Integrated DNA Copy Number and Expression Profiling Identifies IGF1R as a Prognostic Biomarker in Pediatric Osteosarcoma
Abstract
1. Introduction
2. Results
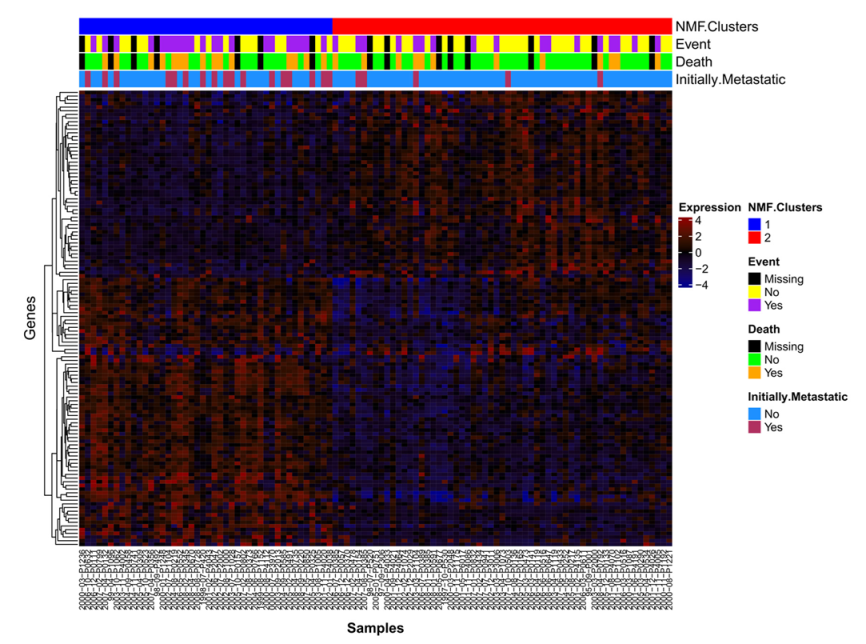
2.1. NMF Clustering of Gene Expression Data Identifies Two Distinct Clusters
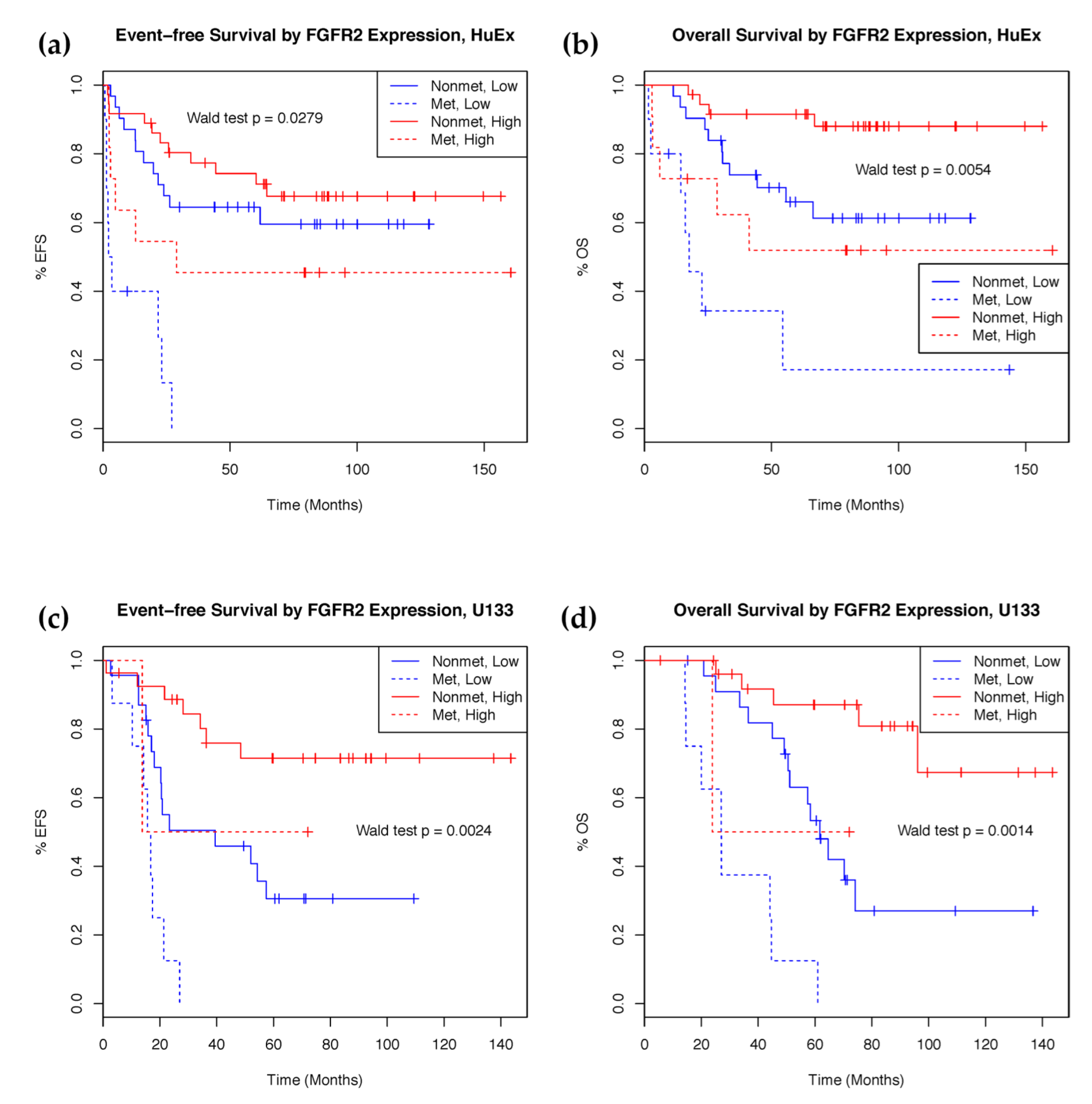
2.2. mRNA Expression Predicts Clinical Outcomes
2.3. Characterization of Osteosarcoma Copy Number Aberrations
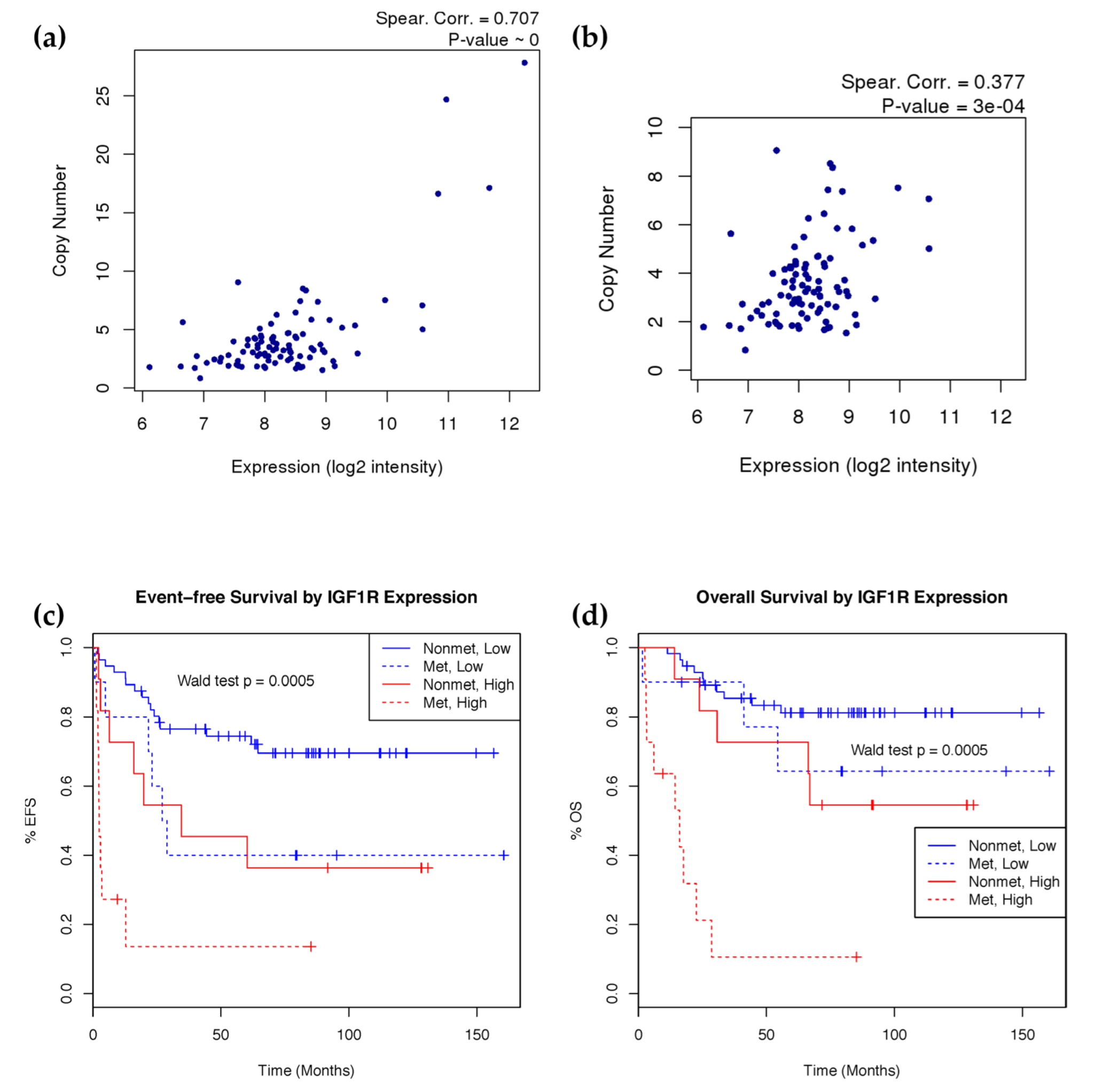
2.4. IGF1R Amplification Is Associated with Higher Expression and Worse Prognosis
3. Discussion
4. Materials and Methods
4.1. Patient Samples
4.2. Single Nucleotide Polymorphism Array Profiling
4.3. mRNA Expression Profiling
4.4. mRNA Clustering
4.5. Differential Expression and Pathway Analysis
4.6. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ries, L.A.G.; Smith, M.A.; Gurney, J.; Linet, M.; Tamra, T.; Young, J.; Bunin, G. Cancer Incidence and Survival among Children and Adolescents: United States SEER Program 1975–1995; National Cancer Institute: Bethesda, MD, USA, 1999.
- Rainusso, N.; Wang, L.L.; Yustein, J.T. The adolescent and young adult with cancer: State of the art-bone tumors. Curr. Oncol. Rep. 2013, 15, 296–307. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.W.; Squire, J.A.; Zielenska, M. The genetics of osteosarcoma. Sarcoma 2012, 2012, 627254. [Google Scholar] [CrossRef] [PubMed]
- Brun, J.; Dieudonne, F.X.; Marty, C.; Muller, J.; Schule, R.; Patino-Garcia, A.; Lecanda, F.; Fromigue, O.; Marie, P.J. FHL2 silencing reduces Wnt signaling and osteosarcoma tumorigenesis in vitro and in vivo. PLoS ONE 2013, 8, e55034. [Google Scholar] [CrossRef] [PubMed]
- Rubin, E.M.; Guo, Y.; Tu, K.; Xie, J.; Zi, X.; Hoang, B.H. Wnt inhibitory factor 1 decreases tumorigenesis and metastasis in osteosarcoma. Mol. Cancer Ther. 2010, 9, 1535–7163. [Google Scholar] [CrossRef] [PubMed]
- Engin, F.; Bertin, T.; Ma, O.; Jiang, M.M.; Wang, L.; Sutton, R.E.; Donehower, L.A.; Lee, B. Notch signaling contributes to the pathogenesis of human osteosarcomas. Hum. Mol. Genet. 2009, 18, 1464–1470. [Google Scholar] [CrossRef]
