Influence of Foods and Nutrition on the Gut Microbiome and Implications for Intestinal Health
Abstract
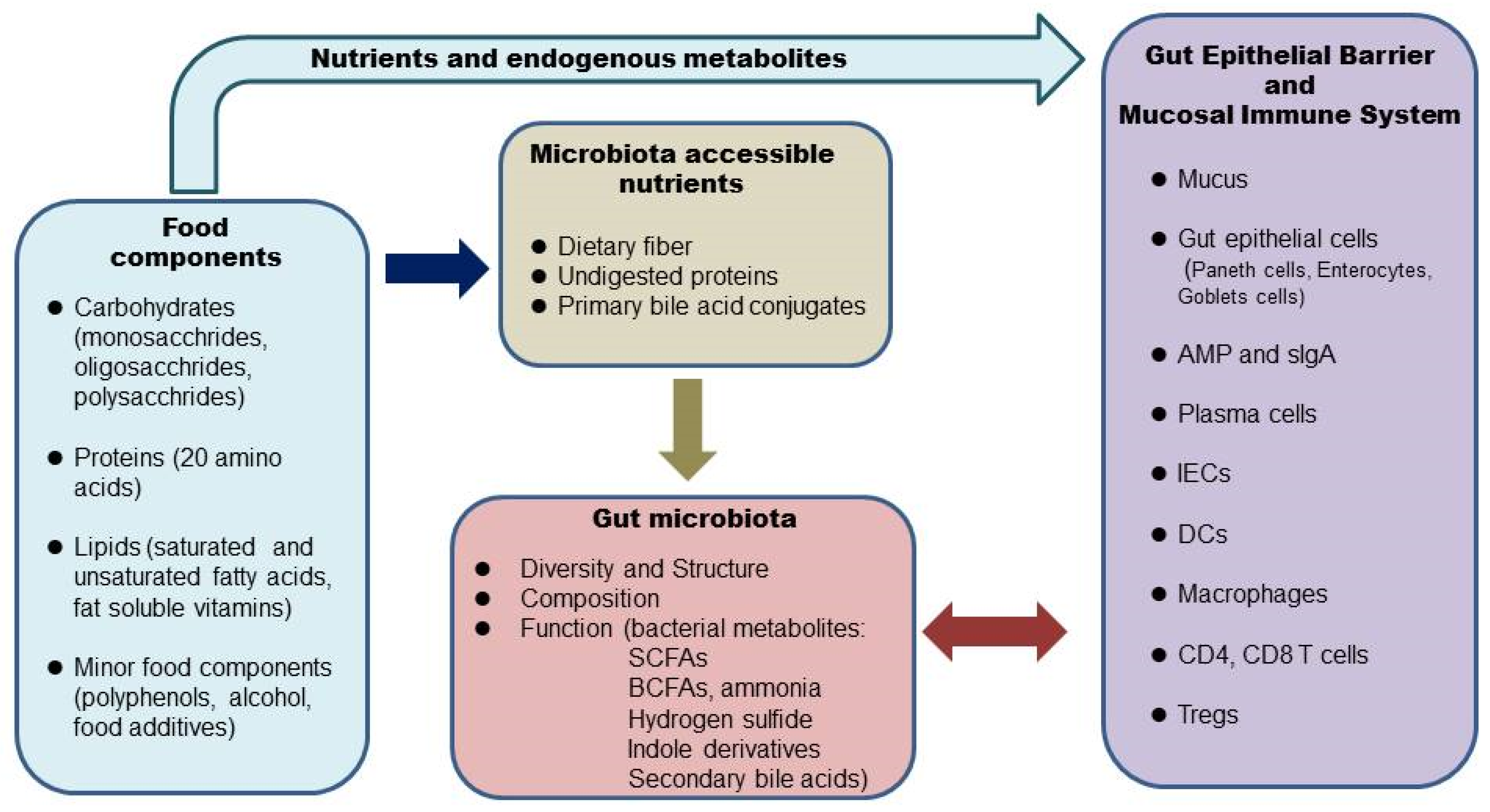
:1. Introduction
2. Diet, the Microbiome, and the Intestinal Barrier
3. Diet, the Microbiome, and the Intestinal Mucosal Immune System
4. Foods and Nutrition on the Gut Microbiome and Intestinal Health
4.1. Dietary Fiber
4.1.1. Dietary Fiber on Gut Microbial Ecology
4.1.2. Dietary Fiber Fermentation by the Gut Microbes Is Type Specific
4.2. Dietary Fat on Gut Microbiome
4.3. Dietary Protein and Certain Amino Acids
4.4. Food Components Negatively Influence the Gut Microbiome and Intestinal Health
5. Dysbiosis in IBD and Gut Microbiome-Targeted Therapies
6. Future Perspectives and Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sender, R.; Fuchs, S.; Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016, 14, e1002533. [Google Scholar] [CrossRef] [PubMed]
- Postler, T.S.; Ghosh, S. Understanding the holobiont: How microbial metabolites affect human health and shape the immune system. Cell Metab. 2017, 26, 110–130. [Google Scholar] [CrossRef] [PubMed]
- Arumugam, M.; Raes, J.; Pelletier, E.; le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Frank, D.N.; St. Amand, A.L.; Feidman, R.A.; Boedeker, E.C.; Harpaz, N.; Pace, N.R. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl. Acad. Sci. USA 2007, 104, 13780–13785. [Google Scholar] [CrossRef]
- Schroeder, B.; Bäckhed, F. Signals from the gut microbiota to distant organs in physiology and disease. Nat. Med. 2016, 22, 1079–1089. [Google Scholar] [CrossRef]
- Singh, R.K.; Chang, H.W.; Yan, D.; Lee, K.M.; Ucmak, D.; Wong, K.; Abrouk, M.; Farahnik, B.; Nakamura, M.; Zhu, T.H.; et al. Influence of diet on the gut microbiome and implications for human health. J. Transl. Med. 2017, 15, 73. [Google Scholar] [CrossRef]
- Smits, S.A.; Leach, J.; Sonnenburg, E.D.; Gonzalez, C.G.; Lichtman, J.S.; Reid, G.; Knight, R.; Manjurano, A.; Changalucha, J.; Elias, J.E.; et al. Seasonal cycling in the gut microbiome of the Hadaz hunter-gatherers of Tanzania. Science 2016, 357, 802–806. [Google Scholar] [CrossRef]
- Jha, A.R.; Davenport, E.R.; Gautam, Y.; Bhandari, D.; Tandukar, S.; Ng, K.M.; Fragiadakis, G.K.; Holmes, S.; Gautam, G.P.; Leach, J.; et al. Gut microbiome transition across a lifestyle gradient in Himalaya. PLoS Biol. 2018, 16, e2005396. [Google Scholar] [CrossRef]
- Deehan, E.C.; Yang, C.; Perez-Muñoz, M.E.; Nguyen, N.K.; Cheng, C.C.; Triador, L.; Zhang, Z.; Bakal, J.A.; Walter, J. Precision microbiome modulation with discrete dietary fiber structures directs short-chain fatty acids production. Cell Host Microbe 2020, 27, 389–404. [Google Scholar] [CrossRef]
- Wastyk, H.; Fragiadakis, G.K.; Perelman, D.; Dahan, D.; Merrill, B.D.; Yu, F.B.; Topf, M.; Gonzalez, C.G.; Treuren, W.V.; Han, S. Gut-microbiota-targeted diets modulate human immune status. Cell 2021, 184, 4137–4153. [Google Scholar] [CrossRef]
- Rinninella, E.; Cintoni, M.; Raoul, P.; Lopetuso, L.R.; Scaldafferri, F.; Pulcini, G.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, G.P. Food components and dietary habits: Keys for a healthy gut microbiota composition. Nutrients 2019, 11, 2393. [Google Scholar] [CrossRef] [PubMed]
- Lane, E.R.; Zisman, T.L.; Suskind, D.L. The microbiota in inflammatory bowel disease: Current and therapeutic insights. J. Inflamm. Res. 2017, 10, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, K.; Kamada, N. Diet-microbiota interactions in inflammatory bowel disease. Nutrients 2021, 13, 1533. [Google Scholar] [CrossRef] [PubMed]
- Staley, C.; Weingarden, A.R.; Khoruts, A.; Sadowsky, M.J. Interaction of gut microbiota with bile acid metabolism and its influence on disease states. Appl. Microbiol. Biotechnol. 2017, 101, 47–64. [Google Scholar] [CrossRef] [PubMed]
- De Santis, S.; Cavalcanti, E.; Mastronardi, M.; Jirillo, E.; Chieppa, M. Nutritional keys for intestinal barrier modulation. Front. Immunol. 2015, 6, 612. [Google Scholar] [CrossRef]
- Gerbe, F.; Legraverend, C.; Jay, P. The intestinal epithelium tuft cells: Specification and function. Cell Mol. Life Sci. 2012, 69, 2907–2917. [Google Scholar] [CrossRef]
