Enhanced Production of Nitrogenated Metabolites with Anticancer Potential in Aristolochia manshuriensis Hairy Root Cultures
Abstract
1. Introduction
2. Results and Discussion
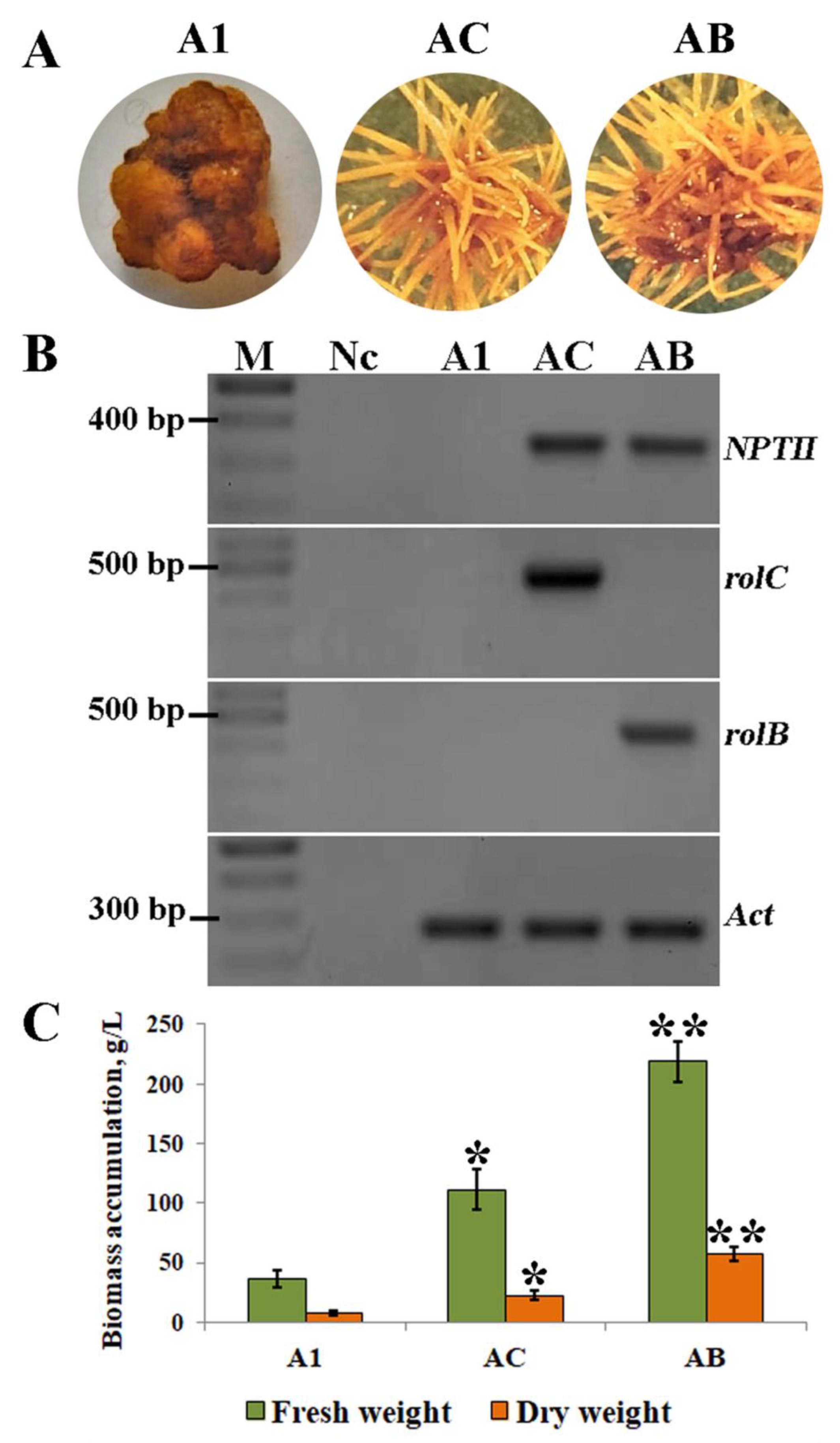
2.1. Effect of Explants on Hairy Root Induction
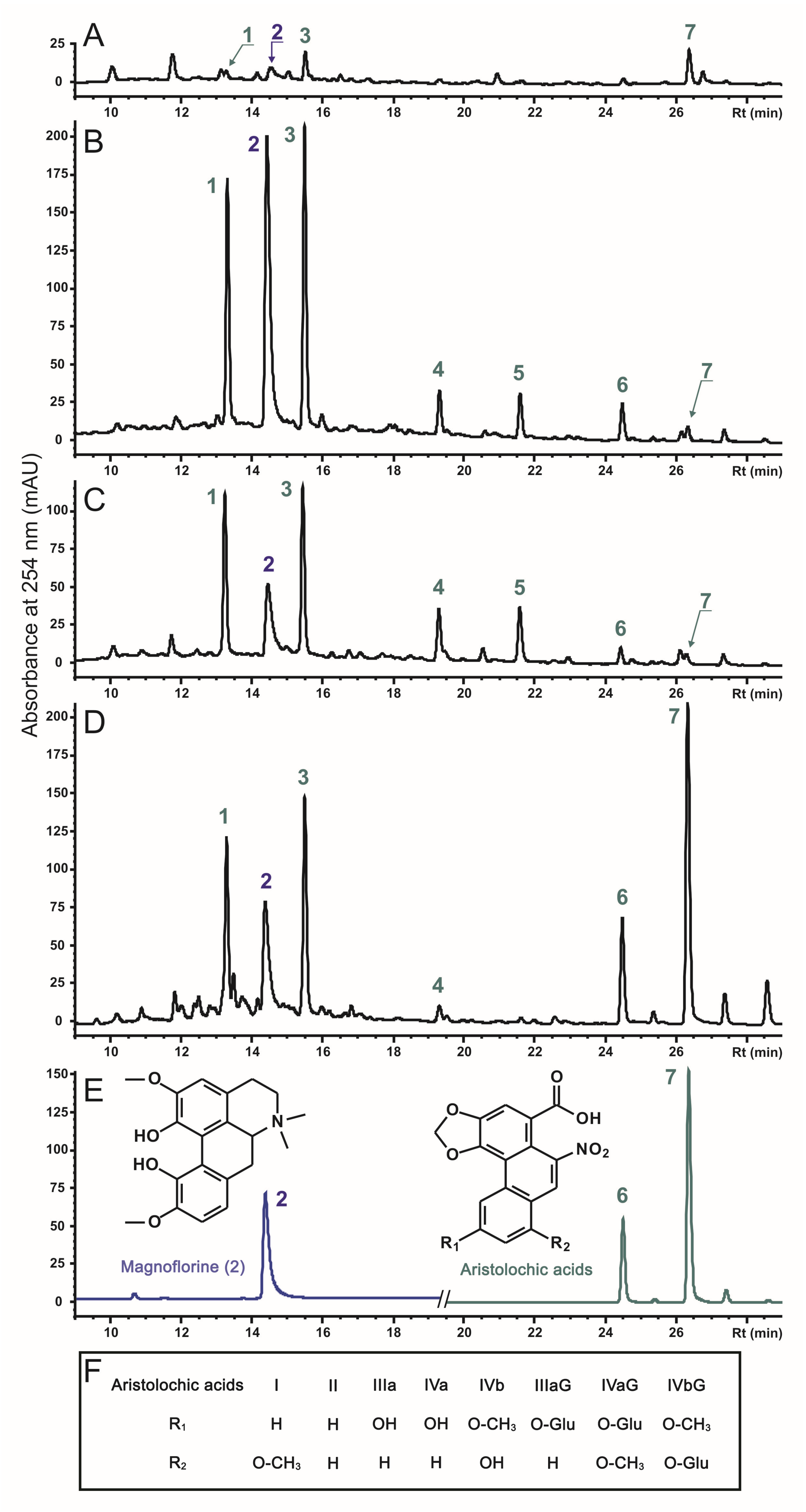
2.2. The Effect of rolC and rolB on Accumulation of Phenanthroic Acid Derivatives in Hairy Root Cultures of A. manshuriensis
2.3. Free Radical Scavenging Activity of A. manshuriensis Cell Cultures Extracts
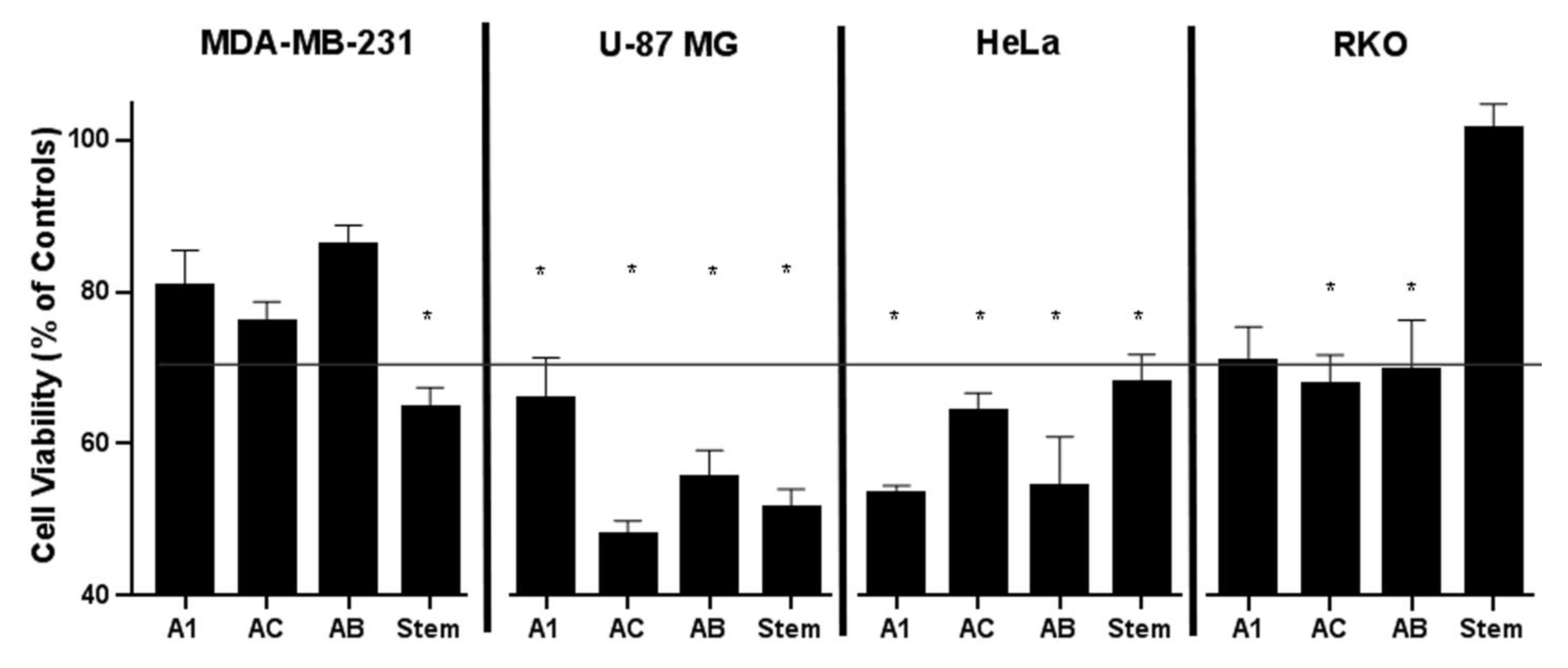
2.4. Cytotoxic Effect of Constituents from A. manshuriensis Cell Cultures
2.5. The Influence of A. manshuriensis Cell Cultures Extract on Cell Cycle of Cancer Cells
3. Materials and Methods
3.1. Plant Material and Tissue Cultures
3.2. DNA Isolation and PCR Analysis
3.3. High-Performance Liquid Chromatography (HPLC) Analysis
3.3.1. Chemicals
3.3.2. Sample Preparation and HPLC Assays
3.4. Antioxidant Activities of Extracts
3.5. Anticancer Activities of Extracts
3.6. Cell Cycle Analysis
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Neinhuis, C.; Wanke, S.; Hilu, K.; Müller, K.; Borsch, T. Phylogeny of Aristolochiaceae based on parsimony, likelihood, and Bayesian analyses of trnL-trnF sequences. Plant. Syst. Evol. 2005, 250, 7–26. [Google Scholar] [CrossRef]
- Bänziger, H.; Disney, R.H.L. Scuttle flies (Diptera: Phoridae) imprisoned by Aristolochia baenzigeri (Aristolochiaceae) in Thailand. Mitt. Schweiz. entomol. Ges. 2006, 79, 29–61. [Google Scholar]
- Murugan, R.; Shivanna, K.R.; Rao, R.R. Pollination biology of Aristolochia tagala, a rare species of medicinal importance. Curr. Sci. 2006, 91, 795–798. [Google Scholar]
- Trujillo, C.G.; Sérsic, A.N. Floral biology of Aristolochia argentina (Aristolochiaceae). Flora Morphol. Distrib. Funct. Ecol. 2006, 201, 374–382. [Google Scholar] [CrossRef]
- Kuo, P.; Li, Y.; Wu, T. Chemical Constituents and Pharmacology of the Aristolochia species. J. Tradit. Complement. Med. 2012, 2, 249–266. [Google Scholar] [CrossRef]
- Lerma-Herrera, M.A.; Beiza-Granados, L.; Ochoa-Zarzosa, A.; López-Meza, J.E.; Navarro-Santos, P.; Herrera-Bucio, R.; Aviña-Verduzco, J.; García-Gutiérrez, H.A. Biological Activities of Organic Extracts of the Genus Aristolochia: A Review from 2005 to 2021. Molecules 2022, 27, 3937. [Google Scholar] [CrossRef] [PubMed]
- Grollman, A.P.; Marcus, D.M. Global hazards of herbal remedies: Lessons from Aristolochia: The lesson from the health hazards of Aristolochia should lead to more research into the safety and efficacy of medicinal plants. EMBO Rep. 2016, 17, 619–625. [Google Scholar] [CrossRef] [PubMed]
- Martena, M.J.; van der Wielen, J.C.A.; van de Laak, L.F.J.; Konings, E.J.M.; de Groot, H.N.; Rietjens, I.M.C.M. Enforcement of the ban on aristolochic acids in Chinese traditional herbal preparations on the Dutch market. Anal. Bioanal. Chem. 2007, 389, 263–275. [Google Scholar] [CrossRef]
- Wu, L.; Sun, W.; Wang, B.; Zhao, H.; Li, Y.; Cai, S.; Xiang, L.; Zhu, Y.; Yao, H.; Song, J.; et al. An integrated system for identifying the hidden assassins in traditional medicines containing aristolochic acids. Sci. Rep. 2015, 5, 11318. [Google Scholar] [CrossRef]
- Nakonechnaya, O.V.; Koren, O.G.; Sidorenko, V.S.; Shabalin, S.A.; Markova, T.O. Poor fruit set due to lack of pollinators in Aristolochia manshuriensis (Aristolochiaceae). Plant. Ecol. Evol. 2021, 154, 39–48. [Google Scholar] [CrossRef]
