Comparative Analysis Based on Physiological and Transcriptomic Data between Juvenile and Adult Tree Peony (Paeonia delavayi)
Abstract
1. Introduction
2. Results
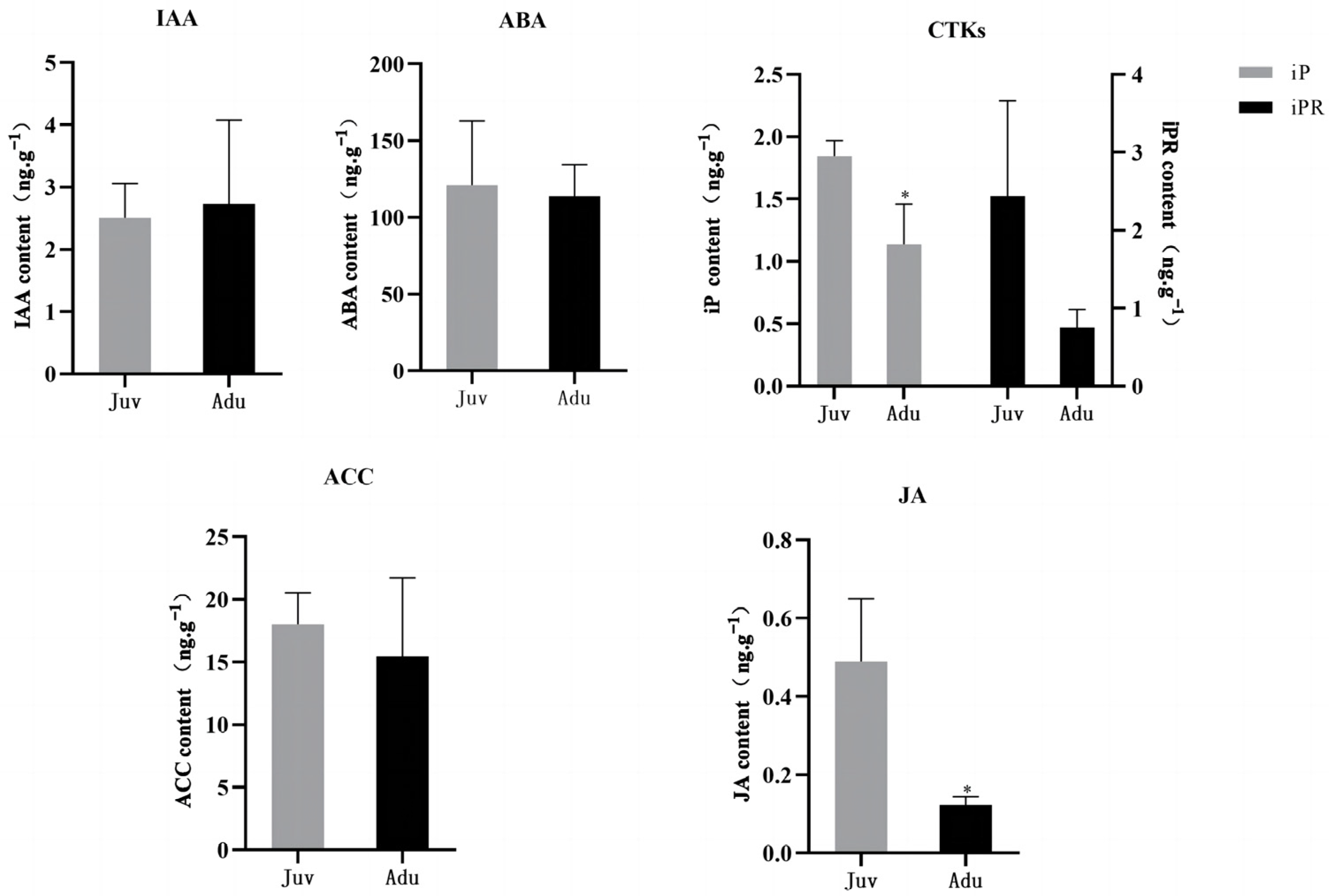
2.1. Physiological Changes between Juvenile and Adult Plants
2.2. Differentially Expressed Genes (DEGs) between Juvenile and Adult Plants and Enriched Pathways of DEGs
2.2.1. Transcriptome Sequencing Assembly and Annotation
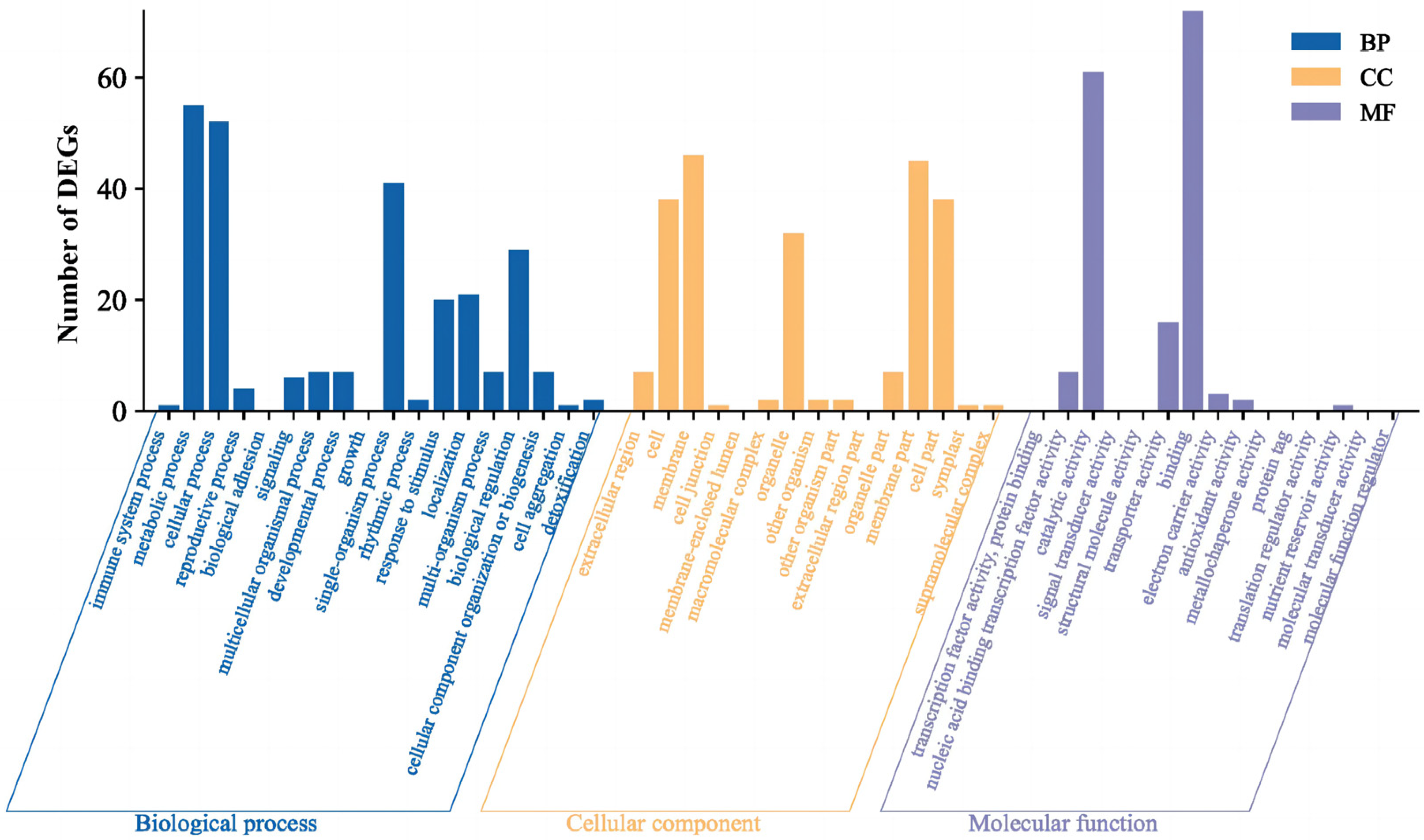
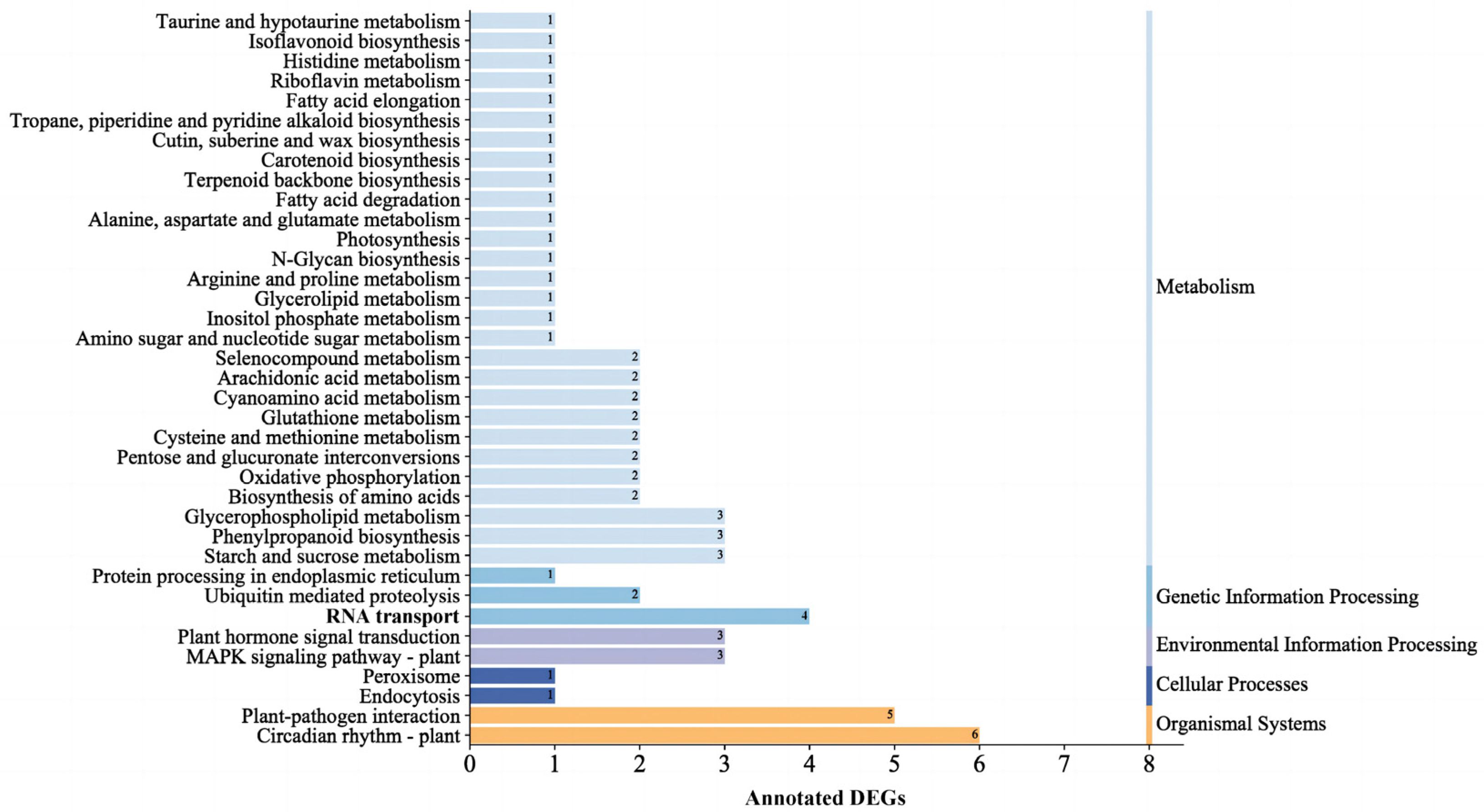
2.2.2. Screening of DEGs and Enrichment Analysis of DEGs
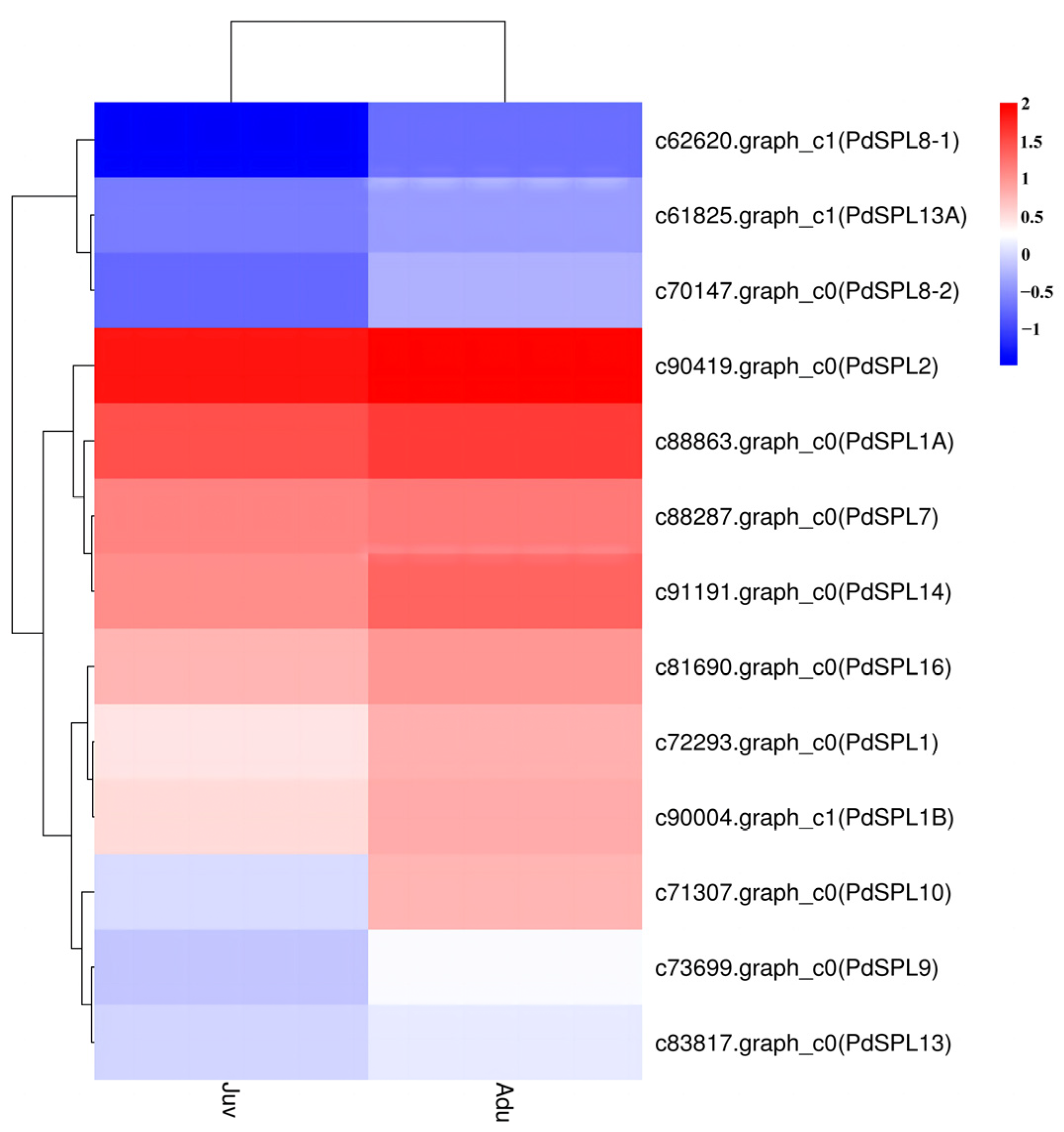
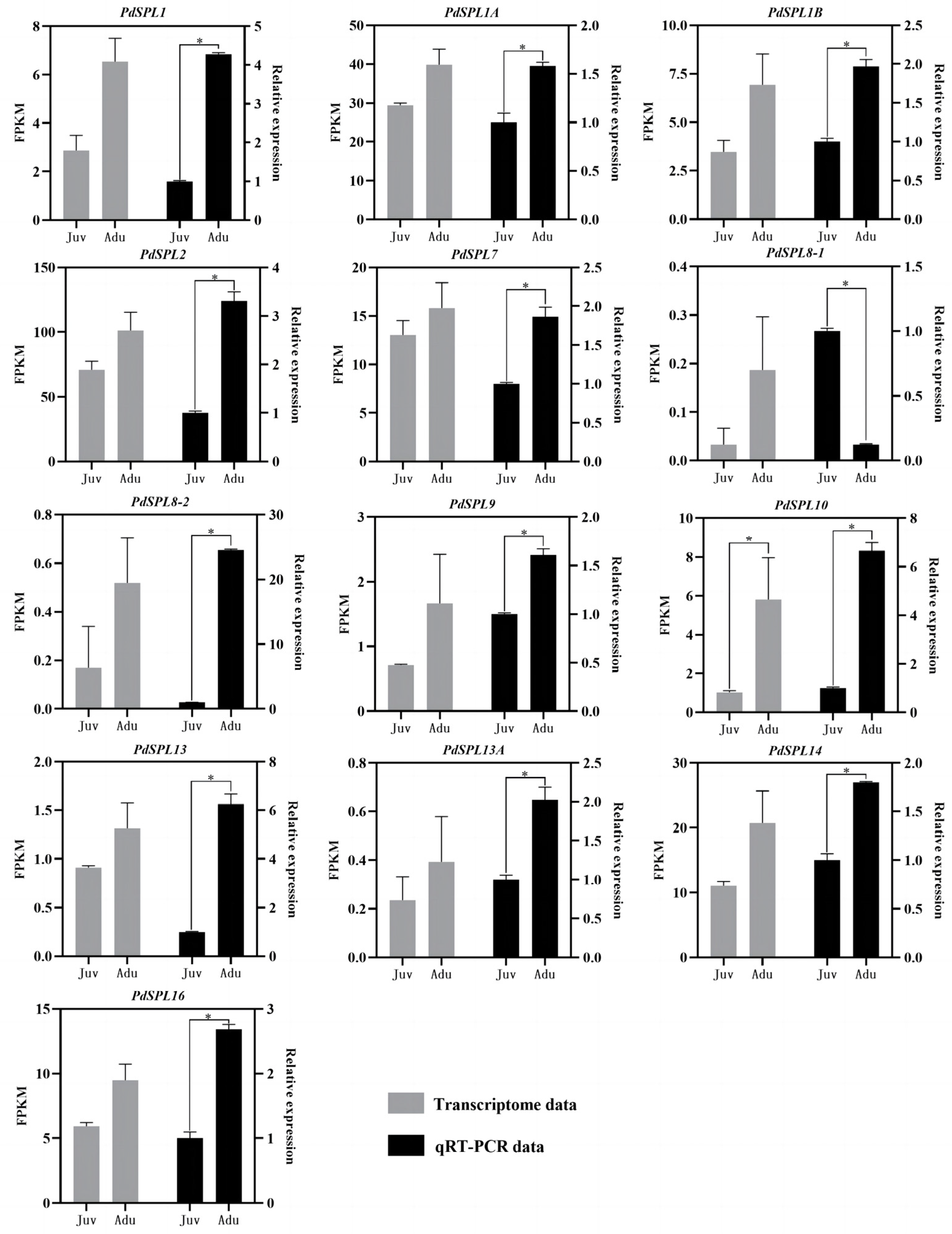
2.3. SPL Genes Expression Analysis
3. Discussion
3.1. Sugar Signaling (Possibly byT6P Pathway) May Mediate Juvenile-to-Adult Transition in P. delavayi
3.2. Hormone Signaling (Possibly in Interactive Way) May Mediate Juvenile-to-Adult Transition in P. delavayi
3.3. Circadian Rhythm Pathway May Mediate Juvenile-to-Adult Process in P. delavayi
3.4. Age Pathway May Mediate Juvenile-to-Adult Transition in P. delavayi
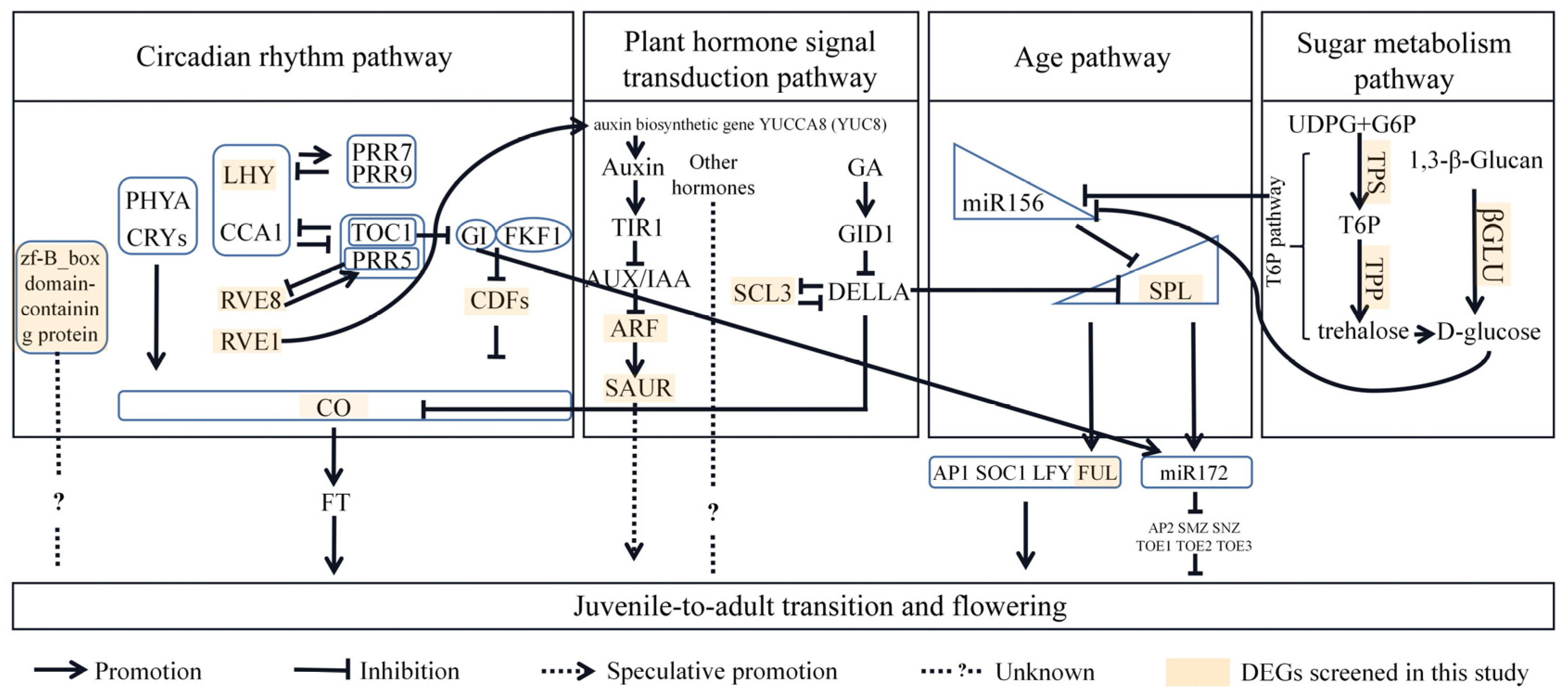
3.5. A Hypothetical Molecular Regulatory Network during Juvenile-to-Adult Transition in P. delavayi
4. Materials and Methods
4.1. Plant Materials
4.2. Methods
4.2.1. Quantitation of Carbohydrates and Plant Hormones
4.2.2. Discovery of DEGs and Pathways
4.2.3. Expression Analysis of SPL Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hong, D.Y. Peonies of the World Part III Phylogeny and Evolution; Kew Publishing Royal Botanic Gardens: Kew, UK, 2021; pp. 215–222. [Google Scholar]
- Yang, Y.; Sun, M.; Li, S.; Chen, Q.; Teixeira da Silva, J.A.; Wang, A.; Yu, X.; Wang, L. Germplasm resources and genetic breeding of Paeonia: A systematic review. Hortic Res. 2020, 7, 107. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Cheng, F.Y.; Wu, J.; He, C.Y. Isolation and functional analysis of Flowering Locus T in tree peonies (PsFT). J. Am. Soc. Hortic. Sci. 2015, 140, 265–271. [Google Scholar] [CrossRef]
- Zhu, F.; Wang, S.; Xue, J.; Li, D.; Ren, X.; Xue, Y.; Zhang, X. Morphological and physiological changes, and the functional analysis of PdSPL9 in the juvenile-to-adult phase transition of Paeonia delavayi. Plant Cell Tissue Organ Cult. 2018, 38, 325–337. [Google Scholar] [CrossRef]
