Two-Component Systems of Streptomyces coelicolor: An Intricate Network to Be Unraveled
Abstract
:1. Streptomyces, the Biotech Gold Mine
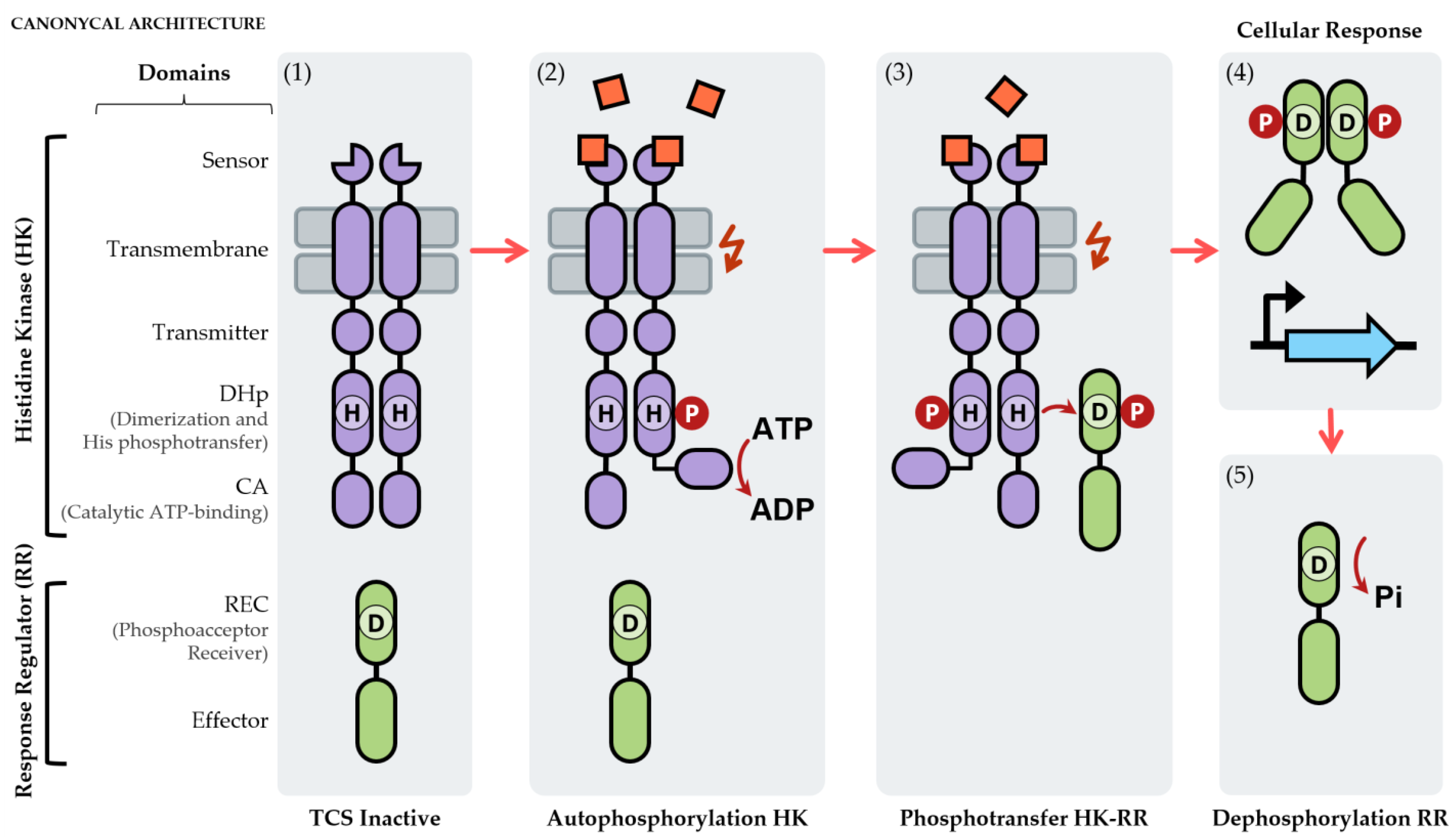
2. Two-Component Systems: Essential Signal Transduction Systems for Bacteria
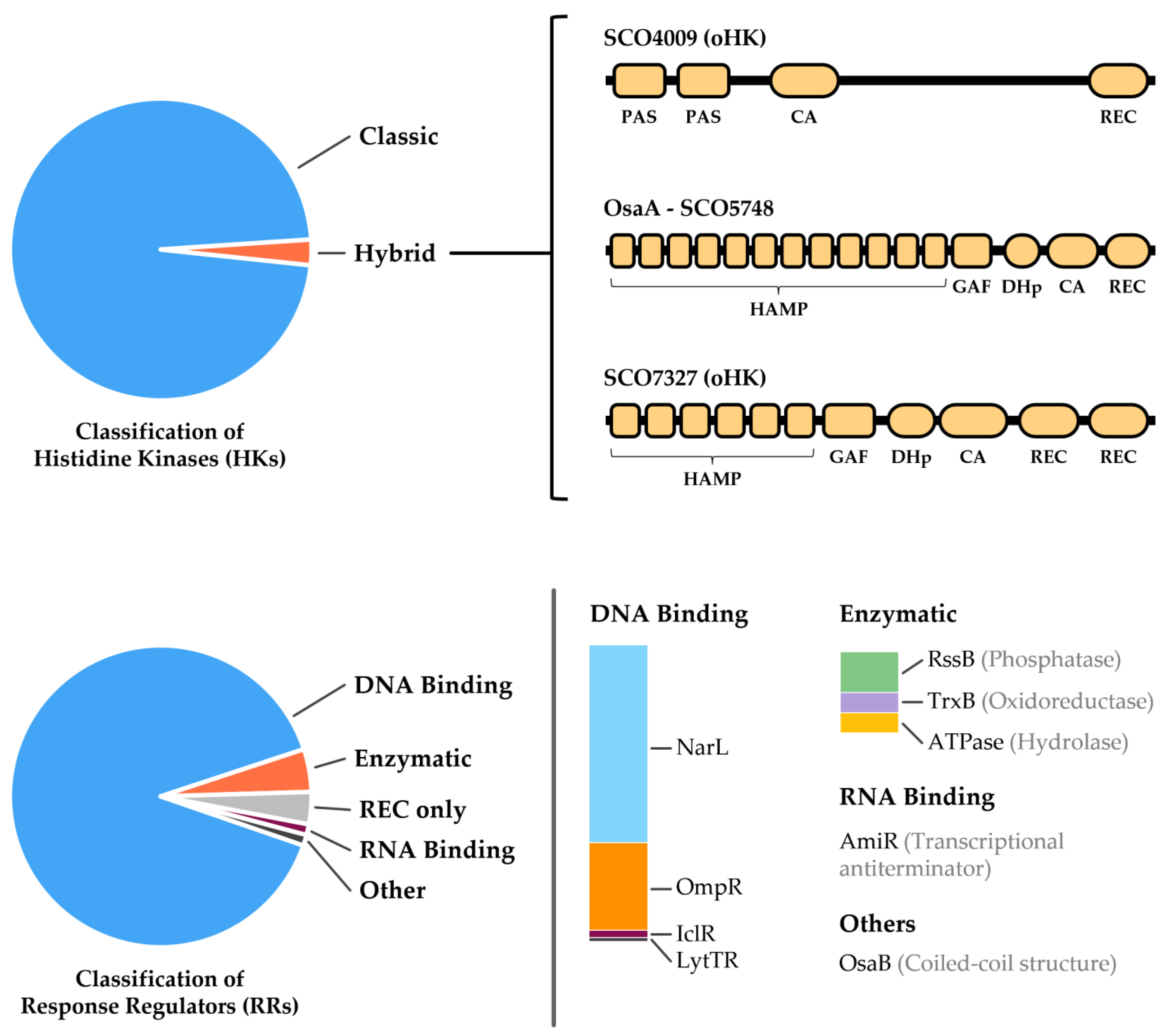
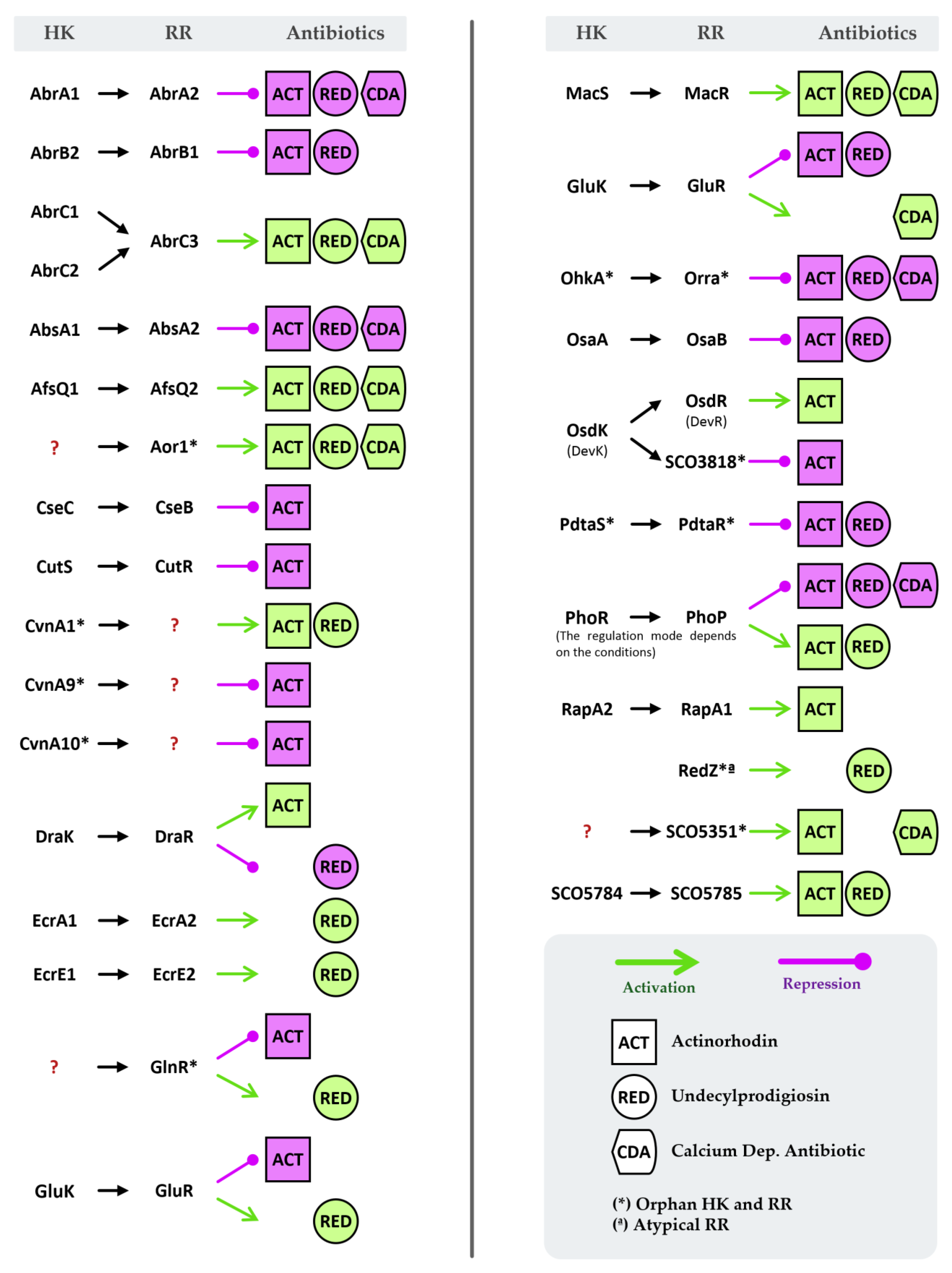
3. Two-Component Systems of Streptomyces coelicolor
3.1. AbrA1/A2 (SCO1744/45)
3.2. AbrB1/B2 (SCO2165/66)
3.3. AbrC1/C2/C3 (SCO4598/97/96)
3.4. AbsA1/A2 (SCO3225/26)
3.5. AfsQ2/Q1 (SCO4906/07)
3.6. Aor1 (SCO2281)
3.7. BldM (SCO4768)
3.8. ChiS/R (SCO5378/77)
3.9. CseC/B (SCO3359/58)
3.10. CssS/R (SCO4155/56)
3.11. CutS/R (SCO5863/62)
3.12. Cvn (Conservons)
3.13. DraK/R (SCO3062/63)
3.14. EcrA1/A2 (SCO2518/17)
3.15. EcrE1/E2 (SCO6421/22)
3.16. GlnR (SCO4159)
3.17. GluK/R (SCO5779/78)
3.18. MacS/R (SCO2121/20)
3.19. MtrB/A (SCO3012/13)
3.20. OhkA (SCO1596) and OrrA (SCO3008)
3.21. OsaA/B (SCO5748/49)
3.22. OsdK/R—DevS/R (SCO0203/04)
3.23. PdtaS (SCO5239) and PdtaR (SCO2013)
3.24. PhoR/P (SCO4229/30)
3.25. RagK/R (SCO4073/72)
3.26. RamR (SCO6685)
3.27. RapA2/A1 (SCO5404/03)
3.28. RedZ (SCO5881)
3.29. SatK/R (SCO3390/89)
3.30. SenS/R (SCO4275/76)
3.31. SitK/R (SCO4667/68)
3.32. VanS/R (SCO3589/90)
3.33. WhiI (SCO6029)
3.34. SCO5282/83
3.35. SCO5351
3.36. SCO5784/85
4. Conclusions
4.1. Regulation Systems Involved in all Cellular Processes
4.2. Molecular Mechanism of Action and its Modulation
4.3. Biotechnological Application
4.4. Future Considerations
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Barka, E.A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Klenk, H.P.; Clement, C.; Ouhdouch, Y.; van Wezel, G.P. Taxonomy, Physiology, and Natural Products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2016, 80, 1–43. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, M.; Kumar, P.; Das, P.; Solanki, R.; Kapur, M.K. Potential applications of extracellular enzymes from Streptomyces spp. in various industries. Arch. Microbiol. 2020, 202, 1597–1615. [Google Scholar] [CrossRef] [PubMed]
- Nett, M.; Ikeda, H.; Moore, B.S. Genomic basis for natural product biosynthetic diversity in the actinomycetes. Nat. Prod. Rep. 2009, 26, 1362–1384. [Google Scholar] [CrossRef] [PubMed]
- Rutledge, P.; Challis, G. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat. Rev. Genet. 2015, 13, 509–523. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, W.; Schumann, P.; Busse, H.J.; Trujillo, M.E.; Kämpfer, P.; Whitman, W.B. Road map of the phyllum Actinobacteria. In Bergey’s Manual of Systematics of Archaea and Bacteria; John Wiley & Sons, Inc., in Association with Bergey’s Manual Trust, 2015. [Google Scholar] [CrossRef]
- Nouioui, I.; Carro, L.; García-López, M.; Meier-Kolthoff, J.P.; Woyke, T.; Kyrpides, N.C.; Pukall, R.; Klenk, H.-P.; Goodfellow, M.; Göker, M. Genome-Based Taxonomic Classification of the Phylum Actinobacteria. Front. Microbiol. 2018, 9, 2007. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kämpfer, P. Streptomyces. In Bergey’s Manual of Systematics of Archaea and Bacteria; John Wiley & Sons, Inc., in Association with Bergey’s Manual Trust, 2015. [Google Scholar] [CrossRef]
- Chater, K.F. Recent advances in understanding Streptomyces. F1000Res 2016, 5, 2795. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kirby, R. Chromosome diversity and similarity within the Actinomycetales. FEMS Microbiol. Lett. 2011, 319, 1–10. [Google Scholar] [CrossRef]
- van Bergeijk, D.A.; Terlouw, B.R.; Medema, M.H.; van Wezel, G.P. Ecology and genomics of Actinobacteria: New concepts for natural product discovery. Nat. Rev. Microbiol. 2020, 18, 546–558. [Google Scholar] [CrossRef]
- Dalisay, D.; Williams, D.E.; Wang, X.L.; Centko, R.; Chen, J.; Andersen, R.J. Marine Sediment-Derived Streptomyces Bacteria from British Columbia, Canada Are a Promising Microbiota Resource for the Discovery of Antimicrobial Natural Products. PLoS ONE 2013, 8, e77078. [Google Scholar] [CrossRef]
- Sabido, E.M.; Tenebro, C.P.; Suarez, A.F.L.; Ong, S.D.C.; Trono, D.J.V.L.; Amago, D.S.; Evangelista, J.J.E.; Reynoso, A.M.Q.; Villalobos, I.G.M.; Alit, L.D.D.; et al. Marine Sediment-Derived Streptomyces Strain Produces Angucycline Antibiotics against Multidrug-Resistant Staphylococcus aureus Harboring SCCmec Type 1 Gene. J. Mar. Sci. Eng. 2020, 8, 734. [Google Scholar] [CrossRef]
- Tenebro, C.P.; Trono, D.; Vicera, C.V.B.; Sabido, E.M.; Ysulat, J.A., Jr.; Macaspac, A.J.M.; Tampus, K.A.; Fabrigar, T.A.P.; Saludes, J.P.; Dalisay, D.S. Multiple strain analysis of {Sabido, 2020 #764} species from Philippine marine sediments reveals intraspecies heterogeneity in antibiotic activities. Sci. Rep. 2021, 11, 17544. [Google Scholar]
- Chevrette, M.G.; Carlson, C.M.; Ortega-Domínguez, H.; Thomas, C.; Ananiev, G.E.; Barns, K.J.; Book, A.J.; Cagnazzo, J.; Carlos, C.; Flanigan, W.; et al. The antimicrobial potential of Streptomyces from insect microbiomes. Nat. Commun. 2019, 10, 516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santamaría, R.I.; Martínez-Carrasco, A.; De La Nieta, R.S.; Torres-Vila, L.M.; Bonal, R.; Martín, J.; Tormo, R.; Reyes, F.; Genilloud, O.; Díaz, M. Characterization of Actinomycetes Strains Isolated from the Intestinal Tract and Feces of the Larvae of the Longhorn Beetle Cerambyx welensii. Microorganisms 2020, 8, 2013. [Google Scholar] [CrossRef] [PubMed]
- Currie, C.R.; Bot, A.N.M.; Boomsma, J.J. Experimental evidence of a tripartite mutualism: Bacteria portect ant fungus gardens from spcecialized parasites. OIKOS 2003, 101, 91–102. [Google Scholar] [CrossRef] [Green Version]
- Currie, C.R.; Scott, J.A.; Summerbell, R.C.; Malloch, D. Fungus-growing ants use antibiotic-producing bacteria to control garden parasites. Nature 1999, 398, 701–704. [Google Scholar] [CrossRef]
- Hulcr, J.; Adams, A.S.; Raffa, K.; Hofstetter, R.W.; Klepzig, K.D.; Currie, C.R. Presence and Diversity of Streptomyces in Dendroctonus and Sympatric Bark Beetle Galleries Across North America. Microb. Ecol. 2011, 61, 759–768. [Google Scholar] [CrossRef]
- Vasanthakumar, A.; Handelsman, J.; Schloss, P.D.; Bauer, L.S.; Raffa, K.F. Gut microbiota of an invasive subcortical beetle, Agrilus planipennis Fairmaire, across various life stages. Environ. Entomol. 2008, 37, 1344–1353. [Google Scholar] [CrossRef]
- Romano-Armada, N.; Yañez-Yazlle, M.F.; Irazusta, V.P.; Rajal, V.B.; Moraga, N.B. Potential of Bioremediation and PGP Traits in Streptomyces as Strategies for Bio-Reclamation of Salt-Affected Soils for Agriculture. Pathogens 2020, 9, 117. [Google Scholar] [CrossRef] [Green Version]
- Tran, T.M.; Ameye, M.; Devlieghere, F.; De Saeger, S.; Eeckhout, M.; Audenaert, K. Streptomyces Strains Promote Plant Growth and Induce Resistance Against Fusarium verticillioides via Transient Regulation of Auxin Signaling and Archetypal Defense Pathways in Maize Plants. Front. Plant Sci. 2021, 12, 755733. [Google Scholar] [CrossRef]
- Wanner, L.A. A Survey of Genetic Variation in Streptomyces Isolates Causing Potato Common Scab in the United States. Phytopathology 2006, 96, 1363–1371. [Google Scholar] [CrossRef] [Green Version]