- Cheung, T.P.; Xue, C.; Leung, K.; Chan, K.; Li, C.G. Aristolochic acids detected in some raw Chinese medicinal herbs and manufactured herbal products--a consequence of inappropriate nomenclature and imprecise labelling? Clin. Toxicol. 2006, 44, 371–378. [Google Scholar] [CrossRef]
- Wang, Y.; Pan, J.X.; Gao, J.J.; Du, K.; Jia, Z.J. The antitumor constituents from stems of Aristolochia manshuriensis. J. Beijing Med. Univ. 2000, 32, 18–21. [Google Scholar]
- Chung, Y.M.; Chang, F.R.; Tseng, T.F.; Hwang, T.L.; Chen, L.C.; Wu, S.F.; Lee, C.L.; Lin, Z.Y.; Chuang, L.Y.; Su, J.H.; et al. A novel alkaloid, aristopyridinone A and anti-inflammatory phenanthrenes isolated from Aristolochia manshuriensis. Bioorg. Med. Chem. Lett. 2011, 21, 1792–1794. [Google Scholar] [CrossRef]
- Zhang, Y.; Jiang, J. Alkaloids from Aristolochia manshuriensis (Aristolochiaceae). Helv. Chim. Acta 2006, 89, 2665–2670. [Google Scholar] [CrossRef]
- Cui, X.; Meng, F.; Pan, X.; Qiu, X.; Zhang, S.; Li, C.; Lu, S. Chromosome-level genome assembly of Aristolochia contorta provides insights into the biosynthesis of benzylisoquinoline alkaloids and aristolochic acids. Hortic. Res. 2022, 9, uhac005. [Google Scholar] [CrossRef] [PubMed]
- Arlt, V.M.; Stiborova, M.; Schmeiser, H.H. Aristolochic acid as a probable human cancer hazard in herbal remedies: A review. Mutagenesis 2002, 17, 265–277. [Google Scholar] [CrossRef]
- Zhang, Y.; Shao, Z.; Han, Y.; Wang, J.; Shi, Y. Study on the attenuation mechanism of Aristolochia manshuriensis compatibility with Zingiberis rhizoma based on different methods of boiling and blending. Int. J. Tradit. Chin. Med. 2019, 6, 1347–1352. [Google Scholar]
- Kang, Y.C.; Chen, M.H.; Lin, C.Y.; Lin, C.Y.; Chen, Y.T. Aristolochic acid-associated urinary tract cancers: An updated meta-analysis of risk and oncologic outcomes after surgery and systematic review of molecular alterations observed in human studies. Ther. Adv. Drug. Saf. 2021, 12, 1–25. [Google Scholar] [CrossRef]
- Hegde, V.R.; Borges, S.; Patel, M.; Das, P.R.; Wu, B.; Gullo, V.P.; Chan, T.M. New potential antitumor compounds from the plant Aristolochia manshuriensis as inhibitors of the CDK2 enzyme. Bioorg. Med. Chem. Lett. 2010, 20, 1344–1346. [Google Scholar] [CrossRef]
- Xu, T.; Kuang, T.; Du, H.; Li, Q.; Feng, T.; Zhang, Y.; Fan, G. Magnoflorine: A review of its pharmacology, pharmacokinetics and toxicity. Pharmacol. Res. 2020, 152, 104632. [Google Scholar] [CrossRef]
- Liang, H.; Chang, X.; Xia, R.N.; Wu, W.; Guo, H.J.; Yang, M. Magnoflorine attenuates cerebral ischemia-induced neuronal injury via autophagy/sirt1/AMPK signaling pathway. Evid. Based. Complement. Alternat. Med. 2022, 2131561. [Google Scholar] [CrossRef]
- Pattar, P.V.; Jayara, M. In vitro Regeneration of plantlets from leaf and nodal explants of Aristolochia indica L. – an important threatened medicinal plant. Asian Pac. J. Trop. Biomed. 2012, 2, S488–S493. [Google Scholar] [CrossRef]
- Sarma, B.; Tanti, B. In vitro regeneration of plantlets from nodal explants of Aristolochia saccata and Aristolochia cathcartii. Eur. J. Biol. Res. 2017, 7, 191–201. [Google Scholar] [CrossRef]
- Remya, M.; Narmatha Bai, V.; Murugesan, S.; Mutharaian, V.N. Changes in bioactive components of Aristolochia tagala. Cham, a rare species of medicinal importance during its in vitro development through direct regeneration. bioRxiv 2016, 037028. [Google Scholar] [CrossRef]
