Integrative Plasma Metabolic and Lipidomic Modelling of SARS-CoV-2 Infection in Relation to Clinical Severity and Early Mortality Prediction
Abstract
1. Introduction
2. Results and Discussion
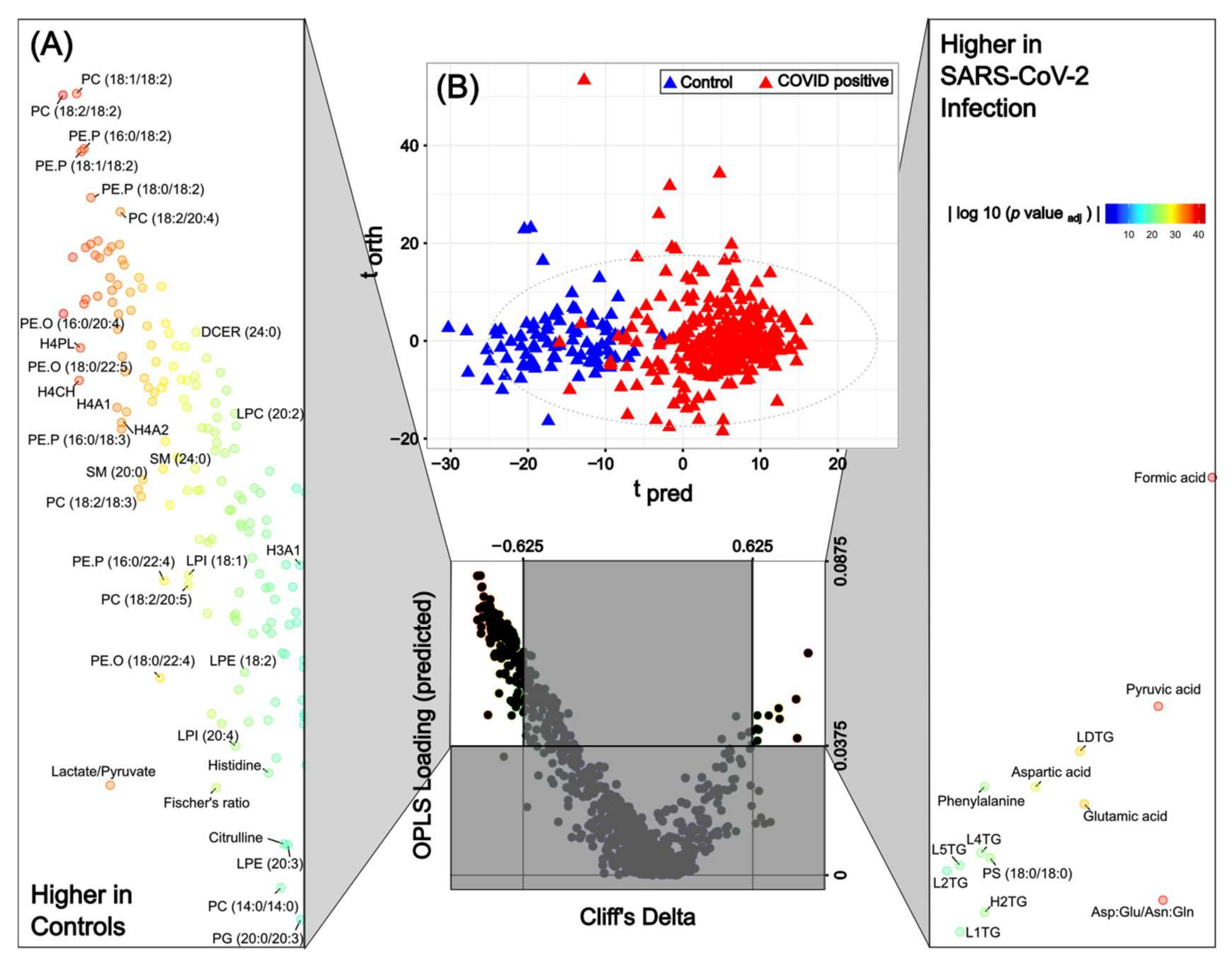
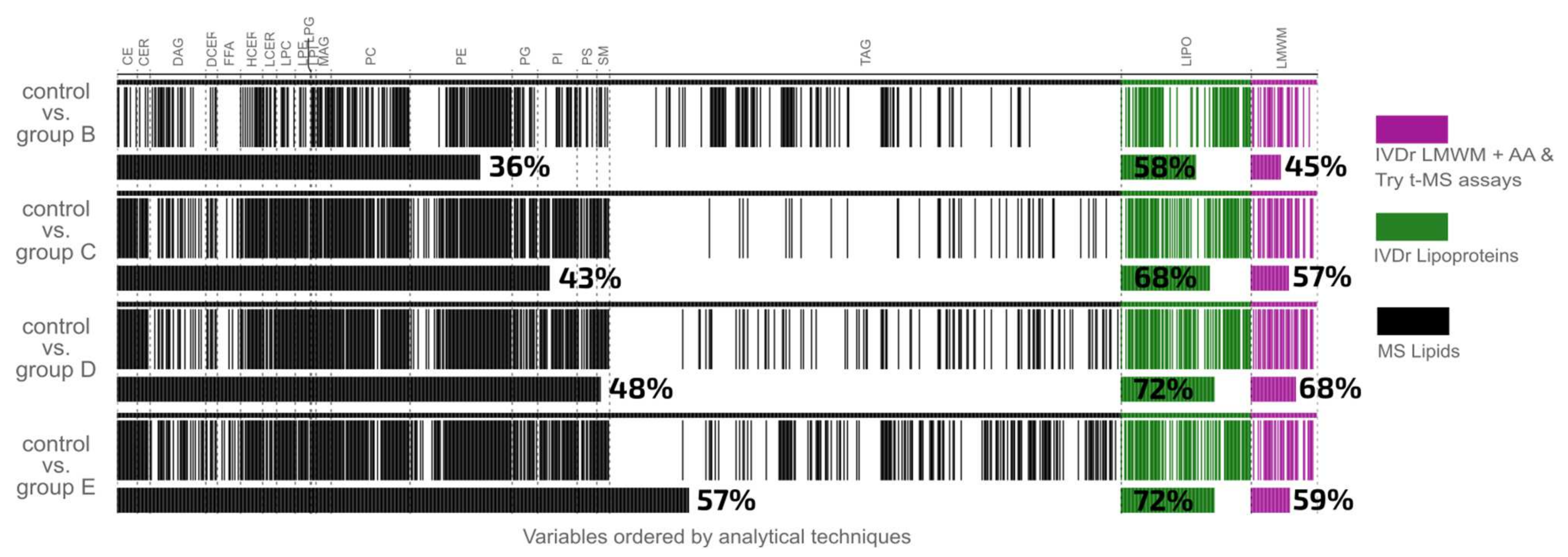
2.1. SARS-CoV-2 Infection Induces Reproducible Metabolic and Lipidomic Consequences across Severity Classes Reflective of Systemic Multi-Organ Effects
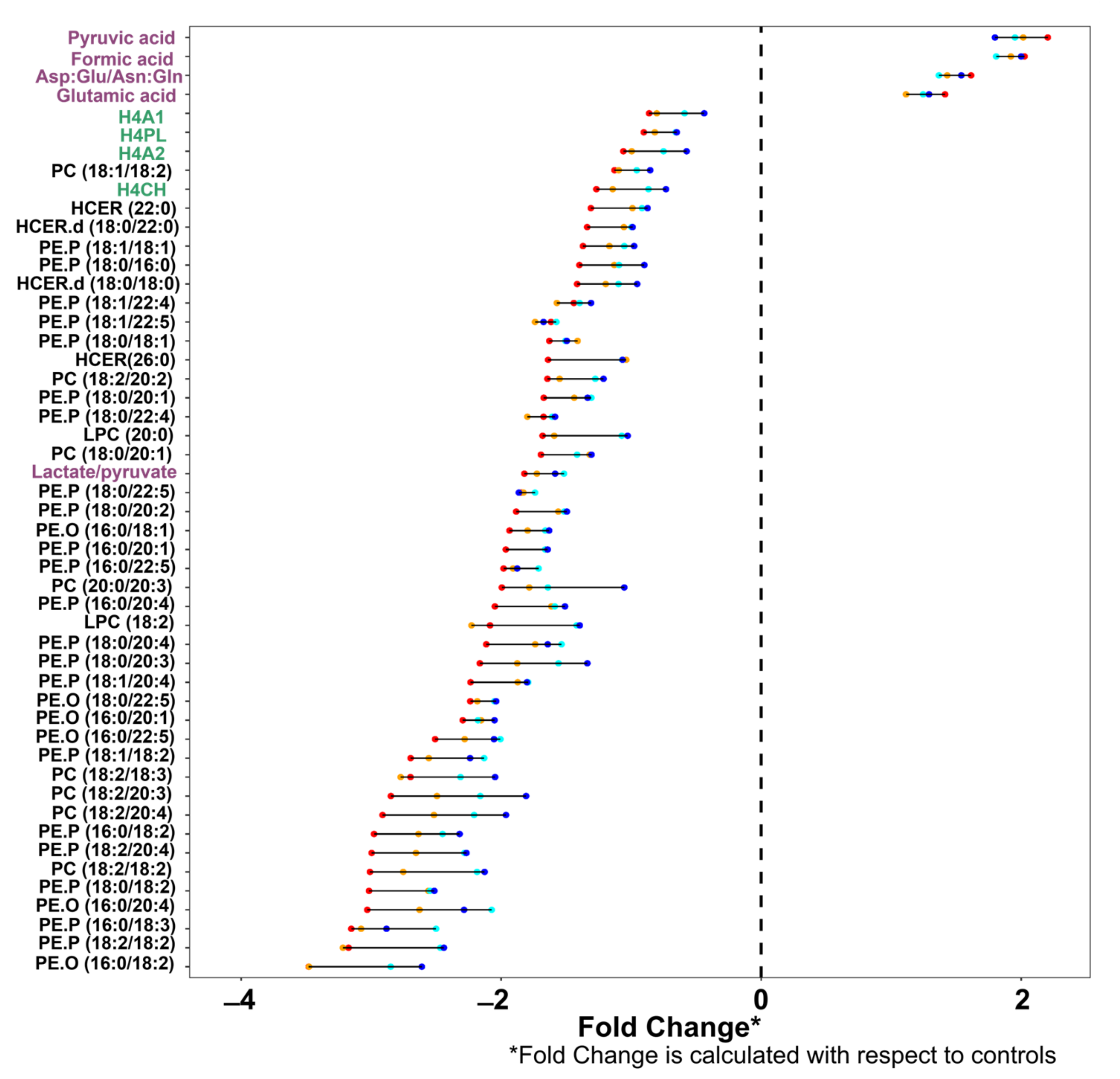
2.2. Metabolite Classes Based on Severity of Infection
2.3. Tryptophan Pathway Metabolism Is Substantially Disrupted in Severe SARS-CoV-2 Infection
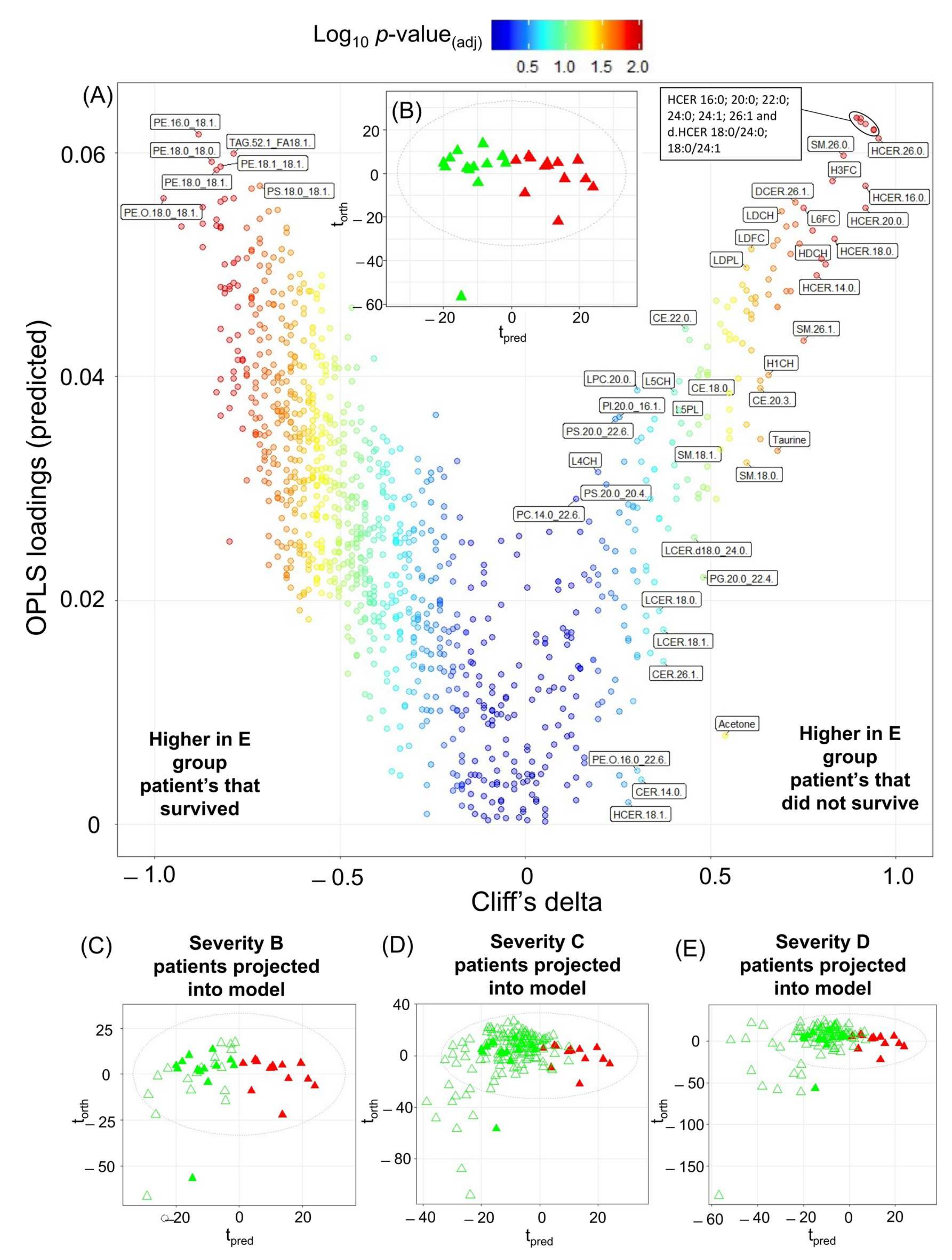
2.4. Mortality Prediction in SARS-CoV-2 Positive Patients
3. Materials and Methods
3.1. Participant Enrolment and Sample Collection
3.2. 1H NMR Spectroscopy Sample Preparation
3.3. 1H NMR Spectroscopy Data Acquisition and Processing Parameters
3.4. Liquid Chromatography Mass Spectrometry (LC-MS)
3.5. LC-MS Lipid Analysis
3.6. Data Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nicholson, J.K. Molecular Phenomic Approaches to Deconvolving the Systemic Effects of SARS-CoV-2 Infection and Post-acute COVID-19 Syndrome. Phenomics 2021, 1, 143–150. [Google Scholar] [CrossRef]
- Holmes, E.; Wist, J.; Masuda, R.; Lodge, S.; Nitschke, P.; Kimhofer, T.; Loo, R.L.; Begum, S.; Boughton, B.; Yang, R.; et al. Incomplete Systemic Recovery and Metabolic Phenoreversion in Post-Acute-Phase Nonhospitalized COVID-19 Patients: Implications for Assessment of Post-Acute COVID-19 Syndrome. J. Proteome Res. 2021, 20, 3315–3329. [Google Scholar] [CrossRef]
- Kimhofer, T.; Lodge, S.; Whiley, L.; Gray, N.; Loo, R.L.; Lawler, N.G.; Nitschke, P.; Bong, S.-H.; Morrison, D.L.; Begum, S.; et al. Integrative Modeling of Quantitative Plasma Lipoprotein, Metabolic, and Amino Acid Data Reveals a Multiorgan Pathological Signature of SARS-CoV-2 Infection. J. Proteome Res. 2020, 19, 4442–4454. [Google Scholar] [CrossRef]
- Lodge, S.; Nitschke, P.; Kimhofer, T.; Coudert, J.D.; Begum, S.; Bong, S.H.; Richards, T.; Edgar, D.; Raby, E.; Spraul, M.; et al. NMR Spectroscopic Windows on the Systemic Effects of SARS-CoV-2 Infection on Plasma Lipoproteins and Metabolites in Relation to Circulating Cytokines. J. Proteome Res. 2021, 20, 1382–1396. [Google Scholar] [CrossRef]
- Gray, N.; Lawler, N.G.; Zeng, A.X.; Ryan, M.; Bong, S.H.; Boughton, B.A.; Bizkarguenaga, M.; Bruzzone, C.; Embade, N.; Wist, J.; et al. Diagnostic Potential of the Plasma Lipidome in Infectious Disease: Application to Acute SARS-CoV-2 Infection. Metabolites 2021, 11, 467. [Google Scholar] [CrossRef] [PubMed]
- Lawler, N.G.; Gray, N.; Kimhofer, T.; Boughton, B.; Gay, M.; Yang, R.; Morillon, A.-C.; Chin, S.-T.; Ryan, M.; Begum, S.; et al. Systemic Perturbations in Amine and Kynurenine Metabolism Associated with Acute SARS-CoV-2 Infection and Inflammatory Cytokine Responses. J. Proteome Res. 2021, 20, 2796–2811. [Google Scholar] [CrossRef] [PubMed]
- Lodge, S.; Nitschke, P.; Kimhofer, T.; Wist, J.; Bong, S.-H.; Loo, R.L.; Masuda, R.; Begum, S.; Richards, T.; Lindon, J.C.; et al. Diffusion and Relaxation Edited Proton NMR Spectroscopy of Plasma Reveals a High-Fidelity Supramolecular Biomarker Signature of SARS-CoV-2 Infection. Anal. Chem. 2021, 93, 3976–3986. [Google Scholar] [CrossRef]
- Lodge, S.; Nitschke, P.; Loo, R.L.; Kimhofer, T.; Bong, S.-H.; Richards, T.; Begum, S.; Spraul, M.; Schaefer, H.; Lindon, J.C.; et al. Low Volume in Vitro Diagnostic Proton NMR Spectroscopy of Human Blood Plasma for Lipoprotein and Metabolite Analysis: Application to SARS-CoV-2 Biomarkers. J. Proteome Res. 2021, 20, 1415–1423. [Google Scholar] [CrossRef] [PubMed]
- Ghini, V.; Meoni, G.; Pelagatti, L.; Celli, T.; Veneziani, F.; Petrucci, F.; Vannucchi, V.; Bertini, L.; Luchinat, C.; Landini, G.; et al. Profiling metabolites and lipoproteins in COMETA, an Italian cohort of COVID-19 patients. PLoS Pathog. 2022, 18, e1010443. [Google Scholar] [CrossRef]
- Thakur, V.; Ratho, R.K.; Kumar, P.; Bhatia, S.K.; Bora, I.; Mohi, G.K.; Saxena, S.K.; Devi, M.; Yadav, D.; Mehariya, S. Multi-Organ Involvement in COVID-19: Beyond Pulmonary Manifestations. J. Clin. Med. 2021, 10, 446. [Google Scholar] [CrossRef]
- Lopes-Pacheco, M.; Silva, P.L.; Cruz, F.F.; Battaglini, D.; Robba, C.; Pelosi, P.; Morales, M.M.; Neves, C.C.; Rocco, P.R.M. Pathogenesis of Multiple Organ Injury in COVID-19 and Potential Therapeutic Strategies. Front. Physiol. 2021, 12, 593223. [Google Scholar] [CrossRef]