- Kirby, R.; Sangal, V.; Tucker, N.P.; Zakrzewska-Czerwińska, J.; Wierzbicka, K.; Herron, P.R.; Chu, C.-J.; Chandra, G.; Fahal, A.H.; Goodfellow, M.; et al. Draft Genome Sequence of the Human Pathogen Streptomyces somaliensis, a Significant Cause of Actinomycetoma. J. Bacteriol. 2012, 194, 3544–3545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quintana, E.T.; Wierzbicka, K.; Mackiewicz, P.; Osman, A.; Fahal, A.; Hamid, M.; Zakrzewska-Czerwinska, J.; Maldonado, L.A.; Goodfellow, M. Streptomyces sudanensis sp. nov., a new pathogen isolated from patients with actinomycetoma. Antonie van Leeuwenhoek 2007, 93, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeño-Tárraga, A.-M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef]
- Hopwood, D.A. Forty years of genetics with Streptomyces: From in vivo through in vitro to in silico. Microbiology 1999, 145, 2183–2202. [Google Scholar] [CrossRef] [Green Version]
- Stock, A.M.; Robinson, V.L.; Goudreau, P.N. Two-component signal transduction. Annu. Rev. Biochem. 2000, 69, 183–215. [Google Scholar] [CrossRef] [Green Version]
- Wang, S. Bacterial two-component systems: Structures and signaling mechanisms. In Prot Phosph Human Health; Huang, C., Ed.; Intech Open: London, UK, 2012. [Google Scholar]
- Bekker, M.; De Mattos, M.J.T.; Hellingwerf, K.J. The Role of Two-Component Regulation Systems in the physiology of the Bacterial Cell. Sci. Prog. 2006, 89, 213–242. [Google Scholar] [CrossRef]
- Jacob-Dubuisson, F.; Mechaly, A.; Betton, J.-M.; Antoine, R. Structural insights into the signalling mechanisms of two-component systems. Nat. Rev. Genet. 2018, 16, 585–593. [Google Scholar] [CrossRef]
- Zschiedrich, C.P.; Keidel, V.; Szurmant, H. Molecular Mechanisms of Two-Component Signal Transduction. J. Mol. Biol. 2016, 428, 3752–3775. [Google Scholar] [CrossRef] [Green Version]
- Krell, T.; Lacal, J.; Busch, A.; Silva-Jiménez, H.; Guazzaroni, M.-E.; Ramos, J.L. Bacterial Sensor Kinases: Diversity in the Recognition of Environmental Signals. Annu. Rev. Microbiol. 2010, 64, 539–559. [Google Scholar] [CrossRef] [PubMed]
- Tierney, A.R.; Rather, P.N. Roles of two-component regulatory systems in antibiotic resistance. Future Microbiol. 2019, 14, 533–552. [Google Scholar] [CrossRef] [PubMed]
- Beier, D. Deviations from the rule: Orphan and atypical response regulators. In Two-Component System in Bacteria; Caister Academic Press: Poole, UK, 2012. [Google Scholar]
- Galperin, M.Y. Structural Classification of Bacterial Response Regulators: Diversity of Output Domains and Domain Combinations. J. Bacteriol. 2006, 188, 4169–4182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Galperin, M.Y. Diversity of structure and function of response regulator output domains. Curr. Opin. Microbiol. 2010, 13, 150–159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Groisman, E.A. Feedback Control of Two-Component Regulatory Systems. Annu. Rev. Microbiol. 2016, 70, 103–124. [Google Scholar] [CrossRef] [PubMed]
- Buschiazzo, A.; Trajtenberg, F. Two-Component Sensing and Regulation: How Do Histidine Kinases Talk with Response Regulators at the Molecular Level? Annu. Rev. Microbiol. 2019, 73, 507–528. [Google Scholar] [CrossRef] [PubMed]
- Laub, M.T.; Goulian, M. Specificity in Two-Component Signal Transduction Pathways. Annu. Rev. Genet. 2007, 41, 121–145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rowland, M.A.; Deeds, E.J. Crosstalk and the evolution of specificity in two-component signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 5550–5555. [Google Scholar] [CrossRef] [Green Version]
- Li, L.; Zhao, Y.; Ma, J.; Tao, H.; Zheng, G.; Chen, J.; Jiang, W.; Lu, Y. The orphan histidine kinase PdtaS-p regulates both morphological differentiation and antibiotic biosynthesis together with the orphan response regulator PdtaR-p in Streptomyces. Microbiol. Res. 2020, 233, 126411. [Google Scholar] [CrossRef]
- Zheng, G.; Liu, P.; He, W.; Tao, H.; Yang, Z.; Sun, C.; Wang, W.; Lu, Y.; Jiang, W. Identification of the cognate response regulator of the orphan histidine kinase OhkA involved in both secondary metabolism and morphological differentiation in Streptomyces coelicolor. Appl. Microbiol. Biotechnol. 2021, 105, 5905–5914. [Google Scholar] [CrossRef]
- Wang, W.; Shu, D.; Chen, L.; Jiang, W.; Lu, Y. Cross-talk between an orphan response regulator and a noncognate histidine kinase in Streptomyces coelicolor. FEMS Microbiol. Lett. 2009, 294, 150–156. [Google Scholar] [CrossRef]
- Yepes, A.; Rico, S.; Rodríguez-García, A.; Santamaria, R.I.; Díaz, M. Novel Two-Component Systems Implied in Antibiotic Production in Streptomyces coelicolor. PLoS ONE 2011, 6, e19980. [Google Scholar] [CrossRef] [Green Version]
- Rico, S.; Yepes, A.; Rodríguez, H.; Santamaría, J.; Antoraz, S.; Krause, E.M.; Díaz, M.; Santamaría, R.I. Regulation of the AbrA1/A2 two-component system in Streptomyces coelicolor and the potential of its deletion strain as a heterologous host for antibiotic production. PLoS ONE 2014, 9, e109844. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sánchez de la Nieta, R.; Antoraz, S.; Alzate, J.F.; Santamaría, R.I.; Díaz, M. Antibiotic Production and Antibiotic Resistance: The Two Sides of AbrB1/B2, a Two-Component System of Streptomyces coelicolor. Front. Microbiol. 2020, 11, 587750. [Google Scholar] [CrossRef] [PubMed]
- Rico, S.; Santamaria, R.I.; Yepes, A.; Rodríguez, H.; Laing, E.; Bucca, G.; Smith, C.P.; Díaz, M. Deciphering the Regulon of Streptomyces coelicolor AbrC3, a Positive Response Regulator of Antibiotic Production. Appl. Environ. Microbiol. 2014, 80, 2417–2428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, R.; Mast, Y.; Wang, J.; Zhang, W.; Zhao, G.; Wohlleben, W.; Lu, Y.; Jiang, W. Identification of two-component system AfsQ1/Q2 regulon and its cross-regulation with GlnR in Streptomyces coelicolor. Mol. Microbiol. 2013, 87, 30–48. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, H.; Rico, S.; Yepes, A.; Franco-Echevarría, E.; Antoraz, S.; Santamaría, R.I.; Díaz, M. The two kinases, AbrC1 and AbrC2, of the atypical two-component system AbrC are needed to regulate antibiotic production and differentiation in Streptomyces coelicolor. Front. Microbiol. 2015, 6, 450. [Google Scholar] [CrossRef]
- Wang, G.; Izawa, M.; Yang, X.; Xu, D.; Tanaka, Y.; Ochi, K. Identification of a Novel Lincomycin Resistance Mutation Associated with Activation of Antibiotic Production in Streptomyces coelicolor A3(2). Antimicrob. Agents Chemother. 2017, 61, e02247-16. [Google Scholar] [CrossRef] [Green Version]
- Adamidis, T.; Riggle, P.; Champness, W. Mutations in a new Streptomyces coelicolor locus which globally block antibiotic biosynthesis but not sporulation. J. Bacteriol. 1990, 172, 2962–2969. [Google Scholar] [CrossRef] [Green Version]
- Anderson, T.; Brian, P.; Riggle, P.; Kong, R.; Champness, W. Genetic suppression analysis of non-antibiotic-producing mutants of the Streptomyces coelicolor absA locus. Microbiology 1999, 145, 2343–2353. [Google Scholar] [CrossRef] [Green Version]