- Sadikovic, B.; Yoshimoto, M.; Chilton-MacNeill, S.; Thorner, P.; Squire, J.A.; Zielenska, M. Identification of interactive networks of gene expression associated with osteosarcoma oncogenesis by integrated molecular profiling. Hum. Mol. Genet. 2009, 18, 1962–1975. [Google Scholar] [CrossRef]
- Khanna, C.; Wan, X.; Bose, S.; Cassaday, R.; Olomu, O.; Mendoza, A.; Yeung, C.; Gorlick, R.; Hewitt, S.M.; Helman, L.J. The membrane-cytoskeleton linker ezrin is necessary for osteosarcoma metastasis. Nat. Med. 2004, 10, 182. [Google Scholar] [CrossRef]
- Sadikovic, B.; Yoshimoto, M.; Al-Romaih, K.; Maire, G.; Zielenska, M.; Squire, J.A. In vitro analysis of integrated global high-resolution DNA methylation profiling with genomic imbalance and gene expression in osteosarcoma. PLoS ONE 2008, 3, e2834. [Google Scholar] [CrossRef]
- Zhang, W.; Qian, J.X.; Yi, H.L.; Yang, Z.D.; Wang, C.F.; Chen, J.Y.; Wei, X.Z.; Fu, Q.; Ma, H. The microRNA-29 plays a central role in osteosarcoma pathogenesis and progression. Mol. Biol. 2012, 46, 622–627. [Google Scholar] [CrossRef]
- Cai, H.; Lin, L.; Cai, H.; Tang, M.; Wang, Z. Prognostic evaluation of microRNA-210 expression in pediatric osteosarcoma. Med. Oncol. 2013, 30, 499. [Google Scholar] [CrossRef]
- Gougelet, A.; Pissaloux, D.; Besse, A.; Perez, J.; Duc, A.; Dutour, A.; Blay, J.Y.; Alberti, L. Micro-RNA profiles in osteosarcoma as a predictive tool for ifosfamide response. Int. J. Cancer 2011, 129, 680–690. [Google Scholar] [CrossRef]
- Zhao, G.; Cai, C.; Yang, T.; Qiu, X.; Liao, B.; Li, W.; Ji, Z.; Zhao, J.; Zhao, H.; Guo, M. MicroRNA-221 induces cell survival and cisplatin resistance through PI3K/Akt pathway in human osteosarcoma. PLoS ONE 2013, 8, e53906. [Google Scholar]
- Gao, Y.; Feng, B.; Han, S.; Zhang, K.; Chen, J.; Li, C.; Wang, R.; Chen, L. The Roles of MicroRNA-141 in Human Cancers: From Diagnosis to Treatment. Cell Physiol. Biochem. 2016, 38, 427–448. [Google Scholar] [CrossRef]
