Genome-Wide Association Study of Salt Tolerance-Related Traits during Germination and Seedling Development in an Intermedium-Spike Barley Collection
Abstract
1. Introduction
2. Results
2.1. Trait Variability and Heritability Estimates of Intermedium-Spike Barley Population
2.2. Means of the Studied Traits and Response to Salinity
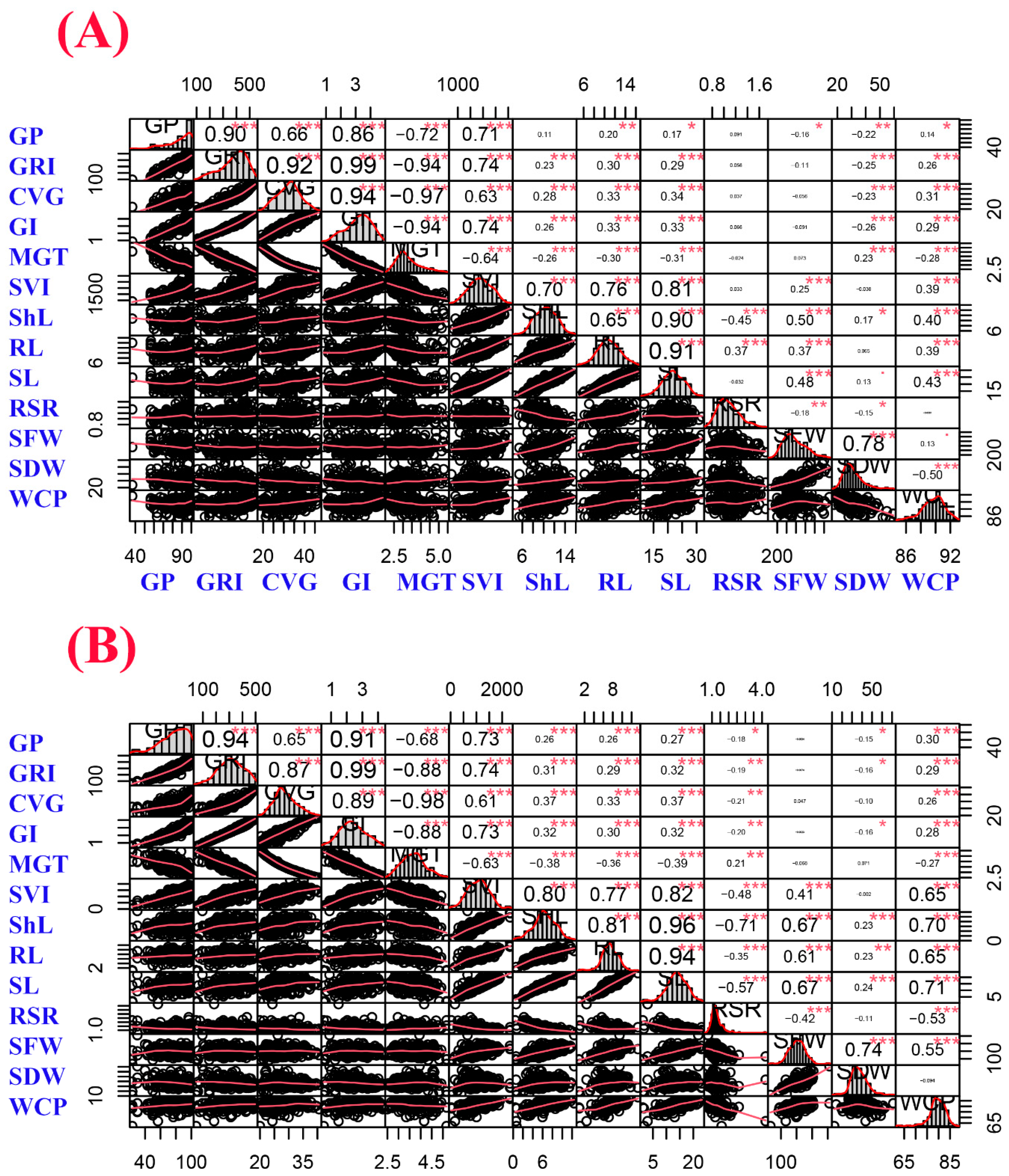
2.3. Phenotypic Correlation among Germination and Seedling Growth Parameters
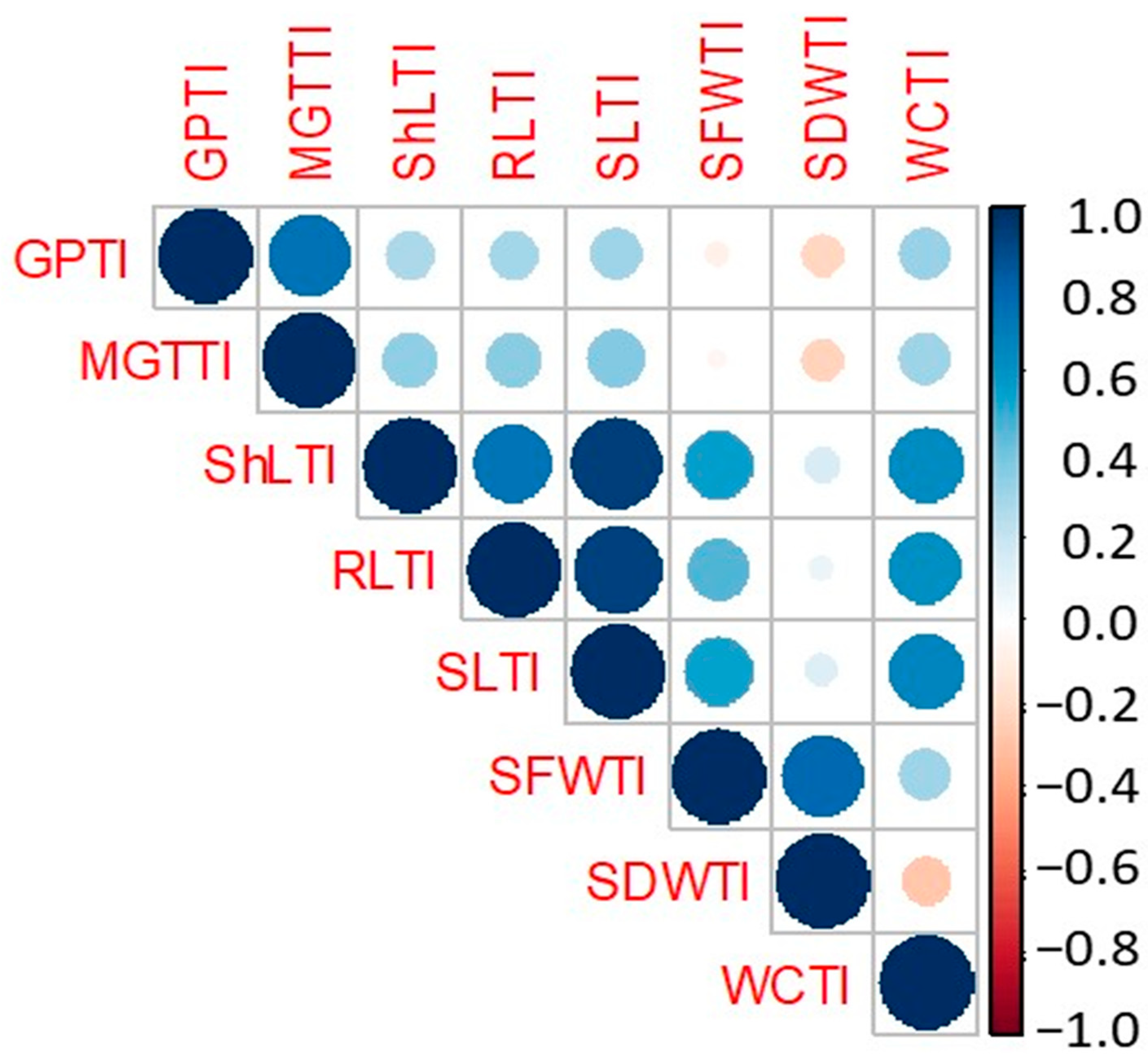
2.4. Salinity Tolerance Indices (STI) and Their Relationships
2.5. QTL Detection
2.6. Candidate Gene Prediction
3. Discussion
4. Materials and Methods
4.1. Plant Material and Genotyping
4.2. Experimental Design and Salinity Stress Treatments
4.3. Evaluation of Germination and Seedling Growth Parameters
- (1)
- Germination percentage (GP in %):
- (2)
- Germination rate index (GRI) was calculated according to Esechie [75] as GRI = (G1 + G2 + … + Gi)/i, where G1 is the germination percentage, calculated daily from day 1 to i = 10. It gives an indication of the percentage of seeds germinating per day during the germination test period.
- (3)
- Coefficient of velocity of germination (CVG) was calculated following Al-Ansari and Ksiksi [76] as follows: CVG = (N1 + N2 + … + Ni) × 100/( N1*T1 + … + Ni*Ti), where N is the number of seeds germinated every day and T is the number of days from seeding corresponding to N. It gives an indication of the speed of germination. CVG values increase when the number of germinated seeds increases and the time required for germination decreases.
- (4)
- Germination index (GI) was calculated according to Benech Arnold et al. [77] as follows: GI = (10 × N1) + (9 × N2) + … + (1 × N10); where N1, N2, …, N10, is the number of seeds germinated on the first, second and subsequent days until the 10th day, and the multipliers (i.e., 10, 9 … etc.) are weights given to the days of the germination. It measures both percentage and speed of germination. High values indicate that seeds germinate early and low values that seeds germinate late.