- Anderson, T.B.; Brian, P.; Champness, W.C. Genetic and transcriptional analysis of absA, an antibiotic gene cluster-linked two-component system that regulates multiple antibiotics in Streptomyces coelicolor. Mol. Microbiol. 2001, 39, 553–566. [Google Scholar] [CrossRef]
- Brian, P.; Riggle, P.J.; Santos, R.A.; Champness, W.C. Global negative regulation of Streptomyces coelicolor antibiotic synthesis mediated by an absA-encoded putative signal transduction system. J. Bacteriol. 1996, 178, 3221–3231. [Google Scholar] [CrossRef] [Green Version]
- Lewis, R.A.; Wahab, A.; Bucca, G.; Laing, E.E.; Möller-Levet, C.S.; Kierzek, A.; Smith, C.P. Genome-wide analysis of the role of the antibiotic biosynthesis regulator AbsA2 in Streptomyces coelicolor A3(2). PLoS ONE 2019, 14, e0200673. [Google Scholar] [CrossRef] [Green Version]
- Ryding, N.J.; Anderson, T.B.; Champness, W.C. Regulation of the Streptomyces coelicolor calcium-dependent antibiotic by absA, encoding a cluster-linked two-component system. J. Bacteriol. 2002, 184, 794–805. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKenzie, N.L.; Nodwell, J.R. Phosphorylated AbsA2 Negatively Regulates Antibiotic Production in Streptomyces coelicolor through Interactions with Pathway-Specific Regulatory Gene Promoters. J. Bacteriol. 2007, 189, 5284–5292. [Google Scholar] [CrossRef] [Green Version]
- Sheeler, N.L.; MacMillan, S.V.; Nodwell, J.R. Biochemical Activities of the absA Two-Component System of Streptomyces coelicolor. J. Bacteriol. 2005, 187, 687–696. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos-Beneit, F.; Rodríguez-García, A.; Martín, J.F. Identification of different promoters in the absA1-absA2 two-component system, a negative regulator of antibiotic production in Streptomyces coelicolor. Mol. Genet. Genom. 2013, 288, 39–48. [Google Scholar] [CrossRef] [PubMed]
- McKenzie, N.L.; Nodwell, J.R. Transmembrane topology of the AbsA1 sensor kinase of Streptomyces coelicolor. Microbiology 2009, 155, 1812–1818. [Google Scholar] [CrossRef] [Green Version]
- McKenzie, N.L.; Thaker, M.; Koteva, K.; Hughes, D.W.; Wright, G.D.; Nodwell, J.R. Induction of antimicrobial activities in heterologous streptomycetes using alleles of the Streptomyces coelicolor gene absA1. J. Antibiot. 2010, 63, 177–182. [Google Scholar] [CrossRef] [Green Version]
- Chen, S.; Zheng, G.; Zhu, H.; He, H.; Chen, L.; Zhang, W.; Jiang, W.; Lu, Y. Roles of two-component system AfsQ1/Q2 in regulating biosynthesis of the yellow-pigmented coelimycin P2 in Streptomyces coelicolor. FEMS Microbiol. Lett. 2016, 363, fnw160. [Google Scholar] [CrossRef] [Green Version]
- Ishizuka, H.; Horinouchi, S.; Kieser, H.M.; Hopwood, D.A.; Beppu, T. A putative two-component regulatory system involved in secondary metabolism in Streptomyces spp. J. Bacteriol. 1992, 174, 7585–7594. [Google Scholar] [CrossRef] [PubMed]
- Shu, D.; Chen, L.; Wang, W.; Yu, Z.; Ren, C.; Zhang, W.; Yang, S.; Lu, Y.; Jiang, W. afsQ1-Q2-sigQ is a pleiotropic but conditionally required signal transduction system for both secondary metabolism and morphological development in Streptomyces coelicolor. Appl. Microbiol. Biotechnol. 2008, 81, 1149–1160. [Google Scholar] [CrossRef]
- Antoraz, S.; Rico, S.; Rodríguez, H.; Sevillano, L.; Alzate, J.F.; Santamaria, R.I.; Díaz, M. The Orphan Response Regulator Aor1 Is a New Relevant Piece in the Complex Puzzle of Streptomyces coelicolor Antibiotic Regulatory Network. Front. Microbiol. 2017, 8, 2444. [Google Scholar] [CrossRef]
- Molle, V.; Buttner, M. Different alleles of the response regulator gene bldM arrest Streptomyces coelicolor development at distinct stages. Mol. Microbiol. 2002, 36, 1265–1278. [Google Scholar] [CrossRef]
- Ryding, N.J.; Bibb, M.J.; Molle, V.; Findlay, K.C.; Chater, K.F.; Buttner, M.J. New Sporulation Loci in Streptomyces coelicolor A3(2). J. Bacteriol. 1999, 181, 5419–5425. [Google Scholar] [CrossRef] [Green Version]
- Bibb, M.J.; Molle, V.; Buttner, M.J. sigma(BldN), an extracytoplasmic function RNA polymerase sigma factor required for aerial mycelium formation in Streptomyces coelicolor A3(2). J. Bacteriol. 2000, 182, 4606–4616. [Google Scholar] [CrossRef] [Green Version]
- Gehring, A.M.; Wang, S.T.; Kearns, D.B.; Storer, N.Y.; Losick, R. Novel Genes That Influence Development in Streptomyces coelicolor. J. Bacteriol. 2004, 186, 3570–3577. [Google Scholar] [CrossRef] [Green Version]
- Al-Bassam, M.M.; Bibb, M.J.; Bush, M.J.; Chandra, G.; Buttner, M.J. Response Regulator Heterodimer Formation Controls a Key Stage in Streptomyces Development. PLoS Genet. 2014, 10, e1004554. [Google Scholar] [CrossRef] [Green Version]
- Homerová, D.; Knirschová, R.; Kormanec, J. Response regulator ChiR regulates expression of chitinase gene, chiC, in Streptomyces coelicolor. Folia Microbiol. 2002, 47, 499–505. [Google Scholar] [CrossRef]
- Kormanec, J.; Ševčíková, B.; Homérová, D. Cloning of a two-component regulatory system probably involved in the regulation of chitinase inStreptomyces cœlicolor A3(2). Folia Microbiol. 2000, 45, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.-J.; Paget, M.; Buttner, M. A signal transduction system in Streptomyces coelicolor that activates the expression of a putative cell wall glycan operon in response to vancomycin and other cell wall-specific antibiotics. Mol. Microbiol. 2002, 44, 1199–1211. [Google Scholar] [CrossRef] [PubMed]
- Paget, M.S.B.; Leibovitz, E.; Buttner, M. A putative two-component signal transduction system regulates sigmaE, a sigma factor required for normal cell wall integrity in Streptomyces coelicolor A3(2). Mol. Microbiol. 1999, 33, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.-J. Construction of a Bioassay System to Identify Extracellular Agents Targeting Bacterial Cell Envelope. Methods Mol. Biol. 2016, 1440, 125–137. [Google Scholar] [CrossRef] [PubMed]
- Hutchings, M.I.; Hong, H.J.; Leibovitz, E.; Sutcliffe, I.C.; Buttner, M.J. The sigma(E) cell envelope stress response of Streptomyces coelicolor is influenced by a novel lipoprotein, CseA. J. Bacteriol. 2006, 188, 7222–7229. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gullón, S.; Vicente, R.L.; Mellado, R.P. A Novel Two-Component System Involved in Secretion Stress Response in Streptomyces lividans. PLoS ONE 2012, 7, e48987. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, H.M.; Chen, M.Y.; Shieh, Y.T.; Bibb, M.J.; Chen, C.W. The cutRS signal transduction system of Streptomyces lividans represses the biosynthesis of the polyketide antibiotic actinorhodin. Mol. Microbiol. 1996, 21, 1075–1085. [Google Scholar]
- Takano, H.; Hashimoto, K.; Yamamoto, Y.; Beppu, T.; Ueda, K. Pleiotropic effect of a null mutation in the cvn1 conservon of Streptomyces coelicolor A3(2). Gene 2011, 477, 12–18. [Google Scholar] [CrossRef]
- Komatsu, M.; Takano, H.; Hiratsuka, T.; Ishigaki, Y.; Shimada, K.; Beppu, T.; Ueda, K. Proteins encoded by the conservon of Streptomyces coelicolor A3(2) comprise a membrane-associated heterocomplex that resembles eukaryotic G protein-coupled regulatory system. Mol. Microbiol. 2006, 62, 1534–1546. [Google Scholar] [CrossRef]