- Maire, G.; Martin, J.W.; Yoshimoto, M.; Chilton-MacNeill, S.; Zielenska, M.; Squire, J.A. Analysis of miRNA-gene expression-genomic profiles reveals complex mechanisms of microRNA deregulation in osteosarcoma. Cancer Genet. 2011, 204, 138–146. [Google Scholar] [CrossRef]
- Won, K.Y.; Kim, Y.W.; Kim, H.-S.; Lee, S.K.; Jung, W.-W.; Park, Y.-K. MicroRNA-199b-5p is involved in the Notch signaling pathway in osteosarcoma. Hum. Pathol. 2013, 44, 1648–1655. [Google Scholar] [CrossRef]
- Miao, J.; Wu, S.; Peng, Z.; Tania, M.; Zhang, C. MicroRNAs in osteosarcoma: Diagnostic and therapeutic aspects. Tumour Biol. 2013, 34, 2093–2098. [Google Scholar] [CrossRef]
- Ottaviani, G.; Jaffe, N. The epidemiology of osteosarcoma. Cancer Treat. Res. 2009, 152, 3–13. [Google Scholar] [CrossRef]
- Goorin, A.M.; Schwartzentruber, D.J.; Devidas, M.; Gebhardt, M.C.; Ayala, A.G.; Harris, M.B.; Helman, L.J.; Grier, H.E.; Link, M.P.; Pediatric Oncology Group. Presurgical chemotherapy compared with immediate surgery and adjuvant chemotherapy for nonmetastatic osteosarcoma: Pediatric Oncology Group Study POG-8651. J. Clin. Oncol. 2003, 21, 1574–1580. [Google Scholar] [CrossRef]
- Bielack, S.S.; Kempf-Bielack, B.; Delling, G.n.; Exner, G.U.; Flege, S.; Helmke, K.; Kotz, R.; Salzer-Kuntschik, M.; Werner, M.; Winkelmann, W. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: An analysis of 1702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J. Clin. Oncol. 2002, 20, 776–790. [Google Scholar] [CrossRef]
- Marina, N.; Gebhardt, M.; Teot, L.; Gorlick, R. Biology and therapeutic advances for pediatric osteosarcoma. Oncologist 2004, 9, 422–441. [Google Scholar] [CrossRef]
- Gualberto, A.; Pollak, M. Emerging role of insulin-like growth factor receptor inhibitors in oncology: Early clinical trial results and future directions. Oncogene 2009, 28, 3009. [Google Scholar] [CrossRef]
- Pappo, A.S.; Vassal, G.; Crowley, J.J.; Bolejack, V.; Hogendoorn, P.C.; Chugh, R.; Ladanyi, M.; Grippo, J.F.; Dall, G.; Staddon, A.P. A phase 2 trial of R1507, a monoclonal antibody to the insulin-like growth factor-1 receptor (IGF-1R), in patients with recurrent or refractory rhabdomyosarcoma, osteosarcoma, synovial sarcoma, and other soft tissue sarcomas: Results of a Sarcoma Alliance for Research Through Collaboration study. Cancer 2014, 120, 2448–2456. [Google Scholar]