- (5)
- Mean germination time (MGT) represents the mean time that seeds require to initiate and end germination. It was calculated according to Orchard [78] as follows:where Ni is the number of the newly germinated seeds at time Ti.MGT = Σ(Ti × Ni)/ΣNi
- (6)
- Seedling vigor index (SVI) was calculated according to Abiri et al. [79] with little modification as a multiplication of the final germination percentage by the total seedling length (shoot and root lengths).
- (7)
- Shoot length (ShL in cm) and root length (RL in cm) were measured manually at the tenth day of germination using a scaled ruler for five seedlings from each replicate at the end of the experiment.
- (8)
- Seedling length (SL in cm) was measured as the total of root and shoot lengths.
- (9)
- Root–shoot ratio (RSR) was calculated as a ratio between root length and shoot length.
- (10)
- Seedlings fresh weight (SFW in g) and seedling dry weight (SDW in mg) were recorded by weighing harvested seedlings at day 10 and, respectively, after drying the fresh weight at 80 °C for 72 h to obtain the SDW using an ultra-micro lab balance (Sartorius AC 1215, Germany).
- (11)
- Water content percentage (WCP in %) was calculated based on the following formula:
4.4. Stress Tolerance Indices (STI)
- (1)
- Germination percentage (GPTI, as germination percentage tolerance index);
- (2)
- Mean germination time (MGTTI, as mean germination time tolerance index);
- (3)
- Shoot length (ShLTI, as shoot length tolerance index);
- (4)
- Root length (RLTI, as root length tolerance index);
- (5)
- Seedling length (SLTI, as seedling length tolerance index);
- (6)
- Seedling fresh weight (SFWTI, as seedling fresh weight tolerance index);
- (7)
- Seedling dry weight (SDWTI, as seedling dry weight tolerance index);
- (8)
- Water content % (WCPTI, as water content percentage tolerance index).