- Veyseh, M.; Webster, P.; Blanco, I. COVID-19-associated inflammatory syndrome in an adult woman with unexplained multiple organ failure: Staying vigilant for COVID-19 complications as the pandemic surges. BMJ Case Rep. 2021, 14, e242034. [Google Scholar] [CrossRef] [PubMed]
- Zaim, S.; Chong, J.H.; Sankaranarayanan, V.; Harky, A. COVID-19 and Multiorgan Response. Curr. Probl. Cardiol. 2020, 45, 100618. [Google Scholar] [CrossRef] [PubMed]
- Groff, D.; Sun, A.; Ssentongo, A.E.; Ba, D.M.; Parsons, N.; Poudel, G.R.; Lekoubou, A.; Oh, J.S.; Ericson, J.E.; Ssentongo, P.; et al. Short-term and Long-term Rates of Postacute Sequelae of SARS-CoV-2 Infection: A Systematic Review. JAMA Netw. Open 2021, 4, e2128568. [Google Scholar] [PubMed]
- Bell, J.; Brown, J.; Nicholson, J.; Sadler, P. Assignment of resonances for ‘acute-phase’ glycoproteins in high resolution proton NMR spectra of human blood plasma. FEBS Lett. 1987, 215, 311–315. [Google Scholar] [CrossRef]
- Xie, Y.; Xu, E.; Bowe, B.; Al-Aly, Z. Long-term cardiovascular outcomes of COVID-19. Nat. Med. 2022, 28, 583–590. [Google Scholar] [CrossRef] [PubMed]
- Nalbandian, A.; Sehgal, K.; Gupta, A.; Madhavan, M.V.; McGroder, C.; Stevens, J.S.; Cook, J.R.; Nordvig, A.S.; Shalev, D.; Sehrawat, T.S.; et al. Post-acute COVID-19 syndrome. Nat. Med. 2021, 27, 601–615. [Google Scholar]
- Jiménez, B.; Holmes, E.; Heude, C.; Tolson, R.F.; Harvey, N.; Lodge, S.L.; Chetwynd, A.J.; Cannet, C.; Fang, F.; Pearce, J.T.M.; et al. Quantitative Lipoprotein Subclass and Low Molecular Weight Metabolite Analysis in Human Serum and Plasma by 1H NMR Spectroscopy in a Multilaboratory Trial. Anal. Chem. 2018, 90, 11962–11971. [Google Scholar] [CrossRef]
- Gray, N.; Lawler, N.G.; Yang, R.; Morillon, A.-C.; Gay, M.C.L.; Bong, S.-H.; Holmes, E.; Nicholson, J.K.; Whiley, L. A simultaneous exploratory and quantitative amino acid and biogenic amine metabolic profiling platform for rapid disease phenotyping via UPLC-QToF-MS. Talanta 2021, 223, 121872. [Google Scholar] [CrossRef]
- Masuda, R.; Lodge, S.; Nitschke, P.; Spraul, M.; Schaefer, H.; Bong, S.-H.; Kimhofer, T.; Hall, D.; Loo, R.L.; Bizkarguenaga, M.; et al. Integrative Modeling of Plasma Metabolic and Lipoprotein Biomarkers of SARS-CoV-2 Infection in Spanish and Australian COVID-19 Patient Cohorts. J. Proteome Res. 2021, 20, 4139–4152.e8. [Google Scholar] [CrossRef]
- Bergamaschi, L.; Mescia, F.; Turner, L.; Hanson, A.L.; Kotagiri, P.; Dunmore, B.J.; Ruffieux, H.; De Sa, A.; Huhn, O.; Morgan, M.D.; et al. Longitudinal analysis reveals that delayed bystander CD8+ T cell activation and early immune pathology distinguish severe COVID-19 from mild disease. Immunity 2021, 54, 1257–1275.e8. [Google Scholar] [CrossRef]
- Valdés, A.; Moreno, L.O.; Rello, S.R.; Orduña, A.; Bernardo, D.; Cifuentes, A. Metabolomics study of COVID-19 patients in four different clinical stages. Sci. Rep. 2022, 12, 1650. [Google Scholar] [CrossRef] [PubMed]
- Rothan, H.A.; Byrareddy, S.N. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J. Autoimmun. 2020, 109, 102433. [Google Scholar] [CrossRef] [PubMed]
- Coperchini, F.; Chiovato, L.; Croce, L.; Magri, F.; Rotondi, M. The cytokine storm in COVID-19: An overview of the involvement of the chemokine/chemokine-receptor system. Cytokine Growth Factor. Rev. 2020, 53, 25–32. [Google Scholar] [CrossRef]
- Leisman, D.E.; Ronner, L.; Pinotti, R.; Taylor, M.D.; Sinha, P.; Calfee, C.S.; Hirayama, A.V.; Mastroiani, F.; Turtle, C.J.; Harhay, M.O.; et al. Cytokine elevation in severe and critical COVID-19: A rapid systematic review, meta-analysis, and comparison with other inflammatory syndromes. Lancet Respir. Med. 2020, 8, 1233–1244. [Google Scholar] [CrossRef] [PubMed]
- Chau, A.S.; Weber, A.G.; Maria, N.I.; Narain, S.; Liu, A.; Hajizadeh, N.; Malhotra, P.; Bloom, O.; Marder, G.; Kaplan, B. The Longitudinal Immune Response to Coronavirus Disease 2019: Chasing the Cytokine Storm. Arthritis Rheumatol. 2021, 73, 23–35. [Google Scholar] [CrossRef]
- Collier, M.E.; Zhang, S.; Scrutton, N.S.; Giorgini, F. Inflammation control and improvement of cognitive function in COVID-19 infections: Is there a role for kynurenine 3-monooxygenase inhibition? Drug Discov. Today 2021, 26, 1473–1481. [Google Scholar] [CrossRef]
- Costanzo, M.; Caterino, M.; Fedele, R.; Cevenini, A.; Pontillo, M.; Barra, L.; Ruoppolo, M. COVIDomics: The Proteomic and Metabolomic Signatures of COVID-19. Int. J. Mol. Sci. 2022, 23, 2414. [Google Scholar] [CrossRef]
