Protonation-State Dependence of Hydration and Interactions in the Two Proton-Conducting Channels of Cytochrome c Oxidase
Abstract
1. Introduction
2. Results
2.1. Conformation Analysis
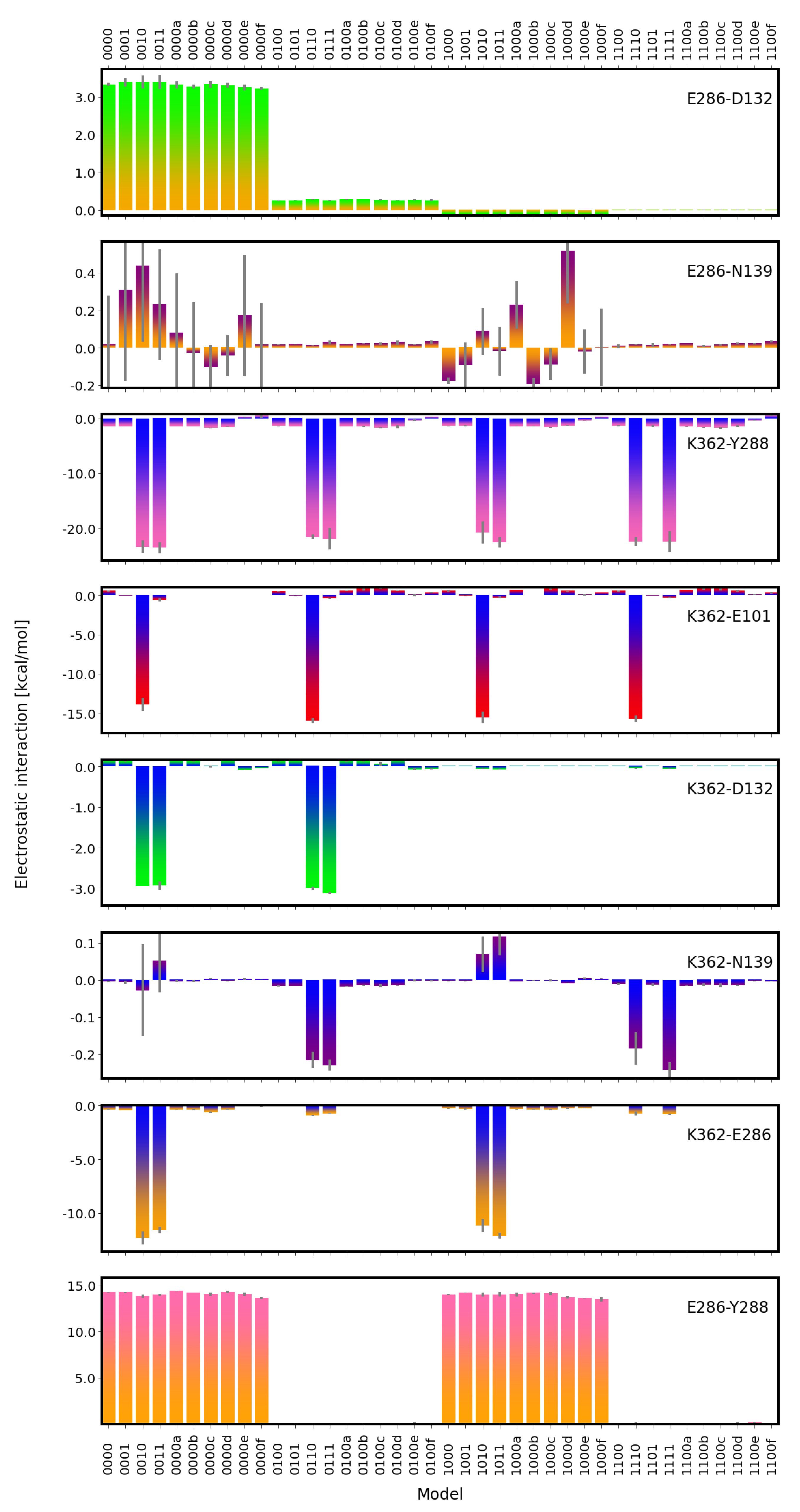
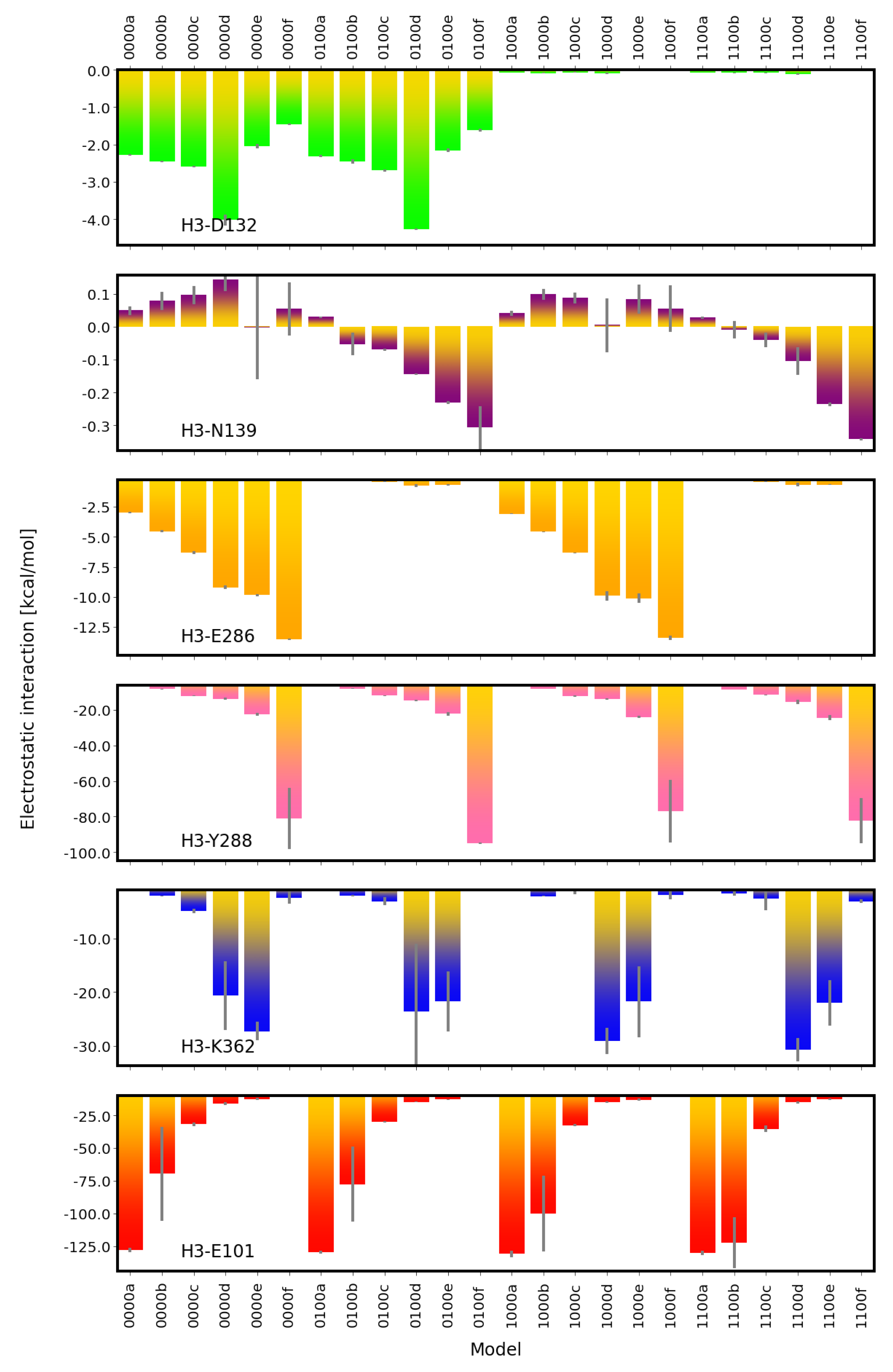
2.2. Electrostatic Interactions