- Yu, Z.; Zhu, H.; Dang, F.; Zhang, W.; Qin, Z.; Yang, S.; Tan, H.; Lu, Y.; Jiang, W. Differential regulation of antibiotic biosynthesis by DraR-K, a novel two-component system in Streptomyces coelicolor. Mol. Microbiol. 2012, 85, 535–556. [Google Scholar] [CrossRef]
- Yu, Z.; Zhu, H.; Zheng, G.; Jiang, W.; Lu, Y. A genome-wide transcriptomic analysis reveals diverse roles of the two-component system DraR-K in the physiological and morphological differentiation of Streptomyces coelicolor. Appl. Microbiol. Biotechnol. 2014, 98, 9351–9363. [Google Scholar] [CrossRef]
- Yeo, K.J.; Han, Y.H.; Eo, Y.; Cheong, H.K. Expression, purification, crystallization and preliminary X-ray analysis of the extracellular sensory domain of DraK histidine kinase from Streptomyces coelicolor. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2013, 69 Pt 8, 909–911. [Google Scholar] [CrossRef] [Green Version]
- Yeo, K.J.; Hong, Y.S.; Jee, J.G.; Lee, J.K.; Kim, H.J.; Park, J.W.; Kim, E.H.; Hwang, E.; Kim, S.Y.; Lee, E.G.; et al. Mechanism of the pH-induced conformational change in the sensor domain of the DraK Histidine kinase via the E83, E105, and E107 residues. PLoS ONE 2014, 9, e107168. [Google Scholar] [CrossRef]
- Yeo, K.J.; Kim, E.H.; Hwang, E.; Han, Y.H.; Eo, Y.; Kim, H.J.; Kwon, O.; Hong, Y.S.; Cheong, C.; Cheong, H.K. pH-dependent structural change of the extracellular sensor domain of the DraK histidine kinase from Streptomyces coelicolor. Biochem. Biophys. Res. Commun. 2013, 431, 554–559. [Google Scholar] [CrossRef]
- Li, Y.Q.; Chen, P.L.; Chen, S.F.; Wu, D.; Zheng, J. A pair of two-component regulatory genes ecrA1/A2 in S. coelicolor. J. Zhejiang Univ. Sci. 2004, 5, 173–179. [Google Scholar] [CrossRef]
- Wang, C.; Ge, H.; Dong, H.; Zhu, C.; Li, Y.; Zheng, J.; Cen, P. A novel pair of two-component signal transduction system ecrE1/ecrE2 regulating antibiotic biosynthesis in Streptomyces coelicolor. Biol. Bratisl. 2007, 62, 511–516. [Google Scholar] [CrossRef]
- Amin, R.; Franz-Wachtel, M.; Tiffert, Y.; Heberer, M.; Meky, M.; Ahmed, Y.; Matthews, A.; Krysenko, S.; Jakobi, M.; Hinder, M.; et al. Post-translational Serine/Threonine Phosphorylation and Lysine Acetylation: A Novel Regulatory Aspect of the Global Nitrogen Response Regulator GlnR in S. coelicolor M145. Front. Mol. Biosci. 2016, 3, 38. [Google Scholar] [CrossRef] [Green Version]
- Amin, R.; Reuther, J.; Bera, A.; Wohlleben, W.; Mast, Y. A novel GlnR target gene, nnaR, is involved in nitrate/nitrite assimilation in Streptomyces coelicolor. Microbiology 2012, 158, 1172–1182. [Google Scholar] [CrossRef] [Green Version]
- Fink, D.; Weißschuh, N.; Reuther, J.; Wohlleben, W.; Engels, A. Two transcriptional regulators GlnR and GlnRII are involved in regulation of nitrogen metabolism in Streptomyces coelicolor A3(2). Mol. Microbiol. 2002, 46, 331–347. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Liu, X.; Wang, J.; Wang, Y.; Zheng, G.; Lu, Y.; Zhao, G.; Wang, J. Insight into the Molecular Mechanism of the Transcriptional Regulation of amtB Operon in Streptomyces coelicolor. Front. Microbiol. 2018, 9, 264. [Google Scholar] [CrossRef]
- Lin, W.; Wang, Y.; Han, X.; Zhang, Z.; Wang, C.; Wang, J.; Yang, H.; Lu, Y.; Jiang, W.; Zhao, G.-P.; et al. Atypical OmpR/PhoB Subfamily Response Regulator GlnR of Actinomycetes Functions as a Homodimer, Stabilized by the Unphosphorylated Conserved Asp-focused Charge Interactions. J. Biol. Chem. 2014, 289, 15413–15425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tiffert, Y.; Franz-Wachtel, M.; Fladerer, C.; Nordheim, A.; Reuther, J.; Wohlleben, W.; Mast, Y. Proteomic analysis of the GlnR-mediated response to nitrogen limitation in Streptomyces coelicolor M145. Appl. Microbiol. Biotechnol. 2011, 89, 1149–1159. [Google Scholar] [CrossRef]
- Tiffert, Y.; Supra, P.; Wurm, R.; Wohlleben, W.; Wagner, R.; Reuther, J. The Streptomyces coelicolor GlnR regulon: Identification of new GlnR targets and evidence for a central role of GlnR in nitrogen metabolism in actinomycetes. Mol. Microbiol. 2008, 67, 861–880. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhao, G.-P. GlnR positively regulates nasA transcription in Streptomyces coelicolor. Biochem. Biophys. Res. Commun. 2009, 386, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Wray, L.V., Jr.; Atkinson, M.R.; Fisher, S.H. Identification and cloning of the glnR locus, which is required for transcription of the glnA gene in Streptomyces coelicolor A3(2). J. Bacteriol. 1991, 173, 7351–7360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wray, L.V., Jr.; Fisher, S.H. The Streptomyces coelicolor glnR gene encodes a protein similar to other bacterial response regulators. Gene 1993, 130, 145–150. [Google Scholar] [CrossRef]
- Xu, Y.; You, D.; Ye, B.-C. Nitrogen regulator GlnR directly controls transcription of genes encoding lysine deacetylases in Actinobacteria. Microbiology 2017, 163, 1702–1710. [Google Scholar] [CrossRef] [PubMed]
- Cen, X.-F.; Wang, J.-Z.; Zhao, G.-P.; Wang, Y.; Wang, J. Molecular evidence for the coordination of nitrogen and carbon metabolisms, revealed by a study on the transcriptional regulation of the agl3EFG operon that encodes a putative carbohydrate transporter in Streptomyces coelicolor. Biochem. Biophys. Res. Commun. 2016, 471, 510–514. [Google Scholar] [CrossRef]
- He, J.-M.; Zhu, H.; Zheng, G.-S.; Liu, P.-P.; Wang, J.; Zhao, G.-P.; Zhu, G.-Q.; Jiang, W.-H.; Lu, Y.-H. Direct Involvement of the Master Nitrogen Metabolism Regulator GlnR in Antibiotic Biosynthesis in Streptomyces. J. Biol. Chem. 2016, 291, 26443–26454. [Google Scholar] [CrossRef] [Green Version]
- Shao, Z.; Deng, W.; Li, S.; He, J.; Ren, S.; Huang, W.; Lu, Y.; Zhao, G.; Cai, Z.; Wang, J. GlnR-Mediated Regulation of ectABCD Transcription Expands the Role of the GlnR Regulon to Osmotic Stress Management. J. Bacteriol. 2015, 197, 3041–3047. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Y.; Zhang, P.; Zhang, J.; Xu, W.; Wang, X.; Wu, L.; Sheng, D.; Ma, W.; Cao, G.; Chen, X.; et al. The developmental regulator MtrA binds GlnR boxes and represses nitrogen metabolism genes in Streptomyces coelicolor. Mol. Microbiol. 2019, 112, 29–46. [Google Scholar] [CrossRef]
- Rodríguez-García, A.; Sola-Landa, A.; Apel, K.; Santos-Beneit, F.; Martín, J.F. Phosphate control over nitrogen metabolism in Streptomyces coelicolor: Direct and indirect negative control of glnR, glnA, glnII and amtB expression by the response regulator PhoP. Nucleic Acids Res. 2009, 37, 3230–3242. [Google Scholar] [CrossRef]
- Santos-Beneit, F.; Rodríguez-García, A.; Martín, J.F. Overlapping binding of PhoP and AfsR to the promoter region of glnR in Streptomyces coelicolor. Microbiol. Res. 2012, 167, 532–535. [Google Scholar] [CrossRef]
