Integrated Analysis Revealed an Inflammatory Cancer-Associated Fibroblast-Based Subtypes with Promising Implications in Predicting the Prognosis and Immunotherapeutic Response of Bladder Cancer Patients
Abstract
1. Introduction
2. Results
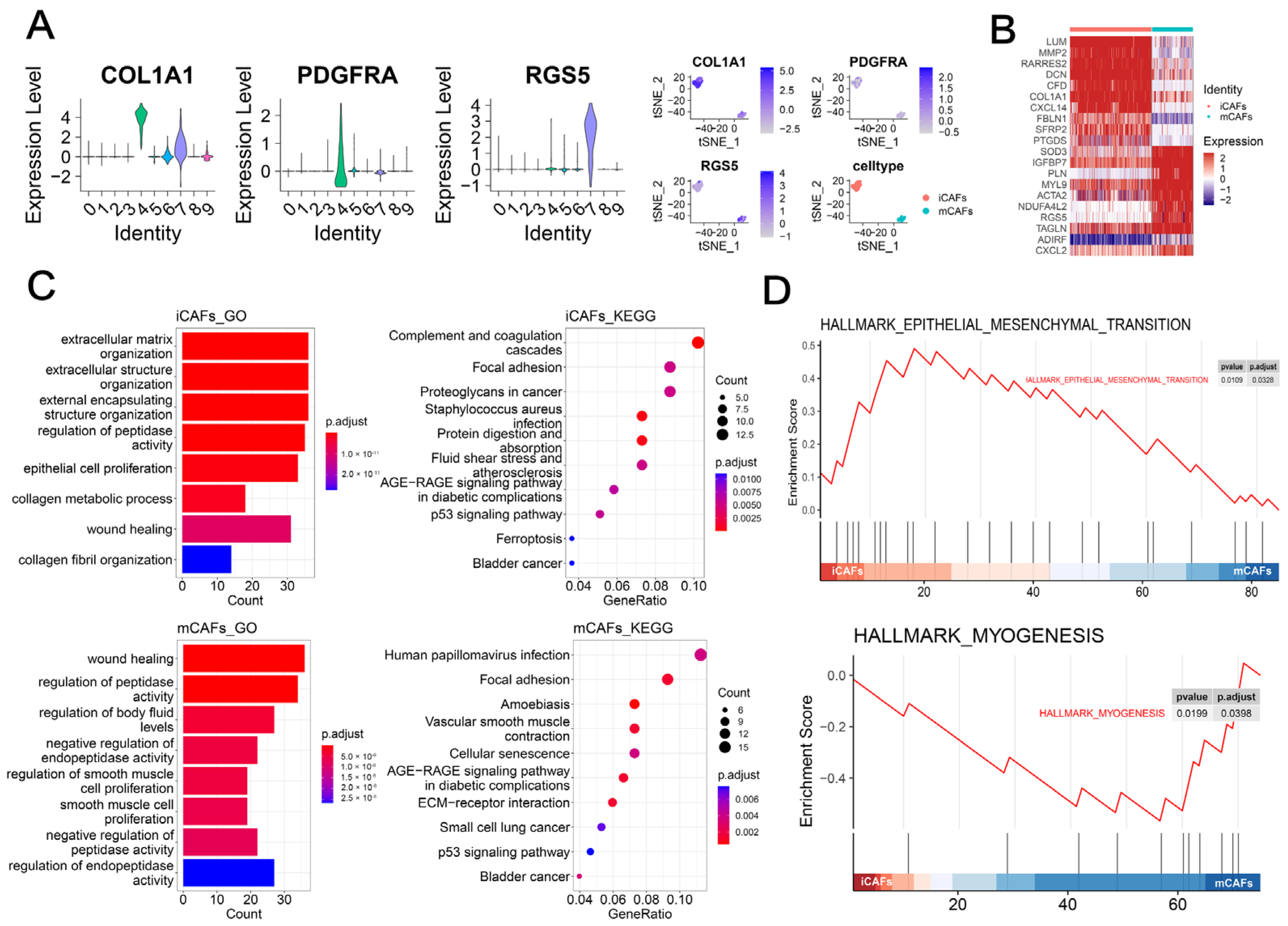
2.1. scRNA-seq Analysis Highlighted the Role of iCAFs in BCa
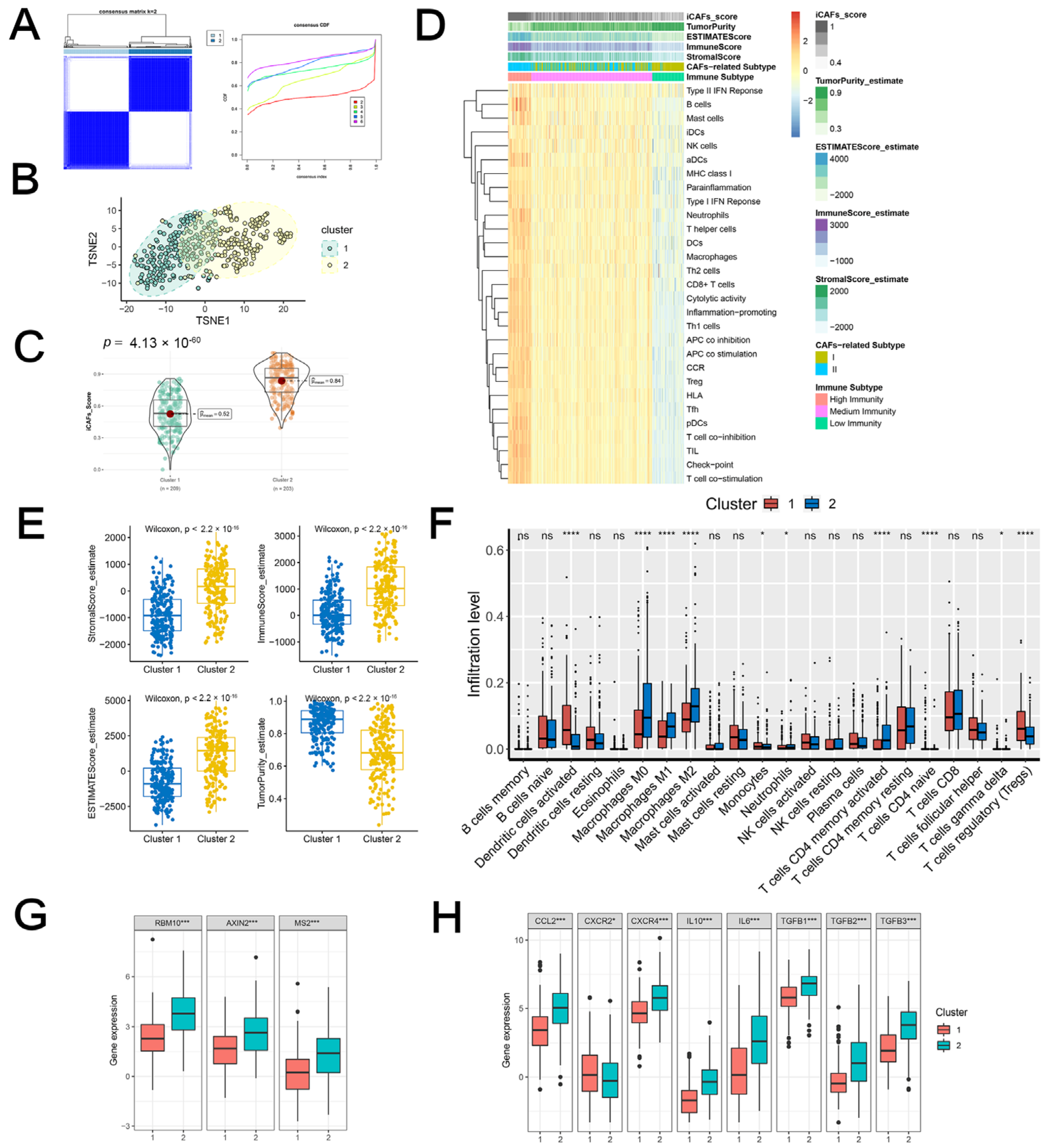
2.2. iCAF-Related Signature Dominated an Inflamed and Immunosuppressive TME of BCa
2.3. iCAF-Related Signature Stratified BCa into Molecular Subtypes with Distinct Biological Features
2.3.1. iCAF-related Subtypes had Distinct Immunogenomic Patterns
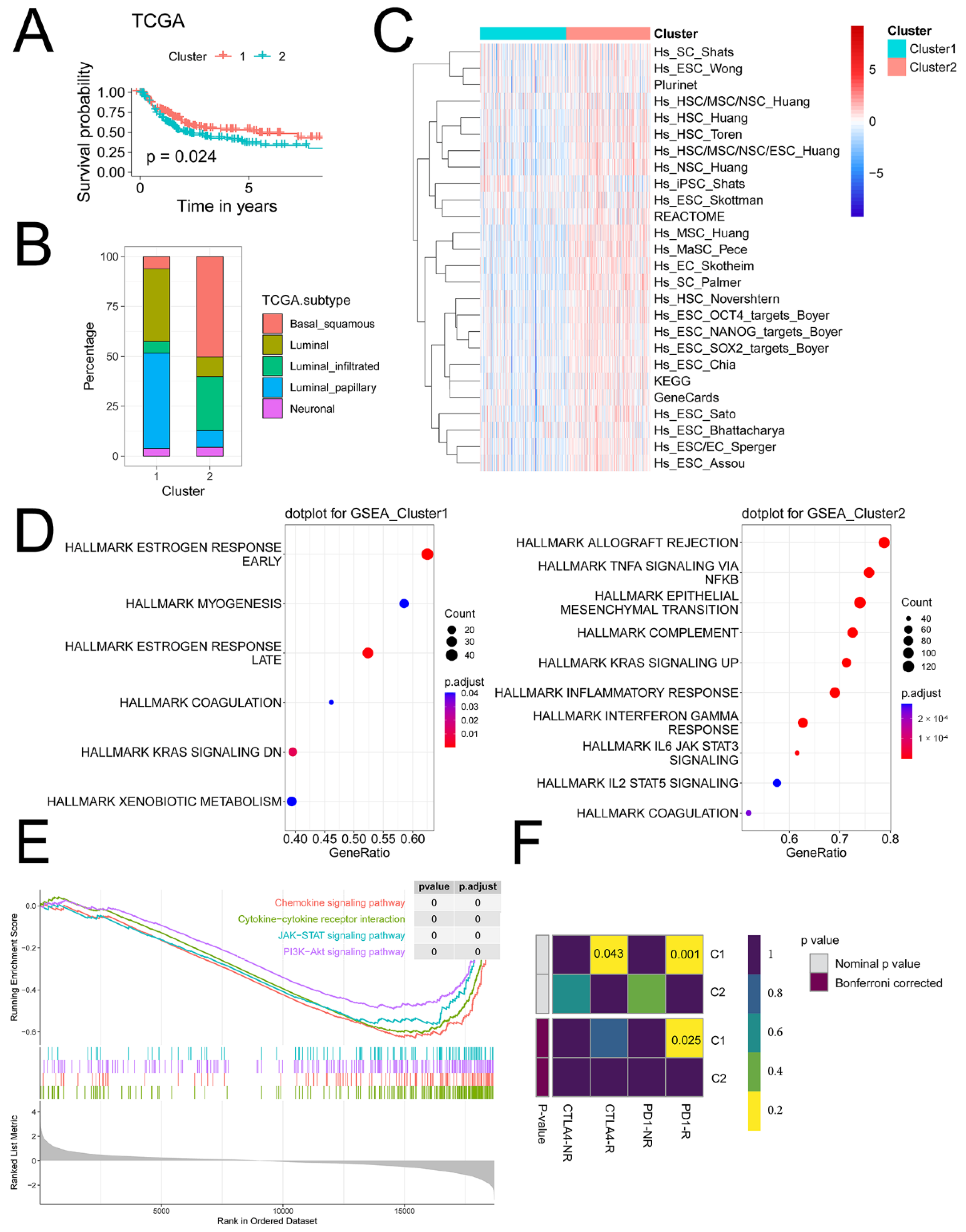
2.3.2. iCAF-related Subtypes had Distinct Dysregulated Pathways
2.3.3. iCAF-related Subtypes Hold Diverse Responses to Immunotherapy and Chemotherapy
2.4. iCAF-related Subtypes Demonstrated Robustness in the External GEO-meta Dataset
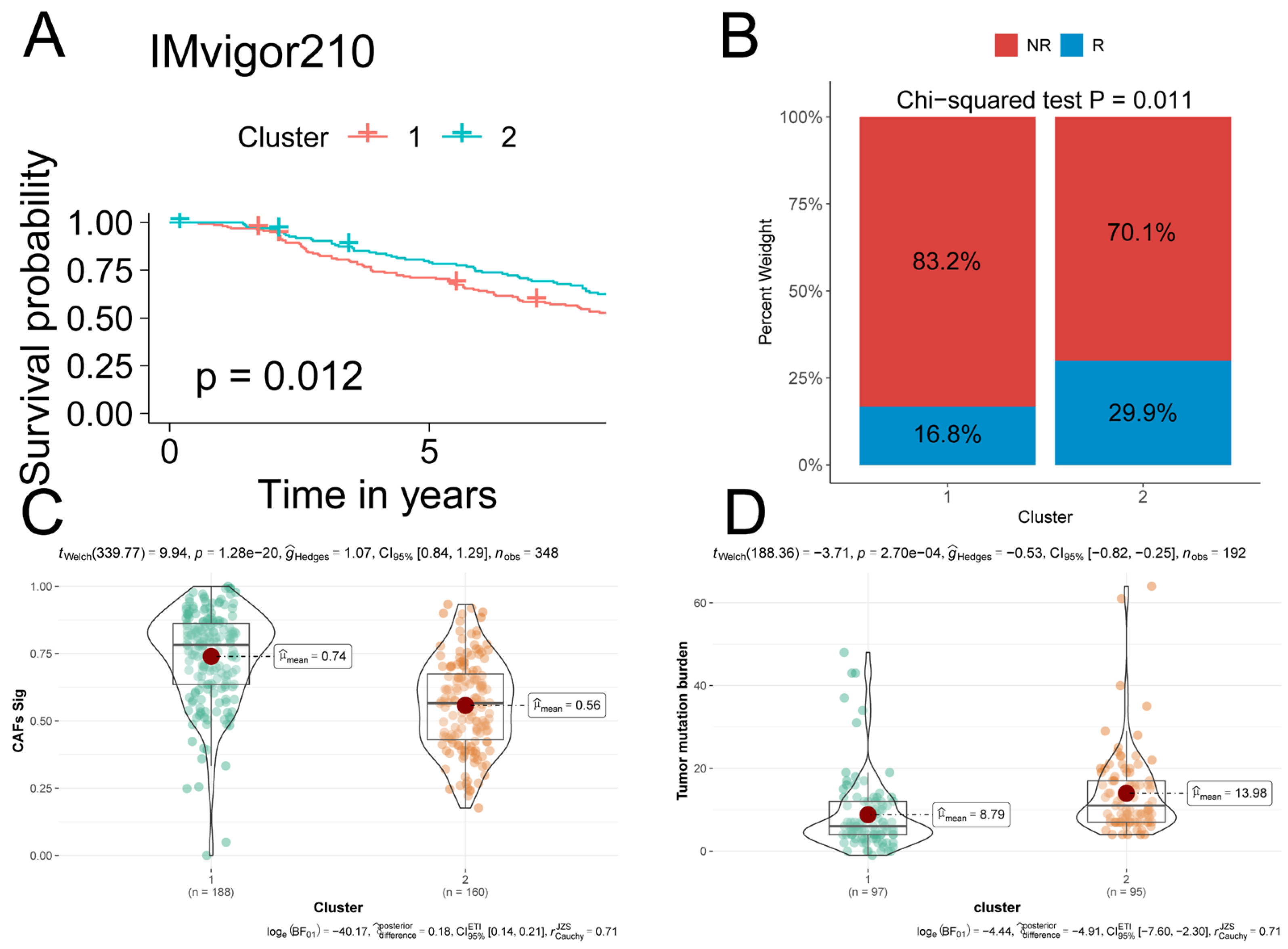
2.5. iCAF-related Subtypes Predicted Prognosis and Immunotherapeutic Response in ICI-Treated Cohorts
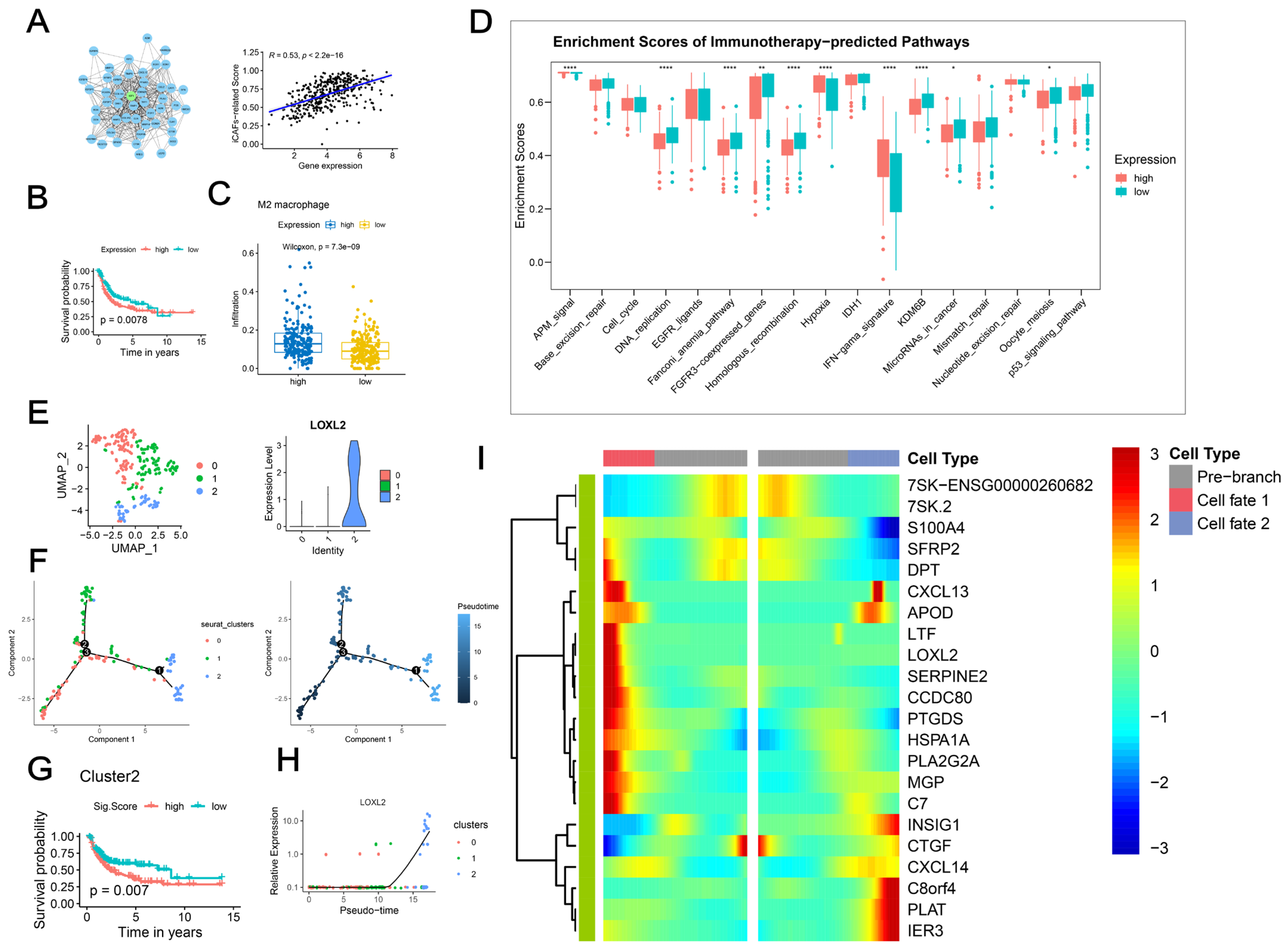
2.6. LOXL2+ iCAFs Predicted Poor Prognosis
2.7. Upregulated LOXL2 in CAFs Promoted the Proliferation, Migration, and Metastasis of BCa Cells
3. Discussion
4. Materials and Methods
4.1. Data Acquisition
4.2. iCAF-related Signature Identification
4.3. Enrichment Analyses
4.4. Unsupervised Clustering and Biological Properties Exploration
4.5. Hug Gene Identification
4.6. Cell Lines, Culture, Transfection, and Reagents
4.7. Quantitative Real-Time Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
4.8. Western Blotting and Antibodies
4.9. Conditioned Medium (CM) Derived from Fibroblasts
4.10. Cell Proliferation Assay
4.11. Wound-Healing and Transwell Assays
4.12. Enzyme-Linked Immunosorbent Assay (ELISA)
4.13. RNA-seq and Analyses