2.3. Channel Hydration
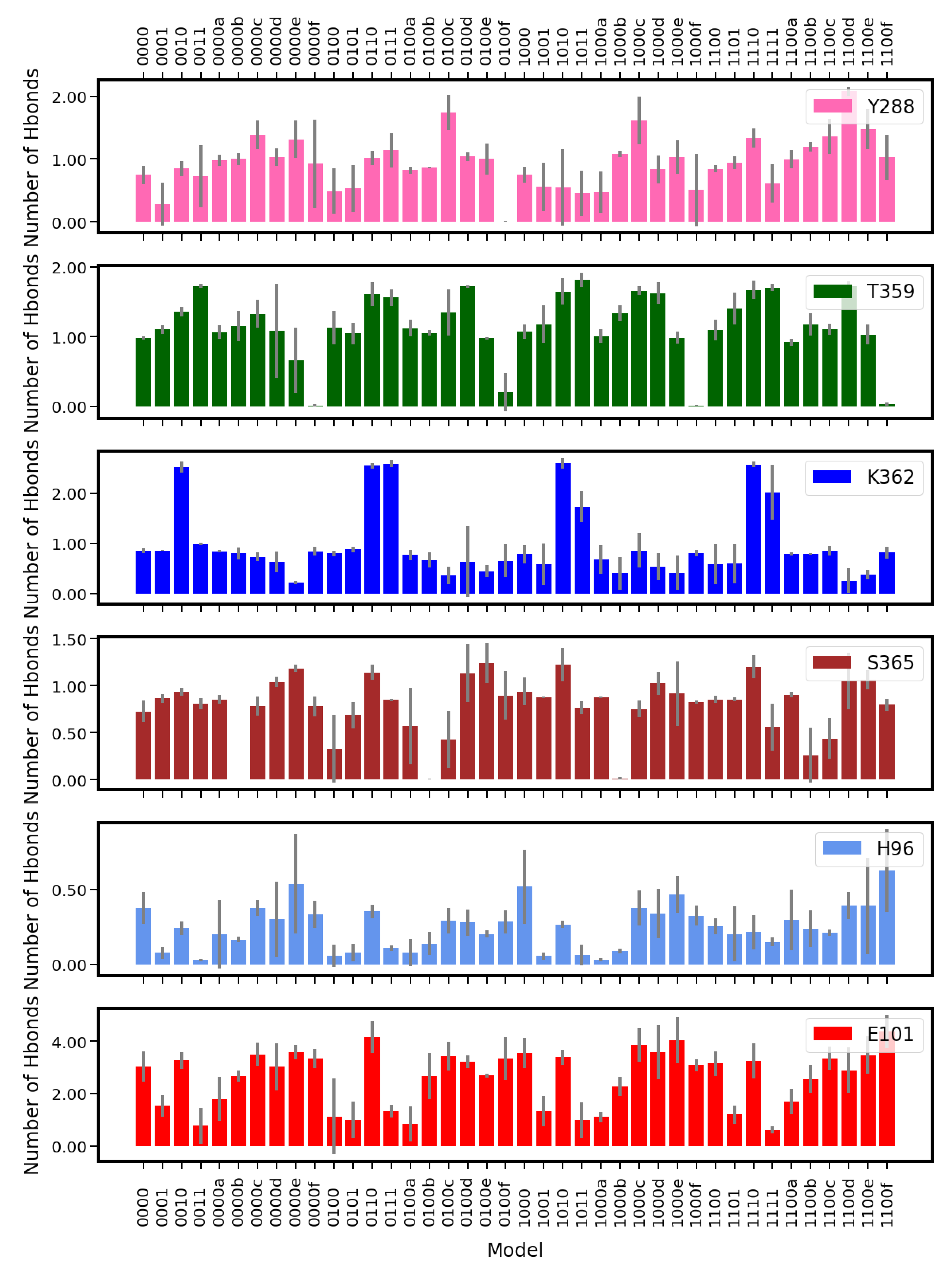
2.4. Hydrogen Bonds
3. Discussion
4. Materials and Methods
4.1. Molecular Dynamics Simulations
Model Setup
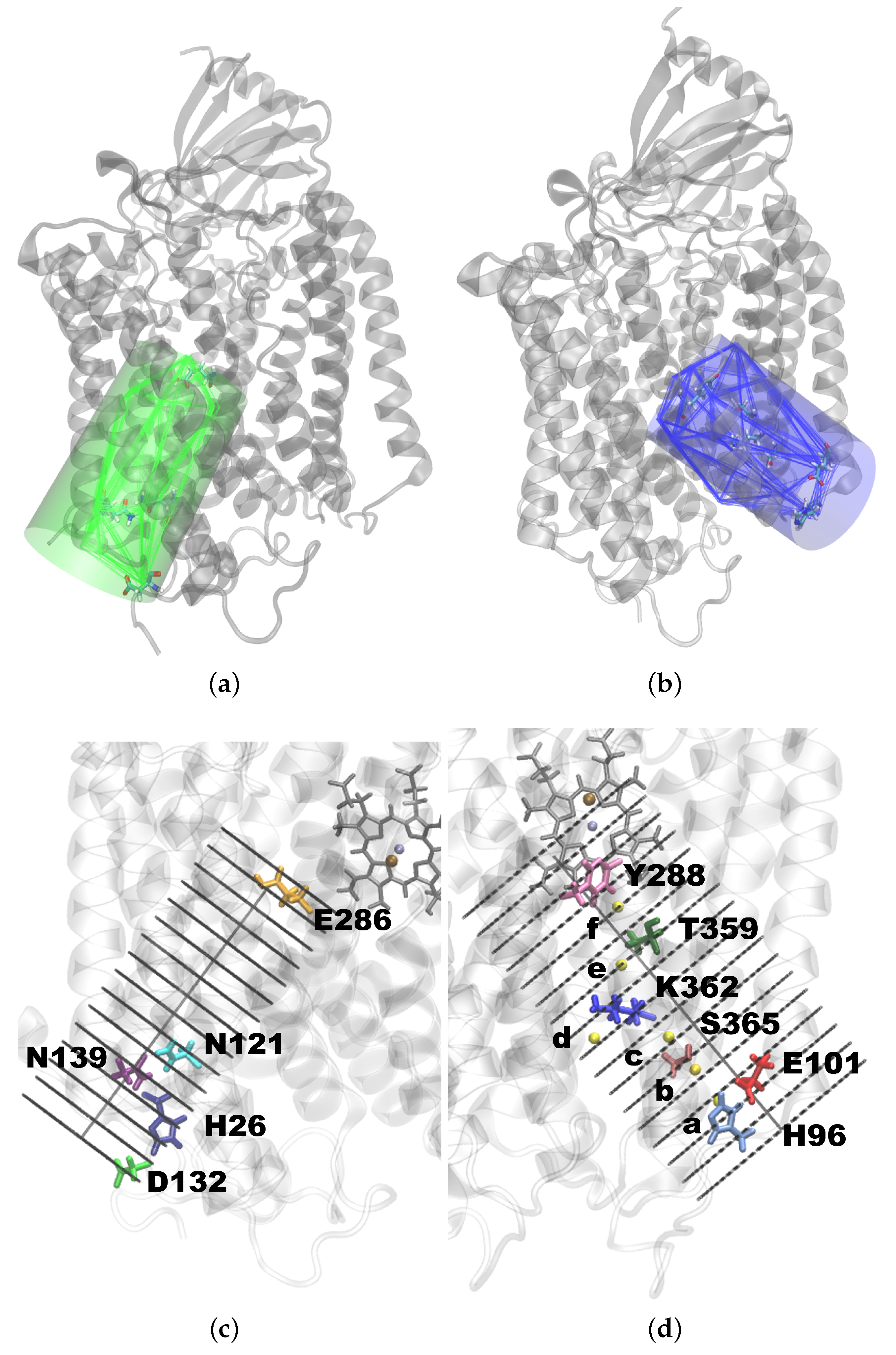
- (a)
- Between H96 and E101 (23.4).
- (b)
- Between E101 and S365 (19.1).