- Man, T.-K.; Lu, X.-Y.; Jaeweon, K.; Perlaky, L.; Harris, C.P.; Shah, S.; Ladanyi, M.; Gorlick, R.; Lau, C.C.; Rao, P.H. Genome-wide array comparative genomic hybridization analysis reveals distinct amplifications in osteosarcoma. BMC Cancer 2004, 4, 45. [Google Scholar] [CrossRef]
- Freeman, S.S.; Allen, S.W.; Ganti, R.; Wu, J.; Ma, J.; Su, X.; Neale, G.; Dome, J.S.; Daw, N.C.; Khoury, J.D. Copy number gains in EGFR and copy number losses in PTEN are common events in osteosarcoma tumors. Cancer Interdiscip. Int. J. Am. Cancer Soc. 2008, 113, 1453–1461. [Google Scholar]
- Kuijjer, M.L.; Rydbeck, H.; Kresse, S.H.; Buddingh, E.P.; Lid, A.B.; Roelofs, H.; Bürger, H.; Myklebost, O.; Hogendoorn, P.C.; Meza-Zepeda, L.A. Identification of osteosarcoma driver genes by integrative analysis of copy number and gene expression data. Genes Chromosomes Cancer 2012, 51, 696–706. [Google Scholar] [CrossRef]
- Lockwood, W.W.; Stack, D.; Morris, T.; Grehan, D.; O’Keane, C.; Stewart, G.L.; Cumiskey, J.; Lam, W.L.; Squire, J.A.; Thomas, D.M. Cyclin E1 is amplified and overexpressed in osteosarcoma. J. Mol. Diagn. 2011, 13, 289–296. [Google Scholar] [CrossRef]
- Bridge, J.A.; Nelson, M.; McComb, E.; McGuire, M.H.; Rosenthal, H.; Vergara, G.; Maale, G.E.; Spanier, S.; Neff, J.R. Cytogenetic findings in 73 osteosarcoma specimens and a review of the literature. Cancer Genet. Cytogenet. 1997, 95, 74–87. [Google Scholar] [CrossRef]
- Mertens, F.; Mandahl, N.; Örndal, C.; Baldetorp, B.; Bauer, H.; Rydholm, A.; Wiebe, T.; Willen, H.; Åkerman, M.; Heim, S. Cytogenetic findings in 33 osteosarcomas. Int. J. Cancer 1993, 55, 44–50. [Google Scholar] [CrossRef]
- Visani, G.; Sapienza, M.R.; Isidori, A.; Tripodo, C.; Laginestra, M.A.; Righi, S.; Sacchetti, C.A.S.; Gazzola, A.; Mannu, C.; Rossi, M. SNPs array karyotyping reveals a novel recurrent 20p13 amplification in primary myelofibrosis. PLoS ONE 2011, 6, e27560. [Google Scholar] [CrossRef]
