Artificial Fluorescent Glucosinolates (F-GSLs) Are Transported by the Glucosinolate Transporters GTR1/2/3
Abstract
1. Introduction
2. Results
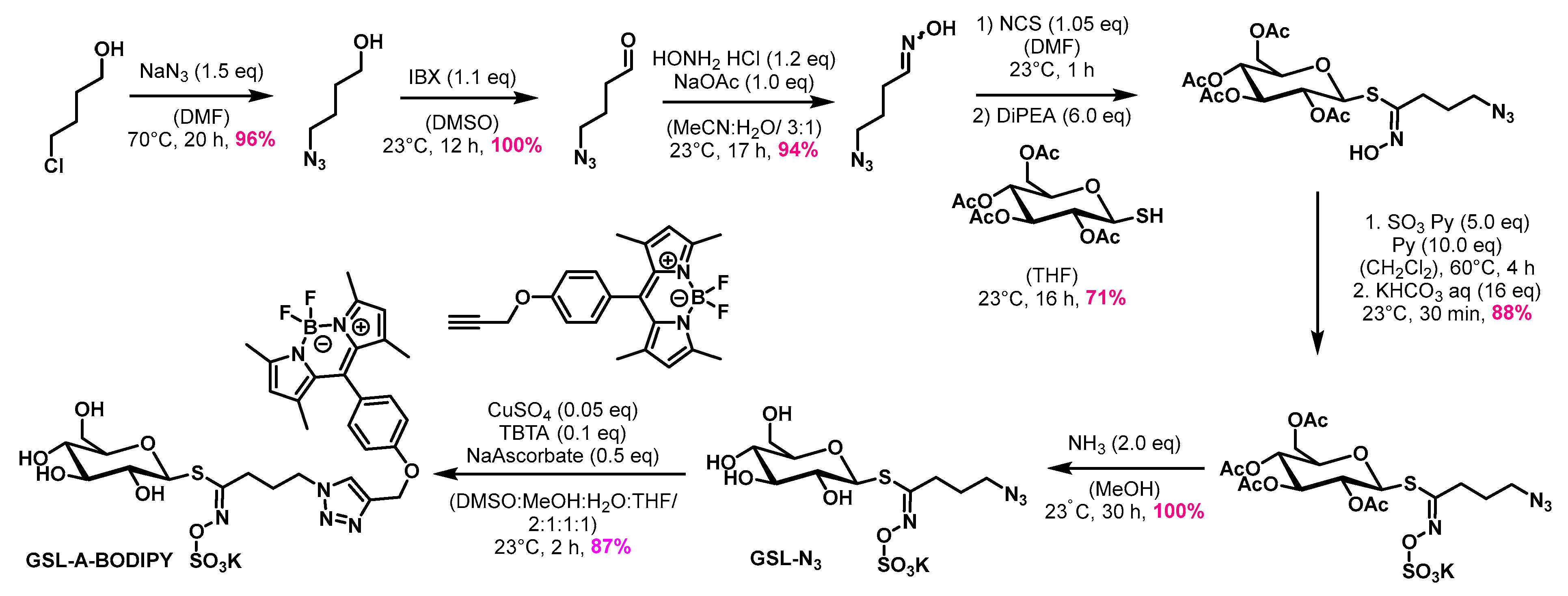
2.1. Synthesis of Seven Artificial Fluorescent Glucosinolates (F-GSLs)