- Nakonechnaya, O.V.; Koren’, O.G.; Zhuravlev, Y.N. Allozyme variation of the relict plant Aristolochia manshuriensis Kom. (Aristolochiaceae). Russ. J. Genet. 2007, 43, 156–164. [Google Scholar] [CrossRef]
- Voronkova, N.M.; Kholina, A.B.; Koldaeva, M.N.; Nakonechnaya, O.V.; Nechaev, V.A. Morphophysiological dormancy, germination, and cryopreservation in Aristolochia contorta seeds. Plant Ecol. Evol. 2018, 151, 77–86. [Google Scholar] [CrossRef]
- Koren, O.G.; Nakonechnaya, O.V.; Zhuravlev, Y.N. Genetic structure of natural populations of the relict species Aristolochia manshuriensis (Aristolochiaceae) in disturbed and intact habitats. Russ. J. Genet. 2009, 45, 678–684. [Google Scholar] [CrossRef]
- Smetanska, I. Production of secondary metabolites using plant cell cultures. In Food Biotechnology; Stahl, U., Donalies, U., Nevoigt, E., Eds.; Springer: Berlin/Heidelberg, Germany, 2008; Volume 111, pp. 187–228. [Google Scholar] [CrossRef]
- Prabha, A.L.; Nandagopalan, V.; Vasanth, K.; Jayabalan, N. Production of aristolochic acid (AA) I and II from Aristolochia indica and Aristolochia bracteolate through in vitro culture. Plant Cell Biotechnol. Mol. Biol. 2012, 13, 131–140. [Google Scholar]
- Zhang, H.; Wang, C. Callus formation of Aristolochia tuberosa and determination of aristolochic acid of callus by HPLC. J. West China Univ. Med. Sci. 1997, 28, 170–173. [Google Scholar]
- Bulgakov, V.P.; Zhuravlev, Y.N.; Fedoreyev, S.A.; Denisenko, V.A.; Veselova, M.; Kulesh, N.I.; Alshevskaya, E.K.; Radchenko, S.V. Constituents of Aristolochia manshuriensis cell suspension culture possessing cardiotonic activity. Fitoterapia 1996, 67, 238–240. [Google Scholar]
- Gutierrez-Valdes, N.; Häkkinen, S.T.; Lemasson, C.; Guillet, M.; Oksman-Caldentey, K.-M.; Ritala, A.; Cardon, F. Hairy root cultures – A versatile tool with multiple applications. Front. Plant Sci. 2020, 11, 33. [Google Scholar] [CrossRef]
- Stepanova, A.Y.; Malunova, M.V.; Gladkov, E.A.; Evsyukov, S.V.; Tereshonok, D.V.; Solov’eva, A.I.; Timiryazev, K.A. Collection of hairy roots as a basis for fundamental and applied research. Molecules 2022, 27, 8040. [Google Scholar] [CrossRef] [PubMed]
- Bulgakov, V.P.; Zhuravlev, Y.N. Generation of Aristolochia manshuriensis Kom. callus tissue cultures. Rastitel’nye Resur. 1989, 25, 266–270. [Google Scholar]
- Bansal, M.; Kumar, A.; Sudhakara Reddy, M. Influence of Agrobacterium rhizogenes strains on hairy root induction and ‘bacoside A’ production from Bacopa monnieri (L.) Wettst. Acta Physiol. Plant. 2014, 36, 2793–2801. [Google Scholar] [CrossRef]
- Panda, B.M.; Mehta, U.J.; Hazra, S. Optimizing culture conditions for establishment of hairy root culture of Semecarpus anacardium L. 3 Biotech 2017, 7, 21. [Google Scholar] [CrossRef]
- Degtyarenko, A.I.; Gorpenchenko, T.Y.; Grigorchuk, V.P.; Bulgakov, V.P.; Shkryl, Y.N. Optimization of the transient Agrobacterium-mediated transformation of Panax ginseng shoots and its use to change the profile of ginsenoside production. Plant Cell Tissue Organ Cult. 2021, 146, 357–373. [Google Scholar] [CrossRef]
- Shahabzadeh, Z.; Heidari, B.; Hafez, R.F. Induction of transgenic hairy roots in Trigonella foenum-graceum co-cultivated with Agrobacterium rhizogenes harboring a GFP gene. J. Crop Sci. Biotechnol. 2013, 16, 263–268. [Google Scholar] [CrossRef]