4.5. Statistical Analyses
4.6. GWAS Analysis
4.7. GWAS’ Significant QTL Annotation
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Atkinson, N.J.; Lilley, C.J.; Urwin, P.E. Identification of genes involved in the response of Arabidopsis to simultaneous biotic and abiotic stresses. Plant Physiol. 2013, 162, 2028–2041. [Google Scholar] [CrossRef] [PubMed]
- Mohanavelu, A.; Naganna, S.R.; Al-Ansari, N. Irrigation Induced Salinity and Sodicity Hazards on Soil and Groundwater: An Overview of Its Causes, Impacts and Mitigation Strategies. Agriculture 2021, 11, 983. [Google Scholar] [CrossRef]
- Manuel, R.; Machado, A.; Serralheiro, R.P.; Alvino, A.; Freire, M.I.; Ferreira, R. Soil Salinity: Effect on Vegetable Crop Growth. Management Practices to Prevent and Mitigate Soil Salinization. Horticulturae 2017, 3, 30. [Google Scholar] [CrossRef]
- Food and Agriculture Organization of the United Nations. Intergovernmental Technical Panel on Soils. Status of the World’s Soil Resources; Food and Agriculture Organization of the United Nations: Rome, Italy, 2015; ISBN 9789251090046. [Google Scholar]
- Kaashyap, M.; Ford, R.; Kudapa, H.; Jain, M.; Edwards, D.; Varshney, R.; Mantri, N. Differential Regulation of Genes Involved in Root Morphogenesis and Cell Wall Modification is Associated with Salinity Tolerance in Chickpea. Sci. Rep. 2018, 8, 4855. [Google Scholar] [CrossRef] [PubMed]
- Mwando, E.; Angessa, T.T.; Han, Y.; Zhou, G.; Li, C. Quantitative Trait Loci Mapping for Vigour and Survival Traits of Barley Seedlings after Germinating under Salinity Stress. Agronomy 2021, 11, 103. [Google Scholar] [CrossRef]
- Xue, W.; Yan, J.; Jiang, Y.; Zhan, Z.; Zhao, G.; Tondelli, A.; Cattivelli, L.; Cheng, J. Genetic dissection of winter barley seedling response to salt and osmotic stress. Mol. Breed. 2019, 39, 137. [Google Scholar] [CrossRef]
- Zhu, M.; Zhou, M.; Shabala, L.; Shabala, S. Linking osmotic adjustment and stomatal characteristics with salinity stress tolerance in contrasting barley accessions. Funct. Plant Biol. 2015, 42, 252–263. [Google Scholar] [CrossRef]
- Arzani, A.; Ashraf, M. Smart Engineering of Genetic Resources for Enhanced Salinity Tolerance in Crop Plants. CRC Crit. Rev. Plant Sci. 2016, 35, 146–189. [Google Scholar] [CrossRef]
- Deinlein, U. Plant salt-tolerance mechanisms. Trends Plant Sci 2014, 19, 371–379. [Google Scholar] [CrossRef]
- Cattivelli, L.; Rizza, F.; Badeck, F.W.; Mazzucotelli, E.; Mastrangelo, A.M.; Francia, E.; Marè, C.; Tondelli, A.; Stanca, A.M. Drought tolerance improvement in crop plants: An integrated view from breeding to genomics. F. Crop. Res. 2008, 105, 1–14. [Google Scholar] [CrossRef]
- Tao, Z.; Kou, Y.; Liu, H.; Li, X.; Xiao, J.; Wang, S. OsWRKY45 alleles play different roles in abscisic acid signalling and salt stress tolerance but similar roles in drought and cold tolerance in rice. J. Exp. Bot. 2011, 62, 4863–4874. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Chen, M.; Li, L.; Xu, Z.; Chen, X.; Guo, J.; Ma, Y. Overexpression of the soybean GmERF3 gene, an AP2/ERF type transcription factor for increased tolerances to salt, drought, and diseases in transgenic tobacco. J. Exp. Bot. 2009, 60, 3781–3796. [Google Scholar] [CrossRef] [PubMed]
- Ying, S.; Zhang, D.F.; Fu, J.; Shi, Y.S.; Song, Y.C.; Wang, T.Y.; Li, Y. Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic Arabidopsis. Planta 2012, 235, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Xu, Y.; Ma, Q.; Xu, W.; Wang, T.; Xue, Y.; Chong, K. Overexpression of an R1R2R3 MYB gene, OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol. 2007, 143, 1739–1751. [Google Scholar] [CrossRef]
- Sayed, M.A.; Nassar, S.M.; Moustafa, E.S.; Said, M.T. Genetic Mapping Reveals Novel Exotic and Elite QTL Alleles for Salinity Tolerance in Barley. Agronomy 2021, 11, 1774. [Google Scholar] [CrossRef]
- Moustafa, E.S.A.; El-Sobky, E.S.E.A.; Farag, H.I.A.; Yasin, M.A.T.; Attia, A.; Rady, M.O.A.; Awad, M.F.; Mansour, E. Sowing date and genotype influence on yield and quality of dual-purpose barley in a salt-affected arid region. Agronomy 2021, 11, 717. [Google Scholar] [CrossRef]
- Ebrahim, F.; Arzani, A.; Rahimmalek, M.; Sun, D.; Peng, J. Salinity tolerance of wild barley Hordeum vulgare ssp. spontaneum. Plant Breed. 2020, 139, 304–316. [Google Scholar] [CrossRef]
- Gharaghanipor, N.; Arzani, A.; Rahimmalek, M.; Ravash, R. Physiological and Transcriptome Indicators of Salt Tolerance in Wild and Cultivated Barley. Front. Plant Sci. 2022, 13, 1039. [Google Scholar] [CrossRef]
- Gorzolka, K.; Kölling, J.; Nattkemper, T.W.; Niehaus, K. Spatio-Temporal Metabolite Profiling of the Barley Germination Process by MALDI MS Imaging. PLoS ONE 2016, 11, e0150208. [Google Scholar] [CrossRef]
- Shelden, M.C.; Dias, D.A.; Jayasinghe, N.S.; Bacic, A.; Roessner, U. Root spatial metabolite profiling of two genotypes of barley (Hordeum vulgare L.) reveals differences in response to short-term salt stress. J. Exp. Bot. 2016, 67, 3731–3745. [Google Scholar] [CrossRef]
