Single-Cell RNA Sequencing Analysis for Oncogenic Mechanisms Underlying Oral Squamous Cell Carcinoma Carcinogenesis with Candida albicans Infection
Abstract
:1. Introduction
2. Results
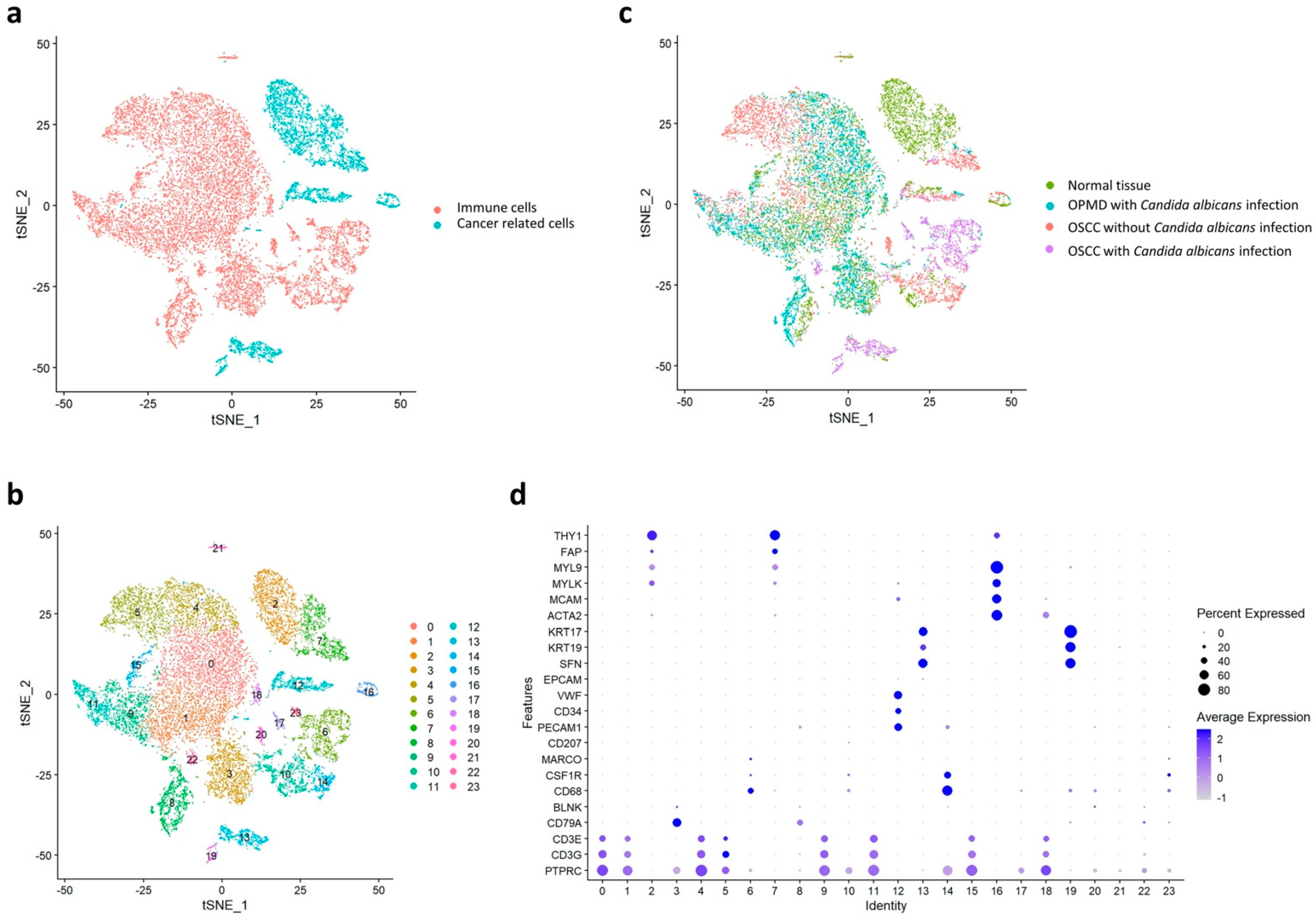
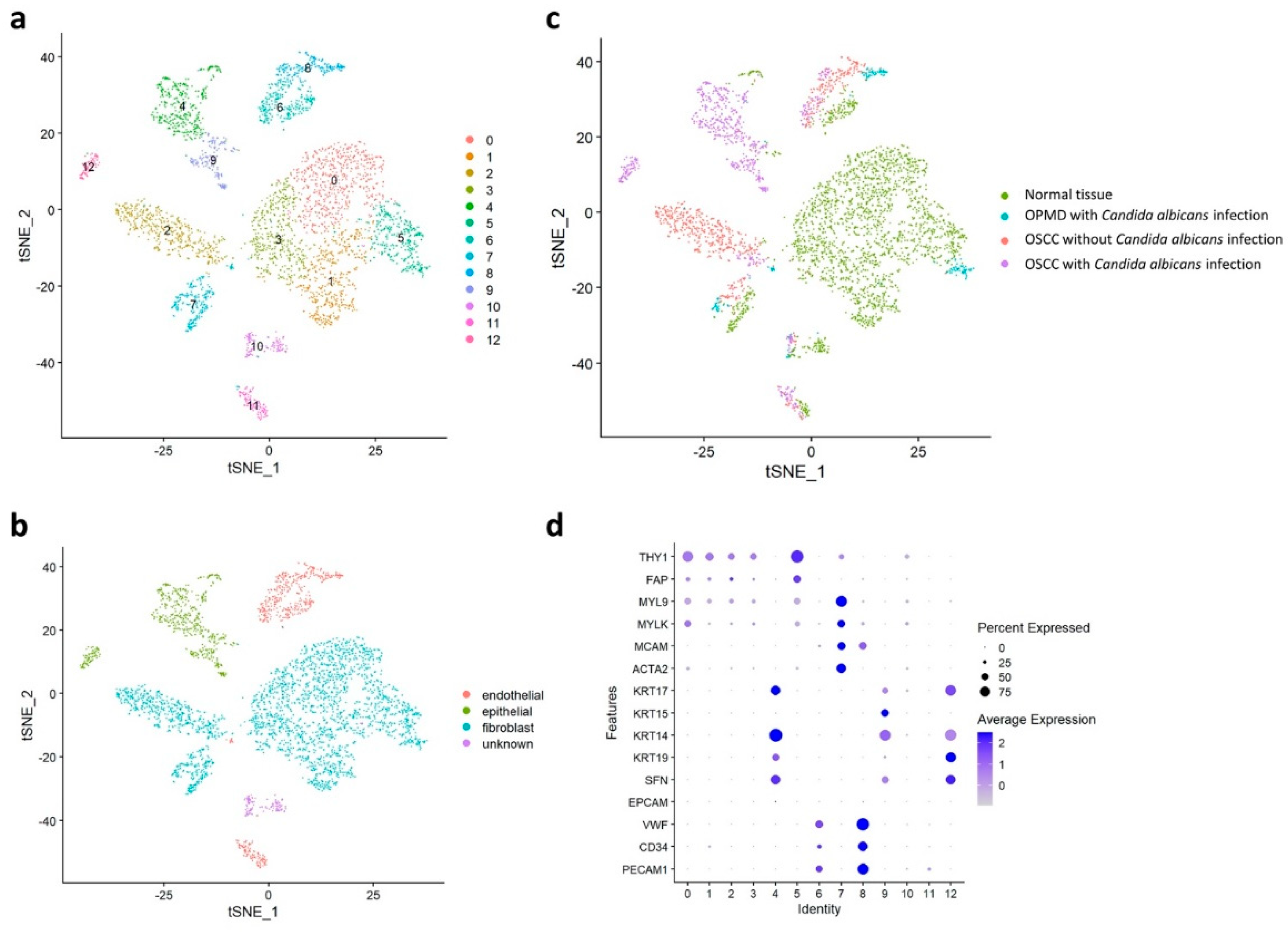
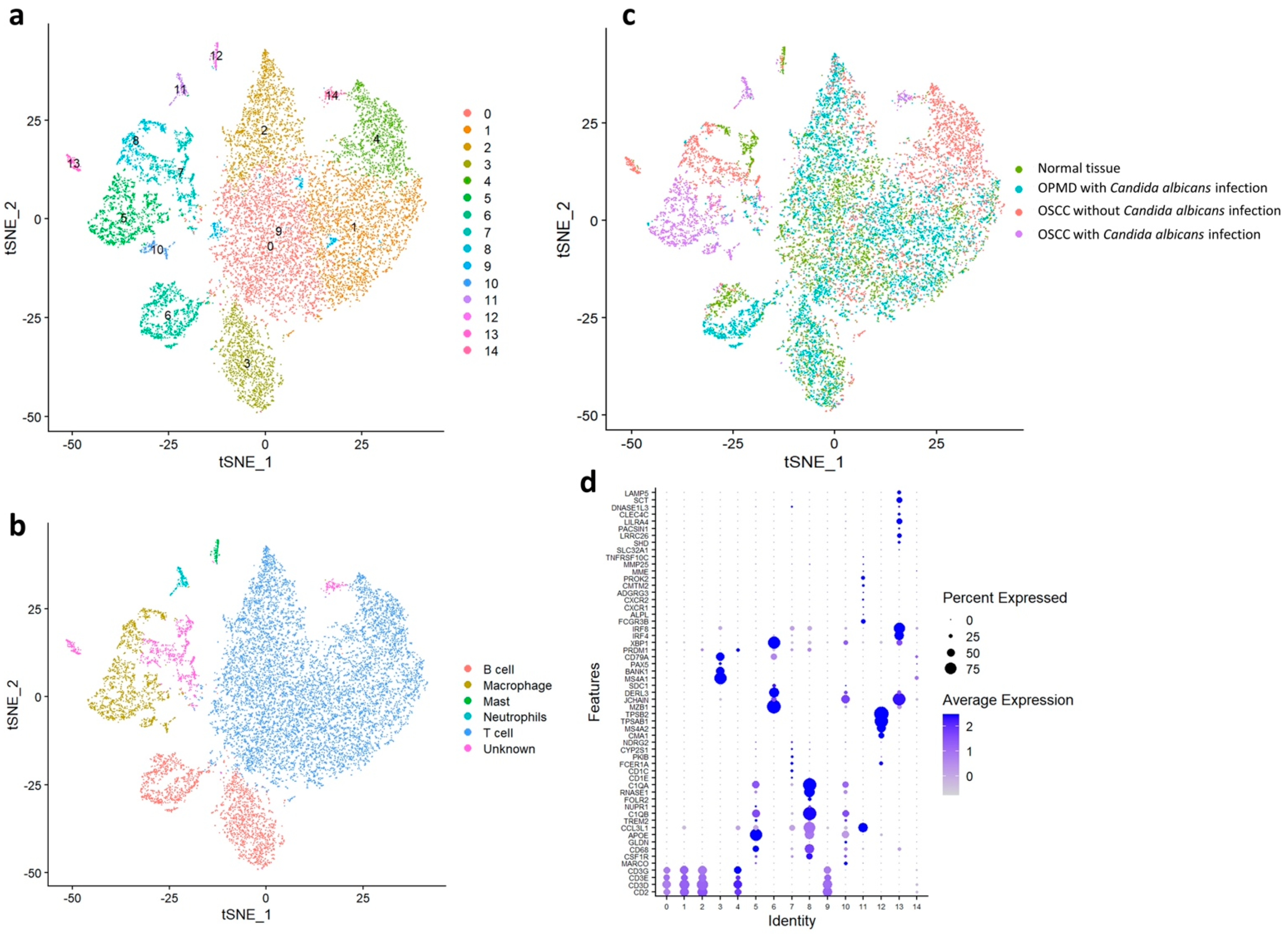
2.1. Single-Cell RNA Sequencing in the Normal Tissue, OPMD Lesion with C. albicans Infection, OSCC Tissue with C. albicans Infection, and OSCC Tissue without C. albicans Infection
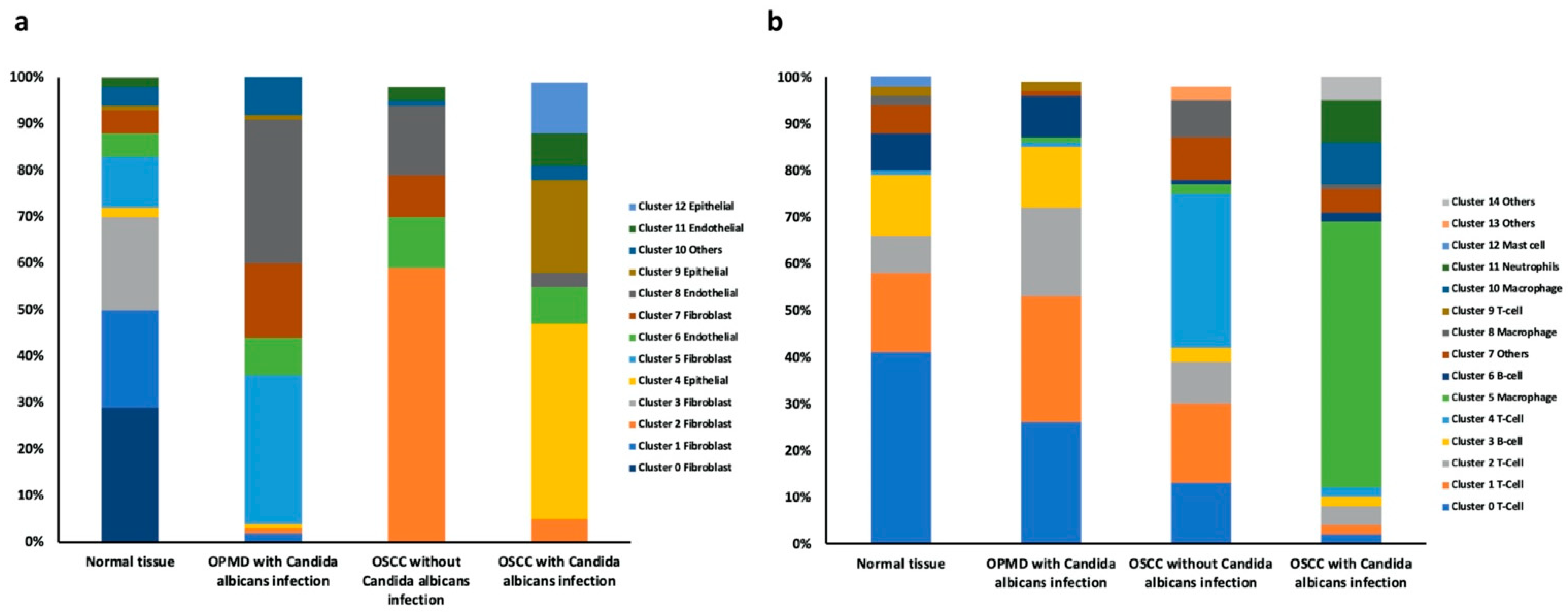
2.2. Cell Composition Change in the Normal Tissue, OPMD Lesion with C. albicans Infection, OSCC Tissue with C. albicans Infection, and OSCC Tissue without C. albicans Infection
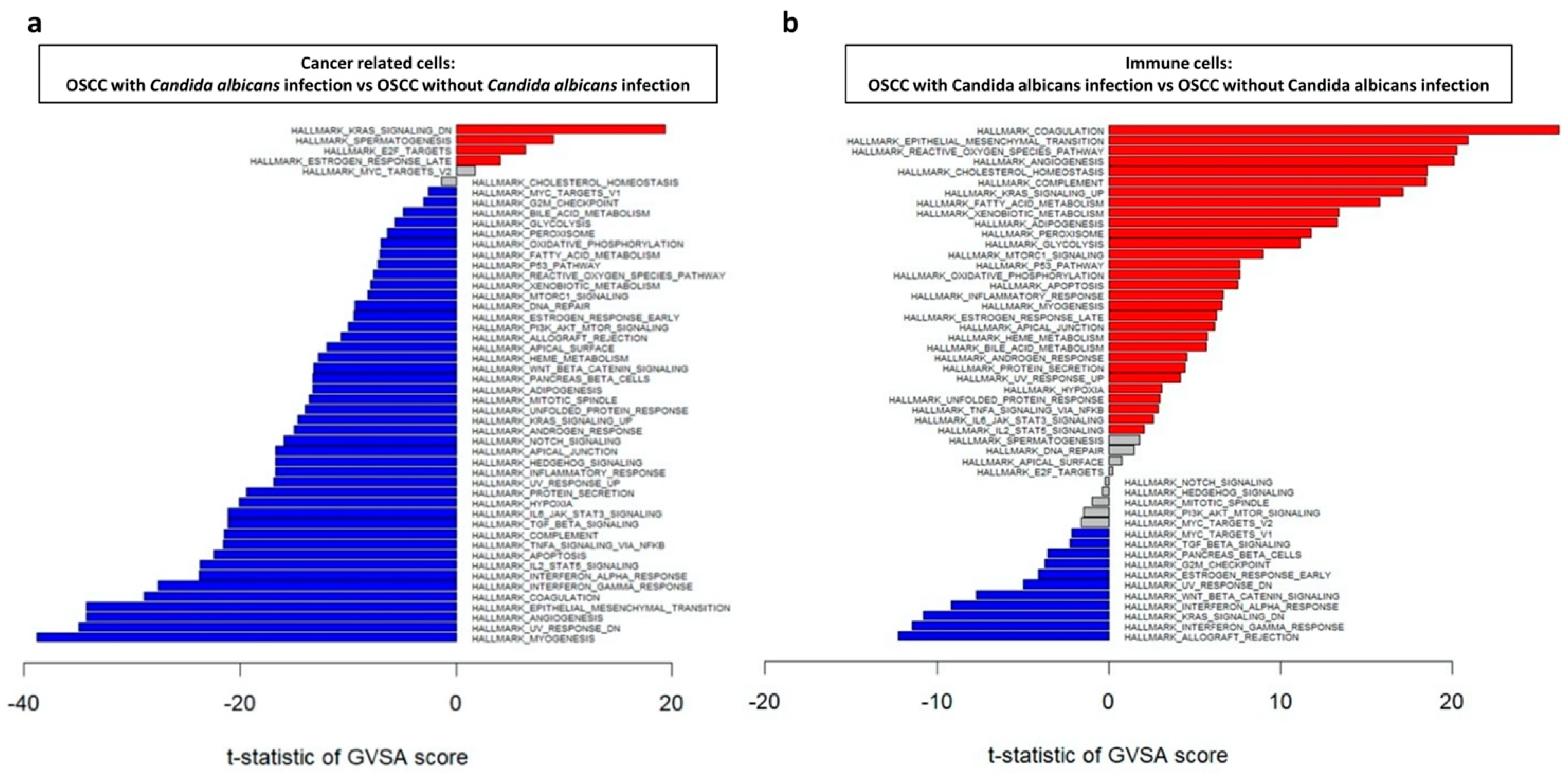
2.3. Regulatory Pathways Involved in OSCC Carcinogenesis with and without C. albicans Infection
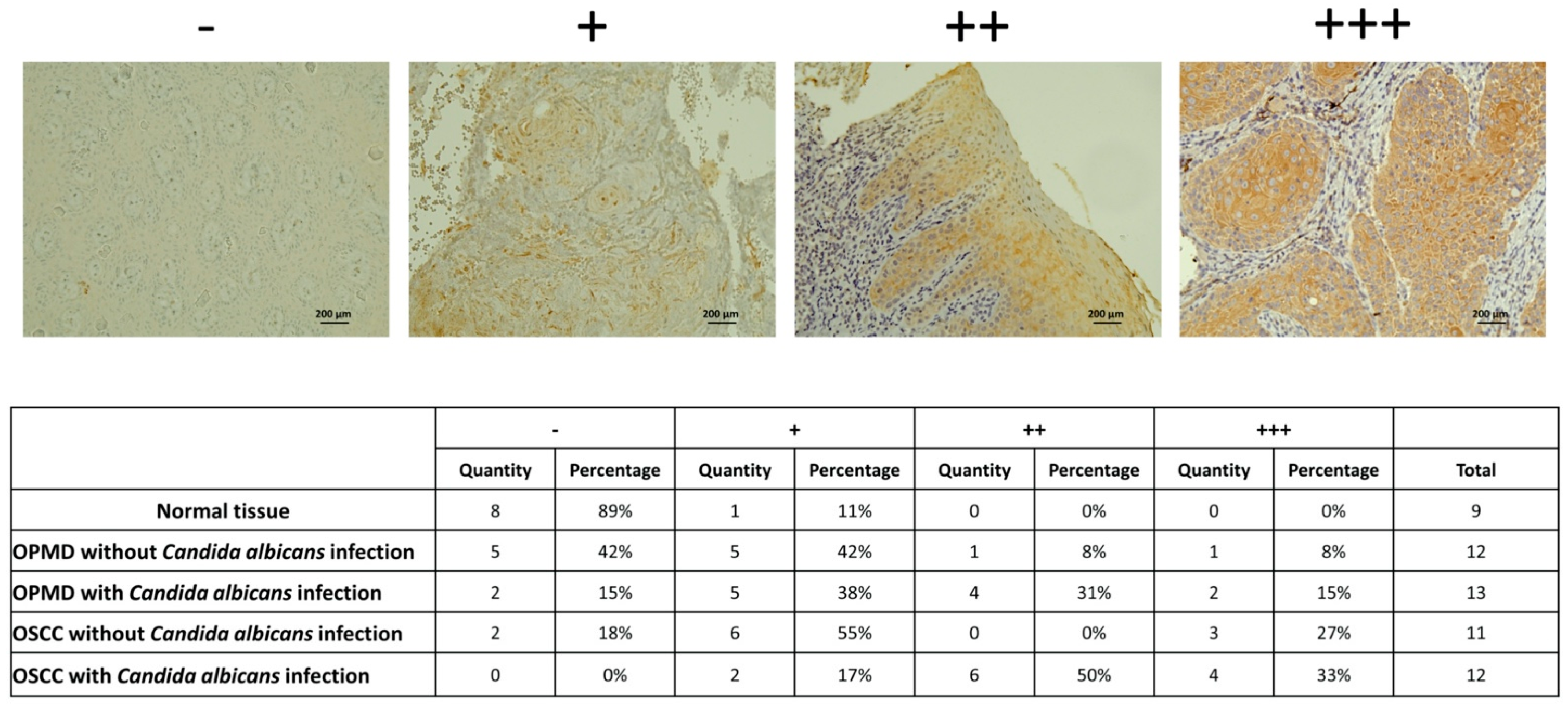
2.4. Stratifin (SFN) as the Specific Biomarker of OSCC with C. albicans Infection
3. Discussion
4. Materials and Methods
4.1. Clinical Patient Specimen Collection
4.2. Candidiasis Detection
4.3. Tissue Dissociation
4.4. 3′ Gene Expression Library Construction on 10× Genomics Platform
4.5. Next-Generation Sequencing
4.6. Bioinformatic Analysis of NGS Data
4.7. Immunohistochemistry Staining
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Sagaert, X.; Vanstapel, A.; Verbeek, S. Tumor Heterogeneity in Colorectal Cancer: What Do We Know So Far? Pathobiology 2018, 85, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Marusyk, A.; Polyak, K. Tumor heterogeneity: Causes and consequences. Biochim. Biophys. Acta 2010, 1805, 105–117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Sayes, N.; Vito, A.; Mossman, K. Tumor Heterogeneity: A Great Barrier in the Age of Cancer Immunotherapy. Cancers 2021, 13, 806. [Google Scholar] [CrossRef] [PubMed]
