BRCA1 the Versatile Defender: Molecular to Environmental Perspectives
Abstract
1. Introduction
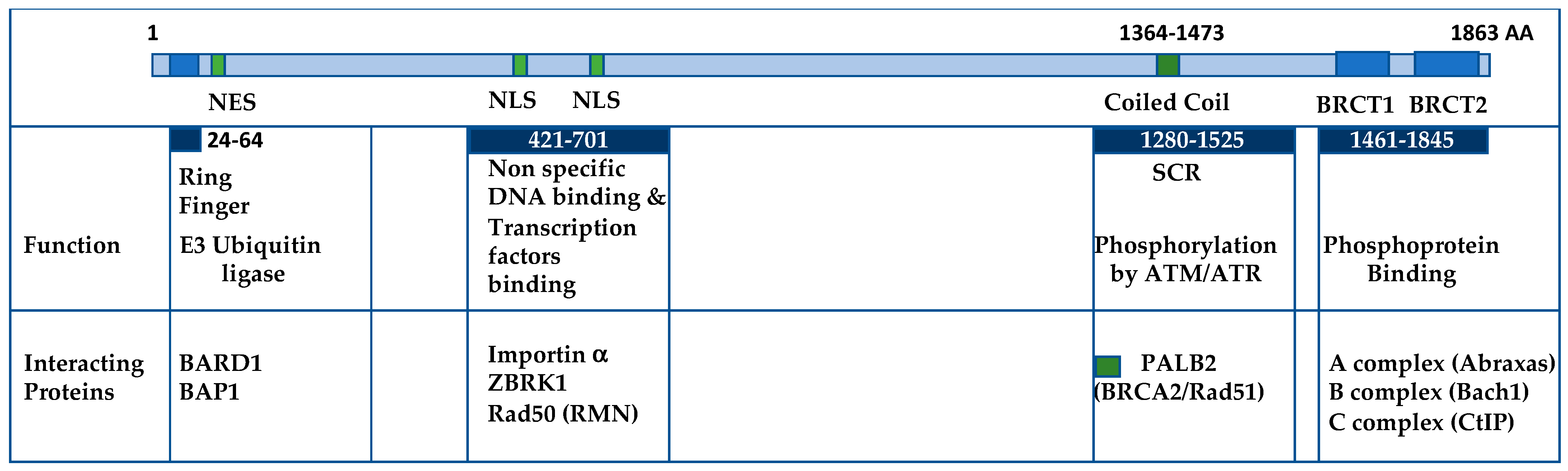
2. BRCA1 Is Involved in Several Complexes
2.1. The BRCA1/BARD1 Complex Functions as a Ubiquitin Ligase
2.2. The BRCA1/ZBRK1/CtIP Complex Functions as a Transcription Co-Repressor
2.3. PALB2 Connects BRCA1 and BRCA2
2.4. The BRCA1 A-, B-, C-Complex
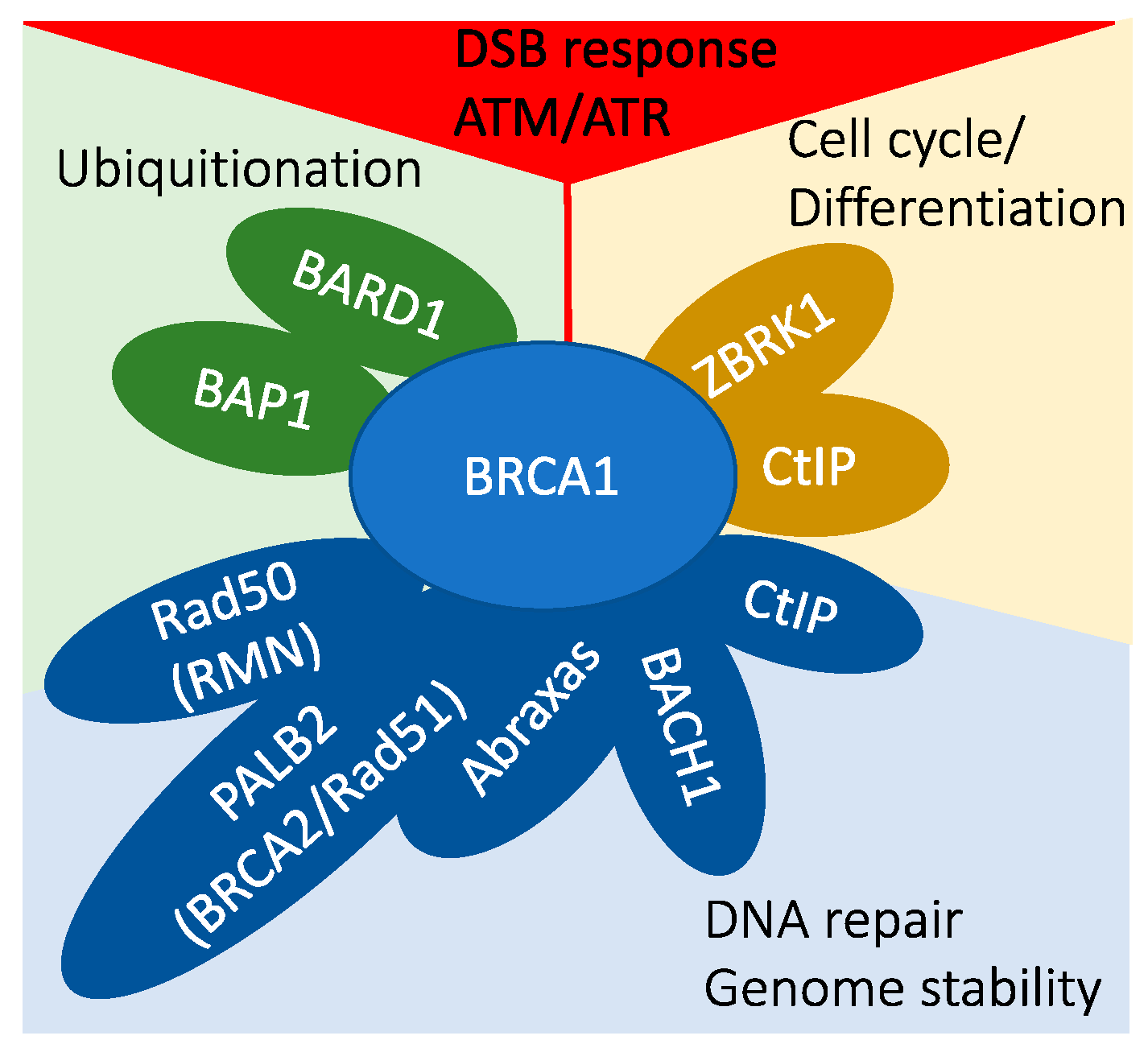
3. BRCA1 Cellular Function
- In DNA damage signaling, BRCA1 is activated through phosphorylation by ATM/ATR kinases, and it plays a role in the G1–S checkpoint response through indirect mechanisms.