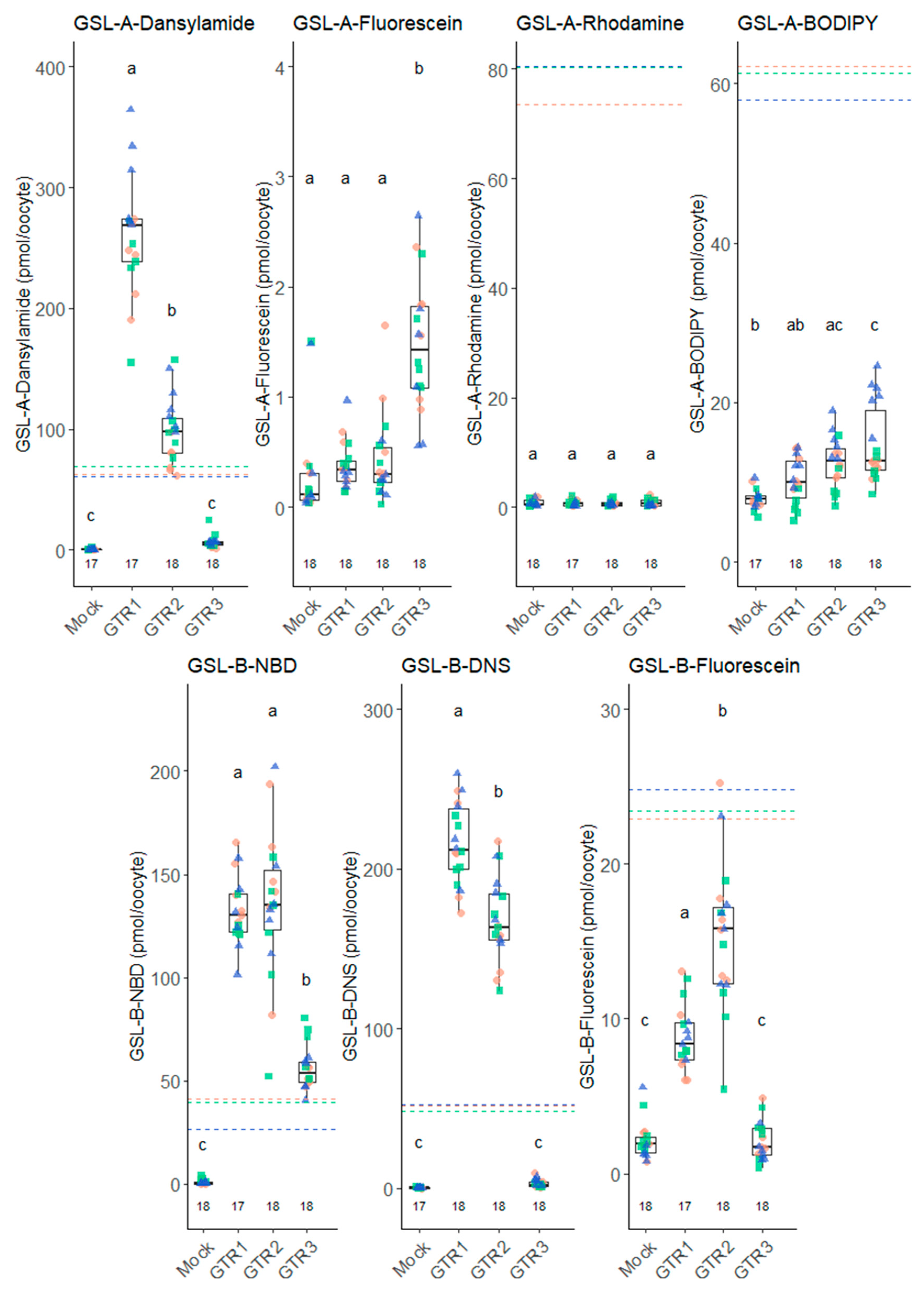
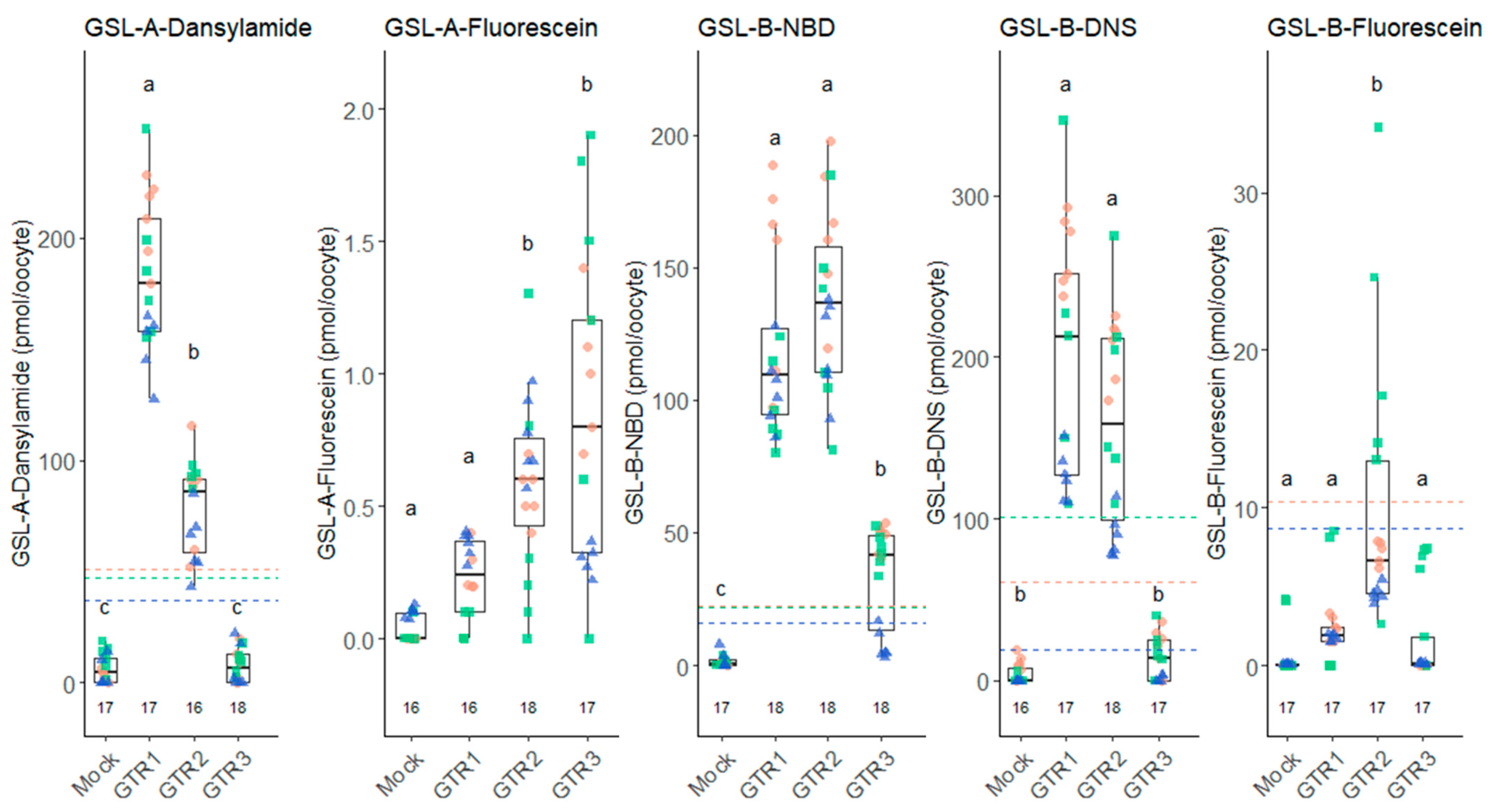
2.2. Transport of Fluorescent Glucosinolates Depends on the Fluorophore Attached
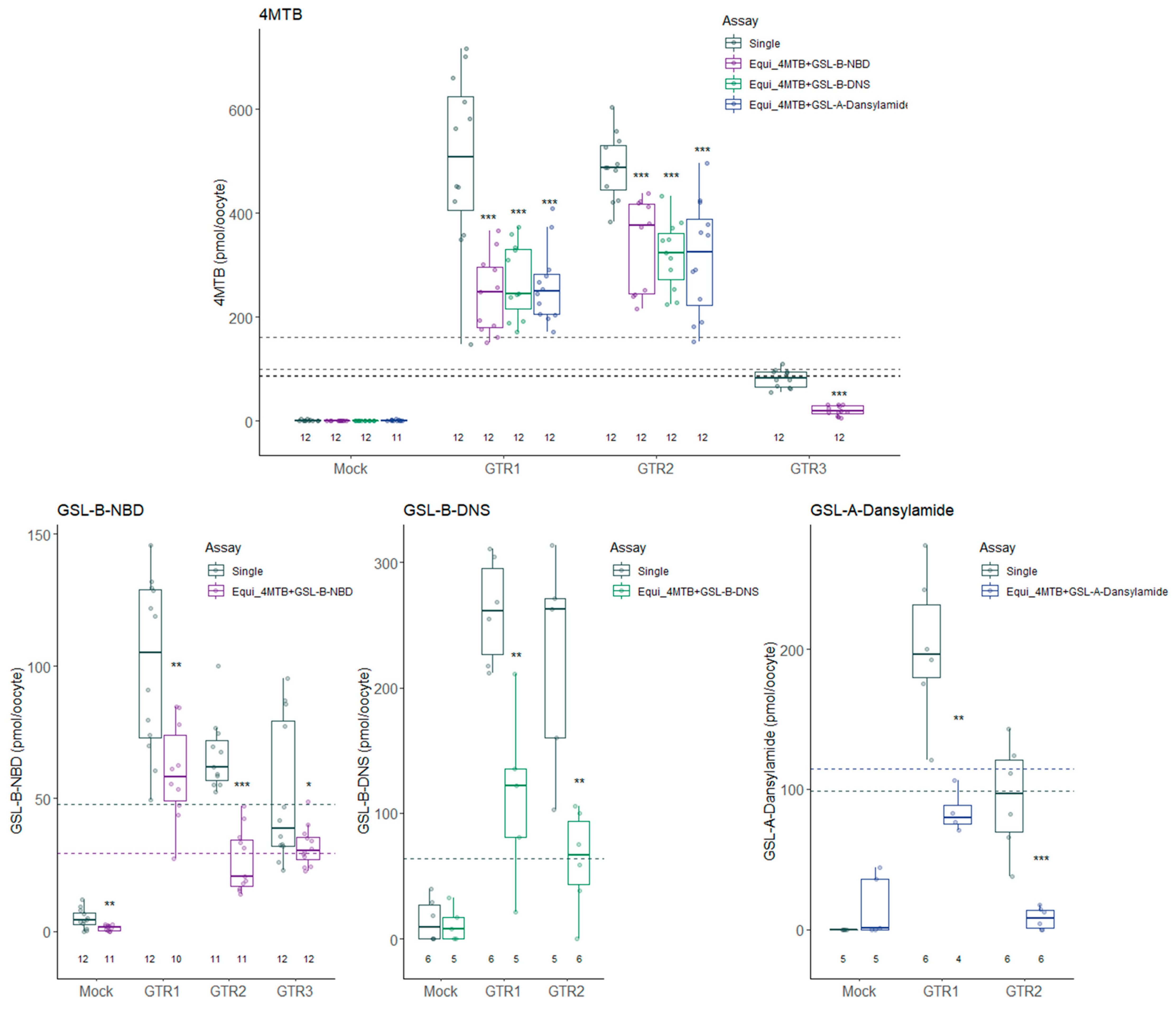
2.3. Natural Glucosinolate Glucoerucin Is in Competition