- Dagogo-Jack, I.; Shaw, A.T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 2018, 15, 81–94. [Google Scholar] [CrossRef]
- Anderson, N.M.; Simon, M.C. The tumor microenvironment. Curr. Biol. 2020, 30, R921–R925. [Google Scholar] [CrossRef] [PubMed]
- Brassart-Pasco, S.; Brézillon, S.; Brassart, B.; Ramont, L.; Oudart, J.B.; Monboisse, J.C. Tumor Microenvironment: Extracellular Matrix Alterations Influence Tumor Progression. Front. Oncol. 2020, 10, 397. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baghban, R.; Roshangar, L.; Jahanban-Esfahlan, R.; Seidi, K.; Ebrahimi-Kalan, A.; Jaymand, M.; Kolahian, S.; Javaheri, T.; Zare, P. Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun. Signal. 2020, 18, 59. [Google Scholar] [CrossRef] [Green Version]
- Zuo, S.; Wei, M.; Wang, S.; Dong, J.; Wei, J. Pan-Cancer Analysis of Immune Cell Infiltration Identifies a Prognostic Immune-Cell Characteristic Score (ICCS) in Lung Adenocarcinoma. Front. Immunol. 2020, 11, 1218. [Google Scholar] [CrossRef]
- Zhang, X.; Shi, M.; Chen, T.; Zhang, B. Characterization of the Immune Cell Infiltration Landscape in Head and Neck Squamous Cell Carcinoma to Aid Immunotherapy. Mol. Ther. Nucleic Acids 2020, 22, 298–309. [Google Scholar] [CrossRef]
- Salazar, N.; Zabel, B.A. Support of Tumor Endothelial Cells by Chemokine Receptors. Front. Immunol. 2019, 10, 147. [Google Scholar] [CrossRef] [Green Version]
- Zhuo, H.; Zhao, Y.; Cheng, X.; Xu, M.; Wang, L.; Lin, L.; Lyu, Z.; Hong, X.; Cai, J. Tumor endothelial cell-derived cadherin-2 promotes angiogenesis and has prognostic significance for lung adenocarcinoma. Mol. Cancer 2019, 18, 34. [Google Scholar] [CrossRef] [PubMed]
- Hida, K.; Maishi, N.; Akiyama, K.; Ohmura-Kakutani, H.; Torii, C.; Ohga, N.; Osawa, T.; Kikuchi, H.; Morimoto, H.; Morimoto, M.; et al. Tumor endothelial cells with high aldehyde dehydrogenase activity show drug resistance. Cancer Sci. 2017, 108, 2195–2203. [Google Scholar] [CrossRef] [PubMed]