- Gupta, S.; Rupasinghe, T.; Callahan, D.L.; Natera, S.H.A.; Smith, P.M.C.; Hill, C.B.; Roessner, U.; Boughton, B.A. Spatio-Temporal Metabolite and Elemental Profiling of Salt Stressed Barley Seeds During Initial Stages of Germination by MALDI-MSI and µ-XRF Spectrometry. Front. Plant Sci. 2019, 10, 1139. [Google Scholar] [CrossRef] [PubMed]
- Askari, H.; Kazemitabar, S.K.; Zarrini, H.N.; Saberi, M.H. Salt tolerance assessment of barley (Hordeum vulgare L.) genotypes at germination stage by tolerance indices. Open Agric. 2016, 1, 37–44. [Google Scholar] [CrossRef]
- Mwando, E.; Han, Y.; Angessa, T.T.; Zhou, G.; Hill, C.B.; Zhang, X.Q.; Li, C. Genome-Wide Association Study of Salinity Tolerance During Germination in Barley (Hordeum vulgare L.). Front. Plant Sci. 2020, 11, 118. [Google Scholar] [CrossRef]
- Mano, Y.; Takeda, K. Mapping quantitative trait loci for salt tolerance at germination and the seedling stage in barley (Hordeum vulgare L.). Euphytica 1997, 94, 263–272. [Google Scholar] [CrossRef]
- Kumar, S.; Dwivedi, S.K.; Singh, S.S.; Jha, S.K.; Elanchezhian, R.; Singh, O.N.; Bhatt, B.P. Identification of drought tolerant rice genotypes by analysing drought tolerance indices and morpho-physiological traits. Sabrao J. Breed. Genet. 2014, 46, 217–230. [Google Scholar]
- Witzel, K.; Weidner, A.; Surabhi, G.K.; Varshney, R.K.; Kunze, G.; Buck-Sorlin, G.H.; BÖrner, A.; Mock, H.P. Comparative analysis of the grain proteome fraction in barley genotypes with contrasting salinity tolerance during germination. Plant. Cell Environ. 2010, 33, 211–222. [Google Scholar] [CrossRef]
- Angessa, T.T.; Zhang, X.Q.; Zhou, G.; Broughton, S.; Zhang, W.; Li, C. Early growth stages salinity stress tolerance in CM72 x Gairdner doubled haploid barley population. PLoS ONE 2017, 12, e0179715. [Google Scholar] [CrossRef]
- Tondelli, A.; Xu, X.; Moragues, M.; Sharma, R.; Schnaithmann, F.; Ingvardsen, C.; Manninen, O.; Comadran, J.; Russell, J.; Waugh, R.; et al. Structural and Temporal Variation in Genetic Diversity of European Spring Two-Row Barley Cultivars and Association Mapping of Quantitative Traits. Plant Genome 2013, 6, plantgenome2013-03. [Google Scholar] [CrossRef]
- Xu, X.; Sharma, R.; Tondelli, A.; Russell, J.; Comadran, J.; Schnaithmann, F.; Pillen, K.; Kilian, B.; Cattivelli, L.; Thomas, W.T.B.; et al. Genome-Wide Association Analysis of Grain Yield-Associated Traits in a Pan-European Barley Cultivar Collection. Plant Genome 2018, 11, 170073. [Google Scholar] [CrossRef]
- Yu, J.; Zhao, W.; Tong, W.; He, Q.; Yoon, M.Y.; Li, F.P.; Choi, B.; Heo, E.B.; Kim, K.W.; Park, Y.J. A Genome-Wide Association Study Reveals Candidate Genes Related to Salt Tolerance in Rice (Oryza sativa) at the Germination Stage. Int. J. Mol. Sci. 2018, 19, 3145. [Google Scholar] [CrossRef]
- Youssef, H.M.; Mascher, M.; Ayoub, M.A.; Stein, N.; Kilian, B.; Schnurbusch, T. Natural diversity of inflorescence architecture traces cryptic domestication genes in barley (Hordeum vulgare L.). Genet. Resour. Crop Evol. 2017, 64, 843–853. [Google Scholar] [CrossRef]
- Youssef, H.M.; Allam, M.; Boussora, F.; Himmelbach, A.; Milner, S.G.; Mascher, M.; Schnurbusch, T. Dissecting the genetic basis of lateral and central spikelet development and grain traits in Intermedium-spike barley (Hordeum vulgare convar. Intermedium). Plants 2020, 9, 1655. [Google Scholar] [CrossRef] [PubMed]
- Mascher, M.; Gundlach, H.; Himmelbach, A.; Beier, S.; Twardziok, S.O.; Wicker, T.; Radchuk, V.; Dockter, C.; Hedley, P.E.; Russell, J.; et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 2017, 544, 427–433. [Google Scholar] [CrossRef]
- Bagues, M.; Sarabi, B.; Ghashghaie, J.; Souli, I.; Nagaz, K. The validity of carbon isotope discrimination as a screening criterion for grain yield in two barley landraces under deficit irrigation with saline water in southern Tunisia. Plant Biotechnol. 2018, 35, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Abbas, G.; Saqib, M.; Akhtar, J.; Haq, M.A. ul Interactive effects of salinity and iron deficiency on different rice genotypes. J. Plant Nutr. Soil Sci. 2015, 178, 306–311. [Google Scholar] [CrossRef]
- Munns, R. Genes and salt tolerance: Bringing them together. New Phytol. 2005, 167, 645–663. [Google Scholar] [CrossRef]
- Soleimani, B.; Sammler, R.; Backhaus, A.; Beschow, H.; Schumann, E.; Mock, H.P.; von Wirén, N.; Seiffert, U.; Pillen, K. Genetic regulation of growth and nutrient content under phosphorus deficiency in the wild barley introgression library S42IL. Plant Breed. 2017, 136, 892–907. [Google Scholar] [CrossRef]
- Bálint, A.F.; Szira, F.; Börner, A.; Galiba, G. Segregation- and association based mapping of loci influencing osmotic tolerance in barley. Acta Biol. Szeged. 2008, 52, 101–102. [Google Scholar]
- Wójcik-Jagła, M.; Rapacz, M.; Tyrka, M.; Kościelniak, J.; Crissy, K.; Zmuda, K. Comparative QTL analysis of early short-time drought tolerance in Polish fodder and malting spring barleys. Theor. Appl. Genet. 2013, 126, 3021–3034. [Google Scholar] [CrossRef]