with the Fluorescent Glucosinolates
3. Discussion
4. Materials and Methods
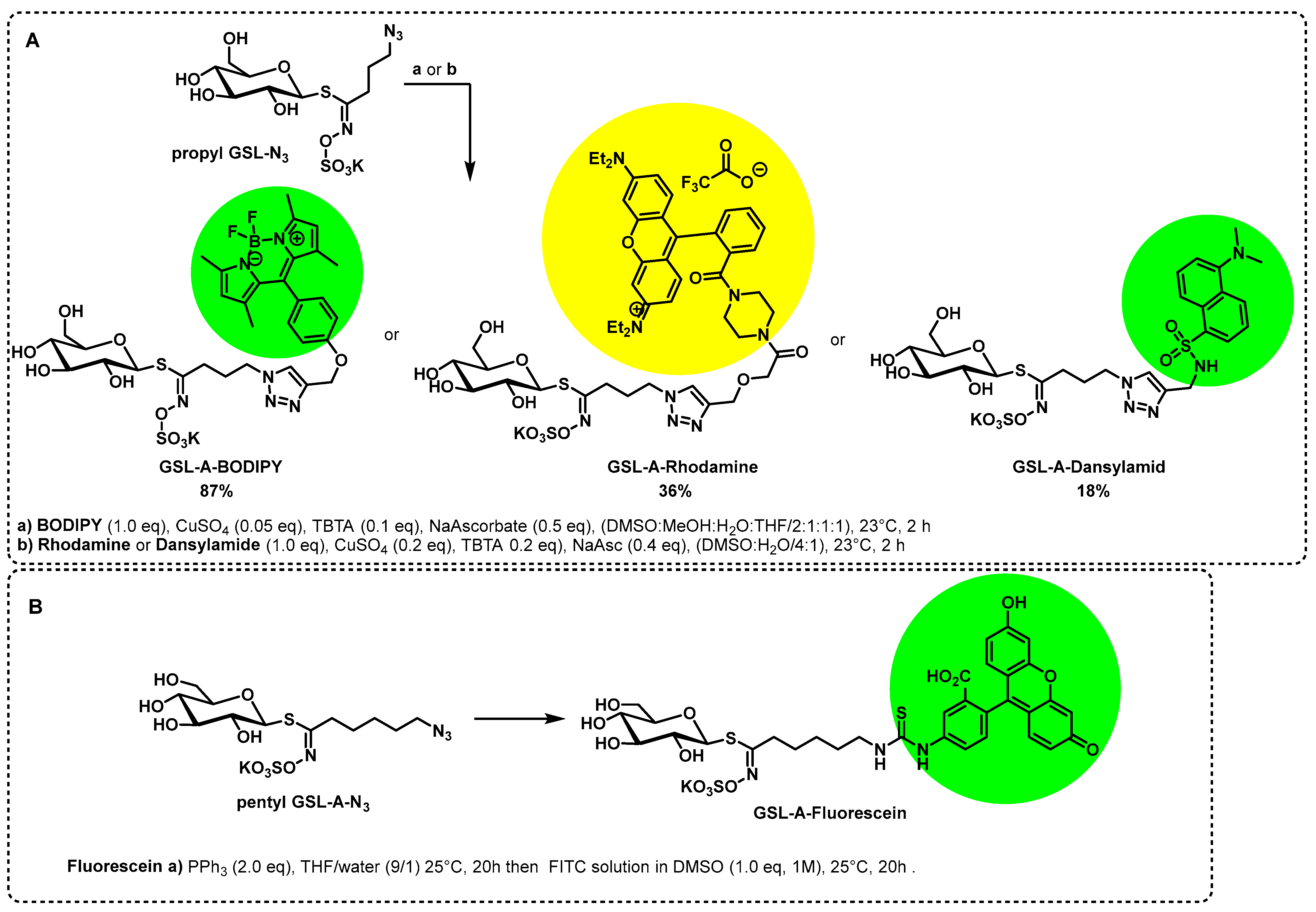
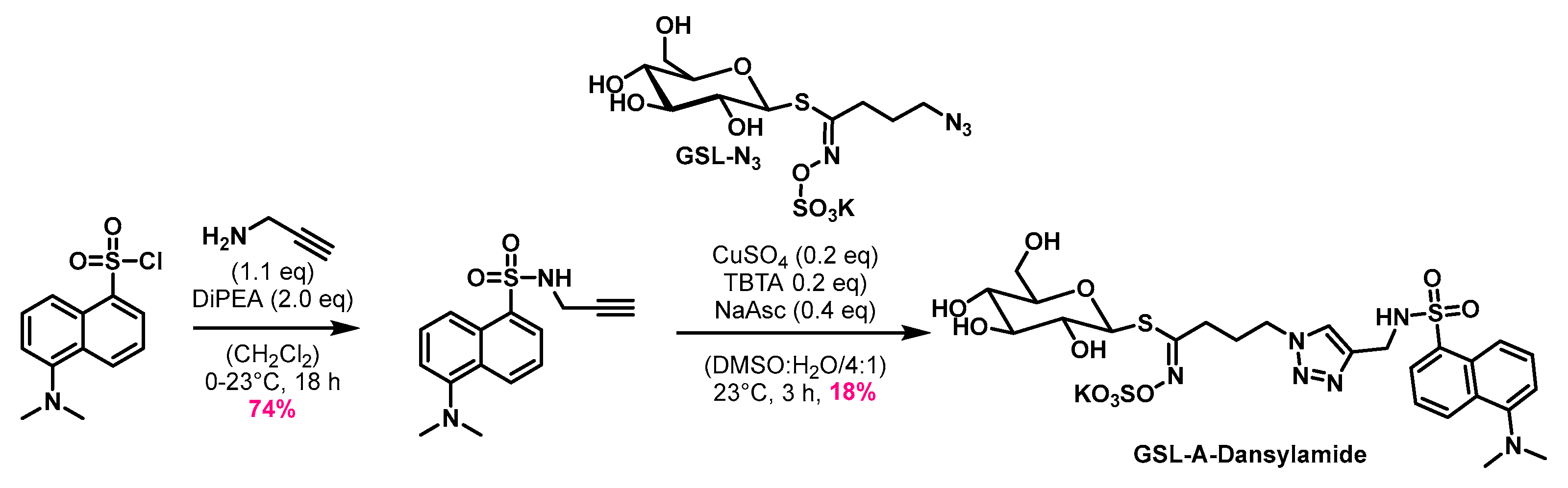
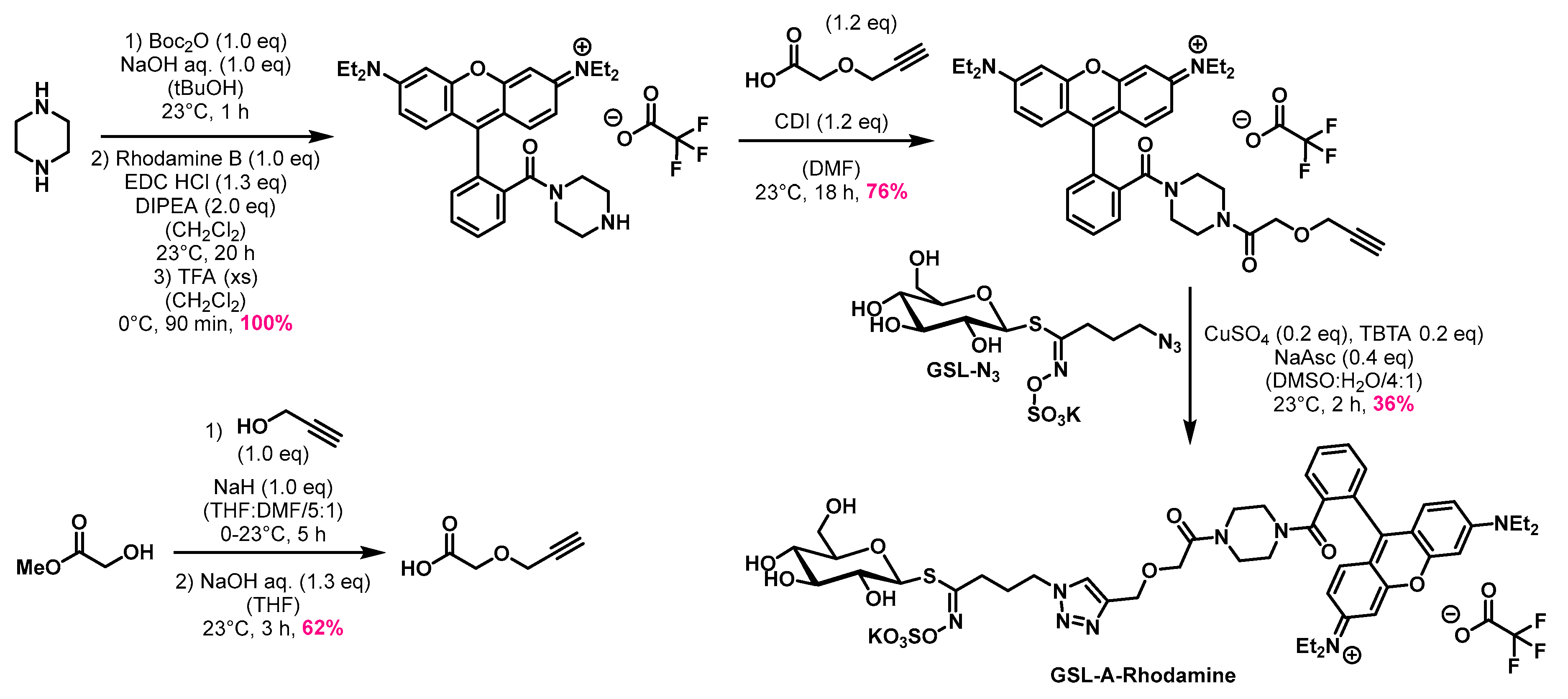
4.1. Synthesis of the Aliphatic Glucosinolates: GSL-A-BODIPY, GSL-A-Dansylamide, and GSL-A-Rhodamine
4.1.1. General Methods
- CAM staining solution [CAM]: 1 g Ce(IV)(SO4)2, 2.5 g (NH4)6Mo4O7 in 100 mL 10% H2SO4
- Ninhydrin staining solution [Ninhydrin]: 1.5 g Ninhydrin in 100 mL abs. EtOH and 3.0 mL HOAc.