- Song, M.; Wang, R.; Zhou, F.; Wang, R.; Zhang, S.; Li, D.; Song, J.; Yang, S.; Yang, Y. SPLs-mediated flowering regulation and hormone biosynthesis and signaling accompany juvenile-adult phase transition in Pyrus. Sci. Hortic. 2020, 48, 109584. [Google Scholar] [CrossRef]
- Vanstraelen, M.; Benkova, E.E. Hormonal interactions in the regulation of plant development. Annu. Rev. Cell Dev. Biol. 2012, 28, 463–487. [Google Scholar] [CrossRef]
- Lebon, G.; Wojnarowiez, G.; Holzapfel, B.; Fontaine, F.; Vaillant-Gaveau, N.; Clément, C. Sugars and flowering in the grapevine (Vitis vinifera L.). J. Exp. Bot. 2008, 59, 2565–2578. [Google Scholar] [CrossRef]
- Yu, S.; Cao, L.; Zhou, C.M.; Zhang, T.Q.; Lian, H.; Sun, Y.; Wu, J.; Huang, J.; Wang, G.; Wang, J.W. Sugar is an endogenous cue for juvenile-to-adult phase transition in plants. Elife 2013, 2, e00269. [Google Scholar] [CrossRef]
- Yu, S.; Wang, J.W. The crosstalk between microRNAs and gibberellin signaling in plants. Plant Cell Physiol. 2020, 61, 1880–1890. [Google Scholar] [CrossRef]
- Day, J.S.; Jameson, P.S.; Gould, K.S. Cytokinin changes during vegetative metamorphosis in Elaeocarpus hookerianus. Aust. J. Plant Physiol. 1995, 22, 67–73. [Google Scholar]
- Xing, L.; Zhang, D.; Li, Y.; Zhao, C.; Zhang, S.; Shen, Y.; An, N.; Han, M. Genome-wide identification of vegetative phase transition-associated microRNAs and target predictions using degradome sequencing in Malus hupehensis. BMC Genom. 2014, 15, 1125. [Google Scholar] [CrossRef]
- Tan, J.; Yi, X.; Luo, L.; Yu, C.; Wang, J.; Cheng, T.; Zhang, Q.; Pan, H. RNA-seq and sRNA-seq analysis in lateral buds and leaves of juvenile and adult roses. Sci. Hortic. 2021, 290, 110513. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Collins, L.J.; Biggs, P.J.; Voelckel, C.; Joly, S. An approach to transcriptome analysis of non-model organisms using short-read sequences. Genome Inform. 2008, 21, 3–14. [Google Scholar] [CrossRef] [PubMed]
- Manuela, D.; Xu, M. Juvenile leaves or adult leaves: Determinants for vegetative phase change in flowering plants. Int. J. Mol. Sci. 2020, 21, 9753. [Google Scholar] [CrossRef]