- Xue, W.; Yan, J.; Zhao, G.; Jiang, Y.; Cheng, J.; Cattivelli, L.; Tondelli, A. A major QTL on chromosome 7HS controls the response of barley seedling to salt stress in the Nure × Tremois population. BMC Genet. 2017, 18, 79. [Google Scholar] [CrossRef]
- Thabet, S.G.; Moursi, Y.S.; Sallam, A.; Karam, M.A.; Alqudah, A.M. Genetic associations uncover candidate SNP markers and genes associated with salt tolerance during seedling developmental phase in barley. Environ. Exp. Bot. 2021, 188, 104499. [Google Scholar] [CrossRef]
- Nayyeripasand, L.; Garoosi, G.A.; Ahmadikhah, A. Genome-Wide Association Study (GWAS) to Identify Salt-Tolerance QTLs Carrying Novel Candidate Genes in Rice During Early Vegetative Stage; 2021; Volume 14, ISBN 0000000151009. [Google Scholar]
- Mwando, E.; Angessa, T.T.; Han, Y.; Li, C. Salinity tolerance in barley during germination—homologs and potential genes. J. Zhejiang Univ. Sci. B 2020, 21, 93–121. [Google Scholar] [CrossRef] [PubMed]
- Dehnavi, A.R.; Zahedi, M.; Ludwiczak, A.; Perez, S.C.; Piernik, A. Effect of salinity on seed germination and seedling development of sorghum (Sorghum bicolor (L.) Moench) genotypes. Agronomy 2020, 10, 859. [Google Scholar] [CrossRef]
- Sayed, M.A.; Tarawneh, R.; Youssef, H.M.; Pillen, K.; Börner, A. Detection and Verification of QTL for Salinity Tolerance at Germination and Seedling Stages Using Wild Barley Introgression Lines. Plants 2021, 10, 2246. [Google Scholar] [CrossRef] [PubMed]
- Abdul Qados, A.M.S. Effect of salt stress on plant growth and metabolism of bean plant Vicia faba (L.). J. Saudi Soc. Agric. Sci. 2011, 10, 7–15. [Google Scholar] [CrossRef]
- Alam, M.A.; Juraimi, A.S.; Rafii, M.Y.; Abdul Hamid, A. Effect of salinity on biomass yield and physiological and stem-root anatomical characteristics of purslane (Portulaca oleracea L.) accessions. Biomed Res. Int. 2015, 2015, 105695. [Google Scholar] [CrossRef]
- Parihar, P.; Singh, S.; Singh, R.; Singh, V.P.; Prasad, S.M. Effect of salinity stress on plants and its tolerance strategies: A review. Environ. Sci. Pollut. Res. 2015, 22, 4056–4075. [Google Scholar] [CrossRef]
- Sayar, R.; Bchini, H.; Mosbahi, M.; Ezzine, M. Effects of salt and drought stresses on germination, emergence and seedling growth of Durum wheat (Triticum durum Desf.). African J. Agric. Res. 2010, 5, 2008–2016. [Google Scholar]
- Zhang, H.; Irving, L.J.; McGill, C.; Matthew, C.; Zhou, D.; Kemp, P. The effects of salinity and osmotic stress on barley germination rate: Sodium as an osmotic regulator. Ann. Bot. 2010, 106, 1027. [Google Scholar] [CrossRef]
- Baǧci, S.A.; Ekiz, H.; Yilmaz, A. Determination of the salt tolerance of some barley genotypes and the characteristics affecting tolerance. Turkish J. Agric. For. 2003, 27, 253–260. [Google Scholar] [CrossRef]
- Allel, D.; Benamar, A.; Badri, M.; Abdelly, C. Evaluation of salinity tolerance indices in North African barley accessions at reproductive stage. Czech J. Genet. Plant Breed. 2019, 55, 61–69. [Google Scholar] [CrossRef]
- Jamshidi, A.; Javanmard, H.R. Evaluation of barley (Hordeum vulgare L.) genotypes for salinity tolerance under field conditions using the stress indices. Ain Shams Eng. J. 2018, 9, 2093–2099. [Google Scholar] [CrossRef]
- Zhao, K.; Tung, C.W.; Eizenga, G.C.; Wright, M.H.; Ali, M.L.; Price, A.H.; Norton, G.J.; Islam, M.R.; Reynolds, A.; Mezey, J.; et al. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. 2011, 2, 467. [Google Scholar] [CrossRef] [PubMed]
- Sbei, H.; Sato, K.; Shehzad, T.; Harrabi, M.; Okuno, K. Detection of QTLs for salt tolerance in Asian barley (Hordeum vulgare L.) by association analysis with SNP markers. Breed. Sci. 2015, 64, 378–388. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Shabala, S.; Ma, Y.; Xu, R.; Zhou, M. Using QTL mapping to investigate the relationships between abiotic stress tolerance (drought and salinity) and agronomic and physiological traits. BMC Genomics 2015, 16, 43. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Zhou, G.; Shabala, S.; Chen, Z.H.; Cai, S.; Li, C.; Zhou, M. Genome-wide association study reveals a new QTL for salinity tolerance in Barley (Hordeum vulgare L.). Front. Plant Sci. 2016, 7, 946. [Google Scholar] [CrossRef]
- Flowers, T.J. Improving crop salt tolerance. J. Exp. Bot. 2004, 55, 307–319. [Google Scholar] [CrossRef]
- Naveed, S.A.; Zhang, F.; Zhang, J.; Zheng, T.Q.; Meng, L.J.; Pang, Y.L.; Xu, J.L.; Li, Z.K. Identification of QTN and candidate genes for Salinity Tolerance at the Germination and Seedling Stages in Rice by Genome-Wide Association Analyses. Sci. Rep. 2018, 8, 6505. [Google Scholar] [CrossRef]
- Browse, J. Jasmonate: An oxylipin signal with many roles in plants. Vitam. Horm. 2005, 72, 431–456. [Google Scholar] [CrossRef]
- Ishiguro, S.; Kawai-Oda, A.; Ueda, J.; Nishida, I.; Okada, K. The defective in anther dehiscience gene encodes a novel phospholipase A1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 2001, 13, 2191–2209. [Google Scholar] [CrossRef]