4.1.2. Synthesis of the GSL-A-N3 and GSL-A-BODIPY
4.1.3. Synthesis of GSL-A-Dansylamide
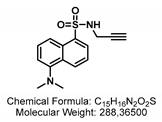
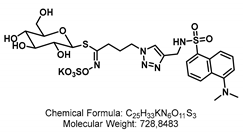
4.1.4. Synthesis of GSL-A-Rhodamine
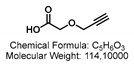
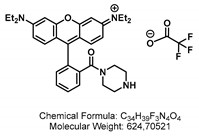
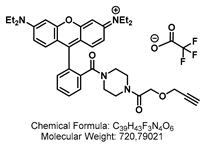
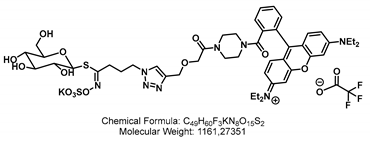
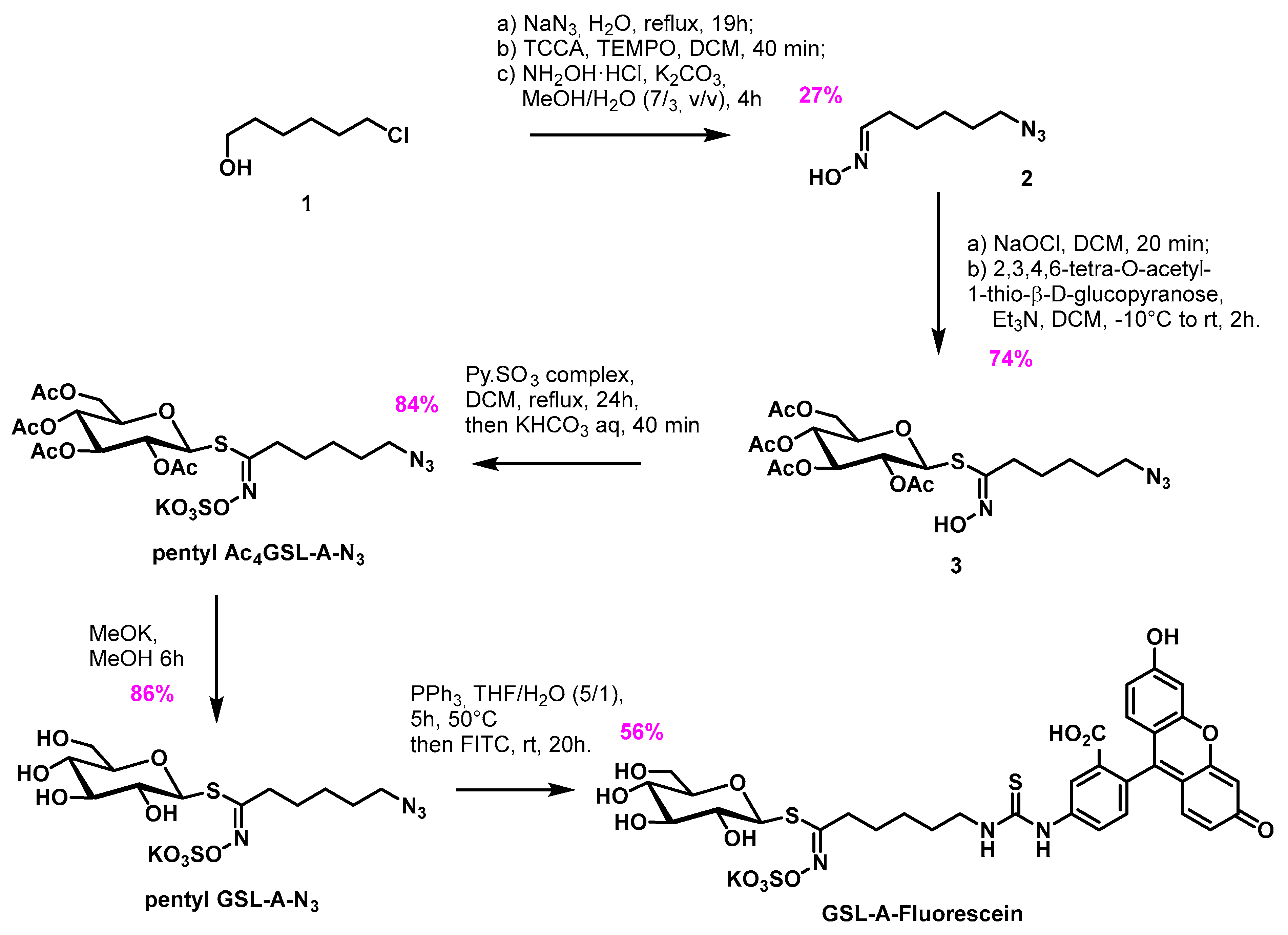
4.2. Synthesis of the Aliphatic Glucosinolate: GLS-A-Fluorescein
4.2.1. General Methods for GLS-A-Fluorescein, GSL-B-DNS, GSL-B-NBD and GSL-B-Fluorescein
4.2.2. Synthesis of GSL-A-Fluorescein
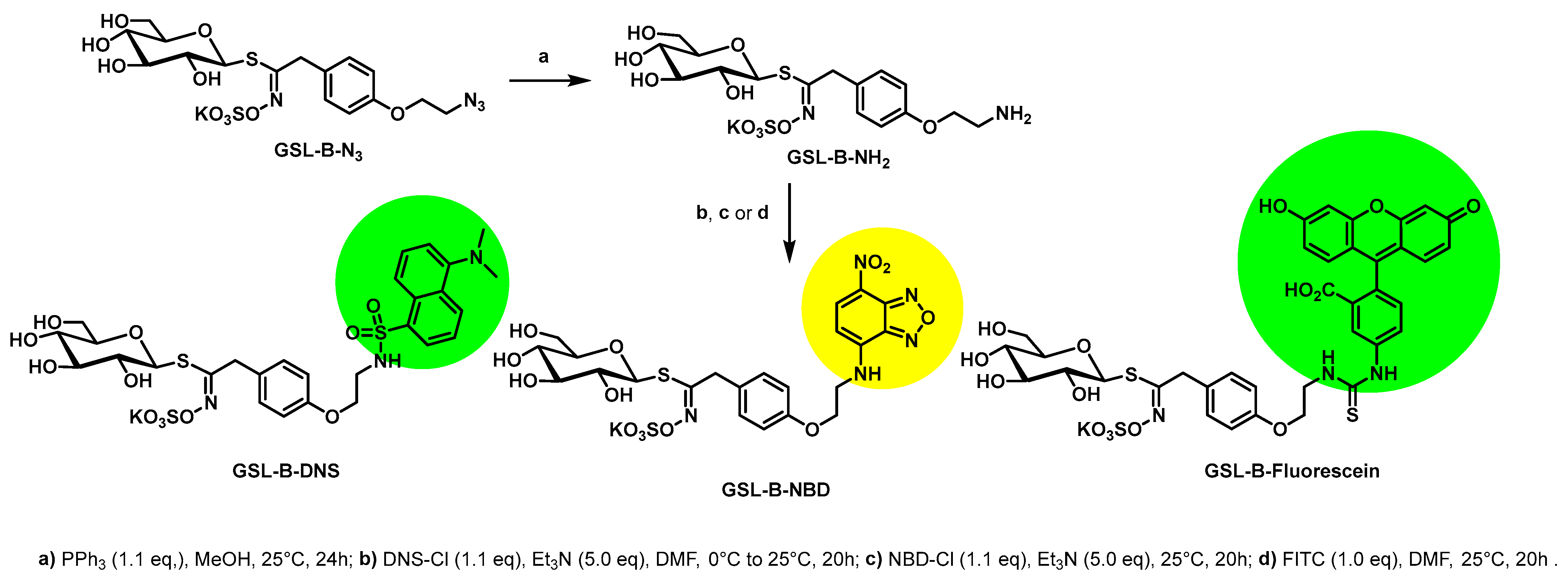
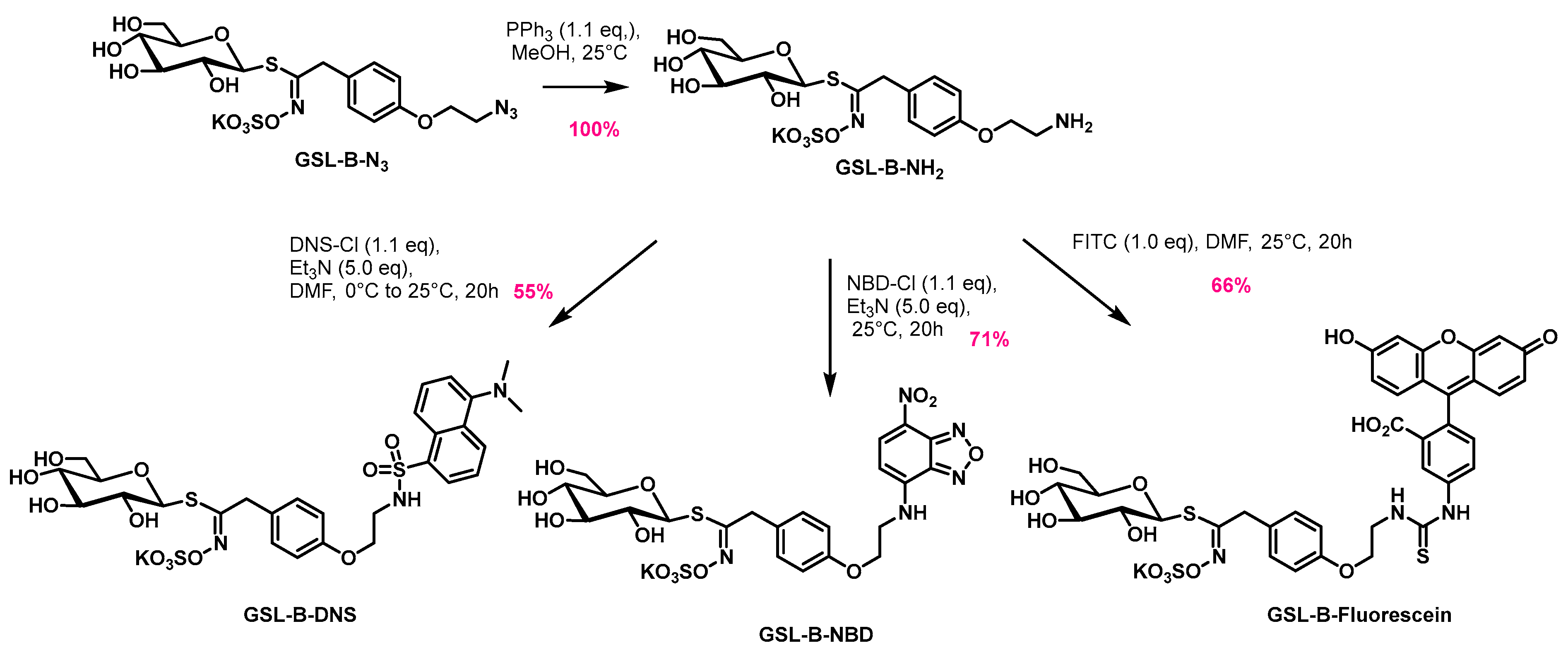
4.3. Synthesis of the Benzylic Glucosinolates: GSL-B-DNS, GSL-B-NBD, and GSL-B-Fluorescein
4.3.1. General Procedure for Synthesis of GSL-B-NH2
4.3.2. Synthesis of GSL-B-DNS
4.3.3. Synthesis of GSL-B-NBD
4.3.4. Synthesis of GSL-B-Fluorescein
4.4. DNA Constructs and cRNA Generation
4.5. Xenopus laevis Oocyte Transport Assays
Measurement on Microplate Reader
4.6. Quantification with LCMS
4.7. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Blažević, I.; Montaut, S.; Burčul, F.; Olsen, C.E.; Burow, M.; Rollin, P.; Agerbirk, N. Glucosinolate structural diversity, identification, chemical synthesis and metabolism in plants. Phytochemistry 2020, 169, 112100. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, V.P.T.; Stewart, J.; Lopez, M.; Ioannou, I.; Allais, F. Glucosinolates: Natural occurrence, biosynthesis, accessibility, isolation, structures, and biological activities. Molecules 2020, 25, 4537. [Google Scholar] [CrossRef] [PubMed]
- Chhajed, S.; Mostafa, I.; He, Y.; Abou-Hashem, M.; El-Domiaty, M.; Chen, S. Glucosinolate biosynthesis and the glucosinolate–myrosinase system in plant defense. Agronomy 2020, 10, 1786. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Madsen, S.R.; Engelen, S.; Jørgensen, M.E.; Olsen, C.E.; Andersen, J.S.; Seynnaeve, D.; Verhoye, T.; Fulawka, R.; Denolf, P.; et al. Reduction of antinutritional glucosinolates in Brassica oilseeds by mutation of genes encoding transporters. Nat. Biotechnol. 2017, 35, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Nega, T. Review on Nutritional Limitations and Opportunities of using Rapeseed Meal and other Rape Seed by-Products in Animal Feeding. J. Nutr. Health Food Eng. JNHFE 2018, 8, 43–48. [Google Scholar] [CrossRef]
- He, Y.; Yang, Z.; Tang, M.; Yang, Q.-Y.; Zhang, Y.; Liu, S. Enhancing canola breeding by editing a glucosinolate transporter gene lacking natural variation. Plant Physiol. 2022, 188, 1848–1851. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Andersen, T.G.; Burow, M.; Madsen, S.R.; Jørgensen, M.E.; Olsen, C.E.; Dreyer, I.; Hedrich, R.; Geiger, D.; Halkier, B.A. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 2012, 488, 531–534. [Google Scholar] [CrossRef]