- Pala, Z.; Shukla, V.; Alok, A.; Kudale, S.; Desai, N. Enhanced production of an anti-malarial compound artesunate by hairy root cultures and phytochemical analysis of Artemisia pallens Wall. 3 Biotech 2016, 6, 182. [Google Scholar] [CrossRef]
- Lee, S.Y.; Xu, H.; Kim, Y.K.; Park, S.U. Rosmarinic acid production in hairy root cultures of Agastache rugosa Kuntze. World J. Microbiol. Biotechnol. 2008, 24, 969–972. [Google Scholar] [CrossRef]
- Saravanakumar, A.; Aslam, A.; Shajahan, A. Development and optimization of hairy root culture systems in Withania somnifera (L.) Dunal for withaferin-A production. Afr. J. Biotechnol. 2012, 11, 16412–16420. [Google Scholar]
- Gai, Q.Y.; Jiao, J.; Luo, M.; Wang, W.; Ma, W.; Zu, Y.G.; Fu, Y.J. Establishment of high-productive Isatis tinctoria L. hairy root cultures: A promising approach for efficient production of bioactive alkaloids. Biochem. Eng. J. 2015, 95, 37–47. [Google Scholar] [CrossRef]
- Zhao, D.; Fu, C.; Chen, Y.; Ma, F. Transformation of Saussurea medusa for hairy roots and jaceosidin production. Plant Cell Rep. 2004, 23, 468–474. [Google Scholar] [CrossRef]
- Sujatha, G.; Zdravković-Korać, S.; Ćalić, D.; Flamini, G.; Ranjitha Kumari, B.D. High-efficiency Agrobacterium rhizogenes-mediated genetic transformation in Artemisia vulgaris: Hairy root production and essential oil analysis. Ind. Crops Prod. 2013, 643–652. [Google Scholar] [CrossRef]
- Chung, I.M.; Rekha, K.; Rajakumar, G.; Thiruvengadam, M. Production of glucosinolates, phenolic compounds and associated gene expression profiles of hairy root cultures in turnip (Brassica rapa ssp. rapa). 3 Biotech 2016, 6, 175. [Google Scholar] [CrossRef] [PubMed]
- Mohebodini, M.; Fathi, R.; Mehri, N. Optimization of hairy root induction in chicory (Cichorium intybus L.) and effects of nanoparticles on secondary metabolites accumulation. Iran. J. Genet. Plant Breed. 2017, 6, 60–68. [Google Scholar] [CrossRef]
- Spena, A.; Schmülling, T.; Koncz, C.; Schell, J.S. Independent and synergistic activity of rol A, B and C loci in stimulating abnormal growth in plants. EMBO J. 1987, 6, 3891–3899. [Google Scholar] [CrossRef]
- Capone, I.; Spanò, L.; Cardarelli, M.; Bellincampi, D.; Petit, A.; Costantino, P. Induction and growth properties of carrot roots with different complements of Agrobacterium rhizogenes T-DNA. Plant Mol. Biol. 1989, 13, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Bulgakov, V.P.; Khodakovskaya, M.V.; Labetskaya, N.V.; Chernoded, G.K.; Zhuravlev, Y.N. The impact of plant rolC oncogene on ginsenoside production by ginseng hairy root cultures. Phytochemistry 1998, 49, 1929–1934. [Google Scholar] [CrossRef]
- Cardarelli, M.; Mariotti, D.; Pomponi, M.; Spanò, L.; Capone, I.; Costantino, P. Agrobacterium rhizogenes T-DNA genes capable of inducing hairy root phenotype. Mol. Gen. Genet. 1987, 209, 475–480. [Google Scholar] [CrossRef]
- Altamura, M.M. Agrobacterium rhizogenes rolB and rolD genes: Regulation and involvement in plant development. Plant Cell Tissue Organ Cult. 2004, 77, 89–101. [Google Scholar] [CrossRef]
- Schmülling, T.; Schell, J.; Spena, A. Single genes from Agrobacterium rhizogenes influence plant development. EMBO J. 1988, 7, 2621–2629. [Google Scholar] [CrossRef] [PubMed]