- Melchor, L.; Saucedo-Cuevas, L.P.; Muñoz-Repeto, I.; Rodríguez-Pinilla, S.M.; Honrado, E.; Campoverde, A.; Palacios, J.; Nathanson, K.L.; García, M.J.; Benítez, J. Comprehensive characterization of the DNA amplification at 13q34 in human breast cancer reveals TFDP1 and CUL4A as likely candidate target genes. Breast Cancer Res. 2009, 11, R86. [Google Scholar] [CrossRef]
- Day, E.; Poulogiannis, G.; McCaughan, F.; Mulholland, S.; Arends, M.J.; Ibrahim, A.E.; Dear, P.H. IRS2 is a candidate driver oncogene on 13q34 in colorectal cancer. Int. J. Exp. Pathol. 2013, 94, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.-T.; You, H.-L.; Weng, S.-W.; Wei, Y.-C.; Eng, H.-L.; Huang, W.-T. Recurrent amplification at 13q34 targets at CUL4A, IRS2, and TFDP1 as an independent adverse prognosticator in Intrahepatic Cholangiocarcinoma. PLoS ONE 2015, 10, e0145388. [Google Scholar] [CrossRef] [PubMed]
- Ulaner, G.A.; Hoffman, A.R.; Otero, J.; Huang, H.Y.; Zhao, Z.; Mazumdar, M.; Gorlick, R.; Meyers, P.; Healey, J.H.; Ladanyi, M. Divergent patterns of telomere maintenance mechanisms among human sarcomas: Sharply contrasting prevalence of the alternative lengthening of telomeres mechanism in Ewing’s sarcomas and osteosarcomas. Genes Chromosomes Cancer 2004, 41, 155–162. [Google Scholar] [CrossRef]
- Wadayama, B.; Toguchida, J.; Shimizu, T.; Ishizaki, K.; Sasaki, M.S.; Kotoura, Y.; Yamamuro, T. Mutation spectrum of the retinoblastoma gene in osteosarcomas. Cancer Res. 1994, 54, 3042–3048. [Google Scholar] [PubMed]
- Araki, N.; Uchida, A.; Kimura, T.; Yoshikawa, H.; Aoki, Y.; Ueda, T.; Takai, S.; Miki, T.; Ono, K. Involvement of the retinoblastoma gene in primary osteosarcomas and other bone and soft-tissue tumors. Clin. Orthop. Relat. Res. 1991, 270, 271–277. [Google Scholar] [CrossRef]
- Kresse, S.H.; Ohnstad, H.O.; Paulsen, E.B.; Bjerkehagen, B.; Szuhai, K.; Serra, M.; Schaefer, K.-L.; Myklebost, O.; Meza-Zepeda, L.A. LSAMP, a novel candidate tumor suppressor gene in human osteosarcomas, identified by array comparative genomic hybridization. Genes Chromosomes Cancer 2009, 48, 679–693. [Google Scholar] [CrossRef]