- (c)
- Between S365 and K362 (15.5).
- (d)
- Just below K362 (12.8).
- (e)
- Between K362 and T359 (10.4).
- (f)
- Between T359 and Y288 (7.3).
| Model | Protonated D-Channel Residue(s) | Protonated K-Channel Residue(s) |
|---|---|---|
| 0000 | - | - |
| 0001 | - | E101 |
| 0010 | - | K362 |
| 0011 | - | K362 and E101 |
| 0000a | - | in position a |
| 0000b | - | in position b |
| 0000c | - | in position c |
| 0000d | - | in position d |
| 0000e | - | in position e |
| 0000f | - | H in position f |
| 0100 | E286 | - |
| 0101 | E286 | E101 |
| 0110 | E286 | K362 |
| 0111 | E286 | K362 and E101 |
| 0100a | E286 | in position a |
| 0100b | E286 | in position b |
| 0100c | E286 | in position c |
| 0100d | E286 | in position d |
| 0100e | E286 | in position e |
| 0100f | E286 | in position f |
| 1000 | D132 | - |
| 1001 | D132 | E101 |
| 1010 | D132 | K362 |
| 1011 | D132 | K362 and E101 |
| 1000a | D132 | in position a |
| 1000b | D132 | in position b |
| 1000c | D132 | in position c |
| 1000d | D132 | in position d |
| 1000e | D132 | in position e |
| 1000f | D132 | in position f |
| 1100 | D132 and E286 | - |
| 1101 | D132 and E286 | E101 |
| 1110 | D132 and E286 | K362 |
| 1111 | D132 and E286 | K362 and E101 |
| 1100a | D132 and E286 | in position a |
| 1100b | D132 and E286 | in position b |
| 1100c | D132 and E286 | in position c |
| 1100d | D132 and E286 | in position d |
| 1100e | D132 and E286 | in position e |
| 1100f | D132 and E286 | in position f |
4.2. Molecular Dynamics Simulations
4.3. Analysis
4.3.1. Conformational Analysis
4.3.2. Electrostatic Interactions
4.3.3. Channel Hydration
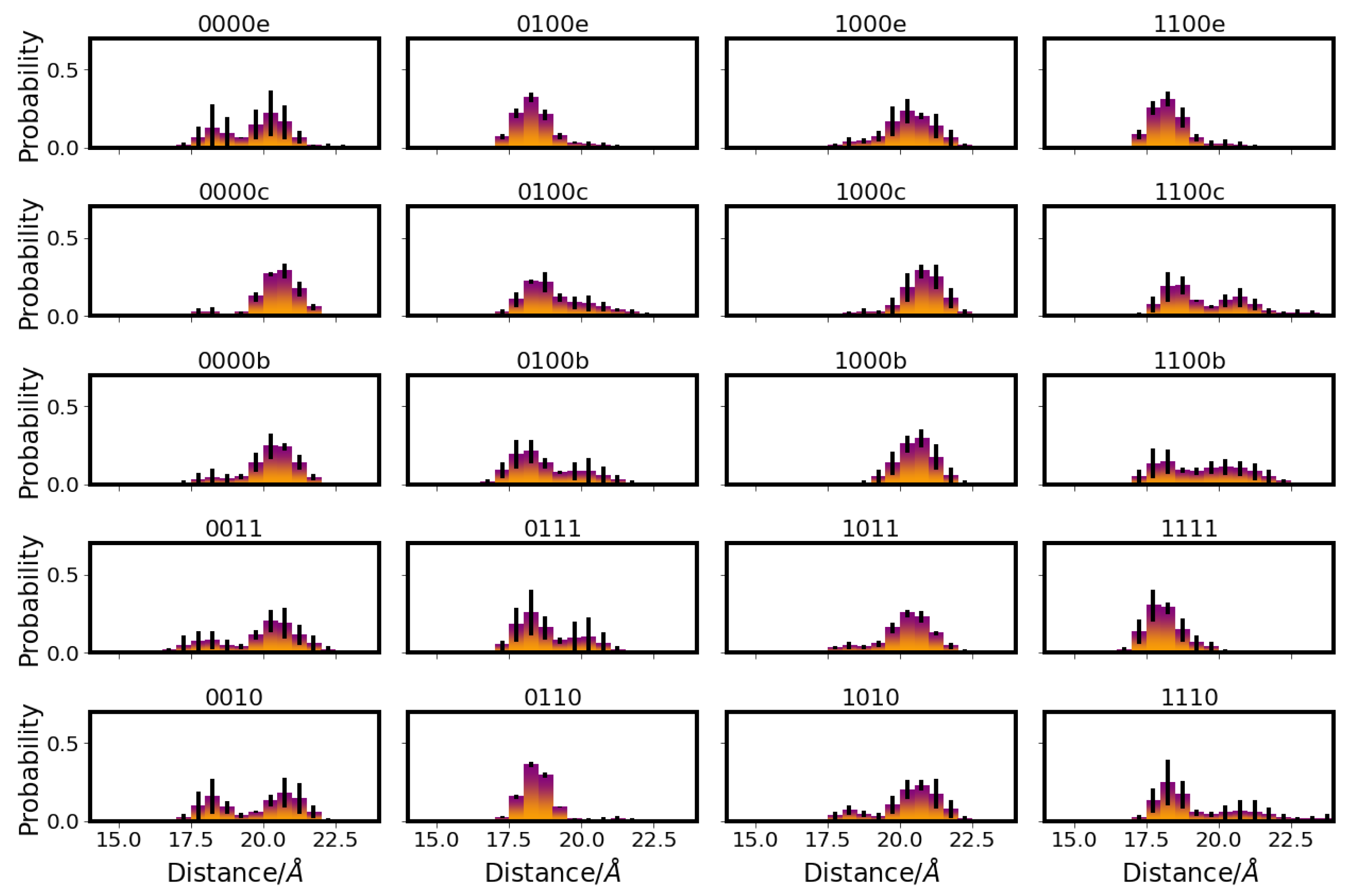
4.3.4. Hydrogen-Bond Probabilities and Hydrogen-Bond Lifetimes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CcO | Cytochrome c Oxidase |
| MD | Molecular dynamics |
| BNC | Bi-nuclear centre |
References
- Babcock, G.T.; Wikström, M. Oxygen activation and the conservation of energy in cell respiration. Nature 1992, 356, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Konstantinov, A.A.; Siletsky, S.; Mitchell, D.; Kaulen, A.; Gennis, R.B. The roles of the two proton input channels in cytochrome c oxidase from Rhodobacter sphaeroides probed by the effects of site-directed mutations on time-resolved electrogenic intraprotein proton transfer. Proc. Nat. Acad. Sci. USA 1997, 94, 9085–9090. [Google Scholar] [CrossRef]
- Ädelroth, P.; Gennis, R.B.; Brzezinski, P. Role of the pathway through K (I-362) in proton transfer in cytochrome c oxidase from R. sphaeroides. Biochemistry 1998, 37, 2470–2476. [Google Scholar] [CrossRef] [PubMed]