- BRCA1 interacts with BARD1 and BAP1, participating in protein ubiquitination processes.
- The BRCA1/ZBRK1/CtIP protein complex is involved in transcriptional regulation.
- A multitude of BRCA1 protein complexes collectively contribute to the maintenance of genomic stability.
3.1. BRCA1 Is a Caretaker and Gate-Keeper
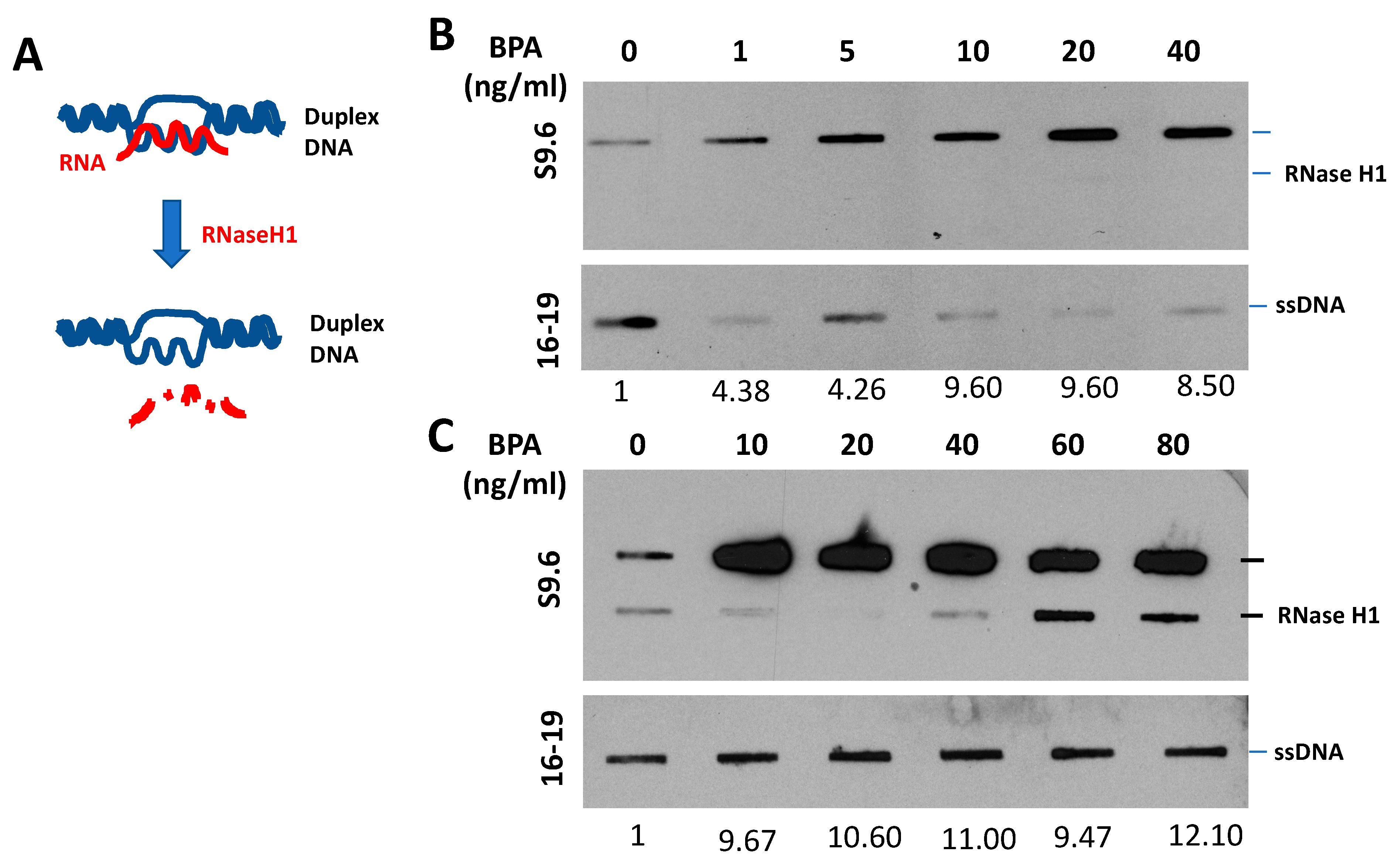
3.2. BRCA1 Prevents R-Loop Formation
3.3. BRCA1 Aids in Cellular Differentiation
4. Lessons Learned from Animal Models
5. Genetic Modifiers for Cancer Incidence
6. Unraveling the Complex Web of Breast Cancer Risk Factors
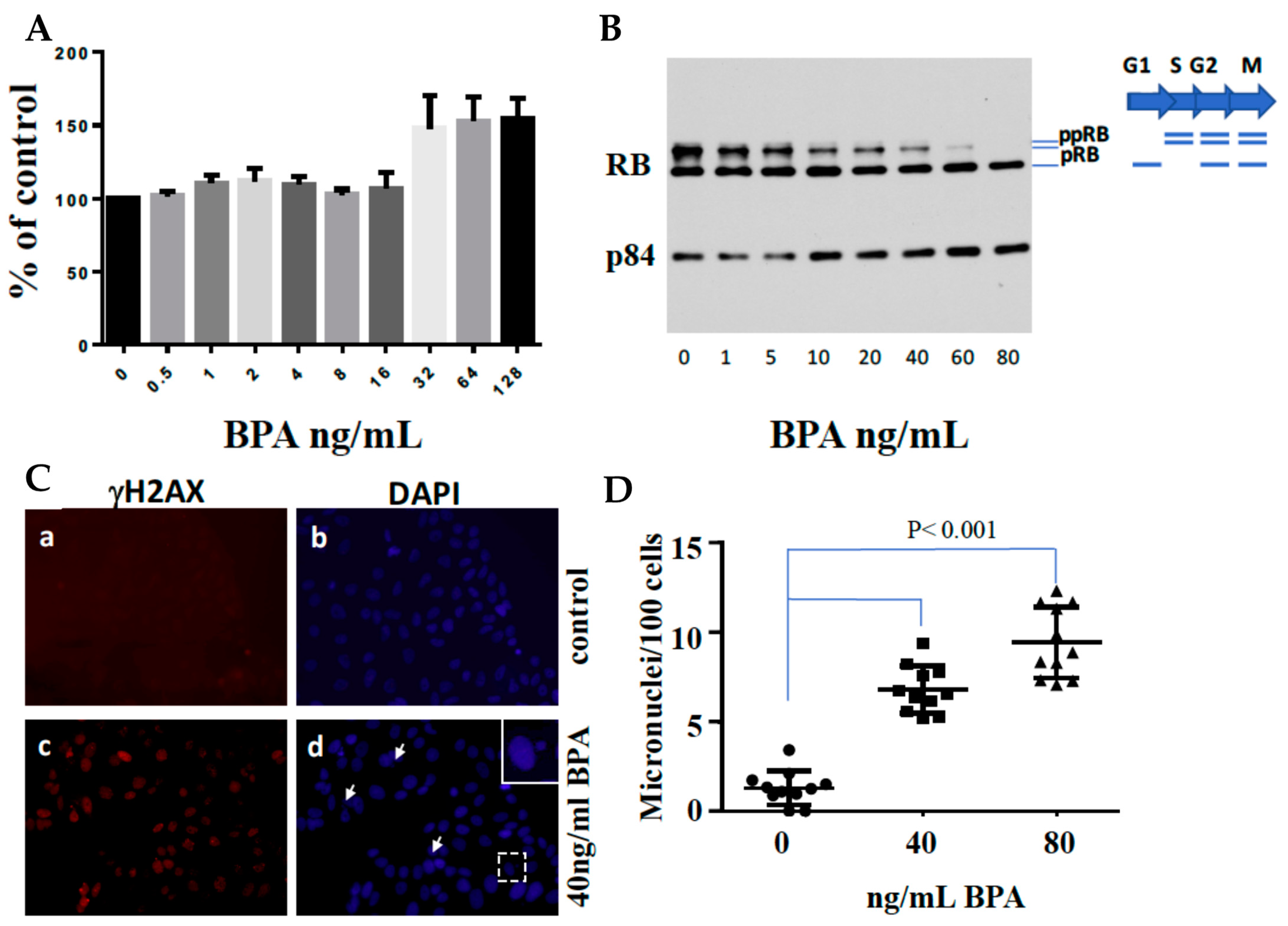
6.1. BRCA1 Deficiency and BPA Response
6.2. BPA’s Estrogenic Behavior
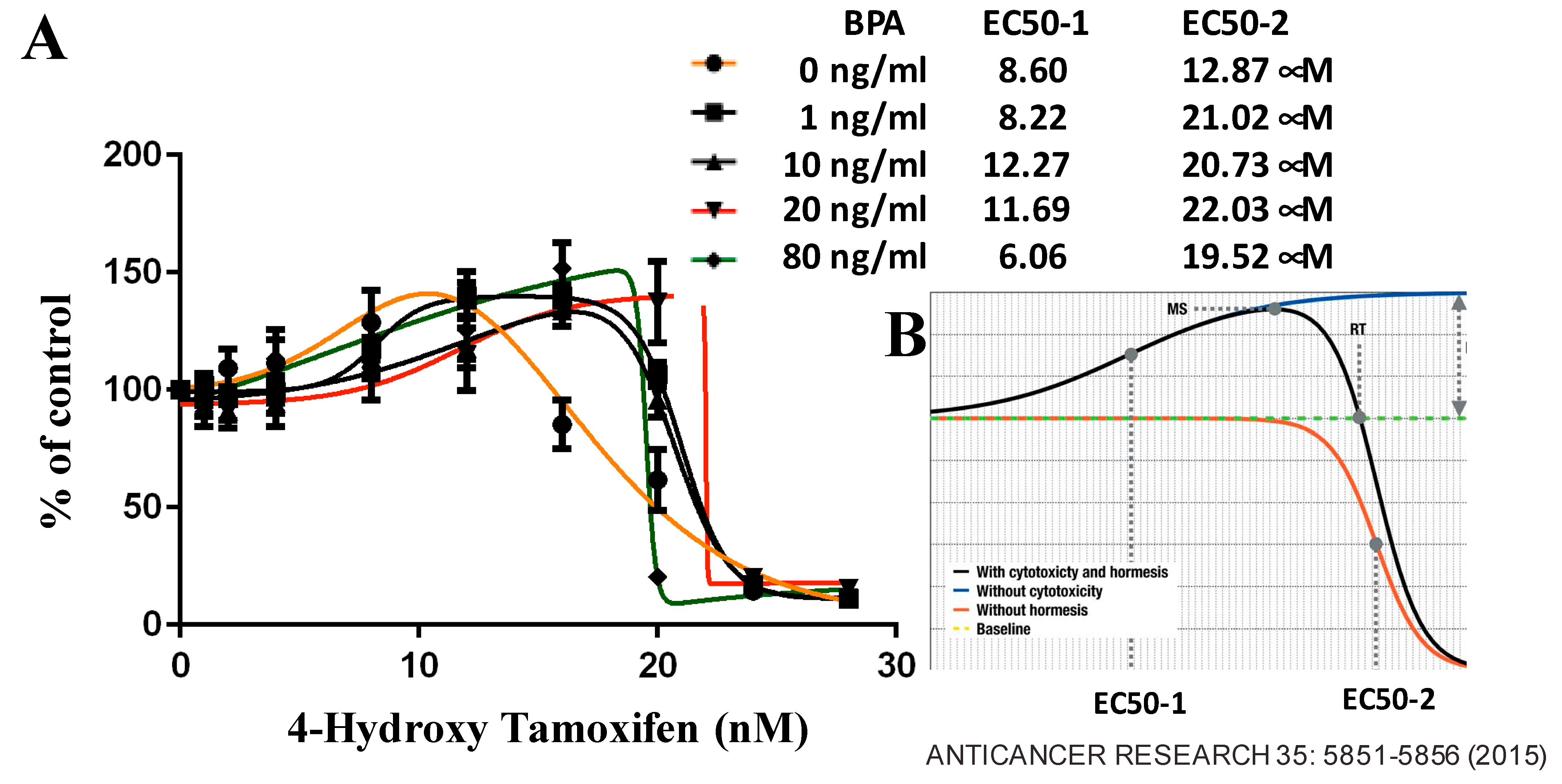
6.3. BPA Effects on ER-Positive Cancer
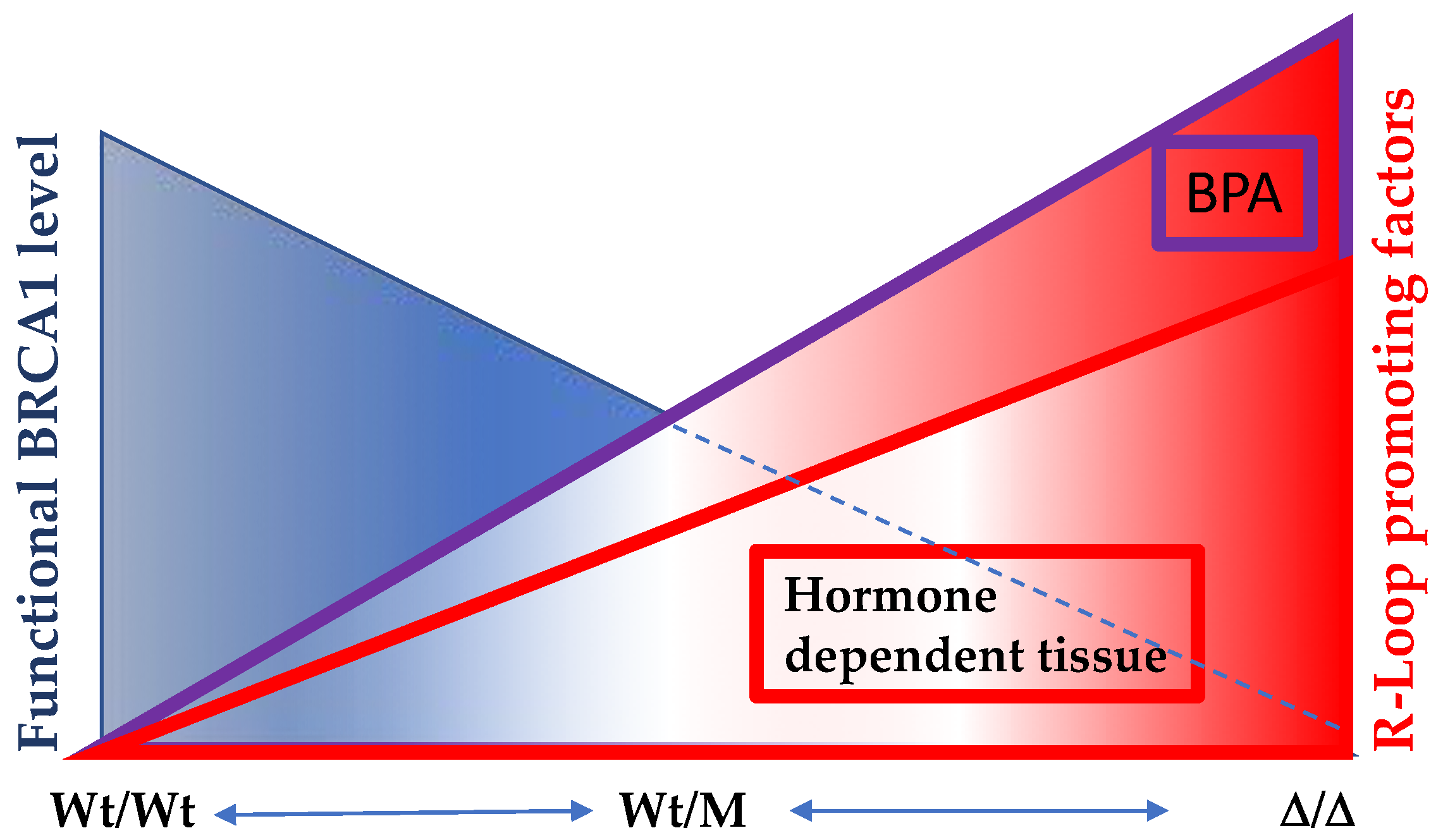
7. A Model for the Tissue Specificity of BRCA Genes
8. Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Miki, Y.; Swensen, J.; Shattuck-Eidens, D.; Futreal, P.A.; Harshman, K.; Tavtigian, S.; Liu, Q.; Cochran, C.; Bennett, L.M.; Ding, W.; et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994, 266, 66–71. [Google Scholar] [CrossRef]
- Chen, Y.; Chen, C.F.; Riley, D.J.; Allred, D.C.; Chen, P.L.; Von Hoff, D.; Osborne, C.K.; Lee, W.H. Aberrant subcellular localization of BRCA1 in breast cancer. Science 1995, 270, 789–791. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Farmer, A.A.; Chen, C.F.; Jones, D.C.; Chen, P.L.; Lee, W.H. BRCA1 is a 220-kDa nuclear phosphoprotein that is expressed and phosphorylated in a cell cycle-dependent manner. Cancer Res. 1996, 56, 3168–3172. [Google Scholar] [PubMed]
- Saurin, A.J.; Borden, K.L.; Boddy, M.N.; Freemont, P.S. Does this have a familiar RING? Trends Biochem. Sci. 1996, 21, 208–214. [Google Scholar] [CrossRef]
- Koonin, E.V.; Altschul, S.F.; Bork, P. BRCA1 protein products…Functional motifs. Nat. Genet. 1996, 13, 266–268. [Google Scholar] [CrossRef]
- Bork, P.; Hofmann, K.; Bucher, P.; Neuwald, A.F.; Altschul, S.F.; Koonin, E.V. A superfamily of conserved domains in DNA damage-responsive cell cycle checkpoint proteins. FASEB J. 1997, 11, 68–76. [Google Scholar] [CrossRef]
- Callebaut, I.; Mornon, J.P. From BRCA1 to RAP1: A widespread BRCT module closely associated with DNA repair. FEBS Lett. 1997, 400, 25–30. [Google Scholar] [CrossRef]
- Scully, R.; Chen, J.; Plug, A.; Xiao, Y.; Weaver, D.; Feunteun, J.; Ashley, T.; Livingston, D.M. Association of BRCA1 with Rad51 in mitotic and meiotic cells. Cell 1997, 88, 265–275. [Google Scholar] [CrossRef]
- Zhong, Q.; Chen, C.F.; Li, S.; Chen, Y.; Wang, C.C.; Xiao, J.; Chen, P.L.; Sharp, Z.D.; Lee, W.H. Association of BRCA1 with the hRad50-hMre11-p95 complex and the DNA damage response. Science 1999, 285, 747–750. [Google Scholar] [CrossRef] [PubMed]
- Struewing, J.P.; Hartge, P.; Wascholder, S.; Baker, S.M.; Berlin, M.; McAdams, M.; Timmerman, M.M.; Brody, L.C.; Tucker, M.A. The Risk of Cancer Associated with Specific Mutations of BRCA1 and BRCA2 among Ashkenazi Jews. N. Engl. J. Med. 1997, 336, 1401–1408. [Google Scholar] [CrossRef]