4.14. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Li, K.; Lin, T.; Xue, W.; Mu, X.; Xu, E.; Yang, X.; Chen, F.; Li, G.; Ma, L.; Wang, G.; et al. Current status of diagnosis and treatment of bladder cancer in China—Analyses of Chinese Bladder Cancer Consortium database. Asian J. Urol. 2015, 2, 63–69. [Google Scholar] [CrossRef]
- Saginala, K.; Barsouk, A.; Aluru, J.S.; Rawla, P.; Padala, S.A.; Barsouk, A. Epidemiology of Bladder Cancer. Med. Sci. 2020, 8, 15. [Google Scholar] [CrossRef]
- Stein, J.P.; Skinner, D.G. Radical cystectomy for invasive bladder cancer: Long-term results of a standard procedure. World J. Urol. 2006, 24, 296–304. [Google Scholar] [CrossRef]
- Bremnes, R.M.; Dønnem, T.; Al-Saad, S.; Al-Shibli, K.; Andersen, S.; Sirera, R.; Camps, C.; Marinez, I.; Busund, L.T. The role of tumor stroma in cancer progression and prognosis: Emphasis on carcinoma-associated fibroblasts and non-small cell lung cancer. J. Thorac. Oncol. 2011, 6, 209–217. [Google Scholar] [CrossRef]
- Mao, X.; Xu, J.; Wang, W.; Liang, C.; Hua, J.; Liu, J.; Zhang, B.; Meng, Q.; Yu, X.; Shi, S. Crosstalk between cancer-associated fibroblasts and immune cells in the tumor microenvironment: New findings and future perspectives. Mol. Cancer 2021, 20, 131. [Google Scholar] [CrossRef]
- Iwai, M.; Tulafu, M.; Togo, S.; Kawaji, H.; Kadoya, K.; Namba, Y.; Jin, J.; Watanabe, J.; Okabe, T.; Hidayat, M.; et al. Cancer-associated fibroblast migration in non-small cell lung cancers is modulated by increased integrin α11 expression. Mol. Oncol. 2021, 15, 1507–1527. [Google Scholar] [CrossRef]
- Chen, Z.; Zhou, L.; Liu, L.; Hou, Y.; Xiong, M.; Yang, Y.; Hu, J.; Chen, K. Single-cell RNA sequencing highlights the role of inflammatory cancer-associated fibroblasts in bladder urothelial carcinoma. Nat. Commun. 2020, 11, 5077. [Google Scholar] [CrossRef]