- Schwarz, S.; Grande, A.V.; Bujdoso, N.; Saedler, H.; Huijser, P. The microRNA regulated SBP-box genes SPL9 and SPL15 control shoot maturation in Arabidopsis. Plant Mol. Biol. 2008, 67, 183–195. [Google Scholar] [CrossRef]
- Gandikota, M.; Birkenbihl, R.P.; Höhmann, S.; Cardon, G.H.; Saedler, H.; Huijser, P. The miRNA156/157 recognition element in the 3′ UTR of the Arabidopsis SBP box gene SPL3 prevents early flowering by translational inhibition in seedlings. Plant J. 2007, 49, 683–693. [Google Scholar] [CrossRef]
- Kim, J.J.; Lee, J.H.; Kim, W.; Jung, H.S.; Huijser, P.; Ahn, J.H. The microRNA156-Squamosa Promoter Binding Protein-Like3 module regulates ambient temperature-responsive flowering via Flowering Locus T in Arabidopsis. Plant Physiol. 2012, 159, 461–478. [Google Scholar] [CrossRef]
- Cardon, G.; Höhmann, S.; Klei, J.; Nettesheim, K.; Saedler, H.; Huijser, P. Molecular characterisation of the Arabidopsis SBP-box genes. Gene 1999, 237, 91–104. [Google Scholar] [CrossRef]
- Hultquist, J.F.; Dorweiler, J.E. Feminized tassels of maize mop1 and ts1 mutants exhibit altered levels of miR156 and specific SBP-box genes. Planta 2008, 229, 99–113. [Google Scholar] [CrossRef]
- Salinas, M.; Xing, S.; Höhmann, S.; Berndtgen, R.; Huijser, P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta 2012, 235, 1171–1184. [Google Scholar] [CrossRef]
- Preston, J.C.; Hileman, L.C. Functional evolution in the plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene family. Front. Plant Sci. 2013, 4, 80. [Google Scholar] [CrossRef] [PubMed]
- Preston, J.C.; Hileman, L.C. SQUAMOSA-PROMOTER BINDING PROTEIN 1 initiates flowering in Antirrhinum majus through the activation of meristem identity genes. Plant J. 2010, 62, 704–712. [Google Scholar] [CrossRef]
- Yamaguchi, A.; Wu, M.F.; Yang, L.; Wu, G.; Poethig, R.S.; Wagner, D. The microRNA-regulated SBP-Box transcription factor SPL3 is a direct upstream activator of LEAFY, FRUITFULL, and APETALA1. Dev Cell. 2009, 17, 268–278. [Google Scholar] [CrossRef]
- Shikata, M.; Koyama, T.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis SBP-box genes SPL10, SPL11 and SPL2 control morphological change in association with shoot maturation in the reproductive phase. Plant Cell Physiol. 2009, 50, 2133–2145. [Google Scholar] [CrossRef]