- Thompson, D.; Easton, D.F. Breast Cancer Linkage Consortium. Cancer Incidence in BRCA1 Mutation Carriers. J. Natl. Cancer Inst. 2002, 94, 1358–1365. [Google Scholar] [CrossRef]
- Hershko, A.; Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 1998, 67, 425–479. [Google Scholar] [CrossRef]
- Wu, L.C.; Wang, Z.W.; Tsan, J.T.; Spillman, M.A.; Phung, A.; Xu, X.L.; Yang, M.C.; Huang, L.Y.; Bowcock, A.M.; Baer, R. Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat. Genet. 1996, 14, 430–440. [Google Scholar] [CrossRef]
- Wu-Baer, F.; Lagrazon, K.; Yuan, W.; Baer, R. The BRCA1/BARD1 heterodimer assembles polyubiquitin chains through an unconventional linkage involving lysine residue K6 of ubiquitin. J. Biol. Chem. 2003, 278, 34743–34746. [Google Scholar] [CrossRef]
- Christensen, D.E.; Brzovic, P.S.; Klevit, R.E. E2-BRCA1 RING interactions dictate synthesis of mono- or specific polyubiquitin chain linkages. Nat. Struct. Mol. Biol. 2007, 14, 941–948. [Google Scholar] [CrossRef]
- Witus, S.R.; Stewart, M.D.; Klevit, R.E. The BRCA1/BARD1 ubiquitin ligase and its substrates. Biochem. J. 2021, 478, 3467–3483. [Google Scholar] [CrossRef]
- Shakya, R.; Reid, L.J.; Reczek, C.R.; Cole, F.; Egli, D.; Lin, C.S.; deRooij, D.G.; Hirsch, S.; Ravi, K.; Hicks, J.B.; et al. BRCA1 tumor suppression depends on BRCT phosphoprotein binding, but not its E3 ligase activity. Science 2011, 334, 525–528. [Google Scholar] [CrossRef]
- Masuda, T.; Xu, X.; Dimitriadis, E.K.; Lahusen, T.; Deng, C.X. ‘DNA binding region’ of BRCA1 affects genetic stability through modulating the intra-S-phase checkpoint. Int. J. Biol. Sci. 2016, 12, 133–143. [Google Scholar] [CrossRef]
- Zheng, L.; Pan, H.; Li, S.; Flesken-Nikitin, A.; Chen, P.L.; Boyer, T.G.; Lee, W.H. Sequence-specific transcriptional corepressor function for BRCA1 through a novel zinc finger protein, ZBRK1. Mol. Cell 2000, 6, 757–768. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, H.; Kajino, K.; Greene, M.I. BRCA1 binds c-Myc and inhibits its transcriptional and transforming activity in cells. Oncogene 1998, 17, 1939–1948. [Google Scholar] [CrossRef]
- MacLachlan, T.K.; Takimoto, R.; El-Deiry, W.S. BRCA1 directs a selective p53-dependent transcriptional response towards growth arrest and DNA repair targets. Mol. Cell Biol. 2002, 22, 4280–4292. [Google Scholar] [CrossRef]
- Abramovitch, S.; Glaser, T.; Ouchi, T.; Werner, H. BRCA1-Sp1 interactions in transcriptional regulation of the IGF-IR gene. FEBS Lett. 2003, 541, 149–154. [Google Scholar] [CrossRef]
- Tan, W.; Zheng, L.; Lee, W.H.; Boyer, T. Functional Dissection of Transcription Factor ZBRK1 Reveals Zinc Fingers with Dual Roles in DNA-binding and BRCA1-dependent Transcriptional Repression. J. Biol. Chem. 2004, 279, 6576–6587. [Google Scholar] [CrossRef]
- Furuta, S.; Wang, J.M.; Wei, S.Z.; Jeng, Y.M.; Jiang, X.Z.; Gu, B.N.; Chen, P.-L.; Lee, E.Y.-H.P.; Lee, W.-H. Removal of BRCA1/CtIP/ZBRK1 repressor complex on ANG1 promoter leads to accelerated mammary tumor growth contributed by prominent vasculature. Cancer Cell 2006, 10, 13–24. [Google Scholar] [CrossRef]