- Brzezinski, P.; Adelroth, P. Proton-controlled electron transfer in cytochrome c oxidase: Functional role of the pathways through Glu 286 and Lys 362. Acta Physiol. Scand. Suppl. 1998, 643, 7–16. [Google Scholar]
- Vygodina, T.; Pecoraro, C.; Mitchell, D.; Gennis, R.; Konstantinov, A. Mechanism of inhibition of electron transfer by amino acid replacement K362M in a proton channel of Rhodobacter sphaeroides cytochrome c oxidase. Biochemistry 1998, 37, 3053–3061. [Google Scholar] [CrossRef] [PubMed]
- Gorriz, R.F.; Imhof, P. Interplay of Hydration and Protonation Dynamics in the K-channel of Cytochrome c Oxidase. Biomolecules 2022, 12, 1615. [Google Scholar] [CrossRef] [PubMed]
- Ghane, T.; Gorriz, R.F.; Wrzalek, S.; Volkenandt, S.; Delatieh, F.; Reidelbach, M.; Imhof, P. Hydrogen-Bonded Network and Water Dynamics in the D-channel of Cytochrome c Oxidase. J. Membr. Biol. 2018, 251, 299–314. [Google Scholar] [CrossRef]
- Qin, L.; Liu, J.; Mills, D.A.; Proshlyakov, D.A.; Hiser, C.; Ferguson-Miller, S. Redox-dependent conformational changes in cytochrome c oxidase suggest a gating mechanism for proton uptake. Biochemistry 2009, 48, 5121–5130. [Google Scholar] [CrossRef]
- Wikström, M.; Gennis, R.B.; Rich, P.R. Structures of the intermediates in the catalytic cycle of mitochondrial cytochrome c oxidase. Biochim. Biophys. Acta BBA Bioenerg. 2023, 1864, 148933. [Google Scholar] [CrossRef]
- Belevich, I.; Bloch, D.A.; Belevich, N.; Wikström, M.; Verkhovsky, M.I. Exploring the proton pump mechanism of cytochrome c oxidase in real time. Proc. Nat. Acad. Sci. USA 2007, 104, 2685–2690. [Google Scholar] [CrossRef]
- Wikstrom, M.; Krab, K.; Sharma, V. Oxygen Activation and Energy Conservation by Cytochrome c Oxidase. Chem. Rev. 2018, 118, 2469–2490. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, K.; Gennis, R.B. Blocking the K-pathway still allows rapid one-electron reduction of the binuclear center during the anaerobic reduction of the aa3-type cytochrome c oxidase from Rhodobacter sphaeroides. Biochim. Biophys. Acta BBA Bioenerg. 2010, 1797, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Magalhāes, P.R.; Oliveira, A.S.F.; Campos, S.R.R.; Soares, C.M.; Baptista, A.M. Effect of a pH Gradient on the Protonation States of Cytochrome c Oxidase: A Continuum Electrostatics Study. J. Chem. Inf. Model. 2017, 57, 256–266. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.; Hiser, C.; Mulichak, A.; Garavito, R.M.; Ferguson-Miller, S. Identification of conserved lipid/detergent-binding sites in a high-resolution structure of the membrane protein cytochrome c oxidase. Proc. Nat. Acad. Sci. USA 2006, 103, 16117–16122. [Google Scholar] [CrossRef] [PubMed]