- Jørgensen, M.E.; Xu, D.; Crocoll, C.; Ernst, H.A.; Ramírez, D.; Motawia, M.S.; Olsen, C.E.; Mirza, O.; Nour-Eldin, H.H.; Halkier, B.A. Origin and evolution of transporter substrate specificity within the NPF family. eLife 2017, 6, e19466. [Google Scholar] [CrossRef]
- Hölzl, G.; Rezaeva, B.R.; Kumlehn, J.; Dörmann, P. Ablation of glucosinolate accumulation in the oil crop Camelina sativa by targeted mutagenesis of genes encoding the transporters GTR1 and GTR2 and regulators of biosynthesis MYB28 and MYB29. Plant Biotechnol. J. 2022. [Google Scholar] [CrossRef]
- Jiang, D.; Lei, J.; Cao, B.; Wu, S.; Chen, G.; Chen, C. Molecular Cloning and Characterization of Three Glucosinolate Transporter (GTR) Genes from Chinese Kale. Genes 2019, 10, 202. [Google Scholar] [CrossRef]
- Nambiar, D.M.; Kumari, J.; Augustine, R.; Kumar, P.; Bajpai, P.K.; Bisht, N.C. GTR1 and GTR2 transporters differentially regulate tissue-specific glucosinolate contents and defence responses in the oilseed crop Brassica juncea. Plant Cell Environ. 2021, 44, 2729–2743. [Google Scholar] [CrossRef]
- Touw, A.J.; Verdecia Mogena, A.; Maedicke, A.; Sontowski, R.; van Dam, N.M.; Tsunoda, T. Both Biosynthesis and Transport Are Involved in Glucosinolate Accumulation During Root-Herbivory in Brassica rapa. Front. Plant Sci. 2019, 10, 1653. [Google Scholar] [CrossRef]
- Jørgensen, M.E.; Crocoll, C.; Halkier, B.A.; Nour-Eldin, H.H. Uptake Assays in Xenopus laevis Oocytes Using Liquid Chromatography-mass Spectrometry to Detect Transport Activity. Bio Protoc. 2017, 7, e2581. [Google Scholar] [CrossRef]
- Jørgensen, M.E.; Nour-Eldin, H.H.; Halkier, B.A. Transport of defense compounds from source to sink: Lessons learned from glucosinolates. Trends Plant Sci. 2015, 20, 508–514. [Google Scholar] [CrossRef]
- Madsen, S.R.; Olsen, C.E.; Nour-Eldin, H.H.; Halkier, B.A. Elucidating the role of transport processes in leaf glucosinolate distribution. Plant Physiol. 2014, 166, 1450–1462. [Google Scholar] [CrossRef]
- Xu, D.; Hanschen, F.S.; Witzel, K.; Nintemann, S.J.; Nour-Eldin, H.H.; Schreiner, M.; Halkier, B.A. Rhizosecretion of stele-synthesized glucosinolates and their catabolites requires GTR-mediated import in Arabidopsis. J. Exp. Bot. 2016, 68, 3205–3214. [Google Scholar] [CrossRef]
- Wulff, N.; Asschenfeldt Ernst, H.; Egevang Jorgensen, M.; Lambertz, S.; Maierhofer, T.; Mussa Belew, Z.; Crocoll, C.; Saddik Motawia, M.; Geiger, D.; Jorgensen, F.S.; et al. An Optimized Screen Reduces the Number of GA Transporters and provides Insights into NPF Substrate Determinants. BioRxiv 2019. [Google Scholar] [CrossRef]
- Longo, A.; Miles, N.W.; Dickstein, R. Genome mining of plant npfs reveals varying conservation of signature motifs associated with the mechanism of transport. Front. Plant Sci. 2018, 9, 1668. [Google Scholar] [CrossRef]
- Parker, J.L.; Newstead, S. Molecular basis of nitrate uptake by the plant nitrate transporter NRT1.1. Nature 2014, 507, 68–72. [Google Scholar] [CrossRef] [PubMed]