- Ahmed, K.M.; Tsai, C.Y.; Lee, W.H. Derepression of HMGA2 via removal of ZBRK1/BRCA1/CtIP complex enhances mammary tumorigenesis. J. Biol. Chem. 2010, 285, 4464–4471. [Google Scholar] [CrossRef]
- Hong, R.X.; Zhang, W.M.; Xia, X.; Zhang, K.; Wang, Y.; Wu, M.J.; Fan, J.; Li, J.; Xia, W.; Xu, F.; et al. Preventing BRCA1/ZBRK1 repressor complex binding to the GOT2 promoter results in accelerated aspartate biosynthesis and promotion of cell proliferation. Mol. Oncol. 2019, 13, 959–977. [Google Scholar] [CrossRef]
- Xia, B.; Sheng, Q.; Nakanishi, K.; Ohashi, A.; Wu, J.; Christ, N.; Liu, X.; Jasin, M.; Couch, F.J.; Livingston, D.M. Control of BRCA2 cellular and clinical functions by a nuclear partner, PALB2. Mol. Cell 2006, 22, 719–729. [Google Scholar] [CrossRef]
- Sy, S.M.; Huen, M.S.; Zhu, Y.; Chen, J. PALB2 regulates recombinational repair through chromatin association and oligomerization. J. Biol. Chem. 2009, 284, 18302–18310. [Google Scholar] [CrossRef]
- Antonis, C.; Antoniou, A.C.; Casadei, S.; Heikkinen, T.; Barrowdale, D.; Pylkäs, K.; Robert, J.; Lee, A.; Subramanian, D.; Leeneer, K.D.; et al. Breast-Cancer Risk in Families with Mutations in PALB2. N. Engl. J. Med. 2014, 371, 497–506. [Google Scholar] [CrossRef]
- Manke, I.A.; Lowery, D.M.; Nguyen, A.; Yaffe, M.B. BRCT repeats as phosphopeptide-binding modules involved in protein targeting. Science 2003, 302, 636–639. [Google Scholar] [CrossRef]
- Yu, X.; Chini, C.C.; He, M.; Mer, G.; Chen, J. The BRCT domain is a phospho-protein binding domain. Science 2003, 302, 639–642. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, M.; Yu, X.; Chen, J.; Songyang, Z. Phosphopeptide binding specificities of BRCA1 COOH-terminal (BRCT) domains. J. Biol. Chem. 2003, 278, 52914–52918. [Google Scholar] [CrossRef]
- Wang, B. BRCA1 tumor suppressor network: Focusing on its tail. Cell Biosci. 2012, 2, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Matsuoka, S.; Ballif, B.A.; Zhang, D.; Smogorzewska, A.; Gygi, S.P.; Elledge, S.J. Abraxas and RAP80 form a BRCA1 protein complex required for the DNA damage response. Science 2007, 316, 1194–1198. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Hurov, K.; Hofmann, K.; Elledge, S.J. NBA1, a new player in the Brca1 A complex, is required for DNA damage resistance and checkpoint control. Genes Dev. 2009, 23, 729–739. [Google Scholar] [CrossRef]
- Hu, X.; Kim, J.A.; Castillo, A.; Huang, M.; Liu, J.; Wang, B. NBA1/MERIT40 and BRE interaction is required for the integrity of two distinct deubiquitinating enzyme BRCC36-containing complexes. J. Biol. Chem. 2011, 286, 11734–11745. [Google Scholar] [CrossRef]
- Castillo, A.; Paul, A.; Sun, B.; Huang, T.H.; Wang, Y.; Yazinski, S.A.; Tyler, J.; Li, L.; You, M.J.; Zou, L.; et al. The BRCA1-interacting protein Abraxas is required for genomic stability and tumor suppression. Cell Rep. 2014, 8, 807–817. [Google Scholar] [CrossRef][Green Version]