- Shehzad, M.; Zhou, Z.; Ditta, A.; Cai, X.; Khan, M.; Xu, Y.; Hou, Y.; Peng, R.; Hao, F.; Wang, K.; et al. Genome-Wide Mining and Identification of Protein Kinase Gene Family Impacts Salinity Stress Tolerance in Highly Dense Genetic Map Developed from Interspecific Cross between G. hirsutum L. and G. darwinii G. Watt. Agronomy 2019, 9, 560. [Google Scholar] [CrossRef]
- Yang, Y.; Wang, Q.; Chen, Q.; Yin, X.; Qian, M.; Sun, X.; Yang, Y. Genome-wide survey indicates diverse physiological roles of the barley (Hordeum vulgare L.) calcium-dependent protein kinase genes. Sci. Rep. 2017, 7, 560. [Google Scholar] [CrossRef] [PubMed]
- Shen, W.; Gómez-Cadenas, A.; Routly, E.L.; Ho, T.H.D.; Simmonds, J.A.; Gulick, P.J. The salt stress-inducible protein kinase gene, Esi47, from the salt-tolerant wheatgrass Lophopyrum elongatum is involved in plant hormone signaling. Plant Physiol. 2001, 125, 1429–1441. [Google Scholar] [CrossRef]
- Qiu, Y.W.; Zhe, F.E.N.G.; Fu, M.M.; Yuan, X.H.; Luo, C.C.; Yu, Y.B.; Feng, Y.Z.; Qi, W.E.I.; Li, F.L. GsMAPK4, a positive regulator of soybean tolerance to salinity stress. J. Integr. Agric. 2019, 18, 372–380. [Google Scholar] [CrossRef]
- Szymańska, K.P.; Polkowska-Kowalczyk, L.; Lichocka, M.; Maszkowska, J.; Dobrowolska, G. SNF1-Related Protein Kinases SnRK2.4 and SnRK2.10 Modulate ROS Homeostasis in Plant Response to Salt Stress. Int. J. Mol. Sci. 2019, 20, 143. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Chen, R.; Jiang, Q.; Sun, X.; Zhang, H.; Hu, Z. GmNAC06, a NAC domain transcription factor enhances salt stress tolerance in soybean. Plant Mol. Biol. 2021, 105, 333–345. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.Z.; Jin, S.H.; Jiang, X.Y.; Dong, R.R.; Li, P.; Li, Y.J.; Hou, B.K. Ectopic expression of UGT75D1, a glycosyltransferase preferring indole-3-butyric acid, modulates cotyledon development and stress tolerance in seed germination of Arabidopsis thaliana. Plant Mol. Biol. 2016, 90, 77–93. [Google Scholar] [CrossRef]
- Chen, N.; Yang, Q.; Pan, L.; Chi, X.; Chen, M.; Hu, D.; Yang, Z.; Wang, T.; Wang, M.; Yu, S. Identification of 30 MYB transcription factor genes and analysis of their expression during abiotic stress in peanut (Arachis hypogaea L.). Gene 2014, 533, 332–345. [Google Scholar] [CrossRef]
- Zhao, Y.; Tian, X.; Wang, F.; Zhang, L.; Xin, M.; Hu, Z.; Yao, Y.; Ni, Z.; Sun, Q.; Peng, H. Characterization of wheat MYB genes responsive to high temperatures. BMC Plant Biol. 2017, 17, 208. [Google Scholar] [CrossRef]
- Du, Y.T.; Zhao, M.J.; Wang, C.T.; Gao, Y.; Wang, Y.X.; Liu, Y.W.; Chen, M.; Chen, J.; Zhou, Y.B.; Xu, Z.S.; et al. Identification and characterization of GmMYB118 responses to drought and salt stress. BMC Plant Biol. 2018, 18, 320. [Google Scholar] [CrossRef]
- Naranjo, M.Á.; Forment, J.; Roldán, M.; Serrano, R.; Vicente, O. Overexpression of Arabidopsis thaliana LTL1, a salt-induced gene encoding a GDSL-motif lipase, increases salt tolerance in yeast and transgenic plants. Plant. Cell Environ. 2006, 29, 1890–1900. [Google Scholar] [CrossRef] [PubMed]
- Mansfeld, R. Das morphologische System der Saatgerste, Hordeum vulgare L. s. l. Der Züchter 1950, 20, 8–24. [Google Scholar] [CrossRef]
- Esechie, H.A. Interaction of Salinity and Temperature on the Germination of Sorghum. J. Agron. Crop Sci. 1994, 172, 194–199. [Google Scholar] [CrossRef]
- Al-Ansari, F.; Ksiksi, T. A Quantitative Assessment of Germination Parameters: The Case of and. Open Ecol. J. 2016, 9, 13–21. [Google Scholar] [CrossRef]
- Benech Arnold, R.L.; Fenner, M.; Edwards, P.J. Changes in germinability, ABA content and ABA embryonic sensitivity in developing seeds of Sorghum bicolor (L.) Moench. induced by water stress during grain filling. New Phytol. 1991, 118, 339–347. [Google Scholar] [CrossRef]
- Orchard, T.J. Estimating the parameters of plant seedling emergence. Seed Sci. Technol. 1977, 5, 61–69. [Google Scholar]
- Abiri, R.; Shaharuddin, N.A.; Maziah, M.; Norhana, Z.; Yusof, B.; Atabaki, N.; Sahebi, M.; Azizi, P. Quantitative assessment of indica rice germination to hydropriming, hormonal priming and polyethylene glycol priming. Chil. J. Agric. Res. 2016, 76, 392–400. [Google Scholar] [CrossRef]
- Fernandez, G.C.J. Effective selection criteria for assessing plant stress tolerance. In Proceedings of the International Symposium on Adaptation of Vegetables and other Food Crops in Temperature and Water Stress, Shanhua, Taiwan, 13–16 August 1992; pp. 257–270. [Google Scholar] [CrossRef]
- SAS Institute. The SAS System for Windows; Release 9.2; SAS Institute: Cary, NC, USA, 2011; Available online: http://www.sciepub.com/reference/166089 (accessed on 4 July 2021).
- Padi, F.K. Genotype environment interaction and yield stability in a cowpea-based cropping system. Euphytica 2007, 158, 11–25. [Google Scholar] [CrossRef]
- Dreissig, S.; Maurer, A.; Sharma, R.; Milne, L.; Flavell, A.J.; Schmutzer, T.; Pillen, K. Natural variation in meiotic recombination rate shapes introgression patterns in intraspecific hybrids between wild and domesticated barley. New Phytol. 2020, 228, 1852–1863. [Google Scholar] [CrossRef]