- Lopatina, T.; Grange, C.; Cavallari, C.; Navarro-Tableros, V.; Lombardo, G.; Rosso, A.; Cedrino, M.; Pomatto, M.A.C.; Koni, M.; Veneziano, F.; et al. Targeting IL-3Rα on tumor-derived endothelial cells blunts metastatic spread of triple-negative breast cancer via extracellular vesicle reprogramming. Oncogenesis 2020, 9, 90. [Google Scholar] [CrossRef] [PubMed]
- Sahai, E.; Astsaturov, I.; Cukierman, E.; DeNardo, D.G.; Egeblad, M.; Evans, R.M.; Fearon, D.; Greten, F.R.; Hingorani, S.R.; Hunter, T.; et al. A framework for advancing our understanding of cancer-associated fibroblasts. Nat. Rev. Cancer 2020, 20, 174–186. [Google Scholar] [CrossRef] [Green Version]
- Liu, T.; Han, C.; Wang, S.; Fang, P.; Ma, Z.; Xu, L.; Yin, R. Cancer-associated fibroblasts: An emerging target of anti-cancer immunotherapy. J. Hematol. Oncol. 2019, 12, 86. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Aupérin, A. Epidemiology of head and neck cancers: An update. Curr. Opin. Oncol. 2020, 32, 178–186. [Google Scholar] [CrossRef]
- Kim, D.; Li, R. Contemporary Treatment of Locally Advanced Oral Cancer. Curr. Treat. Options Oncol. 2019, 20, 32. [Google Scholar] [CrossRef]
- Chiu, S.F.; Ho, C.H.; Chen, Y.C.; Wu, L.W.; Chen, Y.L.; Wu, J.H.; Wu, W.S.; Hung, H.K.; Chiang, W.F. Malignant transformation of oral potentially malignant disorders in Taiwan: An observational nationwide population database study. Medicine 2021, 100, e24934. [Google Scholar] [CrossRef]
- Pritzker, K.P.H.; Darling, M.R.; Hwang, J.T.; Mock, D. Oral Potentially Malignant Disorders (OPMD): What is the clinical utility of dysplasia grade? Expert Rev. Mol. Diagn. 2021, 21, 289–298. [Google Scholar] [CrossRef]