- Cantor, S.B.; Bell, D.W.; Ganesan, S.; Kass, E.M.; Drapkin, R.; Grossman, S.; Wahrer, D.C.; Sgroi, D.C.; Lane, W.S.; Haber, D.A.; et al. BACH1, a novel helicase-like protein, interacts directly with BRCA1 and contributes to its DNA repair function. Cell 2001, 105, 149–160. [Google Scholar] [CrossRef]
- Bridge, W.L.; Vandenberg, C.J.; Franklin, R.J.; Hiom, K. The BRIP1 helicase functions independently of BRCA1 in the Fanconi anemia pathway for DNA crosslink repair. Nat. Genet. 2005, 37, 953–957. [Google Scholar] [CrossRef]
- Levran, O.; Attwooll, C.; Henry, R.T.; Milton, K.L.; Neveling, K.; Rio, P.; Batish, S.D.; Kalb, R.; Velleuer, E.; Barral, S.; et al. The BRCA1-interacting helicase BRIP1 is deficient in Fanconi anemia. Nat. Genet. 2005, 37, 931–933. [Google Scholar] [CrossRef]
- Litman, R.; Peng, M.; Jin, Z.; Zhang, F.; Zhang, J.; Powell, S.; Andreassen, P.R.; Cantor, S.B. BACH1 is critical for homologous recombination and appears to be the Fanconi anemia gene product FANCJ. Cancer Cell 2005, 8, 255–265. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Litman, R.; Jin, Z.; Fong, G.; Cantor, S.B. BACH1 is a DNA repair protein supporting BRCA1 damage response. Oncogene 2006, 25, 2245–2253. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Chen, J. DNA damage-induced cell cycle checkpoint control requires CtIP, a phosphorylation-dependent binding partner of BRCA1 C-terminal domains. Mol. Cell Biol. 2004, 24, 9478–9486. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Fu, S.; Lai, M.; Baer, R.; Chen, J. BRCA1 ubiquitinates its phosphorylation-dependent binding partner CtIP. Genes Dev. 2006, 20, 1721–1726. [Google Scholar] [CrossRef] [PubMed]
- Sartori, A.A.; Lukas, C.; Coates, J.; Mistrik, M.; Fu, S.; Bartek, J.; Baer, R.; Lukas, J.; Jackson, S.P. Human CtIP promotes DNA end resection. Nature 2007, 450, 509–514. [Google Scholar] [CrossRef]
- Limbo, O.; Chahwan, C.; Yamada, Y.; de Bruin, R.A.; Wittenberg, C.; Russell, P. Ctp1 is a cell-cycle-regulated protein that functions with Mre11 complex to control double-strand break repair by homologous recombination. Mol. Cell 2007, 28, 134–146. [Google Scholar] [CrossRef]
- Huertas, P.; Cortes-Ledesma, F.; Sartori, A.A.; Aguilera, A.; Jackson, S.P. CDK targets Sae2 to control DNA-end resection and homologous recombination. Nature 2008, 455, 689–692. [Google Scholar] [CrossRef]
- Yun, M.H.; Hiom, K. CtIP-BRCA1 modulates the choice of DNA double-strand-break repair pathway throughout the cell cycle. Nature 2009, 459, 460–463. [Google Scholar] [CrossRef]
- Nakamura, K.; Kogame, T.; Oshiumi, H.; Shinohara, A.; Sumitomo, Y.; Agama, K.; Pommier, Y.; Tsutsui, K.M.; Tsutsui, K.; Hartsuiker, E.; et al. Collaborative action of Brca1 and CtIP in elimination of covalent modifications from double-strand breaks to facilitate subsequent break repair. PLoS Genet. 2010, 6, e1000828. [Google Scholar] [CrossRef]
- Negrini, S.; Gorgoulis, V.G.; Halazonetis, T.D. Genomic instability—An evolving hallmark of cancer. Nat. Rev. Mol. Cell Biol. 2010, 11, 220–228. [Google Scholar] [CrossRef]