- Pelaseyed, T.; Bergström, J.H.; Gustafsson, J.K.; Ermund, A.; Birchenough, G.M.H.; Schütte, A.; van der Post, S.; Svensson, F.; Rodríguez-Piñeiro, A.M.; Nyström, E.E.L.; et al. The mucus and mucins of the goblet cells and enterocytes provide the first defense line of the gastrointestinal tract and interact with the immune system. Immonol. Rev. 2014, 260, 8–20. [Google Scholar] [CrossRef]
- Desal, M.S.; Seekatz, A.M.; Koropatkin, N.K.; Kamada, N.; Hickey, C.A.; Wolter, M.; Pudlo, N.A.; Kitamoto, S.; Terrapon, N.; Muller, A.; et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 2016, 167, 1339–1353. [Google Scholar]
- Mu, Q.; Kirby, J.; Reilly, C.M.; Luo, X.M. Leaky gut as a danger signal for autoimmune diseases. Front. Immonol. 2017, 8, 598. [Google Scholar] [CrossRef]
- Kalina, U.; Koyama, N.; Hosoda, T.; Nuernberger, H.; Sato, K.; Hoelzer, D.; Herweck, F.; Manigold, T.; Singer, M.V.; Rossol, S.; et al. Enhanced production of IL-18 in butyrate-treated intestinal epithelium by stimulation of the proximal promoter region. Eur. J. Immunol. 2002, 32, 2635–2643. [Google Scholar]
- Zhao, F.; Zhou, G.; Liu, X.; Song, S.; Xu, X.; Hooiveld, G.; Müller, M.; Liu, L.; Kristiansen, K.; Li, C. Dietary protein sources differentially affect the growth of akkermansia muciniphila and maintenance of the gut mucus barrier in mice. Mol. Nutr. Food Res. 2019, 63, 1900589. [Google Scholar] [CrossRef] [PubMed]
- Gruber, L.; Kisling, S.; Lichti, P.; Martin, F.P.; May, S.; Klingenspor, M.; Lichtenegger, M.; Rychlik, M.; Haller, D. High fat diet accelerates pathogenesis of murine crohn’s disease-like ileitis independently of obesity. PLoS ONE 2013, 8, e71161. [Google Scholar] [CrossRef] [PubMed]
- Lam, Y.Y.; Ha, C.W.Y.; Campbell, C.R.; Mitchell, A.J.; Dinudom, A.; Oscarsson, J.; Cook, D.I.; Hunt, N.H.; Caterson, I.D.; Holmes, A.J.; et al. Increased gut permeability and microbioa change associate with mesenteric fat inflammation and metabolic dysfunction in diet-induced obese mice. PLoS ONE 2012, 7, e34233. [Google Scholar] [CrossRef]
- Khan, S.; Waliullah, S.; Godfrey, V.; Khan, M.A.W.; Ramachandran, R.A.; Cantarel, B.; Behrendt, C.; Peng, L.; Hooper, L.V.; Zaki, H. Dietary simple sugars alter microbial ecology in the gut and promote colitis in mice. Sci. Transl. Med. 2020, 12, eaay6218. [Google Scholar] [CrossRef]
- Do, M.H.; Lee, E.; Oh, M.J.; Kim, Y.; Park, H.Y. High-glucose or -fructose diet cause changes of the gut microbiota and metabolic disorders in mice without body weight changes. Nutrients 2018, 10, 761. [Google Scholar] [CrossRef]
- Chassaing, B.; Koren, O.; Goodrich, J.K.; Poole, A.C.; Srinivasan, S.; Ley, R.E.; Gewirtz, A.T. Dietary emulsifiers impact the mouse gut microbiota promoting colitis and metabolic syndrome. Nature 2015, 519, 92–96. [Google Scholar] [CrossRef]
- Panduro, M.; Benoist, C.; Mathis, D. Tissue-Tregs. Annu. Rev. Immunol. 2016, 34, 609–633. [Google Scholar] [CrossRef]
- Tanoue, T.; Atarashi, K.; Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 2016, 16, 295–309. [Google Scholar] [CrossRef]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly-Y, M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef]
- Campbell, C.; McKenney, P.T.; Konstantinovsky, D.; Isaeva, O.I.; Schizas, M.; Verter, J.; Mai, C.; Jin, W.B.; Guo, C.J.; Violante, S. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature 2022, 581, 475–479. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Sun, X.; Oh, S.F.; Wu, M.; Zhang, Y.; Zheng, W.; Geva-Zatorysky, N.; Jupp, R.; Mathis, D.; Benoist, C.; et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 2020, 577, 410–415. [Google Scholar] [CrossRef]
- Lamas, B.; Richard, M.L.; Leducaq, V.; Pham, H.-P.; Michel, M.-L.; da Costa, G.; Bridonneau, C.; Jegou, S.; Hoffmann, T.W.; Natividad, J.M.; et al. CARD9 impacts colitis by altering gut microbiota metabolism of tryptophan into aryl hydrocarbon receptor ligands. Nat. Med. 2016, 22, 598–605. [Google Scholar] [CrossRef] [PubMed]
- Geva-Zatorsky, N.; Sefik, E.; Kua, L.; Pasman, L.; Tan, T.G.; Ortiz-Lopez, A.; Yanortsang, T.B.; Yang, L.; Jupp, R.; Mathis, D. Mining the human gut microbiota for immunomodulatory organisms. Cell 2017, 168, 928–943. [Google Scholar] [CrossRef]
- Fontenot, J.D.; Gavin, M.A.; Rudensky, A.Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 2003, 4, 330–336. [Google Scholar] [CrossRef]