- Mohd Bakri, M.; Mohd Hussaini, H.; Rachel Holmes, A.; David Cannon, R.; Mary Rich, A. Revisiting the association between candidal infection and carcinoma, particularly oral squamous cell carcinoma. J. Oral Microbiol. 2010, 2, 5780. [Google Scholar] [CrossRef] [PubMed]
- Warnakulasuriya, S.; Johnson, N.W.; van der Waal, I. Nomenclature and classification of potentially malignant disorders of the oral mucosa. J. Oral Pathol. Med. 2007, 36, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Mäkinen, A.; Nawaz, A.; Mäkitie, A.; Meurman, J.H. Role of Non-Albicans Candida and Candida Albicans in Oral Squamous Cell Cancer Patients. J. Oral Maxillofac. Surg. 2018, 76, 2564–2571. [Google Scholar] [CrossRef] [Green Version]
- Sankari, S.L.; Gayathri, K.; Balachander, N.; Malathi, L. Candida in potentially malignant oral disorders. J. Pharm. Bioallied. Sci. 2015, 7, S162–S164. [Google Scholar] [CrossRef] [PubMed]
- Sankari, S.L.; Mahalakshmi, K.; Kumar, V.N. A comparative study of Candida species diversity among patients with oral squamous cell carcinoma and oral potentially malignant disorders. BMC Res. Notes 2020, 13, 488. [Google Scholar] [CrossRef] [PubMed]
- Di Cosola, M.; Cazzolla, A.P.; Charitos, I.A.; Ballini, A.; Inchingolo, F.; Santacroce, L. Candida albicans and Oral Carcinogenesis. A Brief Review. J. Fungi 2021, 7, 476. [Google Scholar] [CrossRef] [PubMed]
- Lim, Y.; Totsika, M.; Morrison, M.; Punyadeera, C. Oral Microbiome: A New Biomarker Reservoir for Oral and Oropharyngeal Cancers. Theranostics 2017, 7, 4313–4321. [Google Scholar] [CrossRef]
- Qin, Y.; Zhang, L.; Xu, Z.; Zhang, J.; Jiang, Y.Y.; Cao, Y.; Yan, T. Innate immune cell response upon Candida albicans infection. Virulence 2016, 7, 512–526. [Google Scholar] [CrossRef] [Green Version]