- Jeggo, P.A.; Pearl, L.H.; Carr, A.M. DNA repair, genome stability and cancer: A historical perspective. Nat. Rev. Cancer 2016, 16, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Kinzler, K.W.; Vogelstein, B. Cancer-susceptibility genes. Gatekeepers and caretakers. Nature 1997, 386, 761–763. [Google Scholar] [CrossRef] [PubMed]
- Hatchi, E.; Skourti-Stathaki, K.; Ventz, S.; Pinello, L.; Yen, A.; Kamieniarz-Gdula, K.; Dimitrov, S.; Pathania, S.; McKinney, K.M.; Eaton, M.L.; et al. BRCA1 recruitment to transcriptional pause sites is required for R-loop-driven DNA damage repair. Mol. Cell 2015, 57, 636–647. [Google Scholar] [CrossRef]
- Zhang, X.; Chiang, H.C.; Wang, Y.; Zhang, C.; Smith, S.; Zhao, X.; Nair, S.J.; Michalek, J.; Jatoi, I.; Lautner, M.; et al. Attenuation of RNA polymerase II pausing mitigates BRCA1-associated R-loop accumulation and tumorigenesis. Nat. Commun. 2017, 8, 15908. [Google Scholar] [CrossRef] [PubMed]
- Chiang, H.-C.; Zhang, X.; Li, J.; Zhao, X.; Chen, J.; Wang, H.T.H.; Jatio, I.; Brenner, A.; Hu, Y.; Li, R. BRCA1-associated R-loop affects transcription and differentiation in breast luminal epithelial cells. Nucleic Acids Res. 2019, 47, 5086–5099. [Google Scholar] [CrossRef]
- Furuta, S.; Jiang, X.; Gu, B.; Cheng, E.; Chen, P.L.; Lee, W.H. Depletion of BRCA1 impairs differentiation but enhances proliferation of mammary epithelial cells. Proc. Natl. Sci. Acad. USA 2005, 102, 9176–9181. [Google Scholar] [CrossRef]
- Liu, S.; Ginestier, C.; Charafe-Jauffret, E.; Foco, H.; Kleer, C.G.; Merajver, S.D.; Dontu, G.; Wicha, M.S. BRCA1 regulates human mammary stem/progenitor cell fate. Proc. Natl. Sci. Acad. USA 2008, 105, 1680–1685. [Google Scholar] [CrossRef]
- Buckley, N.E.; Nic An tSaoir, C.B.; Blayney, J.K.; Oram, L.C.; Crawford, N.T.; D’Costa, Z.C.; Quinn, J.E.; Kennedy, R.D.; Harkin, D.P.; Mullan, P.B. BRCA1 is a key regulator of breast differentiation through activation of Notch signalling with implications for anti-endocrine treatment of breast cancers. Nucleic Acids Res. 2013, 41, 8601–8614. [Google Scholar] [CrossRef]
- Ding, L.; Su, Y.; Fassl, A.; Hinohara, K.; Qiu, X.; Harper, N.W.; Huh, S.J.; Bloushtain-Qimron, N.; Jovanović, B.; Ekram, M.; et al. Perturbed myoepithelial cell differentiation in BRCA mutation carriers and in ductal carcinoma in situ. Nat. Commun. 2019, 10, 4182. [Google Scholar] [CrossRef]
- Kim, H.; Lin, Q.; Yun, Z. BRCA1 regulates the cancer stem cell fate of breast cancer cells in the context of hypoxia and histone deacetylase inhibitors. Sci. Rep. 2019, 9, 9702. [Google Scholar] [CrossRef]
- Ding, J.; Deng, C.X. Mouse models of BRCA1 and their application to breast cancer research. Cancer Metastasis Rev. 2013, 32, 25–37. [Google Scholar]
- Liu, C.Y.; Flesken-Nikitin, A.; Li, S.; Zeng, Y.; Lee, W.H. Inactivation of the mouse Brca1 gene leads to failure in the morphogenesis of the egg cylinder in early postimplantation development. Genes Dev. 1996, 10, 1835–1843. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.S.; Cao, L.; Lim, S.C.; Li, C.; Wang, R.H.; Xu, X.; Bachelier, R.; Deng, C.X. Hyperplasia and spontaneous tumor development in the gynecologic system in mice lacking the BRCA1-Delta11 isoform. Mol. Cell Biol. 2006, 26, 6983–6992. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Wagner, K.U.; Larson, D.; Weaver, Z.; Li, C.; Ried, T.; Hennighausen, L.; Wynshaw-Boris, A.; Deng, C.X. Conditional mutation of Brca1 in mammary epithelial cells results in blunted ductal morphogenesis and tumour formation. Nat. Genet. 1999, 22, 37–43. [Google Scholar] [CrossRef]