- Öhlund, D.; Handly-Santana, A.; Biffi, G.; Elyada, E.; Almeida, A.S.; Ponz-Sarvise, M.; Corbo, V.; Oni, T.E.; Hearn, S.A.; Lee, E.J.; et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 2017, 214, 579–596. [Google Scholar] [CrossRef]
- Jiang, Y.; Li, Y.; Zhu, B. T-cell exhaustion in the tumor microenvironment. Cell Death Dis. 2015, 6, e1792. [Google Scholar] [CrossRef]
- Zhang, Z.; Chen, L.; Chen, H.; Zhao, J.; Li, K.; Sun, J.; Zhou, M. Pan-cancer landscape of T-cell exhaustion heterogeneity within the tumor microenvironment revealed a progressive roadmap of hierarchical dysfunction associated with prognosis and therapeutic efficacy. eBioMedicine 2022, 83, 104207. [Google Scholar] [CrossRef]
- Wang, L.; Cao, L.; Wang, H.; Liu, B.; Zhang, Q.; Meng, Z.; Wu, X.; Zhou, Q.; Xu, K. Cancer-associated fibroblasts enhance metastatic potential of lung cancer cells through IL-6/STAT3 signaling pathway. Oncotarget 2017, 8, 76116–76128. [Google Scholar] [CrossRef]
- Li, G.; Satyamoorthy, K.; Meier, F.; Berking, C.; Bogenrieder, T.; Herlyn, M. Function and regulation of melanoma–stromal fibroblast interactions: When seeds meet soil. Oncogene 2003, 22, 3162–3171. [Google Scholar] [CrossRef]
- Linares, J.; Marín-Jiménez, J.A.; Badia-Ramentol, J.; Calon, A. Determinants and Functions of CAFs Secretome During Cancer Progression and Therapy. Front. Cell Dev. Biol. 2021, 8, 621070. [Google Scholar] [CrossRef]
- Peng, L.; Ran, Y.L.; Hu, H.; Yu, L.; Liu, Q.; Zhou, Z.; Sun, Y.M.; Sun, L.C.; Pan, J.; Sun, L.X.; et al. Secreted LOXL2 is a novel therapeutic target that promotes gastric cancer metastasis via the Src/FAK pathway. Carcinogenesis 2009, 30, 1660–1669. [Google Scholar] [CrossRef]