- Yamasaki, H.; Hayashi, M.; Fukazawa, M.; Kobayashi, Y.; Shikanai, T. SQUAMOSA promoter binding protein-like7 is a central regulator for copper homeostasis in Arabidopsis. Plant Cell. 2009, 21, 347–361. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Poethig, R.S. Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 2006, 133, 3539–3547. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Song, C.; Guo, L.; Guo, D.; Xue, X.; Wang, H.; Hou, X. Full-Length transcriptome and transcriptome sequencing unveil potential mechanisms of brassinosteroid-induced flowering delay in tree peony. Horticulturae 2022, 8, 1136. [Google Scholar] [CrossRef]
- Chang, Y.; Hu, T.; Zhang, W.; Zhou, L.; Wang, Y.; Jiang, Z. Transcriptome profiling for floral development in reblooming cultivar ‘High Noon’ of Paeonia suffruticosa. Sci. Data. 2019, 6, 217. [Google Scholar] [CrossRef]
- Wang, S.; Gao, J.; Xue, J.; Xue, Y.; Li, D.; Guan, Y.; Zhang, X. De novo sequencing of tree peony (Paeonia suffruticosa) transcriptome to identify critical genes involved in flowering and floral organ development. BMC Genom. 2019, 20, 572. [Google Scholar] [CrossRef]
- Wang, S.; Ren, X.; Xue, J.; Xue, Y.; Cheng, X.; Hou, X.; Zhang, X. Molecular characterization and expression analysis of the SQUAMOSA PROMOTER BINDING PROTEIN-LIKE gene family in Paeonia suffruticosa. Plant Cell Rep. 2020, 39, 1425–1441. [Google Scholar] [CrossRef]
- Xu, M.; Hu, T.; Zhao, J.; Park, M.Y.; Earley, K.W.; Wu, G.; Yang, L.; Poethig, R.S. Developmental functions of miR156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet. 2016, 12, e1006263. [Google Scholar] [CrossRef] [PubMed]
- Teotia, S.; Tang, G. To bloom or not to bloom: Role of microRNAs in plant flowering. Mol. Plant. 2015, 8, 359–377. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Qi, S.; Zhang, D.; Li, Y.; An, N.; Zhao, C.; Zhao, J.; Shah, K.; Han, M.; Xing, L. Comparative RNA-sequencing-based transcriptome profiling of buds from profusely flowering ‘Qinguan’ and weakly flowering ‘Nagafu no. 2’ apple varieties reveals novel insights into the regulatory mechanisms underlying floral induction. BMC Plant Biol. 2018, 18, 370. [Google Scholar] [CrossRef]