- Weaver, Z.; Montagna, C.; Xu, X.; Howard, T.; Gadina, M.; Brodie, S.G.; Deng, C.X.; Ried, T. Mammary tumors in mice conditionally mutant for Brca1 exhibit gross genomic instability and centrosome amplification yet display a recurring distribution of genomic imbalances that is similar to human breast cancer. Oncogene 2002, 21, 5097–5107. [Google Scholar] [CrossRef]
- Poole, A.J.; Li, Y.; Kim, Y.; Lin, S.C.; Lee, W.H.; Lee, E.Y. Prevention of Brca1-mediated mammary tumorigenesis in mice by a progesterone antagonist. Science 2006, 314, 1467–1470. [Google Scholar] [CrossRef]
- Yang, Y.; Swaminathan, S.; Martin, B.K.; Sharan, S.K. Aberrant splicing induced by missense mutations in BRCA1: Clues from a humanized mouse model. Hum. Mol. Genet. 2003, 12, 2121–2131. [Google Scholar] [CrossRef]
- Chang, S.; Biswas, K.; Martin, B.K.; Stauffer, S.; Sharan, S.K. Expression of human BRCA1 variants in mouse ES cells allows functional analysis of BRCA1 mutations. J. Clin. Investig. 2009, 119, 3160–3171. [Google Scholar] [CrossRef]
- Drost, R.; Bouwman, P.; Rottenberg, S.; Boon, U.; Schut, E.; Klarenbeek, S.; Klijn, C.; van der Heijden, I.; van der Gulden, H.; Wientjens, E.; et al. BRCA1 RING function is essential for tumor suppression but dispensable for therapy resistance. Cancer Cell 2011, 20, 797–809. [Google Scholar] [CrossRef]
- Cao, L.; Xu, X.; Bunting, S.F.; Liu, J.; Wang, R.H.; Cao, L.L.; Wu, J.J.; Peng, T.N.; Chen, J.; Nussenzweig, A.; et al. A selective requirement for 53BP1 inthe biological response to genomic instability induced by Brca1 deficiency. Mol. Cell 2009, 35, 534–541. [Google Scholar] [CrossRef]
- Bunting, S.F.; Callén, E.; Wong, N.; Chen, H.T.; Polato, F.; Gunn, A.; Bothmer, A.; Feldhahn, N.; Fernandez-Capetillo, O.; Cao, L.; et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 2010, 141, 243–254. [Google Scholar] [CrossRef] [PubMed]
- Bouwman, P.; Aly, A.; Escandell, J.M.; Pieterse, M.; Bartkova, J.; van der Gulden, H.; Hiddingh, S.; Thanasoula, M.; Kulkarni, A.; Yang, Q.; et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat. Struct. Mol. Biol. 2010, 17, 688–695. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Cole, F.; Patel, D.S.; Misenko, S.M.; Her, J.; Malhowski, A.; Alhamza, A.; Zheng, H.; Baer, R.; Ludwig, T.; et al. 53BP1 ablation rescues genomic instability in mice expressing ‘RING-less’ BRCA1. EMBO Rep. 2016, 17, 1532–1541. [Google Scholar] [CrossRef]
- Miao, K.; Lei, J.H.; Valecha, M.V.; Zhang, A.; Xu, J.; Wang, L.; Lyu, X.; Chen, S.; Miao, Z.; Zhang, X.; et al. NOTCH1 activation compensates BRCA1 deficiency and promotes triple-negative breast cancer formation. Nat. Commun. 2020, 11, 3256. [Google Scholar] [CrossRef] [PubMed]