- López-Hernández, Y.; Monárrez-Espino, J.; Oostdam, A.-S.H.-V.; Delgado, J.E.C.; Zhang, L.; Zheng, J.; Valdez, J.J.O.; Mandal, R.; González, F.d.L.O.; Moreno, J.C.B.; et al. Targeted metabolomics identifies high performing diagnostic and prognostic biomarkers for COVID-19. Sci. Rep. 2021, 11, 14732. [Google Scholar] [CrossRef]
- Yan, H.; Liang, X.; Du, J.; He, Z.; Wang, Y.; Lyu, M.; Yue, L.; Zhang, F.; Xue, Z.; Xu, L.; et al. Proteomic and metabolomic investigation of serum lactate dehydrogenase elevation in COVID-19 patients. Proteomics 2021, 21, e2100002. [Google Scholar] [CrossRef]
- Walldius, G. The apoB/apoA-I Ratio is a Strong Predictor of Cardiovascular Risk. In Lipoproteins; Frank, S., Kostner, G., Eds.; IntechOpen: London, UK, 2012. [Google Scholar]
- Rafsanjani, M.R.; Pramana, T.Y. Arifin Correlation of Fischer’s Ratio with Liver Fibrosis in Naive Chronic Hepatitis B Patients without Comorbidities. Asian J. Pharmacy Nurs. Med. Sci. 2021, 9, 6786. [Google Scholar] [CrossRef]
- Cihan, M.; Doğan, Ö.; Serdar, C.C.; Yıldırım, A.A.; Kurt, C.; Serdar, M.A. Kynurenine pathway in Coronavirus disease (COVID-19): Potential role in prognosis. J. Clin. Lab. Anal. 2022, 36, e24257. [Google Scholar] [CrossRef] [PubMed]
- Marín-Corral, J.; Rodríguez-Morató, J.; Gomez-Gomez, A.; Pascual-Guardia, S.; Muñoz-Bermúdez, R.; Salazar-Degracia, A.; Pérez-Terán, P.; Restrepo, M.I.; Khymenets, O.; Haro, N.; et al. Metabolic Signatures Associated with Severity in Hospitalized COVID-19 Patients. Int. J. Mol. Sci. 2021, 22, 4794. [Google Scholar] [CrossRef] [PubMed]
- Bruzzone, C.; Bizkarguenaga, M.; Gil-Redondo, R.; Diercks, T.; Arana, E.; de Vicuña, A.G.; Seco, M.; Bosch, A.; Palazón, A.; Juan, I.S.; et al. SARS-CoV-2 Infection Dysregulates the Metabolomic and Lipidomic Profiles of Serum. Iscience 2020, 23, 101645. [Google Scholar] [CrossRef]
- Codo, A.C.; Davanzo, G.G.; de Brito Monteiro, L.; de Souza, G.F.; Muraro, S.P.; Virgilio-da-Silva, J.V.; Prodonoff, J.S.; Carregari, V.C.; de Biagi Junior, C.A.O.; Crunfli, F.; et al. Elevated Glucose Levels Favor SARS-CoV-2 Infection and Monocyte Response through a HIF-1α/Glycolysis-Dependent Axis. Cell Metab. 2020, 32, 437–446.e5. [Google Scholar] [CrossRef]
- McElvaney, O.J.; McEvoy, N.L.; McElvaney, O.F.; Carroll, T.P.; Murphy, M.P.; Dunlea, D.M.; Choileáin, O.N.; Clarke, J.; Eoin O’Connor, H.G. Characterization of the Inflammatory Response to Severe COVID-19 Illness. Am. J. Respir. Crit. Care Med. 2020, 202, 812–821. [Google Scholar] [CrossRef] [PubMed]
- Ruffieux, H.; Hanson, A.L.; Lodge, S.; Lawler, N.G.; Whiley, L.; Gray, N.; Nolan, T.H.; Bergamaschi, L.; Mescia, F.; Turner, L.; et al. A patient-centric characterization of systemic recovery from SARS-CoV-2 infection. Nat. Immunol. 2023, 24, 349–358. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- Grant, R.S.; Passey, R.; Matanovic, G.; Smythe, G.; Kapoor, V. Evidence for increased de novo synthesis of NAD in im-mune-activated RAW264.7 macrophages: A self-protective mechanism? Arch. Biochem. Biophys. 1999, 372, 1–7. [Google Scholar] [CrossRef]
- Páez-Franco, J.C.; Torres-Ruiz, J.; Sosa-Hernández, V.A.; Cervantes-Díaz, R.; Romero-Ramírez, S.; Pérez-Fragoso, A.; Meza-Sánchez, D.E.; Germán-Acacio, J.M.; Maravillas-Montero, J.L.; Mejía-Domínguez, N.R.; et al. Metabolomics analysis reveals a modified amino acid metabolism that correlates with altered oxygen homeostasis in COVID-19 patients. Sci. Rep. 2021, 11, 6350. [Google Scholar] [CrossRef]
- Li, X.; Tu, B.; Zhang, X.; Xu, W.; Chen, J.; Zhao, G.; Xu, B.; Zheng, J.; Yan, Y.; Hao, P.; et al. Dysregulation of glutamine/glutamate metabolism in COVID-19 patients: A metabolism study in African population and mini meta-analysis. J. Med. Virol. 2022, 95, e28150. [Google Scholar] [CrossRef]
- Cengiz, M.; Uysal, B.B.; Ikitimur, H.; Ozcan, E.; Islamoğlu, M.S.; Aktepe, E.; Yavuzer, H.; Yavuzer, S. Effect of oral l-Glutamine supplementation on Covid-19 treatment. Clin. Nutr. Exp. 2020, 33, 24–31. [Google Scholar] [CrossRef]
- Masoodi, M.; Peschka, M.; Schmiedel, S.; Haddad, M.; Frye, M.; Maas, C.; Lohse, A.; Huber, S.; Kirchhof, P.; Nofer, J.-R.; et al. Disturbed lipid and amino acid metabolisms in COVID-19 patients. J. Mol. Med. 2022, 100, 555–568. [Google Scholar] [CrossRef] [PubMed]