- Sakai, R.; Komai, K.; Iizuka-Koga, M.; Yoshimura, A.; Ito, M. Regulatory T cells: Pathophysiological roles and clinical applications. Keio J. Med. 2020, 69, 1–15. [Google Scholar] [CrossRef]
- Mobley, A.R.; Jones, J.M.; Rodriguez, J.; Slavin, J.; Zelman, K.M. Identifying practical solutions to meet America’s fiber needs: Proceedings from the food & fiber summit. Nutrients 2014, 6, 2540–2551. [Google Scholar] [PubMed]
- Deehan, E.; Duar, R.M.; Armet, A.M.; Perez-Muñoz, M.E.; Jin, M.; Water, J. Modulation of the gastrointestinal microbiome with nondigestible fermentable carbohydrates to improve human health. Mirobiol. Spectr. 2017, 5, BAD-0019-2017. [Google Scholar] [CrossRef]
- Hamaker, B.R.; Tuncil, Y.E. A perspective on the complexity of dietary fiber structures and their potential effect on the gut microbiota. J. Mol. Biol. 2014, 426, 3638–3850. [Google Scholar] [CrossRef]
- Sonnenburg, E.D.; Sonnenburg, J.L. Starving our microbial self: The deleterious consequences of a diet deficient in microbiota-accessible carbohydrates. Cell Metab. 2014, 20, 779–786. [Google Scholar] [CrossRef]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Buton, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Sonnnenberg, E.D.; Smits, S.A.; Tikhonov, M.; Higginbottom, S.K.; Wingreen, N.S.; Sonnenberg, J.L. Diet-induced extinctions in the gut microbiota compound over generations. Nature 2016, 529, 212–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tuncil, Y.E.; Thakkar, R.D.; Arioglu-Tuncil, S.; Hamaker, B.R.; Lindemann, S.R. subtle variations in dietary-fiber fine structure differentially influence the composition and metabolic function of gut microbiota. mSphere 2020, 5, e00180-20. [Google Scholar] [CrossRef] [PubMed]
- Thakkar, R.D.; Tuncil, Y.E.; Hamaker, B.R.; Lindemann, S.R. Maize bran particle size governs the community composition and metabolic composition and metabolic output of human gut microbiota in in vitro fermentations. Front. Microboil. 2020, 11, 1009. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Chen, Y.; Wang, W.; Huang, C.; Hu, Y.; Hohnston, L.; Wang, F. Dietary supplementation with fine-grinding wheat bran improves lipid metabolism and inflammatory response via modulating the gut microbiota structure in pregnant sow. Front. Microbiol. 2022, 13, 835950. [Google Scholar] [CrossRef]
- Li, X.; Guo, J.; Ji, K.; Zhang, P. Bamboo shoot fiber prevents obesity in mice by modulating the gut microbiota. Sci. Rep. 2016, 6, 32953. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, J.M. Nutrition and gut health: The impact of specific dietary components-it’s not just five-a-day. Proc. Nutr. Soc. 2021, 80, 9–18. [Google Scholar] [CrossRef]
- Singh, V.; Yoeh, B.S.; Walker, R.E.; Xiao, X.; Saha, P.; Golonka, R.M.; Cai, J.; Bretin, A.C.A.; Cheng, X.; Liu, Q. Microbiota fermentation-NLRP3 axis shapes the impact of dietary fibers on intestinal inflammation. Gut 2019, 68, 1801–1812. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Bäckhed, F.; Fulton, L.; Gordon, J.I. Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 2008, 3, 213–223. [Google Scholar] [CrossRef]
- Hidebrandt, M.A.; Hoffmann, C.; Sherrill-Mix, S.S.; Keilbaugh, S.A.; Hamady, M.; Chen, Y.Y.; Knight, R.; Ahiama, R.S.; Bushman, F.; Wu, G.D. High-fat diet determines the composition of the murine gut microbiome independently of obesity. Gastroenterology 2009, 137, 1716–1724. [Google Scholar] [CrossRef]
- Devkota, S.; Wang, Y.; Musch, M.W.; Leone, V.; Fehlner-Peach, H.; Nadimpalli, A.; Antonopoulos, D.A.; Jabri, B.; Chang, E.B. dietary-fat-induced taurocholic acid promotes pathobiont expansion and colitis in IL10−/− mice. Nature 2012, 487, 104–108. [Google Scholar] [CrossRef]
- Marques, T.M.; Wall, R.; O’Sullivan, O.; Fitzgerald, G.F.; Shanahan, F.; Quigley, E.M.; Cotter, P.D.; Cryan, J.F.; Dinan, T.G.; Ross, R.P.; et al. Dietary trans-10, cis-12-conjugated linoleic acid alters fatty acid metabolism and microbiota composition in mice. Br. J. Nutr. 2015, 113, 728–738. [Google Scholar] [CrossRef] [Green Version]
- Watson, H.; Mitra, S.; Croden, F.C.; Taylor, M.; Wood, H.M.; Perry, S.L.; Spencer, J.A.; Quirke, P.; Toogood, G.J.; Lawton, C.L. A randomized trial of the effect of omega-3 polyunsaturated fatty acid supplements on the human intestinal microbiota. Gut 2018, 67, 1974–1983. [Google Scholar] [CrossRef]