- Tsay, Y.F.; Schroeder, J.I.; Feldmann, K.A.; Crawford, N.M. The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 1993, 72, 705–713. [Google Scholar] [CrossRef]
- Reinders, A.; Sun, Y.; Karvonen, K.L.; Ward, J.M. Identification of amino acids important for substrate specificity in sucrose transporters using gene shuffling. J. Biol. Chem. 2012, 287, 30296–30304. [Google Scholar] [CrossRef] [PubMed]
- Gora, P.J.; Reinders, A.; Ward, J.M. A novel fluorescent assay for sucrose transporters. Plant Methods 2012, 8, 13. [Google Scholar] [CrossRef] [PubMed]
- Glindemann, C.P.; Backenköhler, A.; Strieker, M.; Wittstock, U.; Klahn, P. Synthesis and biochemical evaluation of an artificial, fluorescent glucosinolate (GSL). Chembiochem 2019, 20, 2341–2345. [Google Scholar] [CrossRef] [PubMed]
- Fredy, J.W.; Cutolo, G.; Poret, B.; Nehmé, R.; Hubert-Roux, M.; Gandolfo, P.; Castel, H.; Schuler, M.; Tatibouët, A.; Sabot, C.; et al. Diverted Natural Lossen-type Rearrangement for Bioconjugation through in Situ Myrosinase-Triggered Isothiocyanate Synthesis. Bioconjug. Chem. 2019, 30, 1385–1394. [Google Scholar] [CrossRef] [PubMed]
- Chung, Y.-C.; Cheng, H.-Y.; Wang, W.-T.; Chang, Y.-J.; Lin, S.-M. Transport efficiency of AtGTR1 dependents on the hydrophobicity of transported glucosinolates. Sci. Rep. 2022, 12, 5097. [Google Scholar] [CrossRef]
- Nambiar, D.M.; Kumari, J.; Arya, G.C.; Singh, A.K.; Bisht, N.C. A cell suspension based uptake method to study high affinity glucosinolate transporters. Plant Methods 2020, 16, 75. [Google Scholar] [CrossRef]
- Yang, Z.-L.; Nour-Eldin, H.H.; Hänniger, S.; Reichelt, M.; Crocoll, C.; Seitz, F.; Vogel, H.; Beran, F. Sugar transporters enable a leaf beetle to accumulate plant defense compounds. Nat. Commun. 2021, 12, 2658. [Google Scholar] [CrossRef]
- Chen, S.; Halkier, B.A. In vivo synthesis and purification of radioactive p-hydroxybenzylglucosinolate in Sinapis alba L. Phytochem. Anal. 2000, 11, 174–178. [Google Scholar] [CrossRef]
- Rumin, J.; Bonnefond, H.; Saint-Jean, B.; Rouxel, C.; Sciandra, A.; Bernard, O.; Cadoret, J.-P.; Bougaran, G. The use of fluorescent Nile red and BODIPY for lipid measurement in microalgae. Biotechnol. Biofuels 2015, 8, 42. [Google Scholar] [CrossRef]
- Wang, J.; Guo, X.; Li, L.; Qiu, H.; Zhang, Z.; Wang, Y.; Sun, G. Application of the Fluorescent Dye BODIPY in the Study of Lipid Dynamics of the Rice Blast Fungus Magnaporthe oryzae. Molecules 2018, 23, 1594. [Google Scholar] [CrossRef]
- Martynov, V.I.; Pakhomov, A.A. BODIPY derivatives as fluorescent reporters of molecular activities in living cells. Russ. Chem. Rev. 2021, 90, 1213–1262. [Google Scholar] [CrossRef]
- Branda, N. Degassing Liquids with Freeze-Pump-Thaw Cycling. JoVE Sci. Educ. Database. Org. Chem. 2019. Available online: https://www.jove.com/t/5639/entgasung-von-flssigkeiten-mit-frost-tausalz-pumpe-radfahren (accessed on 2 December 2022).