- Matsuyama, T.; Yoshinaga, S.K.; Shibue, K.; Mak, T.W. Comorbidity-associated glutamine deficiency is a predisposition to severe COVID-19. Cell Death Differ. 2021, 28, 3199–3213. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, S.; Nordqvist, H.; Ambikan, A.T.; Gupta, S.; Sperk, M.; Svensson-Akusjärvi, S.; Mikaeloff, F.; Benfeitas, R.; Saccon, E.; Ponnan, S.M.; et al. Metabolic Perturbation Associated With COVID-19 Disease Severity and SARS-CoV-2 Replication. Mol. Cell. Proteom. 2021, 20, 100159. [Google Scholar] [CrossRef] [PubMed]
- Rubino, F.; Amiel, S.A.; Zimmet, P.; Alberti, G.; Bornstein, S.; Eckel, R.H.; Mingrone, G.; Boehm, B.; Cooper, M.E.; Chai, Z.; et al. New-Onset Diabetes in Covid-19. N. Engl. J. Med. 2020, 383, 789–790. [Google Scholar] [CrossRef]
- Ssentongo, P.; Zhang, Y.; Witmer, L.; Chinchilli, V.M.; Ba, D.M. Association of COVID-19 with diabetes: A systematic review and meta-analysis. Sci. Rep. 2022, 12, 20191. [Google Scholar] [CrossRef]
- Liu, X.; Zheng, Y.; Guasch-Ferré, M.; Ruiz-Canela, M.; Toledo, E.; Clish, C.; Liang, L.; Razquin, C.; Corella, D.; Estruch, R.; et al. High plasma glutamate and low glutamine-to-glutamate ratio are associated with type 2 diabetes: Case-cohort study within the PREDIMED trial. Nutr. Metab. Cardiovasc. Dis. 2019, 29, 1040–1049. [Google Scholar] [CrossRef]
- Samocha-Bonet, D.; Wong, O.; Synnott, E.-L.; Piyaratna, N.; Douglas, A.; Gribble, F.M.; Holst, J.J.; Chisholm, D.J.; Greenfield, J.R. Glutamine Reduces Postprandial Glycemia and Augments the Glucagon-Like Peptide-1 Response in Type 2 Diabetes Patients. J. Nutr. 2011, 141, 1233–1238. [Google Scholar] [CrossRef]
- Albrecht, P.; Lewerenz, J.; Dittmer, S.; Noack, R.; Maher, P.; Methner, A. Mechanisms of Oxidative Glutamate Toxicity: The Glutamate/Cystine Antiporter System xc¯ as a Neuroprotective Drug Target. CNS Neurol. Disord. Drug Targets 2010, 9, 373–382. [Google Scholar] [CrossRef]
- Correia, B.S.B.; Ferreira, V.G.; Piagge, P.M.F.D.; Almeida, M.B.; Assunção, N.A.; Raimundo, J.R.S.; Fonseca, F.L.A.; Carrilho, E.; Cardoso, D.R. 1H qNMR-Based Metabolomics Discrimination of Covid-19 Severity. J. Proteome Res. 2022, 21, 1640–1653. [Google Scholar] [CrossRef] [PubMed]
- Schmelter, F.; Föh, B.; Mallagaray, A.; Rahmöller, J.; Ehlers, M.; Lehrian, S.; von Kopylow, V.; Künsting, I.; Lixenfeld, A.S.; Martin, E.; et al. Metabolic and Lipidomic Markers Differentiate COVID-19 From Non-Hospitalized and Other Intensive Care Patients. Front. Mol. Biosci. 2021, 8, 737039. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Schurink, B.; Roos, E.; Nossent, E.J.; Duitman, J.W.; Vlaar, A.P.; van der Valk, P.; Vaz, F.M.; Yeh, S.-R.; Geeraerts, Z.; et al. Indoleamine 2,3-dioxygenase (IDO)-1 and IDO-2 activity and severe course of COVID-19. J. Pathol. 2022, 256, 256–261. [Google Scholar] [CrossRef] [PubMed]
- Kluck, G.E.G.; Yoo, J.-A.; Sakarya, E.H.; Trigatti, B.L. Good Cholesterol Gone Bad? HDL and COVID-19. Int. J. Mol. Sci. 2021, 22, 10182. [Google Scholar] [CrossRef]
- Wei, C.; Wan, L.; Yan, Q.; Wang, X.; Zhang, J.; Yang, X.; Zhang, Y.; Fan, C.; Li, D.; Deng, Y.; et al. HDL-scavenger receptor B type 1 facilitates SARS-CoV-2 entry. Nat. Metab. 2020, 2, 1391–1400. [Google Scholar] [CrossRef]
- Feingold, K.R. Lipid and Lipoprotein Levels in Patients with COVID-19 Infections. In Endotext; Feingold, K.R., Anawalt, B., Blackman, M.R., Boyce, A., Chrousos, G., Corpas, E., de Herder, W.W., Dhatariya, K., Dungan, K., Hofland, J., et al., Eds.; MDText.com, Inc.: Dartmouth, MA, USA, 2022. [Google Scholar]
- Harbaum, L.; Ghataorhe, P.; Wharton, J.; Jiménez, B.; Howard, L.S.G.; Gibbs, J.S.R.; Nicholson, J.K.; Rhodes, C.J.; Wilkins, M.R. Reduced plasma levels of small HDL particles transporting fibrinolytic proteins in pulmonary arterial hy-pertension. Thorax 2019, 74, 380–389. [Google Scholar] [CrossRef]
- Long, A.T.; Kenne, E.; Jung, R.; Fuchs, T.A.; Renné, T. Contact system revisited: An interface between inflammation, coagulation, and innate immunity. J. Thromb. Haemost. 2016, 14, 427–437. [Google Scholar] [CrossRef]
- Walldius, G.; Jungner, I. Apolipoprotein A-I versus HDL cholesterol in the prediction of risk for myocardial infarction and stroke. Curr. Opin. Cardiol. 2007, 22, 359–367. [Google Scholar] [CrossRef]
- Avogaro, P.; Bon, G.B.; Cazzolato, G.; Quinci, G.B. Are apolipoproteins better discriminators than lipids for atherosclerosis? Lancet 1979, 1, 901–903. [Google Scholar] [CrossRef]
- Walldius, G.; Jungner, I.; Holme, I.; Aastveit, A.H.; Kolar, W.; Steiner, E. High apolipoprotein B, low apolipoprotein A-I, and improvement in the prediction of fatal myocardial in-farction (AMORIS study): A prospective study. Lancet 2001, 358, 2026–2033. [Google Scholar] [CrossRef]