- Pasic, I.; Shlien, A.; Durbin, A.D.; Stavropoulos, D.J.; Baskin, B.; Ray, P.N.; Novokmet, A.; Malkin, D. Recurrent Focal Copy-Number Changes and Loss of Heterozygosity Implicate Two Noncoding RNAs and One Tumor Suppressor Gene at Chromosome 3q13.31 in Osteosarcoma. Cancer Res. 2010, 70, 160–171. [Google Scholar] [CrossRef]
- Yen, C.C.; Chen, W.M.; Chen, T.H.; Chen, W.Y.; Chen, P.C.; Chiou, H.J.; Hung, G.Y.; Wu, H.T.; Wei, C.J.; Shiau, C.Y.; et al. Identification of chromosomal aberrations associated with disease progression and a novel 3q13.31 deletion involving LSAMP gene in osteosarcoma. Int. J. Oncol. 2009, 35, 775–788. [Google Scholar]
- Righi, A.; Gambarotti, M.; Longo, S.; Benini, S.; Gamberi, G.; Cocchi, S.; Vanel, D.; Picci, P.; Bertoni, F.; Simoni, A.; et al. Small cell osteosarcoma: Clinicopathologic, immunohistochemical, and molecular analysis of 36 cases. Am. J. Surg. Pathol. 2015, 39, 691–699. [Google Scholar] [CrossRef]
- Chow, J.P.; Poon, R.Y. The CDK1 inhibitory kinase MYT1 in DNA damage checkpoint recovery. Oncogene 2013, 32, 4778–4788. [Google Scholar] [CrossRef]
- Ueda, T.; Healey, J.H.; Huvos, A.G.; Ladanyi, M. Amplification of the MYC Gene in Osteosarcoma Secondary to Paget’s Disease of Bone. Sarcoma 1997, 1, 131–134. [Google Scholar] [CrossRef]
- Shimizu, T.; Ishikawa, T.; Sugihara, E.; Kuninaka, S.; Miyamoto, T.; Mabuchi, Y.; Matsuzaki, Y.; Tsunoda, T.; Miya, F.; Morioka, H.; et al. c-MYC overexpression with loss of Ink4a/Arf transforms bone marrow stromal cells into osteosarcoma accompanied by loss of adipogenesis. Oncogene 2010, 29, 5687–5699. [Google Scholar] [CrossRef]
- Scionti, I.; Michelacci, F.; Pasello, M.; Hattinger, C.M.; Alberghini, M.; Manara, M.C.; Bacci, G.; Ferrari, S.; Scotlandi, K.; Picci, P.; et al. Clinical impact of the methotrexate resistance-associated genes C-MYC and dihydrofolate reductase (DHFR) in high-grade osteosarcoma. Ann. Oncol. 2008, 19, 1500–1508. [Google Scholar] [CrossRef]