- Moreira, R.A.; Cruz, M.D.C.M.D.; Fagundes, M.C.P.; Pantoja, L.D.A.; Santos, A.S.D. Leaf carbohydrates during flowering and early growth stages of fruitlets in ‘Ponkan’ mandarin tree. Pesqui Agropecu Bras. 2014, 49, 34–39. [Google Scholar] [CrossRef]
- Hu, J.; Liu, Y.; Tang, X.; Rao, H.; Pei, J. Transcriptome profiling of the flowering transition in saffron (Crocus sativus L.). Sci. Rep. 2020, 10, 9680. [Google Scholar] [CrossRef] [PubMed]
- Perrot, T.; Pauly, M.; Ramírez, V. Emerging roles of β-Glucanases in plant development and adaptative responses. Plants 2022, 11, 1119. [Google Scholar] [CrossRef]
- Duan, X.; Zhu, Z.; Yang, Y.; Duan, J.; Jia, Z.; Chen, F.; Sang, Z.; Ma, L. Salicylic acid regulates sugar metabolism that confers freezing tolerance in Magnolia wufengensis during natural cold acclimation. J. Plant Growth Regul. 2022, 41, 227–235. [Google Scholar] [CrossRef]
- Tsai, A.Y.; Gazzarrini, S. Trehalose-6-phosphate and SnRK1 kinases in plant development and signaling: The emerging picture. Front Plant Sci. 2014, 5, 119. [Google Scholar] [CrossRef]
- Ponnu, J.; Wahl, V.; Schmid, M. Trehalose-6-phosphate: Connecting plant metabolism and development. Front Plant Sci. 2011, 2, 70. [Google Scholar] [CrossRef]
- Ponnu, J.; Schlereth, A.; Zacharaki, V.; Działo, M.A.; Abel, C.; Feil, R.; Schmid, M.; Wahl, V. The trehalose 6-phosphate pathway impacts vegetative phase change in Arabidopsis thaliana. Plant J. 2020, 104, 768–780. [Google Scholar] [CrossRef]
- Végvári, G.; Edina, V. Plant hormones, plant growth regulators. Orv. Hetil. 2014, 155, 1011–1018. [Google Scholar] [CrossRef] [PubMed]
- Ivar, W.D.; Trueman, S.J.; Aloisio, X. Maturation and related aspects in clonal forestry—Part I: Concepts, regulation and consequences of phase change. New For. 2014, 45, 449–471. [Google Scholar] [CrossRef]
- Moncaleán, P.; Rodríguez, A.; Fernández, B. Plant growth regulators as putative physiological markers of developmental stage in Prunus persica. Plant Growth Regul. 2002, 36, 27–29. [Google Scholar] [CrossRef]
- Osadchuk, K.; Cheng, C.L.; Irish, E.E. Jasmonic acid levels decline in advance of the transition to the adult phase in maize. Plant Direct. 2019, 3, e00180. [Google Scholar] [CrossRef]