- Masuda, R.; Lodge, S.; Whiley, L.; Gray, N.; Lawler, N.; Nitschke, P.; Bong, S.-H.; Kimhofer, T.; Loo, R.L.; Boughton, B.; et al. Exploration of Human Serum Lipoprotein Supramolecular Phospholipids Using Statistical Heterospectroscopy in n-Dimensions (SHY-n): Identification of Potential Cardiovascular Risk Biomarkers Related to SARS-CoV-2 Infection. Anal. Chem. 2022, 94, 4426–4436. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Shu, T.; Yang, X.; Song, J.-X.; Zhang, M.; Yao, C.; Liu, W.; Huang, M.; Yu, Y.; Yang, Q.; et al. Plasma metabolomic and lipidomic alterations associated with COVID-19. Natl. Sci. Rev. 2020, 7, 1157–1168. [Google Scholar] [CrossRef] [PubMed]
- Almulla, A.F.; Supasitthumrong, T.; Tunvirachaisakul, C.; Algon, A.A.A.; Al-Hakeim, H.K.; Maes, M. The tryptophan catabolite or kynurenine pathway in COVID-19 and critical COVID-19: A systematic review and meta-analysis. BMC Infect. Dis. 2022, 22, 615. [Google Scholar] [CrossRef] [PubMed]
- Gostner, J.M.; Geisler, S.; Stonig, M.; Mair, L.; Sperner-Unterweger, B.; Fuchs, D. Tryptophan Metabolism and Related Pathways in Psychoneuroimmunology: The Impact of Nutrition and Lifestyle. Neuropsychobiology 2020, 79, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Stone, T.W. Endogenous neurotoxins from tryptophan. Toxicon 2001, 39, 61–73. [Google Scholar] [CrossRef]
- Zádori, D.; Klivényi, P.; Vámos, E.; Fülöp, F.; Toldi, J.; Vécsei, L. Kynurenines in chronic neurodegenerative disorders: Future therapeutic strategies. J. Neural Transm. 2009, 116, 1403–1409. [Google Scholar] [CrossRef]
- Kincses, Z.T.; Toldi, J.; Vécsei, L. Kynurenines, neurodegeneration and Alzheimer’s disease. J. Cell. Mol. Med. 2010, 14, 2045–2054. [Google Scholar] [CrossRef]
- Vyavahare, S.; Kumar, S.; Cantu, N.; Kolhe, R.; Bollag, W.B.; McGee-Lawrence, M.E.; Hill, W.D.; Hamrick, M.W.; Isales, C.M.; Fulzele, S. Tryptophan-Kynurenine Pathway in COVID-19-Dependent Musculoskeletal Pathology: A Minireview. Mediat. Inflamm. 2021, 2021, 2911578. [Google Scholar] [CrossRef]
- Chuang, S.-C.; Fanidi, A.; Ueland, P.M.; Relton, C.; Midttun, O.; Vollset, S.E.; Gunter, M.J.; Seckl, M.J.; Travis, R.C.; Wareham, N.; et al. Circulating Biomarkers of Tryptophan and the Kynurenine Pathway and Lung Cancer Risk. Cancer Epidemiol. Biomark. Prev. 2014, 23, 461–468. [Google Scholar] [CrossRef]
- Grant, R.; Coggan, S.; Smythe, G. The Physiological Action of Picolinic Acid in the Human Brain. Int. J. Tryptophan Res. 2009, 2, IJTR.S2469–79. [Google Scholar] [CrossRef]
- Medana, I.M.; Day, N.P.J.; Salahifar-Sabet, H.; Stocker, R.; Smythe, G.; Bwanaisa, L.; Njobvu, A.; Kayira, K.; Turner, G.D.H.; Taylor, T.E.; et al. Metabolites of the Kynurenine Pathway of Tryptophan Metabolism in the Cerebrospinal Fluid of Malawian Children with Malaria. J. Infect. Dis. 2003, 188, 844–849. [Google Scholar] [CrossRef] [PubMed]
- Danlos, F.-X.; Grajeda-Iglesias, C.; Durand, S.; Sauvat, A.; Roumier, M.; Cantin, D.; Colomba, E.; Rohmer, J.; Pommeret, F.; Baciarello, G.; et al. Metabolomic analyses of COVID-19 patients unravel stage-dependent and prognostic biomarkers. Cell Death Dis. 2021, 12, 258. [Google Scholar] [CrossRef] [PubMed]
- Clayton, T.A.; Lindon, J.C.; Cloarec, O.; Antti, H.; Charuel, C.; Hanton, G.; Provost, J.-P.; Le Net, J.-L.; Baker, D.; Walley, R.J.; et al. Pharmaco-metabonomic phenotyping and personalized drug treatment. Nature 2006, 440, 1073–1077. [Google Scholar] [CrossRef]
- Clayton, T.A.; Baker, D.; Lindon, J.C.; Everett, J.R.; Nicholson, J.K. Pharmacometabonomic identification of a significant host-microbiome metabolic interaction affecting human drug metabolism. Proc. Natl. Acad. Sci. USA 2009, 106, 14728–14733. [Google Scholar] [CrossRef]
- McPhail, M.J.; Shawcross, D.L.; Lewis, M.R.; Coltart, I.; Want, E.J.; Antoniades, C.G.; Veselkov, K.; Triantafyllou, E.; Patel, V.; Pop, O.; et al. Multivariate metabotyping of plasma predicts survival in patients with decompensated cirrhosis. J. Hepatol. 2016, 64, 1058–1067. [Google Scholar] [CrossRef] [PubMed]
- Cas, M.D.; Ottolenghi, S.; Morano, C.; Rinaldo, R.; Roda, G.; Chiumello, D.; Centanni, S.; Samaja, M.; Paroni, R. Link between serum lipid signature and prognostic factors in COVID-19 patients. Sci. Rep. 2021, 11, 21633. [Google Scholar] [CrossRef]
- Abusukhun, M.; Winkler, M.S.; Pöhlmann, S.; Moerer, O.; Meissner, K.; Tampe, B.; Hofmann-Winkler, H.; Bauer, M.; Gräler, M.H.; Claus, R.A. Activation of Sphingomyelinase-Ceramide-Pathway in COVID-19 Purposes Its Inhibition for Therapeutic Strategies. Front. Immunol. 2021, 12, 784989. [Google Scholar] [CrossRef]