- Sola-Landa, A.; Rodríguez-García, A.; Amin, R.; Wohlleben, W.; Martín, J.F. Competition between the GlnR and PhoP regulators for the glnA and amtB promoters in Streptomyces coelicolor. Nucleic Acids Res. 2012, 41, 1767–1782. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Cen, X.-F.; Zhao, G.-P.; Wang, J. Characterization of a New GlnR Binding Box in the Promoter of amtB in Streptomyces coelicolor Inferred a PhoP/GlnR Competitive Binding Mechanism for Transcriptional Regulation of amtB. J. Bacteriol. 2012, 194, 5237–5244. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krysenko, S.; Matthews, A.; Busche, T.; Bera, A.; Wohlleben, W. Poly- and Monoamine Metabolism in Streptomyces coelicolor: The New Role of Glutamine Synthetase-Like Enzymes in the Survival under Environmental Stress. Microb. Physiol. 2021, 31, 233–247. [Google Scholar] [CrossRef]
- Martín, J.F.; Sola-Landa, A.; Santos-Beneit, F.; Fernández-Martínez, L.T.; Prieto, C.; Rodríguez-García, A. Cross-talk of global nutritional regulators in the control of primary and secondary metabolism in Streptomyces. Microb. Biotechnol. 2010, 4, 165–174. [Google Scholar] [CrossRef] [Green Version]
- Reuther, J.; Wohlleben, W. Nitrogen Metabolism in Streptomyces coelicolor: Transcriptional and Post-Translational Regulation. Microb. Physiol. 2006, 12, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Jiang, W.; Lu, Y. A Novel Two-Component System, GluR-GluK, Involved in Glutamate Sensing and Uptake in Streptomyces coelicolor. J. Bacteriol. 2017, 199, e00097-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, M.; Xu, W.; Zhu, Y.; Cui, X.; Pang, X. The Response Regulator MacR and its Potential in Improvement of Antibiotic Production in Streptomyces coelicolor. Curr. Microbiol. 2021, 78, 3696–3707. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, P.; Zhu, Y.; Lu, T.; Wang, Y.; Cao, G.; Shi, M.; Chen, X.-L.; Tao, M.; Pang, X. Novel Two-Component System MacRS Is a Pleiotropic Regulator That Controls Multiple Morphogenic Membrane Protein Genes in Streptomyces coelicolor. Appl. Environ. Microbiol. 2019, 85, e02178-18. [Google Scholar] [CrossRef] [Green Version]
- Hoskisson, P.A.; Hutchings, M.I. MtrAB-LpqB: A conserved three-component system in actinobacteria? Trends Microbiol. 2006, 14, 444–449. [Google Scholar] [CrossRef]
- Zhang, P.; Wu, L.; Zhu, Y.; Liu, M.; Wang, Y.; Cao, G.; Chen, X.-L.; Tao, M.; Pang, X. Deletion of MtrA Inhibits Cellular Development of Streptomyces coelicolor and Alters Expression of Developmental Regulatory Genes. Front. Microbiol. 2017, 8, 2013. [Google Scholar] [CrossRef]
- Som, N.F.; Heine, D.; Holmes, N.; Knowles, F.; Chandra, G.; Seipke, R.F.; Hoskisson, P.A.; Wilkinson, B.; Hutchings, M.I. The MtrAB two-component system controls antibiotic production in Streptomyces coelicolor A3(2). Microbiology 2017, 163, 1415–1419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, Y.; Zhang, P.; Zhang, J.; Wang, J.; Lu, Y.; Pang, X. Impact on Multiple Antibiotic Pathways Reveals MtrA as a Master Regulator of Antibiotic Production in Streptomyces spp. and Potentially in Other Actinobacteria. Appl. Environ. Microbiol. 2020, 86, e00097-17. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhang, P.; Lu, T.; Wang, X.; Li, A.; Lu, Y.; Tao, M.; Pang, X. Impact of MtrA on phosphate metabolism genes and the response to altered phosphate conditions in Streptomyces. Environ. Microbiol. 2021, 23, 6907–6923. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Zhu, H.; Zheng, G.; Jiang, W.; Lu, Y. Metabolic engineering of Streptomyces coelicolor for enhanced prodigiosins (RED) production. Sci. China Life Sci. 2017, 60, 948–957. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; He, J.; Zhu, H.; Yu, Z.; Wang, R.; Chen, Y.; Dang, F.; Zhang, W.; Yang, S.; Jiang, W. An Orphan Histidine Kinase, OhkA, Regulates Both Secondary Metabolism and Morphological Differentiation in Streptomyces coelicolor. J. Bacteriol. 2011, 193, 3020–3032. [Google Scholar] [CrossRef] [Green Version]
- Xu, Z.; Wang, Y.; Chater, K.F.; Ou, H.-Y.; Xu, H.H.; Deng, Z.; Tao, M. Large-Scale Transposition Mutagenesis of Streptomyces coelicolor Identifies Hundreds of Genes Influencing Antibiotic Biosynthesis. Appl. Environ. Microbiol. 2017, 83, e02889-16. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Y.; Wang, X.; Zhang, J.; Ni, X.; Zhang, X.; Tao, M.; Pang, X. The regulatory gene wblA is a target of the orphan response regulator OrrA in Streptomyces coelicolor. Environ. Microbiol. 2022, 24, 3081–3096. [Google Scholar] [CrossRef]
- Fowler-Goldsworthy, K.; Gust, B.; Mouz, S.; Chandra, G.; Findlay, K.C.; Chater, K.F. The actinobacteria-specific gene wblA controls major developmental transitions in Streptomyces coelicolor A3(2). Microbiology 2011, 157, 1312–1328. [Google Scholar] [CrossRef] [Green Version]
- Kang, S.-H.; Huang, J.; Lee, H.-N.; Hur, Y.-A.; Cohen, S.N.; Kim, E.-S. Interspecies DNA Microarray Analysis Identifies WblA as a Pleiotropic Down-Regulator of Antibiotic Biosynthesis in Streptomyces. J. Bacteriol. 2007, 189, 4315–4319. [Google Scholar] [CrossRef] [Green Version]
- Nah, H.-J.; Park, J.; Choi, S.; Kim, E.-S. WblA, a global regulator of antibiotic biosynthesis in Streptomyces. J. Ind. Microbiol. Biotechnol. 2021, 48, kuab007. [Google Scholar] [CrossRef]
- Bishop, A.; Fielding, S.; Dyson, P.; Herron, P. Systematic Insertional Mutagenesis of a Streptomycete Genome: A Link between Osmoadaptation and Antibiotic Production. Genome Res. 2004, 14, 893–900. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martínez, L.F.; Bishop, A.; Parkes, L.; Del Sol, R.; Salerno, P.; Sevcikova, B.; Mazurakova, V.; Kormanec, J.; Dyson, P. Osmoregulation in Streptomyces coelicolor: Modulation of SigB activity by OsaC. Mol. Microbiol. 2009, 71, 1250–1262. [Google Scholar] [CrossRef] [PubMed]
- Fischer, M.; Falke, D.; Rönitz, J.; Haase, A.; Damelang, T.; Pawlik, T.; Sawers, R.G. Hypoxia-induced synthesis of respiratory nitrate reductase 2 of Streptomyces coelicolor A3(2) depends on the histidine kinase OsdK in mycelium but not in spores. Microbiology 2019, 165, 905–916. [Google Scholar] [CrossRef] [PubMed]
- Urem, M.; van Rossum, T.; Bucca, G.; Moolenaar, G.F.; Laing, E.; Świątek-Połatyńska, M.A.; Willemse, J.; Tenconi, E.; Rigali, S.; Goosen, N.; et al. OsdR of Streptomyces coelicolor and the Dormancy Regulator DevR of Mycobacterium tuberculosis Control Overlapping Regulons. mSystems 2016, 1, e00014-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Honma, S.; Ito, S.; Yajima, S.; Sasaki, Y. Nitric Oxide Signaling for Actinorhodin Production in Streptomyces coelicolor A3(2) via the DevS/R Two-Component System. Appl. Environ. Microbiol. 2021, 87, e00480-21. [Google Scholar] [CrossRef]
- Sasaki, Y.; Oguchi, H.; Kobayashi, T.; Kusama, S.; Sugiura, R.; Moriya, K.; Hirata, T.; Yukioka, Y.; Takaya, N.; Yajima, S.; et al. Nitrogen oxide cycle regulates nitric oxide levels and bacterial cell signaling. Sci. Rep. 2016, 6, 22038. [Google Scholar] [CrossRef] [Green Version]
- Voskuil, M.; Visconti, K.; Schoolnik, G. Mycobacterium tuberculosis gene expression during adaptation to stationary phase and low-oxygen dormancy. Tuberculosis 2004, 84, 218–227. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, L.; Zheng, G.; Jiang, W.; Deng, Z.; Wang, Z.; Lu, Y. CRISPR/dCas9-Mediated Multiplex Gene Repression in Streptomyces. Biotechnol. J. 2018, 13, e1800121. [Google Scholar] [CrossRef]
