fear and stress: Lamarck resurrected? Front. Genet. 2014, 5, 133. [Google Scholar] [CrossRef] [PubMed]
- Mezo-González, C.E.; García Santillán, J.A.; Kaeffer, B.; Gourdel, M.; Croyal, M.; Bolaños-Jiménez, F. Both maternal and paternal obesity in the rat induce learning deficits in offspring linked to impaired brain glutamatergic signaling. Acta Physiol. 2023, in press. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaeffer, B. Human Breast Milk miRNAs: Their Diversity and Potential for Preventive Strategies in Nutritional Therapy. Int. J. Mol. Sci. 2023, 24, 16106. https://doi.org/10.3390/ijms242216106
Kaeffer B. Human Breast Milk miRNAs: Their Diversity and Potential for Preventive Strategies in Nutritional Therapy. International Journal of Molecular Sciences. 2023; 24(22):16106. https://doi.org/10.3390/ijms242216106
Chicago/Turabian StyleKaeffer, Bertrand. 2023. "Human Breast Milk miRNAs: Their Diversity and Potential for Preventive Strategies in Nutritional Therapy" International Journal of Molecular Sciences 24, no. 22: 16106. https://doi.org/10.3390/ijms242216106
APA StyleKaeffer, B. (2023). Human Breast Milk miRNAs: Their Diversity and Potential for Preventive Strategies in Nutritional Therapy. International Journal of Molecular Sciences, 24(22), 16106. https://doi.org/10.3390/ijms242216106