- Laaksonen, R.; Ekroos, K.; Sysi-Aho, M.; Hilvo, M.; Vihervaara, T.; Kauhanen, D.; Suoniemi, M.; Hurme, R.; März, W.; Scharnagl, H.; et al. Plasma ceramides predict cardiovascular death in patients with stable coronary artery disease and acute coronary syndromes beyond LDL-cholesterol. Eur. Heart J. 2016, 37, 1967–1976. [Google Scholar] [CrossRef]
- Leimanis-Laurens, M.; Wolfrum, E.; Ferguson, K.; Grunwell, J.R.; Sanfilippo, D.; Prokop, J.W.; Lydic, T.A.; Rajasekaran, S. Hexosylceramides and Glycerophosphatidylcholine GPC(36:1) Increase in Multi-Organ Dysfunction Syndrome Patients with Pediatric Intensive Care Unit Admission over 8-Day Hospitalization. J. Pers. Med. 2021, 11, 339. [Google Scholar] [CrossRef]
- Zhang, J.-Y.; Qu, F.; Li, J.-F.; Liu, M.; Ren, F.; Zhang, J.-Y.; Bian, D.-D.; Chen, Y.; Duan, Z.-P.; Zhang, J.-L.; et al. Up-regulation of Plasma Hexosylceramide (d18: 1/18: 1) Contributes to Genotype 2 Virus Replication in Chronic Hepatitis C: A 20-Year Cohort Study. Medicine 2016, 95, e3773. [Google Scholar] [CrossRef]
- Vitner, E.B.; Avraham, R.; Politi, B.; Melamed, S.; Israely, T. Elevation in sphingolipid upon SARS-CoV-2 infection: Possible implications for COVID-19 pathology. Life Sci. Alliance 2022, 5, 689854. [Google Scholar] [CrossRef] [PubMed]
- Smadja, D.M.; Philippe, A.; Bory, O.; Gendron, N.; Beauvais, A.; Gruest, M.; Peron, N.; Khider, L.; Guerin, C.L.; Goudot, G.; et al. Placental growth factor level in plasma predicts COVID-19 severity and in-hospital mortality. J. Thromb. Haemost. 2021, 19, 1823–1830. [Google Scholar] [CrossRef] [PubMed]
- Villoteau, A.; Asfar, M.; Otekpo, M.; Loison, J.; Gautier, J.; Annweiler, C.; on behalf of the GERIA-COVID study group. Elevated C-reactive protein in early COVID-19 predicts worse survival among hospitalized geriatric patients. PLoS ONE 2021, 16, e0256931. [Google Scholar] [CrossRef]
- De Lorenzo, R.; Sciorati, C.; Ramirez, G.A.; Colombo, B.; Lorè, N.I.; Capobianco, A.; Tresoldi, C.; Cirillo, D.M.; Ciceri, F.; Corti, A.; et al. Chromogranin A plasma levels predict mortality in COVID-19. PLoS ONE 2022, 17, e0267235. [Google Scholar] [CrossRef]
- Li, Y.; Deng, Y.; Ye, L.; Sun, H.; Du, S.; Huang, H.; Zeng, F.; Chen, X.; Deng, G. Clinical Significance of Plasma D-Dimer in COVID-19 Mortality. Front. Med. 2021, 8, 638097. [Google Scholar] [CrossRef] [PubMed]
- Ahirwar, A.K.; Takhelmayum, R.; Sakarde, A.; Rathod, B.D.; Jha, P.K.; Kumawat, R.; Gopal, N. The study of serum hsCRP, ferritin, IL-6 and plasma D-dimer in COVID-19: A retrospective study. Horm. Mol. Biol. Clin. Investig. 2022, 43, 337–344. [Google Scholar] [CrossRef]
- D’amora, P.; Silva, I.D.C.G.; Budib, M.A.; Ayache, R.; Silva, R.M.S.; Silva, F.C.; Appel, R.M.; Júnior, S.S.; Pontes, H.B.D.; Alvarenga, A.C.; et al. Towards risk stratification and prediction of disease severity and mortality in COVID-19: Next generation metabolomics for the measurement of host response to COVID-19 infection. PLoS ONE 2021, 16, e0259909. [Google Scholar] [CrossRef] [PubMed]
- Masvekar, R.R.; Kosa, P.; Jin, K.; Dobbs, K.; Stack, M.A.; Castagnoli, R.; Quaresima, V.; Su, H.C.; Imberti, L.; Notarangelo, L.D.; et al. Prognostic value of serum/plasma neurofilament light chain for COVID -19-associated mortality. Ann. Clin. Transl. Neurol. 2022, 9, 622–632. [Google Scholar] [CrossRef]
- Masana, L.; Correig, E.; Ibarretxe, D.; Anoro, E.; Arroyo, J.A.; Jericó, C.; Guerrero, C.; Miret, M.; Näf, S.; Pardo, A.; et al. Low HDL and high triglycerides predict COVID-19 severity. Sci. Rep. 2021, 11, 7217. [Google Scholar] [CrossRef]
- Richard, V.R.; Gaither, C.; Popp, R.; Chaplygina, D.; Brzhozovskiy, A.; Kononikhin, A.; Mohammed, Y.; Zahedi, R.P.; Nikolaev, E.N.; Borchers, C.H. Early Prediction of COVID-19 Patient Survival by Targeted Plasma Multi-Omics and Machine Learning. Mol. Cell. Proteom. 2022, 21, 100277. [Google Scholar] [CrossRef]
- Lu, C.; Wang, Y.; Sheng, Z.; Liu, G.; Fu, Z.; Zhao, J.; Zhao, J.; Yan, X.; Zhu, B.; Peng, S. NMR-based metabonomic analysis of the hepatotoxicity induced by combined exposure to PCBs and TCDD in rats. Toxicol. Appl. Pharmacol. 2010, 248, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Spriet, L.L.; Whitfield, J. Taurine and skeletal muscle function. Curr. Opin. Clin. Nutr. Metab. Care 2015, 18, 96–101. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Orozco-Aguilar, J.; Achiardi, O.; Simon, F.; Cabello-Verrugio, C. SARS-CoV-2/Renin-Angiotensin System: De-ciphering the Clues for a Couple with Potentially Harmful Effects on Skeletal Muscle. Int. J. Mol. Sci. 2020, 21, 7904. [Google Scholar] [CrossRef] [PubMed]