- Chin, T.Y.; Meyer, M.M.; Beevers, L. Abscisic-acid-stimulated rooting of stem cuttings. Planta 1969, 88, 192–196. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.L.; Ogawa, M.; Fleet, C.M.; Zentella, R.; Hu, J.; Heo, J.O.; Lim, J.; Kamiya, Y.; Yamaguchi, S.; Sun, T.P. Scarecrow-like 3 promotes gibberellin signaling by antagonizing master growth repressor DELLA in Arabidopsis. Proc. Natl. Acad. Sci. USA 2011, 108, 2160–2165. [Google Scholar] [CrossRef] [PubMed]
- Sedeer, E.S.; Raili, R.; Ykä, H. Crossing paths: Cytokinin signalling and crosstalk. Development 2013, 140, 1373–1383. [Google Scholar] [CrossRef]
- Ghorbel, M.; Brini, F.; Sharma, A.; Landi, M. Role of jasmonic acid in plants: The molecular point of view. Plant Cell Rep. 2021, 40, 1471–1494. [Google Scholar] [CrossRef]
- Ohri, P.; Bhardwaj, R.; Bali, S.; Kaur, R.; Jasrotia, S.; Khajuria, A.; DParihar, R. The common molecular players in plant hormone crosstalk and signaling. Curr. Protein Pept. Sci. 2015, 16, 369–388. [Google Scholar] [CrossRef]
- Yao, W.; Li, C.; Lin, S.; Ren, L.; Wan, Y.; Zhang, L.; Ding, Y. Morphological characteristics and transcriptome comparisons of the shoot buds from flowering and non-flowering Pleioblastus pygmaeus. Forests 2020, 11, 1229. [Google Scholar] [CrossRef]
- Jones, M.A.; Hu, W.; Litthauer, S.; Lagarias, J.C.; Harmer, S.L. A constitutively active allele of phytochrome B maintains circadian robustness in the absence of light. Plant Physiol. 2015, 169, 814–825. [Google Scholar] [CrossRef]
- Harmer, S.L. The circadian system in higher plants. Annu. Rev. Plant Biol. 2009, 60, 357–377. [Google Scholar] [CrossRef]
- Johansson, M.; Dorothee, S. Time to flower: Interplay between photoperiod and the circadian clock. J. Exp. Bot. 2015, 66, 719–730. [Google Scholar] [CrossRef]
- Rawat, R.; Schwartz, J.; Jones, M.A.; Sairanen, I.; Cheng, Y.; Andersson, C.R.; Zhao, Y.; Ljung, K.; Harmer, S.L. REVEILLE1, a Myb-like transcription factor, integrates the circadian clock and auxin pathways. Proc. Natl. Acad. Sci. USA 2009, 106, 16883–16888. [Google Scholar] [CrossRef] [PubMed]
- Peter, H.; Markus, S. The control of developmental phase transitions in plants. Development 2011, 138, 4117–4129. [Google Scholar] [CrossRef]