- Röder, F.T.; Schmülling, T.; Gatz, C. Efficiency of the tetracycline-dependent gene expression system: Complete suppression and efficient induction of the rolB phenotype in transgenic plants. Mol. Gen. Genet. 1994, 243, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, S.D.; Karimi, M.; Impens, L.; Van Lerberge, E.; Coussens, G.; Aesaert, S.; Rombaut, D.; Holtappels, D.; Ibrahim, H.M.M.; Van Montagu, M.; et al. Efficient CRISPR-mediated base editing in Agrobacterium spp. Proc. Natl. Acad. Sci. USA 2021, 118, e2013338118. [Google Scholar] [CrossRef] [PubMed]
- Shkryl, Y.N.; Veremeichik, G.N.; Bulgakov, V.P.; Tchernoded, G.K.; Mischenko, N.P.; Fedoreyev, S.A.; Zhuravlev, Y.N. Individual and combined effects of the rolA, B, and C genes on anthraquinone production in Rubia cordifolia transformed calli. Biotechnol. Bioeng. 2008, 100, 118–125. [Google Scholar] [CrossRef]
- Priestap, H.A.; Velandia, A.E.; Johnson, J.V.; Barbieri, M.A. Secondary metabolite uptake by the Aristolochia-feeding papilionoid butterfly Battus polydamas. Biochem. Syst. Ecol. 2012, 40, 126–137. [Google Scholar] [CrossRef]
- Michl, J.; Kite, G.C.; Wanke, S.; Zierau, O.; Vollmer, G.; Neinhuis, C.; Simmonds, M.S.J.; Heinrich, M. LC-MS- and 1H NMR-based metabolomic analysis and in vitro toxicological assessment of 43 Aristolochia species. J. Nat. Prod. 2016, 79, 30–37. [Google Scholar] [CrossRef]
- Yu, J.; Ma, C.M.; Wang, X.; Shang, M.Y.; Hattori, M.; Xu, F.; Jing, Y.; Dong, S.W.; Xu, Y.Q.; Zhang, C.Y.; et al. Analysis of aristolochic acids, aristololactams and their analogues using liquid chromatography tandem mass spectrometry. Chin. J. Nat. Med. 2016, 14, 626–640. [Google Scholar] [CrossRef]
- Verma, P.; Khan, S.A.; Masood, N.; Manika, N.; Sharma, A.; Verma, N.; Luqman, S.; Mathur, A.K. Differential rubisco content and photosynthetic efficiency of rol gene integrated Vinca minor transgenic plant: Correlating factors associated with morpho-anatomical changes, gene expression and alkaloid productivity. J. Plant Physiol. 2017, 219, 12–21. [Google Scholar] [CrossRef]
- Palazón, J.; Cusidó, R.M.; Gonzalo, J.; Bonfill, M.; Morales, C.; Piñol, M.T. Relation between the amount of rolC gene product and indole alkaloid accumulation in Catharanthus roseus transformed root cultures. J. Plant Physiol. 1998, 153, 712–718. [Google Scholar] [CrossRef]
- Dilshad, E.; Zafar, S.; Ismail, H.; Waheed, M.T.; Cusido, R.M.; Palazon, J.; Mirza, B. Effect of rol genes on polyphenols biosynthesis in Artemisia annua and their effect on antioxidant and cytotoxic potential of the plant. Appl. Biochem. Biotechnol. 2016, 179, 1456–1468. [Google Scholar] [CrossRef]
- Bulgakov, V.P. Functions of rol genes in plant secondary metabolism. Biotechnol. Adv. 2008, 26, 318–324. [Google Scholar] [CrossRef]
- Sim, H.J.; Kim, J.H.; Lee, K.R.; Hong, J. Simultaneous determination of structurally diverse compounds in different Fangchi species by UHPLC-DAD and UHPLC-ESI-MS/MS. Molecules 2013, 18, 5235–5250. [Google Scholar] [CrossRef]
- Canedo-Téxon, A.; Ramón-Farias, F.; Monribot-Villanueva, J.L. Novel findings to the biosynthetic pathway of magnoflorine and taspine through transcriptomic and metabolomic analysis of Croton draco (Euphorbiaceae). BMC Plant Biol. 2019, 19, 560. [Google Scholar] [CrossRef]
- Tian, X.; Guo, S.; He, K.; Roller, M.; Yang, M.; Liu, Q.; Zhang, L.; Ho, C.T.; Bai, N. Qualitative and quantitative analysis of chemical constituents of Ptychopetalum olacoides Benth. Nat. Prod. Res. 2018, 32, 354–357. [Google Scholar] [CrossRef] [PubMed]
- Furuya, T.; Ikuta, A.; Syōno, K. Alkaloids from callus tissue of Papaver somniferum. Phytochemistry 1972, 11, 3041–3044. [Google Scholar] [CrossRef]