- Saura, P.; Riepl, D.; Frey, D.M.; Wikström, M.; Kaila, V.R.I. Electric fields control water-gated proton transfer in cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 2022, 119, e2207761119. [Google Scholar] [CrossRef]
- Woelke, A.L.; Galstyan, G.; Knapp, E.W. Lysine 362 in cytochrome c oxidase regulates opening of the K-channel via changes in pKA and conformation. Biochim. Biophys. Acta BBA Bioenerg. 2014, 1837, 1998–2003. [Google Scholar] [CrossRef]
- Supekar, S.; Kaila, V.R.I. Dewetting transitions coupled to K-channel activation in cytochrome c oxidase. Chem. Sci. 2018, 32, 6703–6710. [Google Scholar] [CrossRef]
- Stegmaier, V.; Gorriz, R.F.; Imhof, P. Protonation Dynamics in the K-Channel of Cytochrome c Oxidase Estimated from Molecular Dynamics Simulations. Processes 2021, 9, 265. [Google Scholar] [CrossRef]
- Wikström, M.; Verkhovsky, M.I.; Hummer, G. Water-gated mechanism of proton translocation by cytochrome c oxidase. Biochim. Biophys. Acta BBA Bioenerg. 2003, 1604, 61–65. [Google Scholar] [CrossRef]
- Sharma, V.; Enkavi, G.; Vattulainen, I.; Róg, T.; Wikström, M. Proton-coupled electron transfer and the role of water molecules in proton pumping by cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 2015, 112, 2040–2045. [Google Scholar] [CrossRef]
- Wikström, M.; Sharma, V. Proton pumping by cytochrome c oxidase—A 40 year anniversary. Biochim. Biophys. Acta BBA Bioenerg. 2018, 1859, 692–698. [Google Scholar] [CrossRef]
- Liang, R.; Swansson, J.M.J.; Peng, Y.; Wikström, M.; Voth, G.A. Multiscale Simulations reveal key-features of the proton-pumping mechanism in cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 2016, 113, 7420–7425. [Google Scholar] [CrossRef]
- Liang, R.; Swansson, J.M.J.; Wikström, M.; Voth, G.A. Understaning the essential proton-pumping kinetic gates and decoupling mutations in cytochrome c oxidase. Proc. Natl. Acad. Sci. USA 2017, 114, 5924–5929. [Google Scholar] [CrossRef]
- Helabad, M.B.; Ghane, T.; Reidelbach, M.; Woelke, A.-L.; Knapp, E.-W.; Imhof, P. Protonation-State-Dependent Communication in Cytochrome c Oxidase. Biophys. J. 2017, 113, 817–828. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- MacKerell, A.D., Jr.; Bashford, D.; Bellott, M.; Dunbrack, R.L., Jr.; Evanseck, J.D.; Field, M.J.; Fischer, S.; Gao, J.; Guo, H.; Ha, S.; et al. All-Atom Empirical Potential for Molecular Modeling and Dynamics Studies of Proteins. J. Phys. Chem. B. 1998, 102, 3586–3616. [Google Scholar] [CrossRef]
- Sagnella, D.; Voth, G. Structure and dynamics of hydronium in the ion channel gramicidin A. Biophys. J. 1996, 70, 2043–2051. [Google Scholar] [CrossRef]