- Shen, Y.; Liu, J.; Zhang, L.; Dong, S.; Zhang, J.; Liu, Y.; Zhou, H.; Dong, W. Identification of Potential Biomarkers and Survival Analysis for Head and Neck Squamous Cell Carcinoma Using Bioinformatics Strategy: A Study Based on TCGA and GEO Datasets. BioMed Res. Int. 2019, 2019, 7376034. [Google Scholar] [CrossRef] [Green Version]
- Zevallos, J.P.; Mazul, A.L.; Walter, V.; Hayes, D.N. Gene Expression Subtype Predicts Nodal Metastasis and Survival in Human Papillomavirus-Negative Head and Neck Cancer. Laryngoscope 2019, 129, 154–161. [Google Scholar] [CrossRef]
- Suvà, M.L.; Tirosh, I. Single-Cell RNA Sequencing in Cancer: Lessons Learned and Emerging Challenges. Mol. Cell 2019, 75, 7–12. [Google Scholar] [CrossRef] [PubMed]
- Puram, S.V.; Tirosh, I.; Parikh, A.S.; Patel, A.P.; Yizhak, K.; Gillespie, S.; Rodman, C.; Luo, C.L.; Mroz, E.A.; Emerick, K.S.; et al. Single-Cell Transcriptomic Analysis of Primary and Metastatic Tumor Ecosystems in Head and Neck Cancer. Cell 2017, 171, 1611–1624.e1624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prasetyanti, P.R.; Medema, J.P. Intra-tumor heterogeneity from a cancer stem cell perspective. Mol. Cancer 2017, 16, 41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, X.; Kang, B.; Zhang, Z. Understanding tumor ecosystems by single-cell sequencing: Promises and limitations. Genome. Biol. 2018, 19, 211. [Google Scholar] [CrossRef] [PubMed]
- Valihrach, L.; Androvic, P.; Kubista, M. Platforms for Single-Cell Collection and Analysis. Int. J. Mol. Sci. 2018, 19, 807. [Google Scholar] [CrossRef] [Green Version]
- Salomon, R.; Kaczorowski, D.; Valdes-Mora, F.; Nordon, R.E.; Neild, A.; Farbehi, N.; Bartonicek, N.; Gallego-Ortega, D. Droplet-based single cell RNAseq tools: A practical guide. Lab. Chip. 2019, 19, 1706–1727. [Google Scholar] [CrossRef]
- Guo, X.; Zhang, Y.; Zheng, L.; Zheng, C.; Song, J.; Zhang, Q.; Kang, B.; Liu, Z.; Jin, L.; Xing, R.; et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 2018, 24, 978–985. [Google Scholar] [CrossRef]