- Uranga, J.A.; Lόpez-Miranda, V.; Lombό, F.; Abalo, R. Food, nutrients and nutraceuticals affecting the course of inflammatory bowel disease. Pharm. Rep. 2016, 68, 816–826. [Google Scholar] [CrossRef]
- Bakke, D.; Sun, J. Ancient nuclear receptor VDR with new functions: Microbiome and inflammation. Inflamm. Bowel. Dis. 2018, 24, 1149–1154. [Google Scholar] [CrossRef]
- Clark, A.; Mach, N. Role of vitamin D in the hygiene hypothesis: The inter play between vitamin D, vitamin D receptors, gut microbiota, and immune response. Front. Immunol. 2016, 7, 627. [Google Scholar] [CrossRef]
- Kang, S.W.; Kim, S.H.; Lee, N.; Lee, W.W.; Hwang, K.A.; Shin, M.S.; Lee, S.H.; Kim, W.U.; Kang, I. 1,25-Dihyroxyvitamin D3 promotes FOXP3 expression via binding to vitamin D response elements in its conserved noncoding sequence region. J. Immunol. 2012, 188, 5276–5282. [Google Scholar] [CrossRef]
- Lagisshetty, V.; Misharin, A.V.; Liu, N.Q.; Lisse, T.S.; Chun, R.F.; Ouyang, Y.; McLachlan, S.M.; Adams, J.S.; Hewison, M. Vitamin D deficiency in mice impairs colonic antibacterial activity and predisposes to colitis. Endocrinology 2010, 151, 2423–2432. [Google Scholar] [CrossRef]
- Jin, D.; Wu, S.; Zhang, Y.; Lu, R.; Xia, Y.; Dong, H.; Sun, J. Lack of vitamin D receptor causes dysbiosis and changes the functions of the murine intestinal microbiome. Clin. Ther. 2015, 37, 996–1009. [Google Scholar] [CrossRef]
- Wang, J.; Thingholm, L.B.; Skiecevičiené, J.; Rausch, P.; Kummen, M.; Hov, J.R.; Degenhardt, F.; Heinsen, F.A.; Rühlemann, M.C.; Szymczak, S.; et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 2016, 48, 1396–1406. [Google Scholar] [CrossRef]
- Snelson, M.; Clarke, R.E.; Nguyen, T.-V.; Penfold, S.A.; Forbes, J.M.; Tan, S.M.; Coughlan, M.T. Long term high protein diet feeding alters the microbiome and increases intestinal permeability, systemic inflammation and kidney injury in mice. Mol. Nutr. Food. Res. 2021, 65, 2000851. [Google Scholar] [CrossRef]
- Ko, G.J.; Rhee, C.M.; Kalantar-Zadeh, K.; Joshi, S. The effects of high-protein diets on kidney health and longevity. J. Am. Soc. Nephrol. 2020, 8, 1667. [Google Scholar] [CrossRef]
- Windey, K.; de Preter, V.; Verbeke, K. Relevance of protein fermentation to gut health. Mol. Nutr. Food. Res. 2012, 56, 184–196. [Google Scholar] [CrossRef]
- Wu, L.; Tang, Z.; Chen, H.; Ren, Z.; Ding, Q.; Liang, K.; Sun, Z. Mutual interaction between gut microbiota and protein/amino acid metabolism for host mucosal immunity and health. Anim. Nutr. 2021, 7, 11–16. [Google Scholar] [CrossRef]
- Beaumont, M.; Portune, K.J.; Steuer, N.; Lan, A.; Cerrudo, V.; Audebert, M.; Dumont, F.; Mancano, G.; Khodorova, N.; Andriamihaja, M.; et al. Quantity and source of dietary protein influence metabolic production by gut microbiota and rectal mucosa gene expression: A randomized, parallel, double-blind trail in overweight humans. Am. J. Clin. Nutr. 2017, 106, 1005–1019. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, Y.; Xu, Y.; Luo, T.; Ge, Y.; Jiang, Y.; Shi, Y.; Sun, J.; Le, G. Dietary methionine restriction improves the gut microbiota and reduces intestinal permeability and inflammation in high-fat-fed mice. Food Funct. 2019, 10, 5952–5968. [Google Scholar] [CrossRef]
- Mu, C.; Yang, Y.; Luo, Z.; Guan, L.; Zhu, W. The colonic microbiome and epithelial transcriptome are altered in rats fed a high-protein diet compared with a normal protein diet. J. Nutr. 2016, 146, 474–483. [Google Scholar] [CrossRef]
- Mu, C.; Yang, Y.; Luo, Z.; Zhu, W. Temporal microbiota changes of high-protein diet intake in a rat model. Anaerobe 2017, 47, 218–225. [Google Scholar] [CrossRef]
- Wu, S.; Bhat, Z.F.; Gounder, R.S.; Ahmed, I.A.M.; AI-Juhaimi, F.Y.; Ding, Y.; Bekhit, A.E.-D.A. Effect of dietary protein and processing on gut microbiota-a systemic review. Nutrients 2022, 14, 453. [Google Scholar] [CrossRef]
- Zhu, Y.; Shi, X.; Lin, X.; Ye, K.; Xu, X.; Li, C.; Zhou, G. Beef, chicken, and soy proteins in diets induce different gut microbiota and metabolites in rats. Front. Microbiol. 2017, 8, 1395. [Google Scholar] [CrossRef]