- Gorpenchenko, T.Y.; Grigorchuk, V.P.; Fedoreyev, S.A.; Tarbeeva, D.V.; Tchernoded, G.K.; Bulgakov, V.P. Stepharine production in morphogenic cell cultures of Stephania glabra (ROXB.) Miers. Plant Cell Tissue Organ Cult. 2017, 128, 67–76. [Google Scholar] [CrossRef]
- Minami, H.; Kim, J.S.; Ikezawa, N.; Takemura, T.; Katayama, T.; Kumagai, H.; Sato, F. Microbial production of plant benzylisoquinoline alkaloids. Proc. Natl. Acad. Sci. USA 2008, 105, 7393–7398. [Google Scholar] [CrossRef]
- Shkryl, Y.; Tsydeneshieva, Z.; Degtyarenko, A.; Yugay, Y.; Balabanova, L.; Rusapetova, T.; Bulgakov, V. Plant exosomal vesicles: Perspective information nanocarriers in biomedicine. Appl. Sci. 2022, 12, 8262. [Google Scholar] [CrossRef]
- Selma, S.; Ceulemans, E.; Goossens, A.; Lacchini, E. Clustered regularly interspaced short palindromic repeats tools for plant metabolic engineering: Achievements and perspectives. Curr. Opin. Biotechnol. 2023, 79, 102856. [Google Scholar] [CrossRef]
- Kowalczyk, T.; Merecz-Sadowska, A.; Picot, L.; Brčić Karačonji, I.; Wieczfinska, J.; Śliwiński, T.; Sitarek, P. Genetic manipulation and bioreactor culture of plants as a tool for industry and its applications. Molecules 2022, 27, 795. [Google Scholar] [CrossRef] [PubMed]
- Marone, D.; Mastrangelo, A.M.; Borrelli, G.M.; Mores, A.; Laidò, G.; Russo, M.A.; Ficco, D. Specialized metabolites: Physiological and biochemical role in stress resistance, strategies to improve their accumulation, and new applications in crop breeding and management. Plant Physiol. Biochem. 2022, 172, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Tan, H.; Li, Q.; Chen, J.; Gao, S.; Wang, Y.; Chen, W.; Zhang, L. CRISPR/Cas9-mediated efficient targeted mutagenesis of RAS in Salvia miltiorrhiza. Phytochemistry 2018, 148, 63–70. [Google Scholar] [CrossRef] [PubMed]
- Subramaniyan, V.; Saravanan, R.; Baskaran, D.; Ramalalingam, S. In vitro free radical scavenging and anticancer potential of Aristolochia indica L. against MCF-7 cell line. Int. J. Pharm. 2015, 7, 392–396. [Google Scholar]
- Karan, S.K.; Mishra, S.K.; Pal, D.; Mondal, A. Isolation of β-sitosterol and evaluation of antidiabetic activity of Aristolochia indica in alloxan-induced diabetic mice with a reference to in vitro antioxidant activity. J. Med. Plant Res. 2012, 6, 1219–1223. [Google Scholar] [CrossRef]
- Jegadeeswari, P.; Daffodil, E.; Mohan, V.R. Quantification of total phenolics, flavonoid and in vitro antioxidant activity of Aristolochia bracteata Retz. Int. J. Pharm. Pharm. 2014, 6, 747–752. [Google Scholar]
- El Omari, N.; Sayah, K.; Fettach, S.; El Blidi, O.; Bouyahya, A.; Faouzi, M.E.A.; Kamal, R.; Barkiyou, M. Evaluation of in vitro antioxidant and antidiabetic activities of Aristolochia longa extracts. Evid. Based Complement. Altern. Med. 2019, 2019, 7384735. [Google Scholar] [CrossRef]
- Benmehdi, H.; Behilil, A.; Memmou, F.; Amrouche, A. Free radical scavenging activity, kinetic behaviour and phytochemical constituents of Aristolochia clematitis L. roots. Arab. J. Chem. 2017, 10, S1402–S1408. [Google Scholar] [CrossRef]
- Guinnin, F.F.D.; Sacramento, I.T.; Ategbo, J.M.; Agbangnan, C.D.P. Physico-chemical composition and radicalscavenging activity evaluation of the extracts of Aristolochia albida Duch. (Aristolochiaceae) of Benin. J. Appl. Biosci. 2016, 107, 10460–10470. [Google Scholar] [CrossRef]