- Klauda, J.B.; Venable, R.M.; Freites, J.A.; O’Connor, J.W.; Tobias, D.J.; Mondragon-Ramirez, C.; Vorobyov, I.; MacKerell, A.D., Jr.; Pastor, R.W. Update of the CHARMM all-atom additive force field for lipids: Validation on six lipid types. J. Phys. Chem. B 2010, 114, 7830–7843. [Google Scholar] [CrossRef] [PubMed]
- Darden, T.; York, D.; Pedersen, L.G. Particle mesh Ewald: An Nlog(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Ryckaert, J.P.; Ciccotti, G.; Berendsen, H.J.C. Numerical integration of the cartesian equations of motion of a system with constraints: Molecular dynamics of n-alkanes. J. Comp. Phys. 1977, 23, 327–341. [Google Scholar] [CrossRef]
- Phillips, J.C.; Braun, R.; Wang, W.; Gumbart, J.; Tajkhorshid, E.; Villa, E.; Chipot, C.; Skeel, R.D.; Kale, L.; Schulten, K. Scalable molecular dynamics with NAMD. J. Comput. Chem. 2005, 26, 1781–1802. [Google Scholar] [CrossRef] [PubMed]
- Bagga, J.; Heinz, A. JGraph—A Java Based System for Drawing Graphs and Running Graph Algorithms. In Graph Drawing; Lecture Notes in Computer Science; Mutzel, P., Jünger, M., Leipert, S., Eds.; Springer: Berlin/Heidelberg, Germany, 2002; Volume 2265, pp. 459–460. [Google Scholar]
- Gowers, R.J.; Linke, M.; Barnoud, J.; Reddy, T.J.E.; Melo, M.N.; Seyler, S.L.; Domanski, J.; Dotson, D.L.; Buchoux, S.; Kenney, I.M.; et al. MDAnalysis: A Python Package for the Rapid Analysis of Molecular Dynamics Simulations; Technical Report; Los Alamos National Lab. (LANL): Los Alamos, NM, USA, 2019. [Google Scholar]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD—Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Hagberg, A.; Swart, P.; Chult, D.S. Exploring Network Structure, Dynamics, and Function Using NetworkX; Technical Report; Los Alamos National Lab. (LANL): Los Alamos, NM, USA, 2008. [Google Scholar]
- Dijkstra, E. A note on two problems in connexion with graphs. Num. Math. 1959, 1, 269–271. [Google Scholar] [CrossRef]
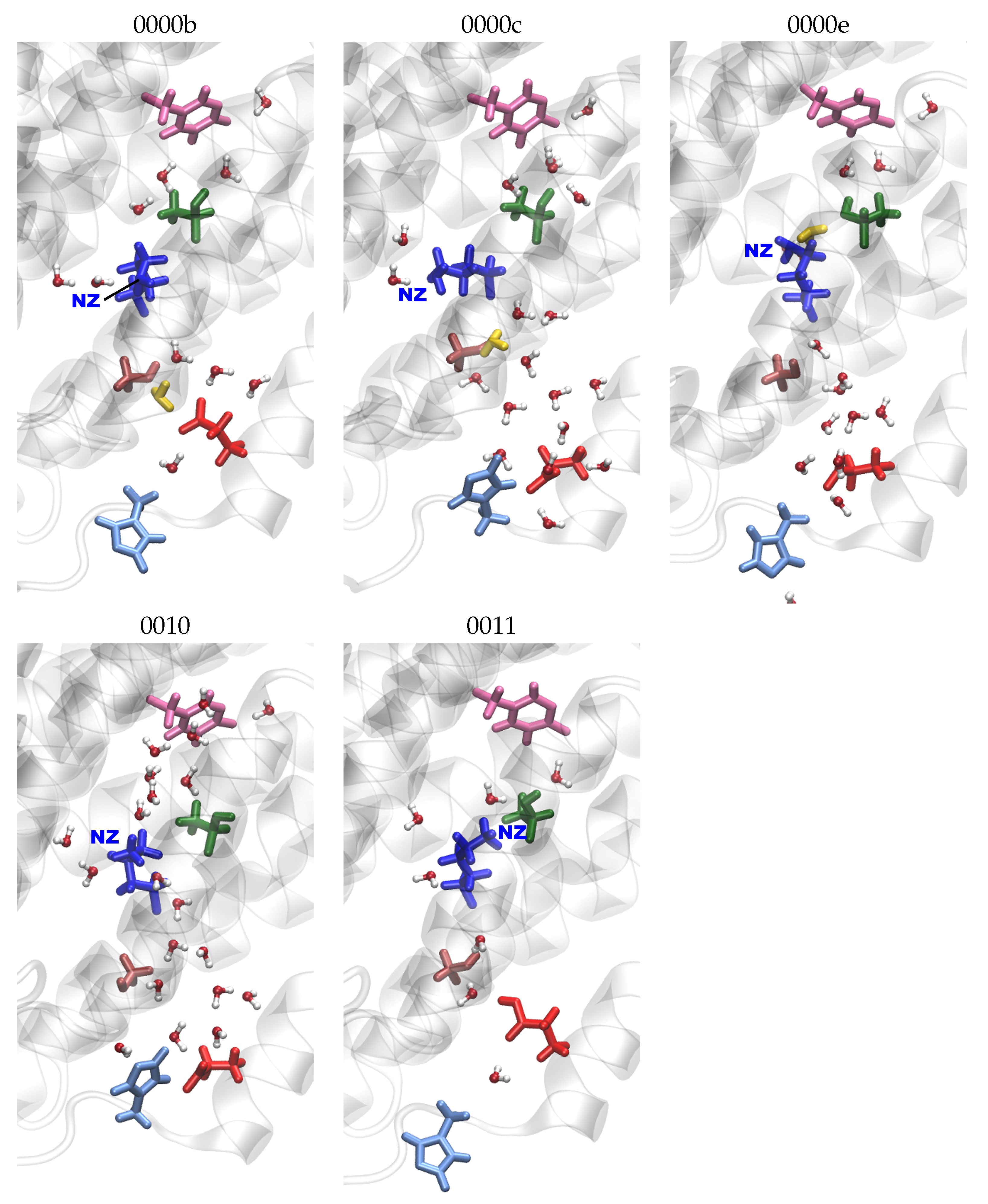


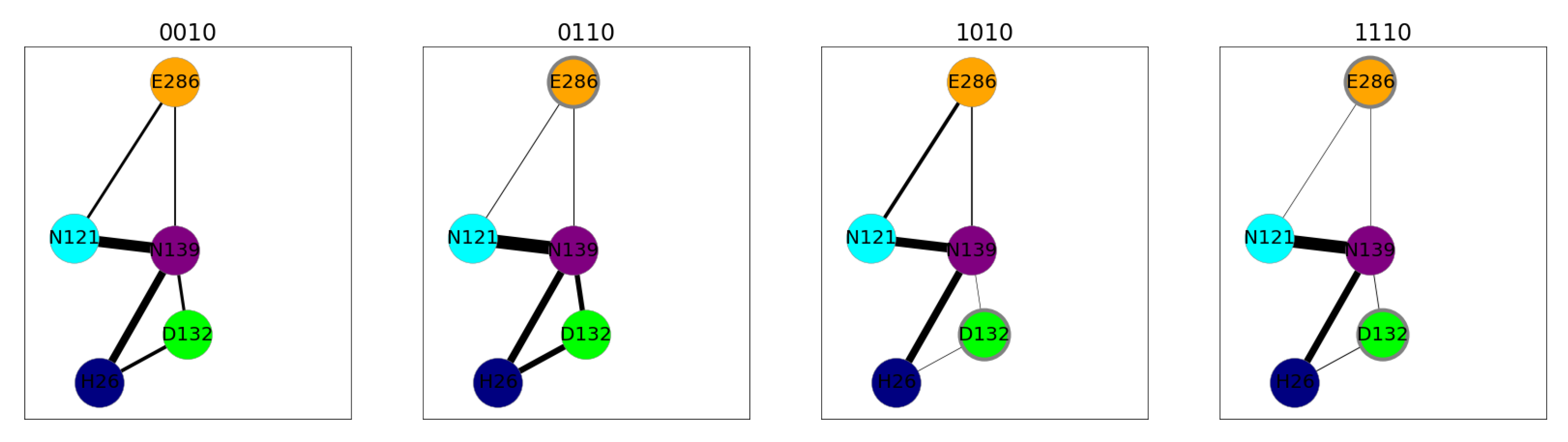
 ), N121 (cyan arrow—left
), N121 (cyan arrow—left  ), D132 (lime diamond
), D132 (lime diamond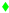  ), N139 (purple arrow—right
), N139 (purple arrow—right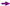  ), and E286 (orange star
), and E286 (orange star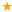  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
 ), N121 (cyan arrow—left
), N121 (cyan arrow—left  ), D132 (lime diamond
), D132 (lime diamond  ), N139 (purple arrow—right
), N139 (purple arrow—right  ), and E286 (orange star
), and E286 (orange star  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
 ), N121 (cyan arrow—left