- Butteiger, D.N.; Hibberd, A.A.; McGraw, N.J.; Napawan, N.; Hall-Porter, J.M.; Krul, E. soy protein compared with milk protein in a western diet increases gut microbial diversity and reduces serum lipids in golden Syrian hamsters. J. Nutr. 2016, 146, 697–705. [Google Scholar] [CrossRef]
- Jensen, B.A.H.; Holm, J.B.; Larsen, I.S.; von Burg, N.; Derer, S.; Sonne, S.B.; Pärregaard, S.I.; Damgarrd, M.V.; Indrelid, S.A.; Rivollier, A. Lysates of Methyloccocus capsulatus Bath induce a lean-like microbiota, intestinal FoxP3+RORγt+IL-17+ Tregs and improve metabolism. Nat. Commun. 2021, 12, 1093. [Google Scholar] [CrossRef] [PubMed]
- Weisstaub, G.; Araya, M. Acute malnutrition in Latin America: The challenge of ending avoidable deaths. J. Pediatr. Gastroenterol. Nutr. 2008, 47, S10–S14. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, T.; Perlot, T.; Rehman, A.; Trichereau, J.; Ishiguro, H.; Paolino, M.; Sigl, V.; Hanada, T.; Hanada, R.; Lipinski, S.; et al. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature 2012, 487, 477–481. [Google Scholar] [CrossRef]
- Islam, J.; Sato, S.; Watanable, K.; Watanable, T.; Ardiansyah; Hirahara, K.; Aoyama, Y.; Tomita, S.; Aso, H.; Komai, M.; et al. Dietary tryptophan alleviates dextran sodium sulfate-induced colitis through aryl hydrocarbon receptor in mice. J. Nutr. Biochem. 2017, 42, 43–50. [Google Scholar] [CrossRef] [PubMed]
- Zelante, T.; Iannitti, R.G.; Cunhua, C.; de Luca, A.; Giovannini, G.; Pieraccini, G.; Zecchi, R.; D’Angelo, C.; Massi-Benedetti, C.; Fallarino, F.; et al. Tryptophan catabolites from microbiota engage an aryl hydrocarbon receptor and balance mucosal reactivity via interleutin-22. Immunity 2013, 39, 372–385. [Google Scholar] [CrossRef]
- Mezrich, J.D.; Fechner, J.H.; Zhang, X.; Johnson, B.P.; Burlingham, W.J.; Bradfield, C.A. An interaction between kynurenine and the aryl hydrocarbon receptor can generate regulatory T cells. J. Immunol. 2010, 185, 3190–3198. [Google Scholar] [CrossRef]
- Singh, N.; Gurav, A.; Sivaprakasam, S.; Brady, E.; Padia, R.; Shi, H.; Thangaralu, M.; Prasad, P.D.; Manicassamy, S.; Munn, D.H.; et al. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity 2014, 40, 128–139. [Google Scholar] [CrossRef]
- Kitsmoto, S.; Alteri, C.J.; Rodgigues, M.; Nagao-Kitamoto, H.; Sugihara, K.; Himpsl, S.D.; Bazzi, M.; Miyoshi, M.; Nishioka, T.; Hayashi, A.; et al. Dietary L-serine confers a competitive fitness advantage to Enterobacteriaceae in the inflamed gut. Nat. Microbiol. 2020, 5, 116–125. [Google Scholar] [CrossRef]
- Ozdal, T.; Sela, D.; Xiao, J.; Boyacioglu, D.; Chen, F.; Capanoglu, E. The reciprocal interactions between polyphenols and the gut microbiota and effects on bioaccessibility. Nutrients 2016, 8, 78. [Google Scholar] [CrossRef]
- Yang, S.; Li, L.; Yu, L.; Sun, L.; Li, K.; Tong, C.; Xu, W.; Cui, G.; Long, M.; Li, P. Selenium-enriched yeast reduces caecal pathological injuries and intervenes changes of the diversity of ceacal microbiota caused by Ochratoxin-A in broilers. Food Chem. Toxicol. 2020, 137, 111139. [Google Scholar] [CrossRef] [PubMed]
- Roca-Saavedra, P.; Mendez-Vilabrille, V.; Miranda, J.M.; Nebot, C.; Cardelle-Cobas, A.; Franco, C.M.; Cepeda, A. Food additives, contaminants and other minor components: Effects on human microbiota-a review. J. Physiol. Biochem. 2018, 74, 69–83. [Google Scholar] [CrossRef] [PubMed]
- Lloyd-Price, J.; Arze, C.; Ananthakrishnan, A.N.; Schirmer, M.; Avila-Pacheco, J.; Poon, T.W.; Andrews, E.; Ajami, N.; Bonham, K.S.; Brislawn, C.J.; et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel diseases. Nature 2019, 569, 655–662. [Google Scholar] [CrossRef] [PubMed]
- Schirmer, M.; Franzos, E.A.; Lloyd-Price, J.; Mciver, L.J.; Schwager, R.; Poon, T.W.; Ananthakrishnan, A.N.; Andrews, E.; Barron, G.; Lake, K.; et al. Dynamics of metatranscription in the inflammatory bowel disease gut microbiome. Nat. Microbiol. 2018, 3, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Franzosa, E.A.; Sirota-Madi, A.; Avila-Pacheco, J.; Fornelos, N.; Haiser, H.J.; Reinker, S.; Vatanen, T.; Hall, A.B.; Mallick, H.; Mciver, L.J.; et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat. Microbiol. 2019, 4, 293–305. [Google Scholar] [CrossRef]