- Okon, E.; Kukula-Koch, W.; Jarzab, A.; Halasa, M.; Stepulak, A.; Wawruszak, A. Advances in chemistry and bioactivity of magnoflorine and magnoflorine-containing extracts. Int. J. Mol. Sci. 2020, 21, 1330. [Google Scholar] [CrossRef]
- Li, C.; Wang, M.H. Potential biological activities of magnoflorine: A compound from Aristolochia debilis sieb. et Zucc. Korean J. Plant Res. 2014, 27, 223–228. [Google Scholar] [CrossRef]
- Mohamed, S.; Hassan, E.; Ibrahim, N. Cytotoxic and antiviral activities of aporphine alkaloids of Magnolia grandiflora L. Nat. Prod. Res. 2010, 24, 1395–1402. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.Y.; Ye, X.L.; Cui, X.L.; He, K.; Jin, Y.N.; Chen, Z.; Li, X.G. Cytotoxicity and antihyperglycemic effect of minor constituents from Rhizoma coptis in HepG2 cells. Fitoterapia 2012, 83, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Okon, E.; Kukula-Koch, W.; Halasa, M.; Jarzab, A.; Baran, M.; Dmoszynska-Graniczka, M.; Angelis, A.; Kalpoutzakis, E.; Guz, M.; Stepulak, A.; et al. Magnoflorine-isolation and the anticancer potential against NCI-H1299 lung, MDA-MB-468 breast, T98G glioma, and TE671 rhabdomyosarcoma cancer cells. Biomolecules 2020, 10, 1532. [Google Scholar] [CrossRef]
- Sun, X.L.; Zhang, X.W.; Zhai, H.J.; Zhang, D.; Ma, S.Y. Magnoflorine inhibits human gastric cancer progression by inducing autophagy, apoptosis and cell cycle arrest by JNK activation regulated by ROS. Biomed. Pharmacother. 2020, 125, 109118. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.; Xiaojun, X.; Peilong, C. Magnoflorine improves sensitivity to doxorubicin (DOX) of breast cancer cells via inducing apoptosis and autophagy through AKT/mTOR and p38 signaling pathways. Biomed. Pharmacother. 2020, 121, 109139. [Google Scholar] [CrossRef] [PubMed]
- Angelova, A.L.; Grekova, S.P.; Heller, A.; Kuhlmann, O.; Soyka, E.; Giese, T.; Aprahamian, M.; Bour, G.; Rüffer, S.; Cziepluch, C.; et al. Complementary induction of immunogenic cell death by oncolytic parvovirus H-1PV and gemcitabine in pancreatic cancer. J. Virol. 2014, 88, 5263–5276. [Google Scholar] [CrossRef] [PubMed]
- Chunmei, L.; Myeong-Hyeon, W. Aristolochia debilis Sieb. et Zucc. induces apoptosis and reactive oxygen species in the HT-29 human colon cancer cell line. Cancer Biother. Radiopharm. 2013, 28, 717–724. [Google Scholar] [CrossRef]
- Bulgakov, V.P.; Tchernoded, G.K.; Veselova, M.V.; Fedoreyev, S.A.; Muzarok, T.I.; Zhuravlev, Y.N. Catechin production in cultured cells of Taxus cuspidata and Taxus baccata. Biotechnol. Lett. 2011, 33, 1879–1883. [Google Scholar] [CrossRef]
- Hernalsteens, J.P.; Van Vliet, F.; De Beuckeleer, M.; Depicker, A.; Engler, G.; Lemmers, M.; Holsters, M.; Van Montagu, M.; Schell, J. The Agrobacterium tumefaciens Ti plasmid as a host vector system for introducing foreign DNA into plant cells. Nature 1980, 287, 654–656. [Google Scholar] [CrossRef]
- Echt, C.S.; Erdahl, L.A.; McCoy, T.J. Genetic segregation of random amplified polymorphic DNA in diploid cultivated alfalfa. Genome 1992, 35, 84–87. [Google Scholar] [CrossRef]
- Gorpenchenko, T.Y.; Grigorchuk, V.P.; Bulgakov, D.V.; Tchernoded, G.K.; Bulgakov, V.P. Tempo-spatial pattern of stepharine accumulation in Stephania glabra morphogenic tissues. Int. J. Mol. Sci. 2019, 20, 808. [Google Scholar] [CrossRef] [PubMed]