- Park, J.S.; Lee, J.H.; Lee, Y.S.; Kim, J.K.; Dong, S.M.; Yoon, D.S. Emerging role of LOXL2 in the promotion of pancreas cancer metastasis. Oncotarget 2016, 7, 42539–42552. [Google Scholar] [CrossRef]
- Tian, J.; Sun, H.X.; Li, Y.C.; Jiang, L.; Zhang, S.L.; Hao, Q. LOXL 2 Promotes The Epithelial-Mesenchymal Transition And Malignant Progression Of Cervical Cancer. Onco Targets Ther. 2019, 12, 8947–8954. [Google Scholar] [CrossRef]
- Peng, D.H.; Rodriguez, B.L.; Diao, L.; Chen, L.; Wang, J.; Byers, L.A.; Wei, Y.; Chapman, H.A.; Yamauchi, M.; Behrens, C.; et al. Collagen promotes anti-PD-1/PD-L1 resistance in cancer through LAIR1-dependent CD8+ T cell exhaustion. Nat. Commun. 2020, 11, 4520. [Google Scholar] [CrossRef]
- Gong, L.; Zhang, Y.; Yang, Y.; Yan, Q.; Ren, J.; Luo, J.; Tiu, Y.C.; Fang, X.; Liu, B.; Lam, R.H.W.; et al. Inhibition of lysyl oxidase-like 2 overcomes adhesion-dependent drug resistance in the collagen-enriched liver cancer microenvironment. Hepatol. Commun. 2022, 6, 3194–3211. [Google Scholar] [CrossRef]
- Wang, L.; Sebra, R.P.; Sfakianos, J.P.; Allette, K.; Wang, W.; Yoo, S.; Bhardwaj, N.; Schadt, E.E.; Yao, X.; Galsky, M.D.; et al. A reference profile-free deconvolution method to infer cancer cell-intrinsic subtypes and tumor-type-specific stromal profiles. Genome Med. 2020, 12, 24. [Google Scholar] [CrossRef]
- Sfakianos, J.P.; Daza, J.; Hu, Y.; Anastos, H.; Bryant, G.; Bareja, R.; Badani, K.K.; Galsky, M.D.; Elemento, O.; Faltas, B.M.; et al. Epithelial plasticity can generate multi-lineage phenotypes in human and murine bladder cancers. Nat. Commun. 2020, 11, 2540. [Google Scholar] [CrossRef]
- Leek, J.T.; Johnson, W.E.; Parker, H.S.; Jaffe, A.E.; Storey, J.D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 2012, 28, 882–883. [Google Scholar] [CrossRef]