- Gamberi, G.; Benassi, M.S.; Bohling, T.; Ragazzini, P.; Molendini, L.; Sollazzo, M.R.; Pompetti, F.; Merli, M.; Magagnoli, G.; Balladelli, A.; et al. C-myc and c-fos in human osteosarcoma: Prognostic value of mRNA and protein expression. Oncology 1998, 55, 556–563. [Google Scholar] [CrossRef]
- Ladanyi, M.; Park, C.K.; Lewis, R.; Jhanwar, S.C.; Healey, J.H.; Huvos, A.G. Sporadic amplification of the MYC gene in human osteosarcomas. Diagn. Mol. Pathol. 1993, 2, 163–167. [Google Scholar] [CrossRef]
- Kuijjer, M.L.; Peterse, E.F.; van den Akker, B.E.; Briaire-de Bruijn, I.H.; Serra, M.; Meza-Zepeda, L.A.; Myklebost, O.; Hassan, A.B.; Hogendoorn, P.C.; Cleton-Jansen, A.-M. IR/IGF1R signaling as potential target for treatment of high-grade osteosarcoma. BMC Cancer 2013, 13, 245. [Google Scholar] [CrossRef]
- Pollak, M. The insulin and insulin-like growth factor receptor family in neoplasia: An update. Nat. Rev. Cancer 2012, 12, 159–169. [Google Scholar] [CrossRef]
- Humbel, R.E. Insulin-like growth factors I and II. Eur. J. Biochem. 1990, 190, 445–462. [Google Scholar] [CrossRef]
- Schmid, C. Insulin-like growth factors. Cell Biol. Int. 1995, 19, 445–457. [Google Scholar] [CrossRef]
- Denley, A.; Cosgrove, L.J.; Booker, G.W.; Wallace, J.C.; Forbes, B.E. Molecular interactions of the IGF system. Cytokine Growth Factor Rev. 2005, 16, 421–439. [Google Scholar] [CrossRef]
- Yu, H.; Mistry, J.; Nicar, M.J.; Khosravi, M.J.; Diamandis, A.; van Doorn, J.; Juul, A. Insulin-like growth factors (IGF-I, free IGF-I, and IGF-II) and insulin-like growth factor binding proteins (IGFBP-2, IGFBP-3, IGFBP-6, and ALS) in blood circulation. J. Clin. Lab. Anal. 1999, 13, 166–172. [Google Scholar] [CrossRef]
- Rikhof, B.; de Jong, S.; Suurmeijer, A.J.; Meijer, C.; van der Graaf, W.T. The insulin-like growth factor system and sarcomas. J. Pathol. 2009, 217, 469–482. [Google Scholar] [CrossRef]
- Burrow, S.; Andrulis, I.L.; Pollak, M.; Bell, R.S. Expression of insulin-like growth factor receptor, IGF-1, and IGF-2 in primary and metastatic osteosarcoma. J. Surg. Oncol. 1998, 69, 21–27. [Google Scholar] [CrossRef]