- Mensor, L.L.; Menezes, F.S.; Leitao, G.G.; Reis, A.S.; Santos, T.S.; Coube, C.S. Screening of Brazilian plant extracts for antioxidant activity by the use of DPPH free radical method. Phytother. Res. 2001, 15, 127–130. [Google Scholar] [CrossRef] [PubMed]
- Shkryl, Y.; Rusapetova, T.; Yugay, Y.; Egorova, A.; Silant’ev, V.; Grigorchuk, V.; Karabtsov, A.; Timofeeva, Y.; Vasyutkina, E.; Kudinova, O.; et al. Biosynthesis and cytotoxic properties of Ag, Au, and bimetallic nanoparticles synthesized using Lithospermum erythrorhizon callus culture extract. Int. J. Mol. Sci. 2021, 22, 9305. [Google Scholar] [CrossRef] [PubMed]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological Statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 1–9. [Google Scholar]

| Stem | Petiole | Leaf | |
|---|---|---|---|
| Control | - | - | - |
| rolA | - | - | - |
| rolC | 10.8 ± 1.9 | 5.9 ± 0.6 ** | - |
| rolB | 20.1 ± 2.3 | 8.3 ± 0.9 ** | - |
| A1 | AC | AB | Stem | |
|---|---|---|---|---|
| Magnoflorine | 0.99 ± 0.102 D | 5.72 ± 0.686 A | 2.76 ± 0.304 C | 3.39 ± 0.475 B |
| AA-I | 0.11 ± 0.010 B | 0.02 ± 0.002 C | 0.02 ± 0.002 C | 1.23 ± 0.139 A |
| AA-II | 0.01 ± 0.001 C | 0.08 ± 0.010 B | 0.06 ± 0.007 B | 0.34 ± 0.027 A |
| AA-IIIa | ND | 0.12 ± 0.016 B | 0.22 ± 0.033 A | 0.04 ± 0.006 C |
| AA-IVa/b | 0.01 ± 0.001 C | 0.15 ± 0.020 B | 0.29 ± 0.034 A | ND |
| AA-IIIa-G | 0.02 ± 0.003 B | 0.77 ± 0.098 A | 0.80 ± 0.119 A | 0.82 ± 0.109 A |
| AA-Iva/b-G | 0.14 ± 0.019 C | 0.99 ± 0.133 AB | 0.94 ± 0.136 B | 1.18 ± 0.154 A |
| Sum of Aas | 0.29 ± 0.037 C | 2.13 ± 0.280 B | 2.32 ± 0.266 B | 3.61 ± 0.513 A |
| Sample | IC50 (µg/mL) |
|---|---|
| Vitamin C | 276 ± 2.29 A |
| A1 | 116 ± 4.71 B |
| AC | 41 ± 1.49 C |
| AB | 40 ± 2.41 C |
| Stem | 44 ± 1.54 C |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shkryl, Y.N.; Tchernoded, G.K.; Yugay, Y.A.; Grigorchuk, V.P.; Sorokina, M.R.; Gorpenchenko, T.Y.; Kudinova, O.D.; Degtyarenko, A.I.; Onishchenko, M.S.; Shved, N.A.; et al. Enhanced Production of Nitrogenated Metabolites with Anticancer Potential in Aristolochia manshuriensis Hairy Root Cultures. Int. J. Mol. Sci. 2023, 24, 11240. https://doi.org/10.3390/ijms241411240
Shkryl YN, Tchernoded GK, Yugay YA, Grigorchuk VP, Sorokina MR, Gorpenchenko TY, Kudinova OD, Degtyarenko AI, Onishchenko MS, Shved NA, et al. Enhanced Production of Nitrogenated Metabolites with Anticancer Potential in Aristolochia manshuriensis Hairy Root Cultures. International Journal of Molecular Sciences. 2023; 24(14):11240. https://doi.org/10.3390/ijms241411240
Chicago/Turabian StyleShkryl, Yury N., Galina K. Tchernoded, Yulia A. Yugay, Valeria P. Grigorchuk, Maria R. Sorokina, Tatiana Y. Gorpenchenko, Olesya D. Kudinova, Anton I. Degtyarenko, Maria S. Onishchenko, Nikita A. Shved, and et al. 2023. "Enhanced Production of Nitrogenated Metabolites with Anticancer Potential in Aristolochia manshuriensis Hairy Root Cultures" International Journal of Molecular Sciences 24, no. 14: 11240. https://doi.org/10.3390/ijms241411240
APA StyleShkryl, Y. N., Tchernoded, G. K., Yugay, Y. A., Grigorchuk, V. P., Sorokina, M. R., Gorpenchenko, T. Y., Kudinova, O. D., Degtyarenko, A. I., Onishchenko, M. S., Shved, N. A., Kumeiko, V. V., & Bulgakov, V. P. (2023). Enhanced Production of Nitrogenated Metabolites with Anticancer Potential in Aristolochia manshuriensis Hairy Root Cultures. International Journal of Molecular Sciences, 24(14), 11240. https://doi.org/10.3390/ijms241411240