- Mariathasan, S.; Turley, S.J.; Nickles, D.; Castiglioni, A.; Yuen, K.; Wang, Y.; Kadel III, E.E.; Koeppen, H.; Astarita, J.L.; Cubas, R. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 2018, 554, 544–548. [Google Scholar] [CrossRef]
- Hugo, W.; Zaretsky, J.M.; Sun, L.; Song, C.; Moreno, B.H.; Hu-Lieskovan, S.; Berent-Maoz, B.; Pang, J.; Chmielowski, B.; Cherry, G.; et al. Genomic and Transcriptomic Features of Response to Anti-PD-1 Therapy in Metastatic Melanoma. Cell 2016, 165, 35–44. [Google Scholar] [CrossRef]
- Kim, J.Y.; Choi, J.K.; Jung, H. Genome-wide methylation patterns predict clinical benefit of immunotherapy in lung cancer. Clin. Epigenet. 2020, 12, 119. [Google Scholar] [CrossRef]
- Pinto, J.P.; Kalathur, R.K.; Oliveira, D.V.; Barata, T.; Machado, R.S.; Machado, S.; Pacheco-Leyva, I.; Duarte, I.; Futschik, M.E. StemChecker: A web-based tool to discover and explore stemness signatures in gene sets. Nucleic Acids Res. 2015, 43, W72–W77. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Y.; Yang, T.; Xing, H.; Wang, Y.; Gao, L.; Guo, X.; Xing, B.; Wang, Y.; Ma, W. Machine learning revealed stemness features and a novel stemness-based classification with appealing implications in discriminating the prognosis, immunotherapy and temozolomide responses of 906 glioblastoma patients. Brief. Bioinform. 2021, 22, bbab032. [Google Scholar] [CrossRef]
- Yao, J.; Liu, Y.; Yang, J.; Li, M.; Li, S.; Zhang, B.; Yang, R.; Zhang, Y.; Cui, X.; Feng, C. Single-Cell Sequencing Reveals that DBI is the Key Gene and Potential Therapeutic Target in Quiescent Bladder Cancer Stem Cells. Front. Genet. 2022, 13, 904536. [Google Scholar] [CrossRef]
- Hao, Y.; Hao, S.; Andersen-Nissen, E.; Mauck, W.M.; Zheng, S.; Butler, A.; Lee, M.J.; Wilk, A.J.; Darby, C.; Zager, M.; et al. Integrated analysis of multimodal single-cell data. Cell 2021, 184, 3573–3587.e3529. [Google Scholar] [CrossRef]