- Zatreanu, D.; Robinson, H.M.R.; Alkhatib, O.; Boursier, M.; Finch, H.; Geo, L.; Grande, D.; Grinkevich, V.; Heald, R.A.; Langdon, S.; et al. Polθ Inhibitors Elicit BRCA-Gene Synthetic Lethality and Target PARP Inhibitor Resistance. Nat. Commun. 2021, 12, 3636. [Google Scholar] [CrossRef]
- Nacson, J.; Di Marcantonio, D.; Wang, Y.; Bernhardy, A.J.; Clausen, E.; Hua, X.; Cai, K.Q.; Martinez, E.; Feng, W.; Callén, E.; et al. BRCA1 mutational complementation induces synthetic viability. Mol. Cell 2020, 78, 951–959. [Google Scholar] [CrossRef] [PubMed]
- Fenichel, P.; Chevalier, N. Brucker-Davis, F. Bisphenol A: An endocrine and metabolic disruptor. Ann. Endocrinol. 2013, 74, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Jones, L.P.; Sampson, A.; Kang, H.J.; Kim, H.J.; Yi, Y.-W.; Kwon, S.Y.; Babus, J.K.; Wang, A.; Bae, I. Loss of BRCA1 leads to an increased sensitivity to Bisphenol A. Toxicol. Lett. 2010, 199, 261–268. [Google Scholar] [CrossRef]
- Stork, C.T.; Bocek, M.; Crossley, M.P.; Sollier, J.; Sanz, L.A.; Chédin, F.; Swigut, T.; Cimprich, K.A. Co-transcriptional R-loops are the main cause of estrogen-induced DNA damage. Elife 2016, 5, e17548. [Google Scholar] [CrossRef]
- Boguslawski, S.J.; Smith, D.E.; Michalak, M.A.; Mickelson, K.E.; Yehle, C.O.; Patterson, W.L.; Carrico, R.J. Characterization of monoclonal antibody to DNA.RNA and its application to immunodetection of hybrids. J. Immunol. Methods 1986, 89, 123–130. [Google Scholar] [CrossRef]
- Yoshimasu, T.; Ohashi, T.; Oura, S.; Kokawa, Y.; Kawago, M.; Hirai, Y.; Miyasaka, M.; Nishiguchi, H.; Kawashima, S.; Yata, Y.; et al. A theoretical model for the Hormetic dose-response curve for anticancer agents. Anticancer Res. 2015, 35, 5851–5855. [Google Scholar] [PubMed]
- Schaefer, M.H.; Serrano, L. Cell type-specific properties and environment shape tissue specificity of cancer genes. Sci. Rep. 2016, 6, 20707. [Google Scholar] [CrossRef] [PubMed]
- Semmler, L.; Reiter-Brennan, C.; Klein, A. BRCA1 and breast cancer: A review of the underlying mechanisms resulting in the tissue-specific tumorigenesis in mutation carriers. J. Breast Cancer 2019, 22, 1–14. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhong, A.X.; Chen, Y.; Chen, P.-L. BRCA1 the Versatile Defender: Molecular to Environmental Perspectives. Int. J. Mol. Sci. 2023, 24, 14276. https://doi.org/10.3390/ijms241814276
Zhong AX, Chen Y, Chen P-L. BRCA1 the Versatile Defender: Molecular to Environmental Perspectives. International Journal of Molecular Sciences. 2023; 24(18):14276. https://doi.org/10.3390/ijms241814276
Chicago/Turabian StyleZhong, Amy X., Yumay Chen, and Phang-Lang Chen. 2023. "BRCA1 the Versatile Defender: Molecular to Environmental Perspectives" International Journal of Molecular Sciences 24, no. 18: 14276. https://doi.org/10.3390/ijms241814276
APA StyleZhong, A. X., Chen, Y., & Chen, P.-L. (2023). BRCA1 the Versatile Defender: Molecular to Environmental Perspectives. International Journal of Molecular Sciences, 24(18), 14276. https://doi.org/10.3390/ijms241814276