- Wang, J.W.; Park, M.Y.; Wang, L.J.; Koo, Y.; Chen, X.Y.; Weigel, D.; Poethig, R.S. MiRNA control of vegetative phase change in trees. PLoS Genet. 2011, 7, e1002012. [Google Scholar] [CrossRef]
- Ma, Z.; Li, W.; Wang, H.; Yu, D. WRKY transcription factors WRKY12 and WRKY13 interact with SPL10 to modulate age-mediated flowering. J. Integr. Plant Biol. 2020, 62, 1659–1673. [Google Scholar] [CrossRef]
- Yao, T.; Park, B.S.; Mao, H.Z.; Seo, J.S.; Ohama, N.; Li, Y.; Yu, N.; Mustafa, N.F.B.; Huang, C.H.; Chua, N.H. Regulation of flowering time by SPL10/MED25 module in Arabidopsis. New Phytol. 2019, 224, 493–504. [Google Scholar] [CrossRef] [PubMed]
- Xia, M.; Zhang, W.; Chang, Y.; Ma, Y.; Deng, Y.; Fan, K.; Zhang, X.; Jiang, Z.; Hu, T. Preliminary investigation on the functional validation and interactions of PoWOX genes in Peony (Paeonia ostii). Horticulturae 2022, 8, 266. [Google Scholar] [CrossRef]
- Wen, S.S.; Chen, L.; Tian, R.N. Micropropagation of tree peony (Paeonia sect. Moutan): A review. Plant Cell Tissue Organ Cult. 2020, 141, 1–14. [Google Scholar] [CrossRef]
- Flachowsky, H.; Le Roux, P.M.; Peil, A.; Patocchi, A.; Richter, K.; Hanke, M.V. Application of a high-speed breeding technology to apple (Malus × domestica) based on transgenic early flowering plants and marker-assisted selection. New Phytol. 2011, 192, 364–377. [Google Scholar] [CrossRef] [PubMed]
- Klocko, A.L.; Ma, C.; Robertson, S.; Esfandiari, E.; Nilsson, O.; Strauss, S.H. FT overexpression induces precocious flowering and normal reproductive development in Eucalyptus. Plant Biotechnol. J. 2016, 14, 808–819. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.J.; Shi, A.G.; Dong, X.C. Plant Physiology Laboratory Instruction, 1st ed.; China Agricultural Science and Technology Press: Beijing, China, 2002; pp. 87–88. [Google Scholar]
- Li, H.S. Principles and Techniques of Plant Physiological Biochemical Experiment, 1st ed.; Higher Education Press: Beijing, China, 2000; pp. 195–197. [Google Scholar]
- Šimura, J.; Antoniadi, I.; Široká, J.; Tarkowská, D.; Strnad, M.; Ljung, K.; Novák, O. Plant hormonomics: Multiple phytohormone profiling by targeted metabolomics. Plant Physiol. 2018, 177, 476–489. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]