- Hänzelmann, S.; Castelo, R.; Guinney, J. GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013, 14, 7. [Google Scholar] [CrossRef]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L. ClusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef]
- Wilkerson, M.D.; Hayes, D.N. ConsensusClusterPlus: A class discovery tool with confidence assessments and item tracking. Bioinformatics 2010, 26, 1572–1573. [Google Scholar] [CrossRef] [PubMed]
- Van der Maaten, L.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- Yoshihara, K.; Shahmoradgoli, M.; Martínez, E.; Vegesna, R.; Kim, H.; Torres-Garcia, W.; Treviño, V.; Shen, H.; Laird, P.W.; Levine, D.A.; et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 2013, 4, 2612. [Google Scholar] [CrossRef]
- Chen, B.; Khodadoust, M.S.; Liu, C.L.; Newman, A.M.; Alizadeh, A.A. Profiling Tumor Infiltrating Immune Cells with CIBERSORT. Methods Mol. Biol. 2018, 1711, 243–259. [Google Scholar] [CrossRef]
- Finotello, F.; Mayer, C.; Plattner, C.; Laschober, G.; Rieder, D.; Hackl, H.; Krogsdam, A.; Loncova, Z.; Posch, W.; Wilflingseder, D.; et al. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 2019, 11, 34. [Google Scholar] [CrossRef]
- Li, T.; Fu, J.; Zeng, Z.; Cohen, D.; Li, J.; Chen, Q.; Li, B.; Liu, X.S. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020, 48, W509–W514. [Google Scholar] [CrossRef]
- Becht, E.; Giraldo, N.A.; Lacroix, L.; Buttard, B.; Elarouci, N.; Petitprez, F.; Selves, J.; Laurent-Puig, P.; Sautès-Fridman, C.; Fridman, W.H.; et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 2016, 17, 218. [Google Scholar] [CrossRef]
- Zeng, D.; Ye, Z.; Shen, R.; Yu, G.; Wu, J.; Xiong, Y.; Zhou, R.; Qiu, W.; Huang, N.; Sun, L.; et al. IOBR: Multi-Omics Immuno-Oncology Biological Research to Decode Tumor Microenvironment and Signatures. Front. Immunol. 2021, 12, 687975. [Google Scholar] [CrossRef]