), N121 (cyan arrow—left  ), D132 (lime diamond
), D132 (lime diamond  ), N139 (purple arrow—right
), N139 (purple arrow—right  ), and E286 (orange star
), and E286 (orange star  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
 ), N121 (cyan arrow—left
), N121 (cyan arrow—left  ), D132 (lime diamond
), D132 (lime diamond  ), N139 (purple arrow—right
), N139 (purple arrow—right  ), and E286 (orange star
), and E286 (orange star  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
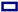
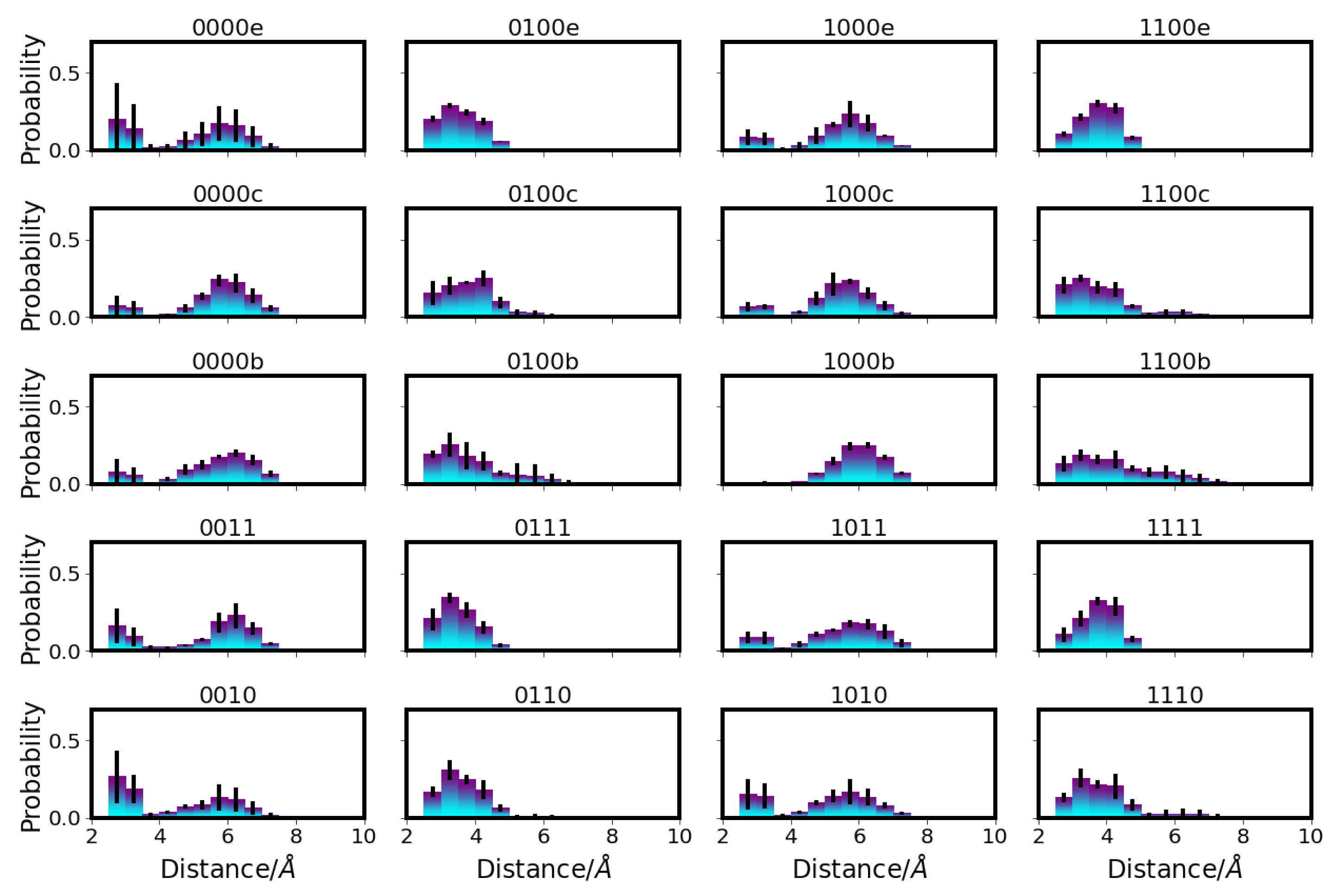
 ), E101 (red square
), E101 (red square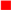  ), S365 (brown triangle—right
), S365 (brown triangle—right  ), K362 (blue cross
), K362 (blue cross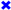 ), T359 (green triangle—left
), T359 (green triangle—left  ), and Y288 (magenta triangle—down
), and Y288 (magenta triangle—down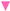  ), and the position of the ion (yellow circle
), and the position of the ion (yellow circle ), if present. Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
), if present. Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
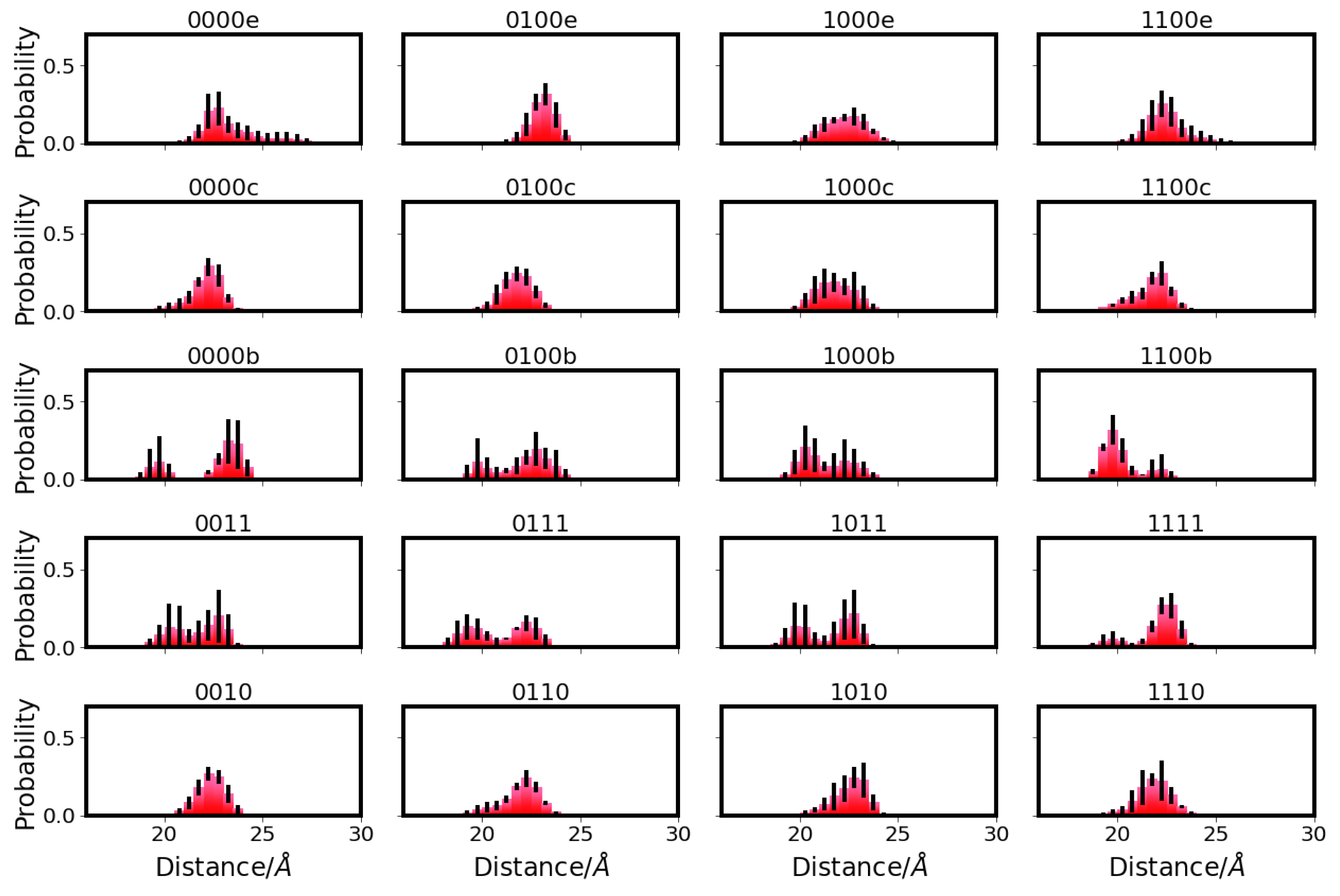
 ), E101 (red square
), E101 (red square  ), S365 (brown triangle—right
), S365 (brown triangle—right  ), K362 (blue cross
), K362 (blue cross ), T359 (green triangle—left
), T359 (green triangle—left  ), and Y288 (magenta triangle—down
), and Y288 (magenta triangle—down  ), and the position of the ion (yellow circle
), and the position of the ion (yellow circle ), if present. Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
), if present. Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.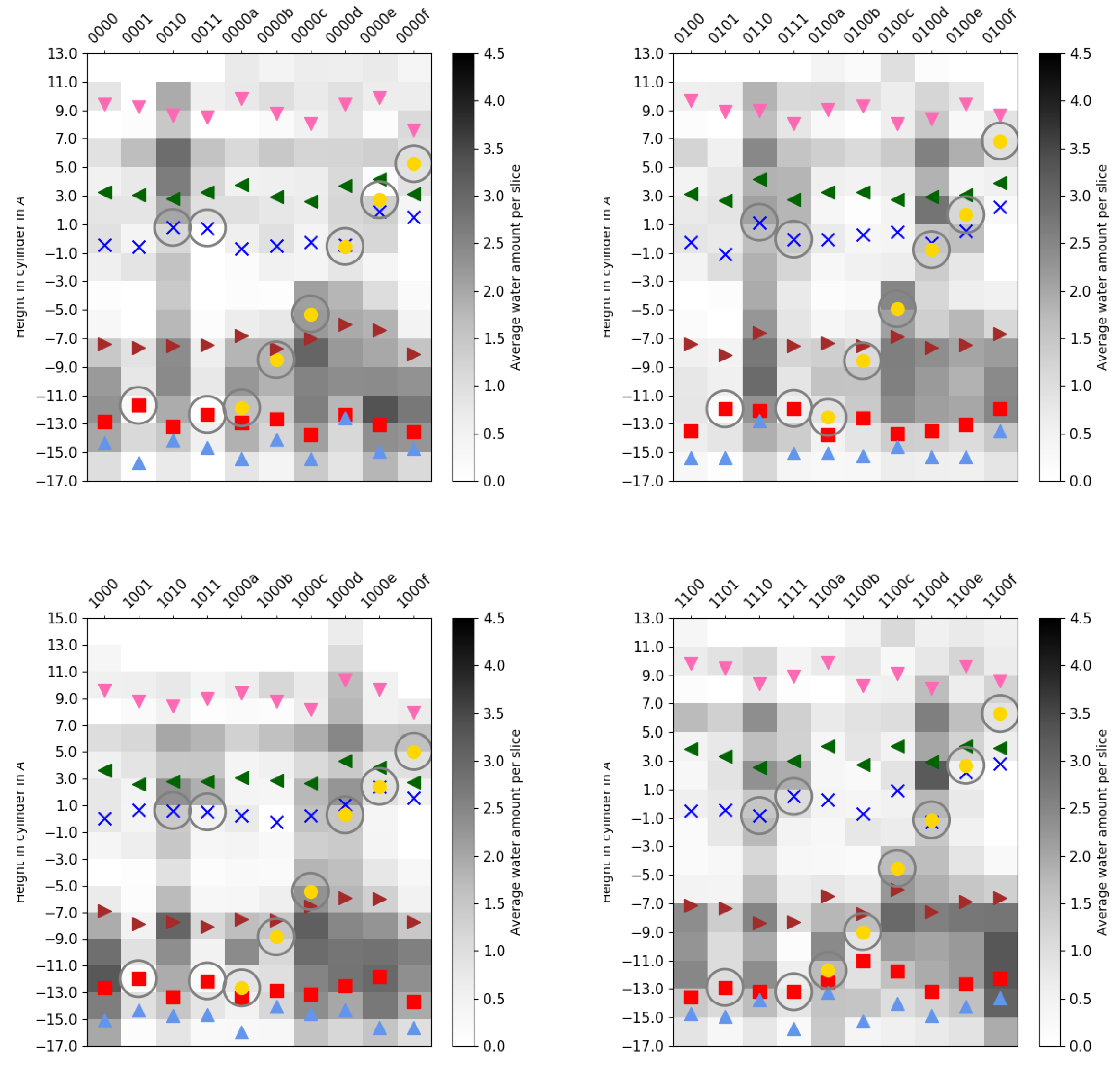
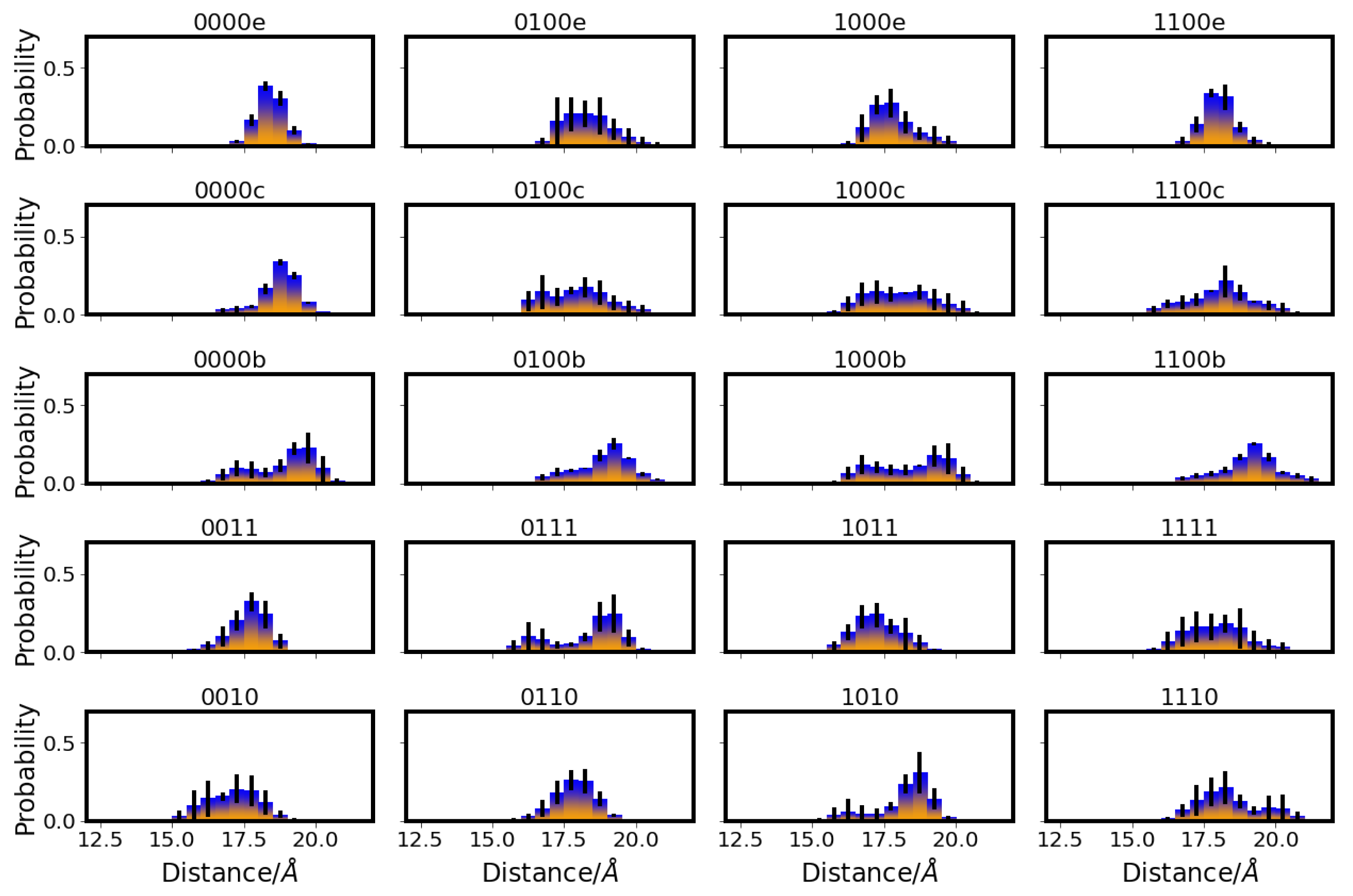
 ), E101 (red square
), E101 (red square  ), S365 (brown triangle—right
), S365 (brown triangle—right  ), K362 (blue cross
), K362 (blue cross  ), T359 (green triangle—left
), T359 (green triangle—left  ), and Y288 (magenta triangle—down
), and Y288 (magenta triangle—down  ), and the position of the ion (yellow circle
), and the position of the ion (yellow circle  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
 ), E101 (red square
), E101 (red square  ), S365 (brown triangle—right
), S365 (brown triangle—right  ), K362 (blue cross
), K362 (blue cross  ), T359 (green triangle—left
), T359 (green triangle—left  ), and Y288 (magenta triangle—down
), and Y288 (magenta triangle—down  ), and the position of the ion (yellow circle
), and the position of the ion (yellow circle  ). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.