- Still, W.C.; Kahn, M.; Mitra, A. Rapid chromatographic technique for preparative separations with moderate resolution. J. Org. Chem. 1978, 43, 2923–2925. [Google Scholar] [CrossRef]
- Phetsang, W.; Pelingon, R.; Butler, M.S.; Kc, S.; Pitt, M.E.; Kaeslin, G.; Cooper, M.A.; Blaskovich, M.A.T. Fluorescent Trimethoprim Conjugate Probes to Assess Drug Accumulation in Wild Type and Mutant Escherichia coli. ACS Infect. Dis. 2016, 2, 688–701. [Google Scholar] [CrossRef]
- Jezowska, M.; Honcharenko, D.; Ghidini, A.; Strömberg, R.; Honcharenko, M. Enabling multiple conjugation to oligonucleotides using “click cycles”. Bioconjug. Chem. 2016, 27, 2620–2628. [Google Scholar] [CrossRef]
- Crocoll, C.; Halkier, B.A.; Burow, M. Analysis and quantification of glucosinolates. Curr. Protoc. Plant Biol. 2016, 1, 385–409. [Google Scholar] [CrossRef]

| No | Plasmid | Organism | Gene |
|---|---|---|---|
| 1 * | pNB1 | Xenopus laevis oocytes | AtGTR1 |
| 2 * | pNB1 | Xenopus laevis oocytes | AtGTR2 |
| 3 ** | pNB1 | Xenopus laevis oocytes | AtGTR3 |
| Fluorophore | Settings | Excitation/Emission |
|---|---|---|
| FITC | Bottom read, gain 75 | 492/524 |
| DNS/Dansylamide | Bottom read, gain 100 | 335/518 |
| NBD | Bottom read, gain 100 | 465/560 |
| Analyte | Retention Time [min] | Q1 [m/z] | Q3 [m/z] | Fragmentor [V] | CE [V] |
|---|---|---|---|---|---|
| 2-propenyl GSL (sinigrin) | 1.72 | 358.0 | 97.0 Qt | 98 | 20 |
| [M-H]− | 358.0 | 259.0 | 98 | 16 | |
| Internal standard (IS) | 358.0 | 75.0 | 98 | 36 | |
| 4MTB GSL (glucoerucin) | 2.56 | 420.0 | 97.0 Qt | 107 | 24 |
| [M-H]− | 420.0 | 259.0 | 107 | 20 | |
| 420.0 | 75.0 | 107 | 40 | ||
| GSL-B-Fluorescein | 3.47 | 856.1 | 518.3 Qt | 150 | 52 |
| [M-H]− | 856.1 | 325.5 | 150 | 92 | |
| GSL-B-DNS | 3.23 | 700.1 | 97.0 Qt | 170 | 40 |
| [M-H]− | 700.1 | 259.0 | 170 | 36 | |
| 700.1 | 275.0 | 170 | 40 | ||
| GSL-B-NBD | 3.28 | 630.0 | 97.0 Qt | 145 | 64 |
| [M-H]− | 630.0 | 259.0 | 145 | 28 | |
| 630.0 | 275.0 | 145 | 28 | ||
| GSL-A-BODIPY | 4.57 | 779.2 | 97.0 Qt | 150 | 40 |
| [M-H]− | 779.2 | 339.2 | 150 | 56 | |
| 779.2 | 537.4 | 150 | 32 | ||
| GSL-A-Dansylamide | 2.72 | 689.1 | 97.0 Qt | 163 | 40 |
| [M-H]− | 689.1 | 249.1 | 163 | 64 | |
| 689.1 | 169.8 | 163 | 96 | ||
| GSL-A-Rhodamine | 3.42 | 1007.5 | 97.0 Qt | 232 | 64 |
| [M-H]− | 1007.5 | 992.5 | 232 | 36 | |
| 1007.5 | 936.9 | 232 | 44 | ||
| GSL-A-Fluorescein | 3.31 | 792.1 | 97.0 Qt | 150 | 52 |
| [M-H]− | 792.1 | 403.2 | 150 | 32 | |
| 792.1 | 302.5 | 150 | 64 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kanstrup, C.; Jimidar, C.C.; Tomas, J.; Cutolo, G.; Crocoll, C.; Schuler, M.; Klahn, P.; Tatibouët, A.; Nour-Eldin, H.H. Artificial Fluorescent Glucosinolates (F-GSLs) Are Transported by the Glucosinolate Transporters GTR1/2/3. Int. J. Mol. Sci. 2023, 24, 920. https://doi.org/10.3390/ijms24020920
Kanstrup C, Jimidar CC, Tomas J, Cutolo G, Crocoll C, Schuler M, Klahn P, Tatibouët A, Nour-Eldin HH. Artificial Fluorescent Glucosinolates (F-GSLs) Are Transported by the Glucosinolate Transporters GTR1/2/3. International Journal of Molecular Sciences. 2023; 24(2):920. https://doi.org/10.3390/ijms24020920
Chicago/Turabian StyleKanstrup, Christa, Claire C. Jimidar, Josip Tomas, Giuliano Cutolo, Christoph Crocoll, Marie Schuler, Philipp Klahn, Arnaud Tatibouët, and Hussam Hassan Nour-Eldin. 2023. "Artificial Fluorescent Glucosinolates (F-GSLs) Are Transported by the Glucosinolate Transporters GTR1/2/3" International Journal of Molecular Sciences 24, no. 2: 920. https://doi.org/10.3390/ijms24020920
APA StyleKanstrup, C., Jimidar, C. C., Tomas, J., Cutolo, G., Crocoll, C., Schuler, M., Klahn, P., Tatibouët, A., & Nour-Eldin, H. H. (2023). Artificial Fluorescent Glucosinolates (F-GSLs) Are Transported by the Glucosinolate Transporters GTR1/2/3. International Journal of Molecular Sciences, 24(2), 920. https://doi.org/10.3390/ijms24020920