- Robertson, A.G.; Kim, J.; Al-Ahmadie, H.; Bellmunt, J.; Guo, G.; Cherniack, A.D.; Hinoue, T.; Laird, P.W.; Hoadley, K.A.; Akbani, R. Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 2017, 171, 540–556.e525. [Google Scholar] [CrossRef]
- Damrauer, J.S.; Hoadley, K.A.; Chism, D.D.; Fan, C.; Tiganelli, C.J.; Wobker, S.E.; Yeh, J.J.; Milowsky, M.I.; Iyer, G.; Parker, J.S. Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proc. Natl. Acad. Sci. USA 2014, 111, 3110–3115. [Google Scholar] [CrossRef]
- Kamoun, A.; de Reyniès, A.; Allory, Y.; Sjödahl, G.; Robertson, A.G.; Seiler, R.; Hoadley, K.A.; Groeneveld, C.S.; Al-Ahmadie, H.; Choi, W.; et al. A Consensus Molecular Classification of Muscle-invasive Bladder Cancer. Eur. Urol. 2020, 77, 420–433. [Google Scholar] [CrossRef] [PubMed]
- Geeleher, P.; Cox, N.; Huang, R.S. pRRophetic: An R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE 2014, 9, e107468. [Google Scholar] [CrossRef] [PubMed]
- Reich, M.; Liefeld, T.; Gould, J.; Lerner, J.; Tamayo, P.; Mesirov, J.P. GenePattern 2.0. Nat. Genet. 2006, 38, 500–501. [Google Scholar] [CrossRef] [PubMed]
- Wen, S.; Hou, Y.; Fu, L.; Xi, L.; Yang, D.; Zhao, M.; Qin, Y.; Sun, K.; Teng, Y.; Liu, M. Cancer-associated fibroblast (CAF)-derived IL32 promotes breast cancer cell invasion and metastasis via integrin β3-p38 MAPK signalling. Cancer Lett. 2019, 442, 320–332. [Google Scholar] [CrossRef]
- Kasashima, H.; Yashiro, M.; Kinoshita, H.; Fukuoka, T.; Morisaki, T.; Masuda, G.; Sakurai, K.; Kubo, N.; Ohira, M.; Hirakawa, K. Lysyl oxidase-like 2 (LOXL2) from stromal fibroblasts stimulates the progression of gastric cancer. Cancer Lett. 2014, 354, 438–446. [Google Scholar] [CrossRef]