- Ottenhoff, M.C.; Ramos, L.A.; Potters, W.; Janssen, M.L.F.; Hubers, D.; Hu, S.; Fridgeirsson, E.A.; Piña-Fuentes, D.; Thomas, R.; van der Horst, I.C.C.; et al. Predicting mortality of individual patients with COVID-19: A multicentre Dutch cohort. BMJ Open 2021, 11, e047347. [Google Scholar] [CrossRef] [PubMed]
- Izcovich, A.; Ragusa, M.A.; Tortosa, F.; Marzio, M.A.L.; Agnoletti, C.; Bengolea, A.; Ceirano, A.; Espinosa, F.; Saavedra, E.; Sanguine, V.; et al. Prognostic factors for severity and mortality in patients infected with COVID-19: A systematic review. PLoS ONE 2020, 15, e0241955. [Google Scholar] [CrossRef]
- Hasan, M.R.; Suleiman, M.; Pérez-López, A. Metabolomics in the Diagnosis and Prognosis of COVID-19. Front. Genet. 2021, 12, 721556. [Google Scholar] [CrossRef]
- Loo, R.L.; Lodge, S.; Kimhofer, T.; Bong, S.-H.; Begum, S.; Whiley, L.; Gray, N.; Lindon, J.C.; Nitschke, P.; Lawler, N.G.; et al. Quantitative In-Vitro Diagnostic NMR Spectroscopy for Lipoprotein and Metabolite Measurements in Plasma and Serum: Recommendations for Analytical Artifact Minimization with Special Reference to COVID-19/SARS-CoV-2 Samples. J. Proteome Res. 2020, 19, 4428–4441. [Google Scholar] [CrossRef]
- Dona, A.C.; Jiménez, B.; Schäfer, H.; Humpfer, E.; Spraul, M.; Lewis, M.R.; Pearce, J.T.M.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. Precision High-Throughput Proton NMR Spectroscopy of Human Urine, Serum, and Plasma for Large-Scale Metabolic Phenotyping. Anal. Chem. 2014, 86, 9887–9894. [Google Scholar] [CrossRef]
- Whiley, L.; Nye, L.C.; Grant, I.; Andreas, N.; Chappell, K.E.; Sarafian, M.H.; Misra, R.; Plumb, R.S.; Lewis, M.R.; Nicholson, J.K.; et al. Ultrahigh-Performance Liquid Chromatography Tandem Mass Spectrometry with Electrospray Ionization Quantification of Tryptophan Metabolites and Markers of Gut Health in Serum and Plasma-Application to Clinical and Epidemiology Cohorts. Anal. Chem. 2019, 91, 5207–5216. [Google Scholar] [CrossRef]
- Gray, N.; Zia, R.; King, A.; Patel, V.C.; Wendon, J.; McPhail, M.J.W.; Coen, M.; Plumb, R.S.; Wilson, I.D.; Nicholson, J.K. High-Speed Quantitative UPLC-MS Analysis of Multiple Amines in Human Plasma and Serum via Precolumn Derivatization with 6-Aminoquinolyl-N-hydroxysuccinimidyl Carbamate: Application to Acetaminophen-Induced Liver Failure. Anal. Chem. 2017, 89, 2478–2487. [Google Scholar] [CrossRef]
- Adams, K.J.; Pratt, B.; Bose, N.; Dubois, L.G.; John-Williams, L.S.; Perrott, K.M.; Ky, K.; Kapahi, P.; Sharma, V.; MacCoss, M.J.; et al. Skyline for Small Molecules: A Unifying Software Package for Quantitative Metabolomics. J. Proteome Res. 2020, 19, 1447–1458. [Google Scholar] [CrossRef]
- Luan, H.; Ji, F.; Chen, Y.; Cai, Z. statTarget: A streamlined tool for signal drift correction and interpretations of quantitative mass spectrometry-based omics data. Anal. Chim. Acta 2018, 1036, 66–72. [Google Scholar] [CrossRef] [PubMed]
- Bylesjö, M.; Rantalainen, M.; Cloarec, O.; Nicholson, J.K.; Holmes, E.; Trygg, J. OPLS discriminant analysis: Combining the strengths of PLS-DA and SIMCA classification. J. Chemom. 2006, 20, 341–351. [Google Scholar] [CrossRef]
- Cliff, N. Dominance statistics: Ordinal analyses to answer ordinal questions. Psychol. Bull. 1993, 114, 494–509. [Google Scholar] [CrossRef]
- Otshudiema, J.O.; Folefack, G.L.T.; Nsio, J.M.; Mbala-Kingebeni, P.; Kakema, C.H.; Kosianza, J.B.; Mfumu, A.K.; Saidi, G.N.; Kabongo, P.M.; Okum, R.; et al. Epidemiological Comparison of Four COVID-19 Waves in the Democratic Republic of the Congo, March 2020–January 2022. J. Epidemiol. Glob. Health 2022, 12, 316–327. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, F.; Myers, J.; Basu, D.; Tintinger, G.; Ueckermann, V.; Mathebula, M.; Ramlall, R.; Spoor, S.; de Villiers, T.; Van der Walt, Z.; et al. Decreased severity of disease during the first global omicron variant covid-19 outbreak in a large hospital in tshwane, south africa. Int. J. Infect. Dis. 2022, 116, 38–42. [Google Scholar] [CrossRef]
| Comparison | Integrated Data Set | Lipids | Lipoproteins | Low mw Metabolites | ||||
|---|---|---|---|---|---|---|---|---|
| AUROC | No. | AUROC | No. | AUROC | No. | AUROC | No. | |
| Control vs. Group B | 0.99 | 404 (39.07%) | 0.98 | 296 (34.22%) | 0.91 | 65 (58.04%) | 0.98 | 27 (47.37%) |