). Residues with an excess proton are marked by a grey circle around the symbols representing the respective residues.


| 00* | 01* | 10* | 11* | ||||
|---|---|---|---|---|---|---|---|
| 0000f | 18.6 ± 0.8 | 0100f | 13.4 ± 2.1 | 1000f | 18.0 ± 1.6 | 1100f | 14.3 ± 0.6 |
| 0000e | 18.5 ± 0.8 | 0100e | 13.7 ± 0.8 | 1000e | 18.7 ± 1.8 | 1100e | 14.3 ± 0.7 |
| 0000d | 19.0 ± 0.3 | 0100d | 13.9 ± 1.4 | 1000d | 17.9 ± 0.5 | 1100d | 15.3 ± 0.9 |
| 0000c | 19.6 ± 1.1 | 0100c | 14.5 ± 0.9 | 1000c | 20.4 ± 0.5 | 1100c | 14.6 ± 0.9 |
| 0000b | 18.0 ± 1.2 | 0100b | 13.5 ± 0.3 | 1000b | 19.0 ± 1.0 | 1100b | 14.8 ± 1.0 |
| 0000a | 18.4 ± 0.5 | 0100a | 14.7 ± 0.3 | 1000a | 18.6 ± 0.4 | 1100a | 13.9 ± 0.5 |
| 0011 | 17.5 ± 0.2 | 0111 | 13.8 ± 1.0 | 1011 | 17.7 ± 0.4 | 1111 | 15.1 ± 0.6 |
| 0010 | 18.4 ± 0.7 | 0110 | 14.4 ± 0.4 | 1010 | 18.8 ± 1.9 | 1110 | 13.9 ± 0.3 |
| 0001 | 17.2 ± 0.6 | 0101 | 13.9 ± 0.7 | 1001 | 19.2 ± 0.7 | 1101 | 15.1 ± 0.4 |
| 0000 | 19.6 ± 0.9 | 0100 | 15.6 ± 0.5 | 1000 | 18.4 ± 1.9 | 1100 | 15.2 ± 0.3 |
| 00* | 01* | 10* | 11* | ||||
|---|---|---|---|---|---|---|---|
| 0000f | 9.1 ± 0.6 | 0100f | 6.6 ± 1.3 | 1000f | 7.1 ± 0.6 | 1100f | 8.5 ± 1.2 |
| 0000e | 7.1 ± 0.3 | 0100e | 10.0 ± 2.6 | 1000e | 9.0 ± 1.1 | 1100e | 9.5 ± 0.5 |
| 0000d | 12.4 ± 3.3 | 0100d | 17.2 ± 1.3 | 1000d | 17.4 ± 4.5 | 1100d | 18.9 ± 0.8 |
| 0000c | 11.5 ± 0.9 | 0100c | 11.8 ± 1.6 | 1000c | 14.9 ± 1.6 | 1100c | 9.9 ± 0.4 |
| 0000b | 6.7 ± 0.1 | 0100b | 5.4 ± 1.1 | 1000b | 5.9 ± 0.6 | 1100b | 6.1 ± 0.5 |
| 0000a | 7.2 ± 0.9 | 0100a | 7.3 ± 1.1 | 1000a | 6.1 ± 1.1 | 1100a | 5.7 ± 1.0 |
| 0011 | 5.6 ± 0.7 | 0111 | 12.1 ± 0.5 | 1011 | 8.1 ± 1.0 | 1111 | 8.1 ± 0.9 |
| 0010 | 18.5 ± 0.7 | 0110 | 19.4 ± 0.9 | 1010 | 15.2 ± 2.9 | 1110 | 16.1 ± 3.8 |
| 0001 | 6.2 ± 0.6 | 0101 | 5.5 ± 4.1 | 1001 | 5.3 ± 1.1 | 1101 | 7.7 ± 2.3 |
| 0000 | 6.5 ± 1.1 | 0100 | 2.7 ± 3.4 | 1000 | 7.4 ± 1.4 | 1100 | 6.2 ± 0.7 |
| 00* | 01* | 10* | 11* | ||||
|---|---|---|---|---|---|---|---|
| 0000f | 0.4 ± 0.4 | 0100f | 0.9 ± 0.2 | 1000f | 0.5 ± 0.3 | 1100f | 0.9 ± 0.0 |
| 0000e | 0.7 ± 0.6 | 0100e | 1.1 ± 0.0 | 1000e | 0.4 ± 0.2 | 1100e | 0.9 ± 0.0 |
| 0000d | 0.5 ± 0.2 | 0100d | 1.0 ± 0.1 | 1000d | 1.0 ± 0.3 | 1100d | 0.9 ± 0.0 |
| 0000c | 0.3 ± 0.2 | 0100c | 0.9 ± 0.2 | 1000c | 0.4 ± 0.1 | 1100c | 1.1 ± 0.1 |
| 0000b | 0.4 ± 0.2 | 0100b | 1.0 ± 0.3 | 1000b | 0.1 ± 0.0 | 1100b | 0.8 ± 0.2 |
| 0000a | 0.5 ± 0.4 | 0100a | 1.1 ± 0.1 | 1000a | 0.8 ± 0.2 | 1100a | 1.0 ± 0.1 |
| 0011 | 0.5 ± 0.3 | 0111 | 1.1 ± 0.1 | 1011 | 0.5 ± 0.2 | 1111 | 0.9 ± 0.1 |
| 0010 | 0.9 ± 0.5 | 0110 | 1.0 ± 0.1 | 1010 | 0.7 ± 0.4 | 1110 | 0.9 ± 0.2 |
| 0001 | 0.8 ± 0.6 | 0101 | 1.1 ± 0.1 | 1001 | 0.3 ± 0.1 | 1101 | 1.0 ± 0.2 |
| 0000 | 0.4 ± 0.3 | 0100 | 1.0 ± 0.0 | 1000 | 0.3 ± 0.1 | 1100 | 0.9 ± 0.2 |
| z | 0000z | 0100z | 1000z | 1100z |
|---|---|---|---|---|
| f | 1.4 ± 0.6 | 1.3 ± 0.4 | 1.5 ± 0.5 | 1.3 ± 0.3 |
| e | 1.0 ± 0.1 | 1.1 ± 0.2 | 1.3 ± 0.3 | 1.4 ± 0.2 |
| c | 2.5 ± 0.1 | 2.6 ± 0.2 | 2.6 ± 0.1 | 2.3 ± 0.2 |
| d | 2.2 ± 0.2 | 2.2 ± 0.4 | 2.0 ± 0.1 | 1.8 ± 0.1 |
| b | 1.2 ± 0.2 | 1.1 ± 0.4 | 0.8 ± 0.4 | 1.3 ± 0.5 |
| a | 1.5 ± 0.2 | 1.0 ± 0.7 | 1.2 ± 0.2 | 1.4 ± 0.2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gorriz, R.F.; Volkenandt, S.; Imhof, P. Protonation-State Dependence of Hydration and Interactions in the Two Proton-Conducting Channels of Cytochrome c Oxidase. Int. J. Mol. Sci. 2023, 24, 10464. https://doi.org/10.3390/ijms241310464
Gorriz RF, Volkenandt S, Imhof P. Protonation-State Dependence of Hydration and Interactions in the Two Proton-Conducting Channels of Cytochrome c Oxidase. International Journal of Molecular Sciences. 2023; 24(13):10464. https://doi.org/10.3390/ijms241310464
Chicago/Turabian StyleGorriz, Rene F., Senta Volkenandt, and Petra Imhof. 2023. "Protonation-State Dependence of Hydration and Interactions in the Two Proton-Conducting Channels of Cytochrome c Oxidase" International Journal of Molecular Sciences 24, no. 13: 10464. https://doi.org/10.3390/ijms241310464
APA StyleGorriz, R. F., Volkenandt, S., & Imhof, P. (2023). Protonation-State Dependence of Hydration and Interactions in the Two Proton-Conducting Channels of Cytochrome c Oxidase. International Journal of Molecular Sciences, 24(13), 10464. https://doi.org/10.3390/ijms241310464