- Chung, W.; Eum, H.H.; Lee, H.O.; Lee, K.M.; Lee, H.B.; Kim, K.T.; Ryu, H.S.; Kim, S.; Lee, J.E.; Park, Y.H.; et al. Single-cell RNA-seq enables comprehensive tumour and immune cell profiling in primary breast cancer. Nat. Commun. 2017, 8, 15081. [Google Scholar] [CrossRef] [Green Version]
- Kim, C.; Gao, R.; Sei, E.; Brandt, R.; Hartman, J.; Hatschek, T.; Crosetto, N.; Foukakis, T.; Navin, N.E. Chemoresistance Evolution in Triple-Negative Breast Cancer Delineated by Single-Cell Sequencing. Cell 2018, 173, 879–893.e813. [Google Scholar] [CrossRef] [Green Version]
- Arya, C.P.; Jaiswal, R.; Tandon, A.; Jain, A. Isolation and identification of oral Candida species in potentially malignant disorder and oral squamous cell carcinoma. Nat. J. Maxillofac. Surg. 2021, 12, 387–391. [Google Scholar] [CrossRef]
- Hafed, L.; Farag, H.; El-Rouby, D.; Shaker, O.; Shabaan, H.A. Candida Albicans Alcohol Dehydrogenase 1 gene in oral dysplasia and oral squamous cell carcinoma. Pol. J. Pathol. 2019, 70, 210–216. [Google Scholar] [CrossRef] [PubMed]
- Pavlova, A.; Sharafutdinov, I. Recognition of Candida albicans and Role of Innate Type 17 Immunity in Oral Candidiasis. Microorganisms 2020, 8, 1340. [Google Scholar] [CrossRef] [PubMed]
- Pellon, A.; Sadeghi Nasab, S.D.; Moyes, D.L. New Insights in Candida albicans Innate Immunity at the Mucosa: Toxins, Epithelium, Metabolism, and Beyond. Front. Cell Infect. Microbiol. 2020, 10, 81. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vadovics, M.; Ho, J.; Igaz, N.; Alföldi, R.; Rakk, D.; Veres, É.; Szücs, B.; Horváth, M.; Tóth, R.; Szücs, A.; et al. Candida albicans Enhances the Progression of Oral Squamous Cell Carcinoma In Vitro and In Vivo. Mbio 2022, 13, e0314421. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, S.A.; Manlove, L.S.; Farrar, M.A. Interleukin-2 and STAT5 in regulatory T cell development and function. Jakstat 2013, 2, e23154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halim, C.E.; Deng, S.; Ong, M.S.; Yap, C.T. Involvement of STAT5 in Oncogenesis. Biomedicines 2020, 8, 316. [Google Scholar] [CrossRef]