- Allenby, N.E.E.; Laing, E.; Bucca, G.; Kierzek, A.; Smith, C.P. Diverse control of metabolism and other cellular processes in Streptomyces coelicolor by the PhoP transcription factor: Genome-wide identification of in vivo targets. Nucleic Acids Res. 2012, 40, 9543–9556. [Google Scholar] [CrossRef]
- Apel, A.K.; Sola-Landa, A.; Rodríguez-García, A.; Martín, J.F. Phosphate control of phoA, phoC and phoD gene expression in Streptomyces coelicolor reveals significant differences in binding of PhoP to their promoter regions. Microbiology 2007, 153, 3527–3537. [Google Scholar] [CrossRef]
- Fernández-Martinez, L.T.; Santos-Beneit, F.; Martín, J.F. Is PhoR-PhoP partner fidelity strict? PhoR is required for the activation of the pho regulon in Streptomyces coelicolor. Mol. Genet. Genom. 2012, 287, 565–573. [Google Scholar] [CrossRef]
- Lejeune, C.; Abreu, S.; Chaminade, P.; Dulermo, T.; David, M.; Werten, S.; Virolle, M.-J. Impact of Phosphate Availability on Membrane Lipid Content of the Model Strains, Streptomyces lividans and Streptomyces coelicolor. Front. Microbiol. 2021, 12, 623919. [Google Scholar] [CrossRef]
- Martín-Martín, S.; Rodríguez-García, A.; Santos-Beneit, F.; Franco-Domínguez, E.; Sola-Landa, A.; Martín, J.F. Self-control of the PHO regulon: The PhoP-dependent protein PhoU controls negatively expression of genes of PHO regulon in Streptomyces coelicolor. J. Antibiot. 2017, 71, 113–122. [Google Scholar] [CrossRef]
- Millan-Oropeza, A.; Henry, C.; Lejeune, C.; David, M.; Virolle, M.-J. Expression of genes of the Pho regulon is altered in Streptomyces coelicolor. Sci. Rep. 2020, 10, 8492. [Google Scholar] [CrossRef]
- Rodríguez-García, A.; Barreiro, C.; Santos-Beneit, F.; Sola-Landa, A.; Martín, J.F. Genome-wide transcriptomic and proteomic analysis of the primary response to phosphate limitation in Streptomyces coelicolor M145 and in a Delta phoP mutant. Proteomics 2007, 7, 2410–2429. [Google Scholar] [CrossRef]
- Santos-Beneit, F.; Barriuso-Iglesias, M.; Fernández-Martínez, L.T.; Martínez-Castro, M.; Sola-Landa, A.; Rodríguez-García, A.; Martín, J.F. The RNA Polymerase Omega Factor RpoZ Is Regulated by PhoP and Has an Important Role in Antibiotic Biosynthesis and Morphological Differentiation in Streptomyces coelicolor. Appl. Environ. Microbiol. 2011, 77, 7586–7594. [Google Scholar] [CrossRef] [Green Version]
- Santos-Beneit, F.; Rodríguez-García, A.; Franco-Domínguez, E.; Martín, J.F. Phosphate-dependent regulation of the low- and high-affinity transport systems in the model actinomycete Streptomyces coelicolor. Microbiology 2008, 154, 2356–2370. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos-Beneit, F.; Rodríguez-García, A.; Martín, J.F. Complex transcriptional control of the antibiotic regulator afsS in Streptomyces: PhoP and AfsR are overlapping, competitive activators. J. Bacteriol. 2011, 193, 2242–2251. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos-Beneit, F.; Rodríguez-García, A.; Sola-Landa, A.; Martín, J.F. Cross-talk between two global regulators in Streptomyces: PhoP and AfsR interact in the control of afsS, pstS and phoRP transcription. Mol. Microbiol. 2009, 72, 53–68. [Google Scholar] [CrossRef] [PubMed]
- Sola-Landa, A.; Moura, R.S.; Martín, J.F. The two-component PhoR-PhoP system controls both primary metabolism and secondary metabolite biosynthesis in Streptomyces lividans. Proc. Natl. Acad. Sci. USA 2003, 100, 6133–6138. [Google Scholar] [CrossRef]
- Sola-Landa, A.; Rodríguez-García, A.; Apel, A.K.; Martín, J.F. Target genes and structure of the direct repeats in the DNA-binding sequences of the response regulator PhoP in Streptomyces coelicolor. Nucleic Acids Res. 2008, 36, 1358–1368. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sola-Landa, A.; Rodríguez-García, A.; Franco-Domínguez, E.; Martín, J.F. Binding of PhoP to promoters of phosphate-regulated genes in Streptomyces coelicolor: Identification of PHO boxes. Mol. Microbiol. 2005, 56, 1373–1385. [Google Scholar] [CrossRef] [PubMed]
- Thomas, L.; Hodgson, D.A.; Wentzel, A.; Nieselt, K.; Ellingsen, T.E.; Moore, J.; Morrissey, E.; Legaie, R.; The STREAM Consortium; Wohlleben, W.; et al. Metabolic Switches and Adaptations Deduced from the Proteomes of Streptomyces coelicolor Wild Type and phoP Mutant Grown in Batch Culture. Mol. Cell. Proteom. 2012, 11, M111.013797. [Google Scholar] [CrossRef] [Green Version]
- Barreiro, C.; Martínez-Castro, M. Regulation of the phosphate metabolism in Streptomyces genus: Impact on the secondary metabolites. Appl. Microbiol. Biotechnol. 2019, 103, 1643–1658. [Google Scholar] [CrossRef]
- Martín, J.F.; Liras, P. Molecular Mechanisms of Phosphate Sensing, Transport and Signalling in Streptomyces and Related Actinobacteria. Int. J. Mol. Sci. 2021, 22, 1129. [Google Scholar] [CrossRef]
- Martín, J.F.; Rodríguez-García, A.; Liras, P. The master regulator PhoP coordinates phosphate and nitrogen metabolism, respiration, cell differentiation and antibiotic biosynthesis: Comparison in Streptomyces coelicolor and Streptomyces avermitilis. J. Antibiot. 2017, 70, 534–541. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martín, J.F.; Santos-Beneit, F.; Rodríguez-García, A.; Sola-Landa, A.; Smith, M.; Ellingsen, T.E.; Nieselt, K.; Burroughs, N.J.; Wellington, E.M.H. Transcriptomic studies of phosphate control of primary and secondary metabolism in Streptomyces coelicolor. Appl. Microbiol. Biotechnol. 2012, 95, 61–75. [Google Scholar] [CrossRef]
- Paolo, S.S.; Huang, J.; Cohen, S.N.; Thompson, C.J. rag genes: Novel components of the RamR regulon that trigger morphological differentiation in Streptomyces coelicolor. Mol. Microbiol. 2006, 61, 1167–1186. [Google Scholar] [CrossRef]
- Nguyen, K.; Willey, J.; Nguyen, L.; Nguyen, L.T.; Viollier, P.; Thompson, C.J. A central regulator of morphological differentiation in the multicellular bacterium Streptomyces coelicolor. Mol. Microbiol. 2002, 46, 1223–1238. [Google Scholar] [CrossRef]
- O’Connor, T.J.; Kanellis, P.; Nodwell, J.R. The ramC gene is required for morphogenesis in Streptomyces coelicolor and expressed in a cell type-specific manner under the direct control of RamR. Mol. Microbiol. 2002, 45, 45–57. [Google Scholar] [CrossRef]
- Lu, Y.; Wang, W.; Shu, D.; Zhang, W.; Chen, L.; Qin, Z.; Yang, S.; Jiang, W. Characterization of a novel two-component regulatory system involved in the regulation of both actinorhodin and a type I polyketide in Streptomyces coelicolor. Appl. Microbiol. Biotechnol. 2007, 77, 625–635. [Google Scholar] [CrossRef]
- White, J.; Bibb, M. bldA dependence of undecylprodigiosin production in Streptomyces coelicolor A3(2) involves a pathway-specific regulatory cascade. J. Bacteriol. 1997, 179, 627–633. [Google Scholar] [CrossRef] [Green Version]