- SAS Institute. The SAS System for Windows; Release 9.4; SAS Institute: Cary, NC, USA, 2013. [Google Scholar]
- Bonferroni, C. Teoria Statistica Delle Classi e Calcolo Delle Probabilita, 8th ed.; Pubblicazioni del R Istituto Superiore di Scienze Economiche e Commericiali di Firenze: Firenze, Italy, 1936; pp. 3–62. [Google Scholar]
| Trait | Control | 150 mM NaCl | Combined ANOVA | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| MS (df = 207) | CV | R2 | Hb | MS | CV | R2 | Hb | T (df = 1) | A (df = 207) | A × T (df = 207) | |
| GP | 502.1 ** | 6.5 | 0.89 | 93.6 | 980.8 ** | 16.3 | 0.75 | 83.8 | 33,750.1 ** | 1177.7 ** | 305.2 ** |
| GRI | 33,928 ** | 8.8 | 0.94 | 96.9 | 31,024 ** | 18.1 | 0.84 | 90.6 | 1,659,664 ** | 54,132 ** | 10,821 ** |
| CVG | 123.5 ** | 10.4 | 0.85 | 90.9 | 54.5 ** | 9.1 | 0.80 | 87.7 | 4496.0 ** | 135.6 ** | 42.4 ** |
| GI | 2.5 ** | 7.8 | 0.95 | 97.6 | 2.1 ** | 18.5 | 0.84 | 90.2 | 163.25 ** | 3.87 ** | 0.75 ** |
| MGT | 1.5 ** | 9.4 | 0.89 | 93.9 | 0.9 ** | 9.5 | 0.78 | 86.3 | 40.0 ** | 1.9 ** | 0.6 ** |
| SVI | 677,900 ** | 12.8 | 0.85 | 91.5 | 531,966 ** | 19.4 | 0.86 | 91.6 | 195,406,315 ** | 958,912 ** | 250,954 ** |
| ShL | 12.8 ** | 11.5 | 0.82 | 89.2 | 15.7 ** | 13.6 | 0.91 | 95.1 | 4406.6 ** | 19.7 ** | 8.8 ** |
| RL | 14.2 ** | 12.3 | 0.79 | 87.0 | 10.7 ** | 11.6 | 0.88 | 93.2 | 4377.8 ** | 17.7 ** | 7.1 ** |
| SL | 44.6 ** | 10.7 | 0.81 | 88.4 | 47.4 ** | 11.2 | 0.91 | 95.0 | 17,569 ** | 64.6 ** | 27.3 ** |
| RSR | 0.1 ** | 11.1 | 0.79 | 86.4 | 0.4 ** | 14.0 | 0.87 | 92.8 | 5.4 ** | 0.3 ** | 0.2 ** |
| SFW | 10,748 ** | 13.1 | 0.79 | 86.5 | 7743 ** | 24.8 | 0.61 | 67.8 | 2,484,890 ** | 13,005 ** | 5486 ** |
| SDW | 161.0 ** | 13.9 | 0.82 | 89.2 | 221.5 ** | 22.3 | 0.61 | 68.5 | 17,361.2 ** | 295.4 ** | 87.1 ** |
| WCP | 6.8 ** | 1.1 | 0.78 | 85.7 | 38.2 ** | 3.1 | 0.76 | 84.1 | 24,654.2 ** | 24.8 ** | 20.3 ** |
| Trait | Control | Salinity | R% | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Mean | Std Error | Minimum | Maximum | Mean | Std Error | Minimum | Maximum | ||
| GP | 87.77 | 0.55 | 30.0 | 100.0 | 77.37 | 0.83 | 10.00 | 100.00 | −11.85 |
| GRI | 370.89 | 4.38 | 77.8 | 555.6 | 297.95 | 4.43 | 11.11 | 511.11 | −19.66 |
| CVG | 32.13 | 0.28 | 17.5 | 50.0 | 28.33 | 0.19 | 16.67 | 47.37 | −11.82 |
| GI | 3.17 | 0.04 | 0.7 | 5.0 | 2.45 | 0.04 | 0.17 | 4.67 | −22.81 |
| MGT | 3.27 | 0.03 | 2.0 | 5.7 | 3.63 | 0.02 | 2.11 | 6.00 | 10.95 |
| SVI | 1880.99 | 20.56 | 644.0 | 3306.7 | 1089.60 | 18.19 | 11.00 | 2483.33 | −42.07 |
| ShL | 10.23 | 0.09 | 4.1 | 17.5 | 6.47 | 0.10 | 0.20 | 13.00 | −36.75 |
| RL | 11.10 | 0.10 | 4.0 | 18.0 | 7.35 | 0.08 | 0.60 | 14.37 | −33.75 |
| SL | 21.33 | 0.17 | 8.3 | 35.5 | 13.82 | 0.17 | 0.80 | 24.83 | −35.19 |
| RSR | 1.11 | 0.01 | 0.6 | 1.8 | 1.24 | 0.02 | 0.58 | 5.00 | 11.94 |
| SFW | 291.00 | 2.70 | 159.2 | 554.9 | 201.76 | 2.61 | 30.70 | 609.60 | −30.67 |
| SDW | 30.01 | 0.32 | 14.1 | 64.4 | 37.47 | 0.44 | 10.00 | 123.20 | 24.86 |
| WCP | 89.63 | 0.07 | 83.2 | 93.0 | 80.74 | 0.16 | 51.12 | 93.46 | −9.92 |
| Index | MS (df = 207) | CV | R2 | Hb | Means | Std Error | Minimum | Maximum |
|---|---|---|---|---|---|---|---|---|
| GPTI | 0.24 ** | 17.3 | 0.83 | 89.9 | 0.90 | 0.012 | 0.06 | 1.30 |
| MGTTI | 0.23 ** | 10.8 | 0.92 | 95.9 | 0.90 | 0.012 | 0.27 | 1.91 |
| ShLTI | 0.29 ** | 18.9 | 0.90 | 94.7 | 0.65 | 0.013 | 0.03 | 1.78 |
| RLTI | 0.21 ** | 17.3 | 0.88 | 93.5 | 0.68 | 0.011 | 0.06 | 1.81 |
| SLTI | 0.21 ** | 16.0 | 0.90 | 94.7 | 0.66 | 0.011 | 0.05 | 1.65 |
| SFWTI | 0.24 ** | 26.4 | 0.77 | 85.2 | 0.71 | 0.013 | 0.11 | 2.10 |
| SDWTI | 1.01 ** | 26.9 | 0.81 | 88.1 | 1.29 | 0.026 | 0.28 | 5.08 |
| WCTI | 0.01 ** | 3.3 | 0.78 | 85.6 | 0.90 | 0.002 | 0.59 | 1.02 |
| Trait | Treatment | Marker | Chr | Position bp | Raw_p | Log10_Raw_p | Estimate |
|---|---|---|---|---|---|---|---|
| QTL detected under control treatment | |||||||
| GP | Control | 6_7696839 | 6H | 7,696,839 | 1.64 × 10−6 | 5.79 | 8.98 |
| CVG | Control | 1_546649042 | 1H | 546,649,042 | 2.07 × 10−7 | 6.68 | 6.86 |
| GI | Control | 1_546649042 | 1H | 546,649,042 | 6.68 × 10−7 | 6.18 | 0.94 |
| MGT | Control | 2_708864195 | 2H | 708,864,195 | 9.67 × 10−7 | 6.01 | 0.5 |
| SFW | Control | 7_14906512 | 7H | 14,906,512 | 6.61 × 10−9 | 8.18 | 52.82 |
| SDW | Control | 7_49898546 | 7H | 49,898,546 | 4.90 × 10−10 | 9.31 | 9.28 |
| QTL detected under salinity treatment | |||||||
| GP | Salinity | 5_586496095 | 5H | 586,496,095 | 1.43 × 10−6 | 5.84 | −13.86 |
| CVG | Salinity | 1_546373322 | 1H | 546,373,322 | 3.76 × 10−6 | 5.42 | 4.58 |
| GI | Salinity | 1_546373322 | 1H | 546,373,322 | 1.02 × 10−6 | 5.99 | 0.94 |
| SVI | Salinity | 5_3766117 | 5H | 3,766,117 | 8.43 × 10−8 | 7.07 | 353.02 |
| ShL | Salinity | 2_151892989 | 2H | 151,892,989 | 1.10 × 10−6 | 5.96 | 2.66 |
| ShL | Salinity | 5_560970990 | 5H | 560,970,990 | 2.27 × 10−7 | 6.64 | −2.66 |
| RL | Salinity | 2_420745590 | 2H | 420,745,590 | 8.71 × 10−8 | 7.06 | −1.68 |