- Wu, Y.; Zhou, B.P. TNF-alpha/NF-kappaB/Snail pathway in cancer cell migration and invasion. Br. J. Cancer 2010, 102, 639–644. [Google Scholar] [CrossRef] [Green Version]
- Hao, Y.; Baker, D.; Ten Dijke, P. TGF-β-Mediated Epithelial-Mesenchymal Transition and Cancer Metastasis. Int. J. Mol. Sci. 2019, 20, 2767. [Google Scholar] [CrossRef] [Green Version]
- Deng, J.; Gao, G.; Wang, L.; Wang, T.; Yu, J.; Zhao, Z. Stratifin expression is a novel prognostic factor in human gliomas. Pathol. Res. Pract. 2011, 207, 674–679. [Google Scholar] [CrossRef]
- Hu, Y.; Zeng, Q.; Li, C.; Xie, Y. Expression profile and prognostic value of SFN in human ovarian cancer. Biosci. Rep. 2019, 39, BSR20190100. [Google Scholar] [CrossRef] [Green Version]
- Yang, L.; Zhang, Z.; Sun, Y.; Pang, S.; Yao, Q.; Lin, P.; Cheng, J.; Li, J.; Ding, G.; Hui, L.; et al. Integrative analysis reveals novel driver genes and molecular subclasses of hepatocellular carcinoma. Aging 2020, 12, 23849–23871. [Google Scholar] [CrossRef] [PubMed]
- Boominathan, L. Some facts and thoughts: p73 as a tumor suppressor gene in the network of tumor suppressors. Mol. Cancer 2007, 6, 27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Safavieh, M.; Coarsey, C.; Esiobu, N.; Memic, A.; Vyas, J.M.; Shafiee, H.; Asghar, W. Advances in Candida detection platforms for clinical and point-of-care applications. Crit. Rev. Biotechnol. 2017, 37, 441–458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Simone, M.; Rossetti, G.; Pagani, M. Chromium 10× Single-Cell 3’ mRNA Sequencing of Tumor-Infiltrating Lymphocytes. Methods Mol. Biol. 2019, 1979, 87–110. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Dawood, M.; Johnson, C.A.; Schmitz, R. RNA Sequencing in B-Cell Lymphomas. Methods Mol. Biol. 2019, 1956, 283–303. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hsieh, Y.-P.; Wu, Y.-H.; Cheng, S.-M.; Lin, F.-K.; Hwang, D.-Y.; Jiang, S.-S.; Chen, K.-C.; Chen, M.-Y.; Chiang, W.-F.; Liu, K.-J.; et al. Single-Cell RNA Sequencing Analysis for Oncogenic Mechanisms Underlying Oral Squamous Cell Carcinoma Carcinogenesis with Candida albicans Infection. Int. J. Mol. Sci. 2022, 23, 4833. https://doi.org/10.3390/ijms23094833
Hsieh Y-P, Wu Y-H, Cheng S-M, Lin F-K, Hwang D-Y, Jiang S-S, Chen K-C, Chen M-Y, Chiang W-F, Liu K-J, et al. Single-Cell RNA Sequencing Analysis for Oncogenic Mechanisms Underlying Oral Squamous Cell Carcinoma Carcinogenesis with Candida albicans Infection. International Journal of Molecular Sciences. 2022; 23(9):4833. https://doi.org/10.3390/ijms23094833
Chicago/Turabian StyleHsieh, Yi-Ping, Yu-Hsueh Wu, Siao-Muk Cheng, Fang-Kuei Lin, Daw-Yang Hwang, Shih-Sheng Jiang, Ken-Chung Chen, Meng-Yen Chen, Wei-Fan Chiang, Ko-Jiunn Liu, and et al. 2022. "Single-Cell RNA Sequencing Analysis for Oncogenic Mechanisms Underlying Oral Squamous Cell Carcinoma Carcinogenesis with Candida albicans Infection" International Journal of Molecular Sciences 23, no. 9: 4833. https://doi.org/10.3390/ijms23094833
APA StyleHsieh, Y.-P., Wu, Y.-H., Cheng, S.-M., Lin, F.-K., Hwang, D.-Y., Jiang, S.-S., Chen, K.-C., Chen, M.-Y., Chiang, W.-F., Liu, K.-J., Huynh, N. C.-N., Huang, W.-T., & Huang, T.-T. (2022). Single-Cell RNA Sequencing Analysis for Oncogenic Mechanisms Underlying Oral Squamous Cell Carcinoma Carcinogenesis with Candida albicans Infection. International Journal of Molecular Sciences, 23(9), 4833. https://doi.org/10.3390/ijms23094833