- Guthrie, E.P.; Flaxman, C.S.; White, J.; Hodgson, D.A.; Bibb, M.J.; Chater, K.F. A response-regulator-like activator of antibiotic synthesis from Streptomyces coelicolor A3(2) with an amino-terminal domain that lacks a phosphorylation pocket. Microbiology 1998, 144, 727–738. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Tian, X.; Wang, J.; Yang, H.; Fan, K.; Xu, G.; Yang, K.; Tan, H. Autoregulation of antibiotic biosynthesis by binding of the end product to an atypical response regulator. Proc. Natl. Acad. Sci. USA 2009, 106, 8617–8622. [Google Scholar] [CrossRef] [Green Version]
- Gongerowska-Jac, M.; Szafran, M.J.; Mikołajczyk, J.; Szymczak, J.; Bartyńska, M.; Gierlikowska, A.; Biały, S.; Elliot, M.A.; Jakimowicz, D. Global Chromosome Topology and the Two-Component Systems in Concerted Manner Regulate Transcription in Streptomyces. mSystems 2021, 6, e01142-21. [Google Scholar] [CrossRef]
- Busche, T.; Winkler, A.; Wedderhoff, I.; Ruckert, C.; Kalinowski, J.; Ortíz de Orué Lucana, D. Deciphering the Transcriptional Response Mediated by the Redox-Sensing System HbpS-SenS-SenR from Streptomycetes. PLoS ONE 2016, 11, e0159873. [Google Scholar] [CrossRef] [Green Version]
- Hong, H.-J.; Hutchings, M.I.; Neu, J.M.; Wright, G.D.; Paget, M.S.B.; Buttner, M.J. Characterization of an inducible vancomycin resistance system in Streptomyces coelicolor reveals a novel gene (vanK) required for drug resistance. Mol. Microbiol. 2004, 52, 1107–1121. [Google Scholar] [CrossRef] [PubMed]
- Hong, H.-J.; Hutchings, M.I.; Buttner, M.J. Vancomycin Resistance VanS/VanR Two-Component Systems. Adv. Exp. Med. Biol. 2008, 631, 200–213. [Google Scholar] [CrossRef]
- Hutchings, M.I.; Hong, H.-J.; Buttner, M.J. The vancomycin resistance VanRS two-component signal transduction system of Streptomyces coelicolor. Mol. Microbiol. 2006, 59, 923–935. [Google Scholar] [CrossRef] [PubMed]
- Koteva, K.; Hong, H.-J.; Wang, X.D.; Nazi, I.; Hughes, D.; Naldrett, M.J.; Buttner, M.; Wright, G. A vancomycin photoprobe identifies the histidine kinase VanSsc as a vancomycin receptor. Nat. Chem. Biol. 2010, 6, 327–329. [Google Scholar] [CrossRef] [PubMed]
- Kwun, M.J.; Novotna, G.; Hesketh, A.R.; Hill, L.; Hong, H.-J. In Vivo Studies Suggest that Induction of VanS-Dependent Vancomycin Resistance Requires Binding of the Drug to d -Ala- d -Ala Termini in the Peptidoglycan Cell Wall. Antimicrob. Agents Chemother. 2013, 57, 4470–4480. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lockey, C.; Edwards, R.J.; Roper, D.I.; Dixon, A.M. The Extracellular Domain of Two-component System Sensor Kinase VanS from Streptomyces coelicolor Binds Vancomycin at a Newly Identified Binding Site. Sci. Rep. 2020, 10, 5727. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novotna, G.B.; Kwun, M.J.; Hong, H.J. In Vivo Characterization of the Activation and Interaction of the VanR-VanS Two-Component Regulatory System Controlling Glycopeptide Antibiotic Resistance in Two Related Streptomyces Species. Antimicrob. Agents Chemother. 2016, 60, 1627–1637. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hong, H.-J.; Hutchings, M.I.; Hill, L.M.; Buttner, M.J. The Role of the Novel Fem Protein VanK in Vancomycin Resistance in Streptomyces coelicolor. J. Biol. Chem. 2005, 280, 13055–13061. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maciunas, L.J.; Porter, N.; Lee, P.J.; Gupta, K.; Loll, P.J. Structures of full-length VanR from Streptomyces coelicolor in both the inactive and activated states. Acta Crystallogr. Sect. D Struct. Biol. 2021, 77, 1027–1039. [Google Scholar] [CrossRef]
- Read, N.; Howlett, R.; Smith, M.C.M. An operon encoding enzymes for synthesis of a putative extracellular carbohydrate attenuates acquired vancomycin resistance in Streptomyces coelicolor. Microbiology 2019, 165, 208–223. [Google Scholar] [CrossRef]
- Santos-Beneit, F. Genome sequencing analysis of Streptomyces coelicolor mutants that overcome the phosphate-depending vancomycin lethal effect. BMC Genom. 2018, 19, 457. [Google Scholar] [CrossRef]
- Santos-Beneit, F.; Fernández-Martínez, L.T.; Rodríguez-García, A.; Martín-Martín, S.; Ordóñez-Robles, M.; Yagüe, P.; Manteca, A.; Martín, J.F. Transcriptional response to vancomycin in a highly vancomycin-resistant Streptomyces coelicolor mutant. Future Microbiol. 2014, 9, 603–622. [Google Scholar] [CrossRef]
- Santos-Beneit, F.; Martín, J.F. Vancomycin resistance in Streptomyces coelicolor is phosphate-dependent but is not mediated by the PhoP regulator. J. Glob. Antimicrob. Resist. 2013, 1, 109–113. [Google Scholar] [CrossRef]
- Chater, K.F. A Morphological and Genetic Mapping Study of White Colony Mutants of Streptomyces coelicolor. J. Gen. Microbiol. 1972, 72, 9–28. [Google Scholar] [CrossRef]
- Chater, K.F. Construction and Phenotypes of Double Sporulation Deficient Mutants in Streptomyces coelicolor A3(2). J. Gen. Microbiol. 1975, 87, 312–325. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, Y.; Fowler, K.; Findlay, K.; Tan, H.; Chater, K.F. An Unusual Response Regulator Influences Sporulation at Early and Late Stages in Streptomyces coelicolor. J. Bacteriol. 2007, 189, 2873–2885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, G.; Tian, Y.; Hu, K.; Zhu, Y.; Chater, K.F.; Feng, C.; Liu, G.; Tan, H. Importance and regulation of inositol biosynthesis during growth and differentiation of Streptomyces. Mol. Microbiol. 2012, 83, 1178–1194. [Google Scholar] [CrossRef] [PubMed]
- Ainsa, J.A.; Parry, H.D.; Chater, K.F. A response regulator-like protein that functions at an intermediate stage of sporulation in Streptomyces coelicolor A3(2). Mol. Microbiol. 1999, 34, 607–619. [Google Scholar] [CrossRef] [PubMed]
- Arroyo-Pérez, E.E.; González-Cerón, G.; Soberón-Chávez, G.; Georgellis, D.; Servin-Gonzalez, L. A Novel Two-Component System, Encoded by the sco5282/sco5283 Genes, Affects Streptomyces coelicolor Morphology in Liquid Culture. Front. Microbiol. 2019, 10, 1568. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.; Zhu, Y.; Zhang, P.; Sheng, D.; Cao, G.; Pang, X. SCO5351 is a pleiotropic factor that impacts secondary metabolism and morphological development in Streptomyces coelicolor. FEMS Microbiol. Lett. 2018, 365, fny150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rozas, D.; Gullón, S.; Mellado, R.P. A Novel Two-Component System Involved in the Transition to Secondary Metabolism in Streptomyces coelicolor. PLoS ONE 2012, 7, e31760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Becerril, A.; Álvarez, S.; Braña, A.F.; Rico, S.; Díaz, M.; Santamaria, R.I.; Salas, J.A.; Méndez, C. Uncovering production of specialized metabolites by Streptomyces argillaceus: Activation of cryptic biosynthesis gene clusters using nutritional and genetic approaches. PLoS ONE 2018, 13, e0198145. [Google Scholar] [CrossRef]
- Ganesh, I.; Kim, T.W.; Na, J.-G.; Eom, G.T.; Hong, S.H. Engineering Escherichia coli to Sense Non-native Environmental Stimuli: Synthetic Chimera Two-component Systems. Biotechnol. Bioprocess Eng. 2019, 24, 12–22. [Google Scholar] [CrossRef]
| Histidine Kinase (HK) | Response Regulator (RR) | Organization (5′→3′) | ||||
|---|---|---|---|---|---|---|
| Gene | Name | Family | Gene | Name | Family | |
| SCO0203 | OsdK/DevS | Classic | SCO0204 | OsdR/DevR | NarL | RR-HK |
| SCO0211 | - | Classic | oHK | |||