- Britton, G.J.; Contijoch, E.J.; Mogno, I.; Vennaro, O.H.; Llewellyn, S.R.; Ng, R.; Li, Z.; Mortha, A.; Merad, M.; Das, A.; et al. Microbiotas from humans with inflammatory bowel disease alter the balance of gut Th17 and RORγt+ regulatory T cells and exacerbate colitis in mice. Immunity 2019, 50, 212–224. [Google Scholar] [CrossRef]
- Fritsch, J.; Garces, L.; Quintero, M.A.; Pignac-Kobinger, J.; Santander, A.M.; Fernández, I.; Ban, Y.J.; Kwon, D.; Phillips, M.C.; Knight, K.; et al. Low-fat, high-fiber diet reduces markers of inflammation and dysbiosis and improves quality of life in patents with ulcerative colitis. Clin. Gastroenterol. Hepatol. 2021, 19, 1189–1199. [Google Scholar] [CrossRef]
- Cox, S.R.; Prince, A.C.; Myers, C.E.; Irving, P.M.; Lindsay, J.O.; Lomer, M.C.; Whelan, K. Fermentable carbohydrates [FODMAPs] exacerbate functional gastrointestinal symptoms in patients with inflammatory bowel disease: A randomized, double-blinded, placebo-controlled, cross-over, re-challenge trial. J. Crohns Colitis. 2017, 11, 1420–1429. [Google Scholar] [CrossRef]
- Cox, S.R.; Lindsay, J.O.; Fromentin, S.; Stagg, A.J.; McCarthy, N.E.; Galleron, N.; Ibraim, S.B.; Roume, H.; Levenez, F.; Pons, N.; et al. Effects of low FODMAP diet on symptoms, fecal microbiome, and markers of inflammation in patients with quiescent inflammatory bowel disease in a randomized trial. Gastroenterology 2020, 158, 176–188. [Google Scholar] [CrossRef]
- Gibson, P.R.; Halmos, E.P.; Muir, J.G. FODMAPS, prebiotics and gut health-the FODMAP hypothesis revisited. Aliment. Pharmacol. Ther. 2020, 52, 233–246. [Google Scholar] [CrossRef]
- Li, P.; Xiao, N.; Zeng, L.; Xiao, J.; Huang, J.; Xu, Y.; Chen, Y.; Ren, Y.; Du, B. Structural characteristics of a mannoglucan isolated from Chinese yam and its treatment effects against gut microbiota dysbiosis and DSS-induced colitis in mice. Carbohydr. Polym. 2020, 250, 116958. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, N.; Kan, J.; Zhang, X.; Wu, X.; Sun, R.; Tang, S.; Liu, J.; Qian, C.; Jin, C. Structural characterization of water-soluble polysaccharide from Arctium lappa and its effects on colitis mice. Carbohr. Polym. 2019, 213, 89–99. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Chen, H.; Kan, J.; Gou, Y.; Liu, J.; Zhang, X.; Wu, X.; Tang, S.; Sun, R.; Qian, C.; et al. Anti-inflammatory properties and gut microbiota modulation of an alkali-soluble polysaccharide from purple sweet potato in DSS-induced colitis mice. Int. J. Biol. Macromol. 2020, 153, 708–722. [Google Scholar] [CrossRef]
- Gou, Y.; Sun, J.; Liu, J.; Chen, H.; Kan, J.; Qian, C.; Zhang, N.; Jin, C. Structural characterization of a water-soluble purple sweet potato polysaccharide and its effect on intestinal inflammation in mice. J. Funct. Foods 2019, 61, 103502. [Google Scholar] [CrossRef]
- Li, Q.; Wu, W.; Fang, X.; Chen, H.; Han, Y.; Liu, R.; Niu, B.; Gao, H. Structural characterization of a polysaccharide from bamboo (Phyllostachys edulis) shoot and its prevention effect on colitis mouse. Food Chem. 2022, 387, 132807. [Google Scholar] [CrossRef]
- Jin, M.; Wang, Y.; Yang, X.; Yin, H.; Nie, S.; Wu, X. Structural characterization of a polysaccharide extracted from noni (Morinda citrifolia L.) and its protective effect against DSS-induced bowel disease in mice. Food Hydrocol. 2019, 90, 189–197. [Google Scholar] [CrossRef]
- Ren, Y.; Geng, Y.; Du, Y.; Li, W.; Lu, Z.M.; Xu, H.Y.; Xu, G.H.; Shi, J.S.; Xu, Z.H. Polysaccharide of Hericium erinaceus attenuates colitis in C57BL/6 mice via regulation of oxidative stress, inflammation-related signaling pathways and modulating the composition of the gut microbiota. J. Nutr. Biochem. 2018, 57, 67–76. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Yuan, S.; Ye, J.; Wang, X.; Zhang, X.; Shen, J.; Yuan, M.; Liao, W. Polysacchride from flammuliana velutipes improves colitis via regulation of colonic microbial dysbiosis and inflammatory responses. Int. J. Biol. Macromol. 2020, 149, 1252–1261. [Google Scholar] [CrossRef] [PubMed]
- Kanwal, S.; Joseph, T.P.; Aliya, S.; Song, S.; Saleem, M.Z.; Nisar, M.A.; Wang, Y.; Meyiah, A.; Ma, Y. Attenuation of DSS induced colitis by Dictyophora indusiata polysaccharide (DIP) via modulation of gut microbiota and inflammatory related signaling pathways. J. Funct. Foods 2020, 64, 103641. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, A.C. What is the healthy microbiota composition? A changing ecosystem across age, environment, diet, and disease. Mciroorganisms 2019, 7, 14. [Google Scholar] [CrossRef]