| SL | Salinity | 5_561126360 | 5H | 561,126,360 | 2.58 × 10−7 | 6.59 | −5.12 |
| SL | Salinity | 7_637877064 | 7H | 637,877,064 | 2.22 × 10−7 | 6.65 | 3.04 |
| SFW | Salinity | 1_487884122 | 1H | 487,884,122 | 3.79 × 10−11 | 10.42 | −63.42 |
| SFW | Salinity | 3_599466174 | 3H | 599,466,174 | 1.49 × 10−6 | 5.83 | −42.16 |
| SDW | Salinity | 1_15066059 | 1H | 15,066,059 | 5.23 × 10−8 | 7.28 | 8.84 |
| WCP | Salinity | 7_629887676 | 7H | 629,887,676 | 2.47 × 10−6 | 5.61 | −2.86 |
| QTL detected for salinity tolerance indices | |||||||
| ShLTI | Index | 2_420745590 | 2H | 420,745,590 | 2.25 × 10−10 | 9.65 | −22.08 |
| RLTI | Index | 2_698868743 | 2H | 698,868,743 | 4.14 × 10−8 | 7.38 | −15.96 |
| SLTI | Index | 2_420745590 | 2H | 420,745,590 | 1.48 × 10−9 | 8.83 | −17.58 |
| SFWTI | Index | 7_70461482 | 7H | 70,461,482 | 6.25 × 10−7 | 6.20 | −14.92 |
| Marker | Candidate Gene | Chr. | Start | End | Function Description |
|---|---|---|---|---|---|
| m1_15066059 | HORVU1Hr1G007420 | 1H | 15,064,891 | 15,064,891 | Cytochrome P450 superfamily protein |
| m1_466766698 | HORVU1Hr1G065250 | 1H | 466,760,732 | 466,760,732 | Potassium channel AKT2 |
| m1_487884122 | HORVU1Hr1G069990 | 1H | 487,882,055 | 487,882,055 | U-box domain-containing protein 16 |
| m1_528149209 | HORVU1Hr1G081810 | 1H | 528,147,129 | 528,147,129 | 26S proteasome non-ATPase regulatory subunit 1 homolog A |
| m1_546373322 | HORVU1Hr1G090570HORVU1Hr1G090580 | 1H1H | 546,411,012546,421,876 | 546,412,822546,423,244 | Phospholipase A1-II 6 |
| m2_419658044 | HORVU2Hr1G062320 | 2H | 419,657,547 | 419,657,547 | tRNA dimethylallyltransferase 2 |
| m2_698868743 | HORVU2Hr1G102970 | 2H | 698,868,231 | 698,868,231 | F-box/RNI-like superfamily protein |
| m2_710957942 | HORVU2Hr1G106970 | 2H | 710,953,055 | 710,953,055 | Homeobox-leucine zipper protein family |
| m3_166063769 | HORVU3Hr1G032440 | 3H | 166,062,751 | 166,062,751 | Two-component response regulator-like APRR2 |
| m3_361778117 | HORVU3Hr1G050400 | 3H | 361,773,216 | 361,773,216 | Peptidyl-prolyl cis-trans isomerase-like 1 |
| m3_599466174 | HORVU3Hr1G082580 | 3H | 599,464,312 | 599,464,312 | Protein NRT1/ PTR FAMILY 5.10 |
| m3_603625697 | HORVU3Hr1G083460 | 3H | 603,625,804 | 603,625,804 | DOF zinc finger protein 1 |
| m3_648172294 | HORVU3Hr1G094870 | 3H | 648,170,663 | 648,170,663 | histidine kinase 3 |
| m3_662456881 | HORVU3Hr1G099770 | 3H | 662,455,277 | 662,455,277 | hydroxyethylthiazole kinase family protein |
| m4_631432489 | HORVU4Hr1G085320 | 4H | 631,514,701 | 631,519,979 | NAC domain protein |
| m4_608376198 | HORVU4Hr1G078670 | 4H | 608,370,056 | 608,370,056 | Mitochondrial substrate carrier family protein |
| m5_3766117 | HORVU5Hr1G001090 | 5H | 3,762,210 | 3,762,210 | BEL1-like homeodomain 6 |
| m5_659835561 | HORVU5Hr1G121610 | 5H | 659,835,141 | 659,835,141 | B-cell receptor-associated protein 31-like |
| m6_7696839 | HORVU6Hr1G003300 | 6H | 7,696,550 | 7,696,550 | nitrate reductase 1 |
| m6_32557121 | HORVU6Hr1G014910 | 6H | 32,555,240 | 32,555,240 | D.melanogaster polytene |
| m7_49898546 | HORVU7Hr1G027930 | 7H | 49,898,118 | 49,898,118 | Raffinose synthase family protein |
| m7_70461482 | HORVU7Hr1G034050 | 7H | 70,456,816 | 70,456,816 | Pentatricopeptide repeat-containing protein |
| m7_161050366 | HORVU7Hr1G047740 | 7H | 161,049,541 | 161,049,541 | L-gulonolactone oxidase 2 |
| m7_167285781 | HORVU7Hr1G048720 | 7H | 167,284,758 | 167,284,758 | Heme A synthase |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sayed, M.A.; Maurer, A.; Schmutzer, T.; Schnurbusch, T.; Börner, A.; Hansson, M.; Pillen, K.; Youssef, H.M. Genome-Wide Association Study of Salt Tolerance-Related Traits during Germination and Seedling Development in an Intermedium-Spike Barley Collection. Int. J. Mol. Sci. 2022, 23, 11060. https://doi.org/10.3390/ijms231911060
Sayed MA, Maurer A, Schmutzer T, Schnurbusch T, Börner A, Hansson M, Pillen K, Youssef HM. Genome-Wide Association Study of Salt Tolerance-Related Traits during Germination and Seedling Development in an Intermedium-Spike Barley Collection. International Journal of Molecular Sciences. 2022; 23(19):11060. https://doi.org/10.3390/ijms231911060
Chicago/Turabian StyleSayed, Mohammed A., Andreas Maurer, Thomas Schmutzer, Thorsten Schnurbusch, Andreas Börner, Mats Hansson, Klaus Pillen, and Helmy M. Youssef. 2022. "Genome-Wide Association Study of Salt Tolerance-Related Traits during Germination and Seedling Development in an Intermedium-Spike Barley Collection" International Journal of Molecular Sciences 23, no. 19: 11060. https://doi.org/10.3390/ijms231911060
APA StyleSayed, M. A., Maurer, A., Schmutzer, T., Schnurbusch, T., Börner, A., Hansson, M., Pillen, K., & Youssef, H. M. (2022). Genome-Wide Association Study of Salt Tolerance-Related Traits during Germination and Seedling Development in an Intermedium-Spike Barley Collection. International Journal of Molecular Sciences, 23(19), 11060. https://doi.org/10.3390/ijms231911060