| Control vs. Group C | 0.99 | 483 (46.71%) | 0.97 | 372 (43.00%) | 0.95 | 81 (72.32%) | 0.99 | 35 (61.40%) |
| Control vs. Group D | 0.99 | 537 (51.93%) | 0.99 | 414 (47.86%) | 0.97 | 84 (75.00%) | 1.00 | 39 (68.42%) |
| Control vs. Group E | 1.00 | 608 (58.80%) | 0.99 | 490 (56.65%) | 1.00 | 83 (74.12%) | 1.00 | 34 (59.65%) |
| Group B vs. Group C | NS (0.51) | X | NS (0.49) | X | NS (0.58) | X | NS (0.58) | X |
| Group B vs. Group D | 0.62 | 0 (0.00%) | 0.64 | 0 (0.00%) | 0.64 | 0 (0.00%) | 0.76 | 4 (0.07%) |
| Group B vs. Group E | 0.77 | 6 (0.01%) | 0.79 | 4 (0.01%) | 0.70 | 0 (0.00%) | 0.78 | 5 (0.09%) |
| Group C vs. Group D | 0.61 | 12 (0.01%) | NS (0.59) | x | NS (0.58) | x | 0.65 | 15 (26.32%) |
| Group C vs. Group E | 0.73 | 175 (0.17%) | 0.72 | 81 (0.09%) | 0.72 | 58 (0.52%) | 0.74 | 18 (31.58%) |
| Group D vs. Group E | 0.64 | x | NS (0.59) | x | NS (0.59) | x | 0.62 | 1 (0.02%) |
| Rank | Control vs. B Severity | Control vs. C Severity | Control vs. D Severity | Control vs. E Severity |
|---|---|---|---|---|
| 1 | Pyruvic acid | Pyruvic acid | Pyruvic acid | Neopterin/tryptophan |
| 2 | Formic acid | Formic acid | Formic acid | Pyruvic acid |
| 3 | Asp:Glu/Asn:Gln | Asp:Glu/Asn:Gln | Asp:Glu/Asn:Gln | Formic acid |
| 4 | PC (16:0/18:2) | H4PL | H4A1 | Asp:Glu/Asn:Gln |
| 5 | PC (18:1/18:2) | PC (18:1/18:2) | H4PL | TPA1 |
| 6 | PC (20:0/20:4) | H4CH | H4A2 | TPA2 |
| 7 | HCER (22:0) | HCER.d (18:0/18:0) | PC (18:1/18:2) | H4A1 |
| 8 | HCER.d (18:0/18:0) | PE.P (18:0/18:1) | H4CH | H4PL |
| 9 | PE.P (18:1/20:1) | PC (18:2/20:2) | PC (18:2/20:2) | H4A2 |
| 10 | PC (18:2/20:2) | Lactate/pyruvate | LPC (20:0) | PC (18:1/18:2) |
| 11 | PE.P (18:0/20:1) | PE.P (18:0/22:5) | PE.P (18:0/22:5) | H4CH |
| 12 | LPC (18:2) | PE.O (16:0/18:1) | PE.P (16:0/22:5) | PC (18:0/20:1) |
| 13 | PE.P (18:0/20:4) | PE.P (16:0/20:1) | PE.P (16:0/20:4) | PE.P (18:0/22:5) |
| 14 | PE.O (18:0/22:5) | PE.P (16:0/22:5) | LPC (18:2) | PE.P (16:0/22:5) |
| 15 | PE.O (16:0/20:1) | LPC (18:2) | PE.P (18:1/20:4) | PC (20:0/20:3) |
| 16 | PE.P (18:1/18:2) | PE.P (18:0/20:4) | PE.P (18:1/18:2) | PE.P (16:0/20:4) |
| 17 | PC (18:2/18:3) | PE.O (16:0/20:1) | PC (18:2/18:3) | LPC (18:2) |
| 18 | PC (18:2/20:4) | PE.P (18:1/18:2) | PC (18:2/20:3) | PE.P (18:1/20:4) |
| 19 | PE.P (16:0/18:2) | PC (18:2/18:3) | PE.P (16:0/18:2) | PE.P (18:1/18:2) |
| 20 | PE.P (18:2/20:4) | PE.P (16:0/18:2) | PE.P (18:2/20:4) | PC (18:2/20:3) |
| 21 | PC (18:2/18:2) | PE.P (18:2/20:4) | PC (18:2/18:2) | PC (18:2/20:4) |
| 22 | PE.P (18:0/18:2) | PC (18:2/18:2) | PE.P (18:0/18:2) | PE.P (16:0/18:2) |
| 23 | PE.O (16:0/20:4) | PE.P (18:0/18:2) | PE.O (16:0/20:4) | PE.P (18:2/20:4) |
| 24 | PE.P (16:0/18:3) | PE.O (16:0/20:4) | PE.P (18:2/18:2) | PC (18:2/18:2) |
| 25 | PE.O (16:0/18:2) | PE.O (16:0/18:2) | PE.O (16:0/18:2) | PE.O (16:0/20:4) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lodge, S.; Lawler, N.G.; Gray, N.; Masuda, R.; Nitschke, P.; Whiley, L.; Bong, S.-H.; Yeap, B.B.; Dwivedi, G.; Spraul, M.; et al. Integrative Plasma Metabolic and Lipidomic Modelling of SARS-CoV-2 Infection in Relation to Clinical Severity and Early Mortality Prediction. Int. J. Mol. Sci. 2023, 24, 11614. https://doi.org/10.3390/ijms241411614
Lodge S, Lawler NG, Gray N, Masuda R, Nitschke P, Whiley L, Bong S-H, Yeap BB, Dwivedi G, Spraul M, et al. Integrative Plasma Metabolic and Lipidomic Modelling of SARS-CoV-2 Infection in Relation to Clinical Severity and Early Mortality Prediction. International Journal of Molecular Sciences. 2023; 24(14):11614. https://doi.org/10.3390/ijms241411614
Chicago/Turabian StyleLodge, Samantha, Nathan G. Lawler, Nicola Gray, Reika Masuda, Philipp Nitschke, Luke Whiley, Sze-How Bong, Bu B. Yeap, Girish Dwivedi, Manfred Spraul, and et al. 2023. "Integrative Plasma Metabolic and Lipidomic Modelling of SARS-CoV-2 Infection in Relation to Clinical Severity and Early Mortality Prediction" International Journal of Molecular Sciences 24, no. 14: 11614. https://doi.org/10.3390/ijms241411614
APA StyleLodge, S., Lawler, N. G., Gray, N., Masuda, R., Nitschke, P., Whiley, L., Bong, S.-H., Yeap, B. B., Dwivedi, G., Spraul, M., Schaefer, H., Gil-Redondo, R., Embade, N., Millet, O., Holmes, E., Wist, J., & Nicholson, J. K. (2023). Integrative Plasma Metabolic and Lipidomic Modelling of SARS-CoV-2 Infection in Relation to Clinical Severity and Early Mortality Prediction. International Journal of Molecular Sciences, 24(14), 11614. https://doi.org/10.3390/ijms241411614