- Hamada, T.; Keum, N.; Nishihara, R.; Ogino, S. Molecular pathological epidemiology: New developing frontiers of big data science to study etiologies and pathogenesis. J. Gastroenterol. 2017, 52, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Imchen, M.; Moopantakath, J.; Kumavath, R. Gut microbiota and antibiotics: Dysbiosis and antimicrobial resistance. Comp. Gut Microbiota 2022, 1, 374–386. [Google Scholar] [CrossRef]
- Blázquez, J.; Rodríguez-Beltrán, J.; Matic, I. Antibiotic-inducing genetic variation: How it arises and how it can be prevented. Annu. Rev. Microbiol. 2018, 72, 209–230. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Martins dos Santos, V.A.; Ott, S.J.; Moya, A. Gut microbiota disturbance during antibiotic therapy: A multi-omic approach. Gut Microbes 2014, 5, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Lanklema, J.M.; Cranendonk, D.R.; Belzer, C.; de vos, A.F.; de vos, W.M.; van der Poll, T.; Wiersinga, W.J. Antibiotic-induced gut microbiota disruption during human endotoxemia: A randomized controlled study. Gut 2017, 66, 1623–1630. [Google Scholar] [CrossRef]
- Suez, J.; Zmora, N.; Zilberman-Schapira, G.; Mor, U.; Dori-Bachash, M.; Bashiardes, S.; Zur, M.; Regev-Lehavi, D.; Brik, R.B.Z.; Federic, S.; et al. Post-antibiotic gut mucosal microbiome reconstitution is impaired by probiotics and improved by autologous FMT. Cell 2018, 174, 1406–1423. [Google Scholar] [CrossRef]
- Van Nood, E.; Vrieze, A.; Nieuwdrop, M.; Niuwdorp, M.; Fuentes, S.; Zoetendal, E.G.; de Vos, W.M.; Visser, C.E.; Kuijper, E.J.; Bartelsman, J.; et al. Duodenal infusion of donor feces for recurrent Clostridium difficile. N. Engl. J. Med. 2013, 368, 407–415. [Google Scholar] [CrossRef]
- Kootte, R.S.; Levin, E.; Salojävi, J.; Smits, L.P.; Hartstra, A.V.; Udayappan, S.D.; Hermes, G.; Bouter, K.E.; Koopen, A.M.; Hoist, J.J.; et al. Improvement of insulin sensitivity after lean donor feces in metabolic syndrome is driven by baseline intestinal microbiota composition. Cell Metab. 2017, 26, 611–619. [Google Scholar] [CrossRef]
- Moayyedi, P.; Surette, M.G.; Kim, P.T.; Libertucci, J.; Wolfe, M.; Onishi, C.; Armstrong, D.; Marshall, J.K.; Kassam, Z.; Reinisch, W.; et al. Fecal microbiota transplantation induces remission in patients with active ulcerative colitis in a randomized controlled trial. Gastroenterology 2015, 149, 102–109. [Google Scholar] [CrossRef]
- Costello, S.P.; Huges, P.A.; Waters, O.; Bryant, R.V.; Vincent, A.D.; Blatchford, P.; Katsikeros, R.; Makanyanga, J.; Campaniello, C.; Mavrangelos, C.; et al. Effects of fecal microbiota transplantation on 8-week ulcerative colitis: A randomized clinical trial. JAMA 2019, 321, 156–164. [Google Scholar] [CrossRef]
- Bullman, S.; Eggermont, A.; Johnston, C.D.; Zitvogel, L. Harnessing the microbiome to restore immunotherapy response. Nat. Cancer 2021, 2, 1301–1304. [Google Scholar] [CrossRef] [PubMed]
- Kwak, S.; Choi, J.; Hink, T.; Reske, K.A.; Blount, K.; Jones, C.; Bost, M.H.; Sun, X.; Burnham, C.A.N.; Dubberke, E.R.; et al. Impact of investigational microbiota therapeutic RBX2660 on the gut microbiome and resistome revealed by a placebo-controlled clinical trial. Microbiome 2020, 8, 125. [Google Scholar] [CrossRef] [PubMed]
- Ratner, M. Microbial cocktails raise bar for C. DIFF. treatment. Nat. Biotechnol. 2020, 38, 1366–1367. [Google Scholar] [CrossRef] [PubMed]
- Tanoue, T.; Morita, S.; Plichta, D.R.; Skelly, A.N.; Suda, W.; Sugiura, Y.; Narushima, S.; Vlamarks, H.; Motoo, I.; Sugita, K.; et al. A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 2019, 565, 600–605. [Google Scholar] [CrossRef]
- Inamura, K.; Hamada, T.; Bullman, S.; Ugai, T.; Yachida, S.; Ogino, S. Cancer as microenvironmental, systemic and environmental diseases: Opportunity for transdisciplinary microbiomics science. Gut 2022. on-line ahead of print. [Google Scholar] [CrossRef]
- Colombo, F.; ILLescas, O.; Noci, S.; Minnai, F.; Pintarelli, G.; Pettinicchio, A.; Vannelli, A.; Sorrentino, L.; Battaglia, L.; Cosimelli, M. Gut microbiota composition in colorectal cancer patients is genetically regulated. Sci. Rep. 2022, 12, 11424. [Google Scholar] [CrossRef]