| ID | Gene Annotation | Gene Name | J-FPKM | A-FPKM | Fold Change |
|---|---|---|---|---|---|
| c74517.graph_c0 | AP1/FUL-like protein | PdFUL | 5.55 | 32.18 | 5.80 |
| c83958.graph_c0 | CONSTANS-LIKE 2 | PdCOL2 | 16.15 | 52.53 | 3.25 |
| c85008.graph_c0 | FLOWERING-promoting factor 1 | PdFPF1 | 2.73 | 44.99 | 16.48 |
| c89148.graph_c0 | protein LHY | PdLHY | 1.97 | 38.62 | 19.60 |
| c71307.graph_c0 | promoter-binding protein SPL10 | PdSPL10 | 1.03 | 5.81 | 5.64 |
| KEGG Pathway | ID | Gene Annotation | Up/Down-Regulated | Fold Change |
|---|---|---|---|---|
| Circadian rhythm-plant | c70450.graph_c0 | REVEILLE 1 | up | 4.12 |
| c70837.graph_c0 | B-box zinc finger protein 19 | up | 4.49 | |
| c83340.graph_c1 | Cyclic dof factor 3 | up | 5.07 | |
| c83958.graph_c0 | CONSTANS-LIKE 2 | up | 3.25 | |
| c84340.graph_c1 | REVEILLE 8 | up | 3.41 | |
| c89148.graph_c0 | LHY | up | 19.57 | |
| Plant hormone signal transduction | c41494.graph_c0 | Scarecrow-like protein 3 | up | 3.08 |
| c53132.graph_c0 | Auxin-responsive protein SAUR32-like | up | 2.79 | |
| c86094.graph_c1 | Auxin response factor 7 | up | 2.66 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhai, X.; Feng, Y.; Zhang, X.; Guo, X. Comparative Analysis Based on Physiological and Transcriptomic Data between Juvenile and Adult Tree Peony (Paeonia delavayi). Int. J. Mol. Sci. 2023, 24, 10906. https://doi.org/10.3390/ijms241310906
Zhai X, Feng Y, Zhang X, Guo X. Comparative Analysis Based on Physiological and Transcriptomic Data between Juvenile and Adult Tree Peony (Paeonia delavayi). International Journal of Molecular Sciences. 2023; 24(13):10906. https://doi.org/10.3390/ijms241310906
Chicago/Turabian StyleZhai, Xiaoli, Yan Feng, Xiuxin Zhang, and Xianfeng Guo. 2023. "Comparative Analysis Based on Physiological and Transcriptomic Data between Juvenile and Adult Tree Peony (Paeonia delavayi)" International Journal of Molecular Sciences 24, no. 13: 10906. https://doi.org/10.3390/ijms241310906
APA StyleZhai, X., Feng, Y., Zhang, X., & Guo, X. (2023). Comparative Analysis Based on Physiological and Transcriptomic Data between Juvenile and Adult Tree Peony (Paeonia delavayi). International Journal of Molecular Sciences, 24(13), 10906. https://doi.org/10.3390/ijms241310906