| SCO0422 | - | Classic | SCO0421 | - | NarL | HK-RR |
| SCO0551 | - | Classic | SCO0552 | - | OmpR | RR-HK |
| SCO0588 | CvnA11 | Classic | oHK | |||
| SCO0599 | - | Classic | oHK | |||
| SCO0676 | - | Classic | oHK | |||
| SCO0767 | - | Classic | oHK | |||
| SCO0871 | - | Classic | SCO0870 | - | CheY | RR-HK-RR |
| SCO0872 | - | RssB | ||||
| SCO0946 | - | Classic | oHK | |||
| SCO1071 | - | Classic | SCO1070 | - | NarL | HK-RR |
| SCO1137 | - | Classic | SCO1136 | - | IclR | HK-RR |
| SCO1160 | CvnA3 | Classic | oHK | |||
| SCO1217 | - | Classic | oHK | |||
| SCO1220 | - | LytTR | oRR | |||
| SCO1259 | - | Classic | SCO1260 | - | NarL | HK-RR |
| SCO1369 | - | Classic | SCO1370 | - | NarL | HK-RR |
| SCO1402 | CvnA4 | Classic | oHK | |||
| SCO1596 | OhkA | Classic | oHK | |||
| SCO1630 | CvnA9 | Classic | oHK | |||
| SCO1654 | - | NarL | oRR | |||
| SCO1744 | AbrA1 | Classic | SCO1745 | AbrA2 | NarL | HK-RR |
| SCO1802 | - | Classic | SCO1801 | - | NarL | HK-RR |
| SCO2013 | PdtaR | AmiR | oRR | |||
| SCO2121 | MacS | Classic | SCO2120 | MacR | NarL | HK-RR |
| SCO2142 | - | Classic | SCO2143 | - | OmpR | HK-RR |
| SCO2152 | - | CheY | oRR | |||
| SCO2166 | AbrB2 | Classic | SCO2165 | AbrB1 | NarL | HK-RR |
| SCO2215 | - | Classic | SCO2216 | - | NarL | HK-RR |
| SCO2281 | Aor1 | NarL | oRR | |||
| SCO2307 | - | Classic | SCO2308 | - | NarL | HK-RR |
| SCO2359 | - | Classic | SCO2358 | - | NarL | HK-RR |
| SCO2452 | - | Classic | oHK | |||
| SCO2518 | EcrA1 | Classic | SCO2517 | EcrA2 | NarL | HK-RR |
| SCO2800 | - | Classic | SCO2801 | - | OmpR | RR-HK |
| SCO2879 | CvnA12 | Classic | oHK | |||
| SCO3008 | OrrA | NarL | oRR | |||
| SCO3012 | MtrB | Classic | SCO3013 | MtrA | OmpR | RR-HK |
| SCO3062 | DraK | Classic | SCO3063 | DraR | OmpR | RR-HK |
| SCO3119 | - | Classic | oHK | |||
| SCO3134 | - | NarL | oRR | |||
| SCO3144 | - | NarL | oRR | |||
| SCO3225 | AbsA1 | Classic | SCO3226 | AbsA2 | NarL | HK-RR |
| SCO3284 | - | Classic | oHK | |||
| SCO3359 | CseC | Classic | SCO3358 | CseB | OmpR | RR-HK |
| SCO3390 | SatK | Classic | SCO3389 | SatR | NarL | HK-RR |
| SCO3589 | VanS | Classic | SCO3590 | VanR | OmpR | RR-HK |
| SCO3639 | - | Classic | SCO3638 | - | NarL | HK-RR |
| SCO3641 | - | Classic | SCO3640 | - | NarL | HK-RR |
| SCO3654 | - | Classic | SCO3653 | - | NarL | HK-RR |
| SCO3740 | - | Classic | SCO3741 | - | OmpR | RR-HK |
| SCO3750 | - | Classic | oHK | |||
| SCO3757 | - | Classic | SCO3756 | - | NarL | HK-RR |
| SCO3796 | - | Classic | oHK | |||
| SCO3818 | - | NarL | oRR | |||
| SCO3948 | - | Classic | oHK | |||
| SCO4009 | - | Hybrid | oHK | |||
| SCO4021 | - | Classic | SCO4020 | - | OmpR | RR-HK |
| SCO4073 | RagK | Classic | SCO4072 | RagR | NarL | HK-RR |
| SCO4120 | - | Classic | oHK | |||
| SCO4124 | - | Classic | SCO4123 | - | NarL | HK-RR |
| SCO4155 | CssS | Classic | SCO4156 | CssR | OmpR | RR-HK |
| SCO4159 | GlnR | OmpR | oRR | |||
| SCO4201 | - | RssB | oRR | |||
| SCO4229 | PhoR | Classic | SCO4230 | PhoP | OmpR | HK-RR |
| SCO4275 | SenS | Classic | SCO4276 | SenR | NarL | HK-RR |
| SCO4362 | - | Classic | SCO4363 | - | NarL | HK-RR |
| SCO4597 | AbrC2 | Classic | SCO4596 | AbrC3 | NarL | HK-HK-RR |
| SCO4598 | AbrC1 | Classic | ||||
| SCO4667 | SitK | Classic | SCO4668 | SitR | NarL | HK-RR |
| SCO4768 | BldM | NarL | oRR | |||
| SCO4791 | - | Classic | SCO4792 | - | NarL | HK-RR |
| SCO4906 | AfsQ2 | Classic | SCO4907 | AfsQ1 | OmpR | RR-HK |
| SCO5006 | - | ATPase | oRR | |||
| SCO5040 | - | Classic | oHK | |||
| SCO5104 | - | Classic | oHK | |||
| SCO5131 | - | Classic | SCO5132 | - | NarL | HK-RR |
| SCO5239 | PdtaS | Classic | oHK | |||
| SCO5282 | - | Classic | SCO5283 | - | OmpR | RR-HK |
| SCO5289 | CvnA5 | Classic | oHK | |||
| SCO5304 | - | Classic | oHK | |||
| SCO5351 | - | CheY | oRR | |||
| SCO5378 | ChiS | Classic | SCO5377 | ChiR | NarL | HK-RR |
| SCO5404 | RapA2 | Classic | SCO5403 | RapA1 | OmpR | RR-HK |
| SCO5435 | - | Classic | SCO5434 | - | IclR | HK-RR |
| SCO5454 | - | Classic | SCO5455 | - | NarL | HK-RR |
| SCO5460 | - | Classic | oHK | |||
| SCO5540 | CvnA2 | Classic | oHK | |||
| SCO5544 | CvnA1 | Classic | oHK | |||
| SCO5683 | - | Classic | SCO5684 | - | NarL | HK-RR |
| SCO5748 | OsaA | Hybrid | SCO5749 | OsaB | OsaB | HK-RR |
| SCO5779 | GluK | Classic | SCO5778 | GluR | OmpR | RR-HK |
| SCO5784 | - | Classic | SCO5785 | - | NarL | HK-RR |
| SCO5824 | - | Classic | SCO5825 | - | NarL | HK-RR |
| SCO5829 | - | Classic | SCO5828 | - | NarL | HK-RR |
| SCO5863 | CutS | Classic | SCO5862 | CutR | OmpR | RR-HK |
| SCO5871 | KdepD | Classic | SCO5872 | KdpE | OmpR | HK-RR |
| SCO5881 | RedZ | NarL | oRR | |||
| SCO6029 | WhiI | NarL | oRR | |||
| SCO6069 | CvnA6 | Classic | oHK | |||
| SCO6139 | - | Classic | SCO6140 | - | NarL | HK-RR |
| SCO6163 | - | Classic | SCO6162 | - | NarL | HK-RR |
| SCO6253 | - | Classic | SCO6254 | - | NarL | HK-RR |
| SCO6268 | - | Classic | oHK | |||
| SCO6353 | - | Classic | SCO6354 | - | OmpR | RR-HK |
| SCO6362 | - | Classic | SCO6363 | - | NarL | HK-RR |
| SCO6364 | - | OmpR | oRR | |||
| SCO6369 | - | Classic | oHK | |||
| SCO6421 | EcrE1 | Classic | SCO6422 | EcrE2 | NarL | HK-RR |
| SCO6424 | - | Classic | oHK | |||
| SCO6484 | - | Classic | oHK | |||
| SCO6668 | - | Classic | SCO6667 | - | NarL | HK-RR |
| SCO6685 | RamR | NarL | oRR | |||
| SCO6794 | CvnA7 | Classic | oHK | |||
| SCO6943 | CvnA8 | Classic | oHK | |||
| SCO7009 | - | Classic | oHK | |||
| SCO7076 | - | Classic | SCO7075 | - | OmpR | RR-HK |
| SCO7089 | - | Classic | SCO7088 | - | NarL | HK-RR |
| SCO7220 | - | Classic | oHK | |||
| SCO7231 | - | Classic | SCO7230 | - | OmpR | RR-HK |
| SCO7297 | - | Classic | SCO7298 | - | TrxB | RR-HK |
| SCO7327 | - | Hybrid | oHK | |||
| SCO7354 | - | Classic | oHK | |||
| SCO7422 | CvnA10 | Classic | oHK | |||
| SCO7463 | CvnA13 | Classic | oHK | |||
| SCO7534 | - | Classic | SCO7533 | - | OmpR | RR-HK |
| SCO7649 | - | Classic | SCO7648 | - | NarL | HK-RR |
| SCO7711 | - | Classic | SCO7712 | - | NarL | HK-RR |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sánchez de la Nieta, R.; Santamaría, R.I.; Díaz, M. Two-Component Systems of Streptomyces coelicolor: An Intricate Network to Be Unraveled. Int. J. Mol. Sci. 2022, 23, 15085. https://doi.org/10.3390/ijms232315085
Sánchez de la Nieta R, Santamaría RI, Díaz M. Two-Component Systems of Streptomyces coelicolor: An Intricate Network to Be Unraveled. International Journal of Molecular Sciences. 2022; 23(23):15085. https://doi.org/10.3390/ijms232315085
Chicago/Turabian StyleSánchez de la Nieta, Ricardo, Ramón I. Santamaría, and Margarita Díaz. 2022. "Two-Component Systems of Streptomyces coelicolor: An Intricate Network to Be Unraveled" International Journal of Molecular Sciences 23, no. 23: 15085. https://doi.org/10.3390/ijms232315085
APA StyleSánchez de la Nieta, R., Santamaría, R. I., & Díaz, M. (2022). Two-Component Systems of Streptomyces coelicolor: An Intricate Network to Be Unraveled. International Journal of Molecular Sciences, 23(23), 15085. https://doi.org/10.3390/ijms232315085