- Hu, S.; Vila, A.V.; Gacesa, R.; Collij, V.; Stevens, C.; Fu, J.M.; Wrong, I.; Talkowski, M.E.; Rivas, M.A.; Imhann, F.; et al. Whole exome sequencing analyses reveal gene-microbiota interactions in the context of IBD. Gut 2021, 70, 285–296. [Google Scholar] [CrossRef]
- Ogino, S.; Nowak, J.A.; Hamada, T.; Milner, D.A.; Nishihara, R. Insights into pathogenic interactions among environment, host, and tumor at the crossroads of molecular pathology and epidermiology. Annu. Rev. Pathol. 2019, 14, 83–103. [Google Scholar] [CrossRef]
- Kausar, T.; Hanan, E.; Ayob, O.; Praween, B.; Azad, Z. A review on functional ingredients in red meat. Bioinformation 2019, 15, 358–363. [Google Scholar] [CrossRef]
| Polysaccharide | Food Source | Animal Model and Experimental Design | Changes in the Gut Microbiome and Inflammation | Reference |
|---|---|---|---|---|
| CYP-1 | Chinese yam, the rhizome of Dioscorea opposite Thunb | DSS-induced colitis in C57BL/6 mice 3% DSS treat 1 wk followed by CYP-1 for 7 d | ↑mRNA of ZO, claudin-1, occludin, connexin-43 ↑α-diversity ↑Firmicutes/Bacteroidetes ratio ↓Alistipes, Helicobacter ↓serum LPS, IL-18, TNF-α, IL-1β | [91] |
| ALP-1 | Root of Burdock Arctium lappa | DSS-induced colitis in ICR mice Pretreat with ALP-1 (300 mg/kg) for 7 d followed by 3% DSS for 7 d | ↑total gut bacteria ↑F/B ratio ↓Proteobacteria ↓Staphyloccus ↓colon and serum IL-1β, IL-6, TNF-α, ↑IL-10 | [92] |
| ASPP | Purple sweet potato | DSS-induced colitis in ICR mice 400 mg/kg with 2.5% DSS for 7 d | higher total bacteria higher diversity ↑ Firmicutes/Bacteroidetes ratio, ↓ Proteobacteria ↑acetate, propionate ↓IL-1β, IL-6, TNF-α Changes in 25 functional pathways | [93] |
| WPSPP-1 | Purple sweet potato | DSS-induced colitis in ICR mice 400 mg/kg with 4% DSS for 7 d | higher total bacteria ↑ Firmicutes/Bacteroidetes ratio, ↑acetate, propionate, butyrate ↓ Proteobacteria ↓IL-1β, IL-6, TNF-α, ↑IL-10 | [94] |
| BSDF-1 | Bamboo shoot Phyllostachys edulis | DSS-induced colitis in C57BL/6 mice Pretreat (100, 200, 400 mg/kg) for 7 d followed by 4% DSS for 7 d | ↑ mRNA of occludens-1, claudin-1, occluding ↓Parabaceroides, Mucispirillum ↓Helicobacter, Bacteroides ↓Streptococcus ↑Prevotella, Alitipes, Anaerostipes ↑Odoribacter, Bifidobacterium ↑butyricimonas, Lactobaccillus ↓NF-κB, NLR | [95] |
| NFP | Noni fruit Morinda citrifolia L. | DSS-induced colitis in C57BL.6 mice 10 mg/kg with 2% DSS for 11 d | redistribution of ZO-1, occludin in colonic epithelial cells thicker mucus layer more goblet cells | [96] |
| HECP | Mushroom Hericium erinaceus | DSS-induced colitis in C57BL/6 mice 250, 500 mg/kg with 2% DSS for 7 d | ↓Verrucomicrobia ↓Actinobacteria ↓amino acids metabolism pathways ↓NF-κB, Akt, MAPK ↓oxidative stress | [97] |
| FVP | Mushroom Flammuliana velutipes | DSS-induced colitis in Spragur Dawley rats Pretreat with 50, 100, 200 mg/kg followed by 4.5% DSS for 7 d | ↑ α-diversity ↑ F/B ratio ↑Ruminal butyrivibrios ↑Roseburia, S24-7 ↓Helicobacteraceae ↑butyrate, isovaleric acid, valeric acids | [98] |
| DIP | Mushroom Dictyophora indusiata | DSS-induce colitis in BALB/c mice Pretreat with 10, 33 mg/kg DIP for 7 d followed by 3.5% DSS for 7 d | ↑total gut bacteria ↑ F/B ratio ↓Enterobacteriales ↓Gammaproteobacteria ↓inflammatory cytokines ↓oxidative stress ↑mucins, goblet cells | [99] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, P. Influence of Foods and Nutrition on the Gut Microbiome and Implications for Intestinal Health. Int. J. Mol. Sci. 2022, 23, 9588. https://doi.org/10.3390/ijms23179588
Zhang P. Influence of Foods and Nutrition on the Gut Microbiome and Implications for Intestinal Health. International Journal of Molecular Sciences. 2022; 23(17):9588. https://doi.org/10.3390/ijms23179588
Chicago/Turabian StyleZhang, Ping. 2022. "Influence of Foods and Nutrition on the Gut Microbiome and Implications for Intestinal Health" International Journal of Molecular Sciences 23, no. 17: 9588. https://doi.org/10.3390/ijms23179588
APA StyleZhang, P. (2022). Influence of Foods and Nutrition on the Gut Microbiome and Implications for Intestinal Health. International Journal of Molecular Sciences, 23(17), 9588. https://doi.org/10.3390/ijms23179588