- Chen, H.; Chen, G. Dissecting Immunosuppressive Cell Communication Patterns Reveals JunB Proto-Oncogene (JUNB) Shaping a Non-Inflamed Tumor Microenvironment. Front. Genet. 2022, 13, 883583. [Google Scholar] [CrossRef]
- Ihaka, R.; Gentleman, R. R: A language for data analysis and graphics. J. Comput. Graph. Stat. 1996, 5, 299–314. [Google Scholar]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, H.; Yang, W.; Xue, X.; Li, Y.; Jin, Z.; Ji, Z. Integrated Analysis Revealed an Inflammatory Cancer-Associated Fibroblast-Based Subtypes with Promising Implications in Predicting the Prognosis and Immunotherapeutic Response of Bladder Cancer Patients. Int. J. Mol. Sci. 2022, 23, 15970. https://doi.org/10.3390/ijms232415970
Chen H, Yang W, Xue X, Li Y, Jin Z, Ji Z. Integrated Analysis Revealed an Inflammatory Cancer-Associated Fibroblast-Based Subtypes with Promising Implications in Predicting the Prognosis and Immunotherapeutic Response of Bladder Cancer Patients. International Journal of Molecular Sciences. 2022; 23(24):15970. https://doi.org/10.3390/ijms232415970
Chicago/Turabian StyleChen, Hualin, Wenjie Yang, Xiaoqiang Xue, Yingjie Li, Zhaoheng Jin, and Zhigang Ji. 2022. "Integrated Analysis Revealed an Inflammatory Cancer-Associated Fibroblast-Based Subtypes with Promising Implications in Predicting the Prognosis and Immunotherapeutic Response of Bladder Cancer Patients" International Journal of Molecular Sciences 23, no. 24: 15970. https://doi.org/10.3390/ijms232415970
APA StyleChen, H., Yang, W., Xue, X., Li, Y., Jin, Z., & Ji, Z. (2022). Integrated Analysis Revealed an Inflammatory Cancer-Associated Fibroblast-Based Subtypes with Promising Implications in Predicting the Prognosis and Immunotherapeutic Response of Bladder Cancer Patients. International Journal of Molecular Sciences, 23(24), 15970. https://doi.org/10.3390/ijms232415970