- Cao, Y.; Roth, M.; Piperdi, S.; Montoya, K.; Sowers, R.; Rao, P.; Geller, D.; Houghton, P.; Kolb, E.A.; Gill, J.; et al. Insulin-like growth factor 1 receptor and response to anti-IGF1R antibody therapy in osteosarcoma. PLoS ONE 2014, 9, e106249. [Google Scholar] [CrossRef]
- Wagner, L.M.; Fouladi, M.; Ahmed, A.; Krailo, M.D.; Weigel, B.; DuBois, S.G.; Doyle, L.A.; Chen, H.; Blaney, S.M. Phase II study of cixutumumab in combination with temsirolimus in pediatric patients and young adults with recurrent or refractory sarcoma: A report from the Children’s Oncology Group. Pediatr. Blood Cancer 2015, 62, 440–444. [Google Scholar] [CrossRef]
- Wilkie, A.O.M.; Patey, S.J.; Kan, S.-h.; van den Ouweland, A.M.W.; Hamel, B.C.J. FGFs, their receptors, and human limb malformations: Clinical and molecular correlations. Am. J. Med. Genet. 2002, 112, 266–278. [Google Scholar] [CrossRef]
- Mendoza, S.; David, H.; Gaylord, G.M.; Miller, C.W. Allelic loss at 10q26 in osteosarcoma in the region of the BUB3 and FGFR2 genes. Cancer Genet. Cytogenet. 2005, 158, 142–147. [Google Scholar] [CrossRef]
- Walkley, C.R.; Qudsi, R.; Sankaran, V.G.; Perry, J.A.; Gostissa, M.; Roth, S.I.; Rodda, S.J.; Snay, E.; Dunning, P.; Fahey, F.H.; et al. Conditional mouse osteosarcoma, dependent on p53 loss and potentiated by loss of Rb, mimics the human disease. Genes Dev. 2008, 22, 1662–1676. [Google Scholar] [CrossRef]
- Diez de Medina, S.G.; Chopin, D.; El Marjou, A.; Delouvee, A.; LaRochelle, W.J.; Hoznek, A.; Abbou, C.; Aaronson, S.A.; Thiery, J.P.; Radvanyi, F. Decreased expression of keratinocyte growth factor receptor in a subset of human transitional cell bladder carcinomas. Oncogene 1997, 14, 323–330. [Google Scholar] [CrossRef][Green Version]
- Naimi, B.; Latil, A.; Fournier, G.; Mangin, P.; Cussenot, O.; Berthon, P. Down-regulation of (IIIb) and (IIIc) isoforms of fibroblast growth factor receptor 2 (FGFR2) is associated with malignant progression in human prostate. Prostate 2002, 52, 245–252. [Google Scholar] [CrossRef]
- Amann, T.; Bataille, F.; Spruss, T.; Dettmer, K.; Wild, P.; Liedtke, C.; Muhlbauer, M.; Kiefer, P.; Oefner, P.J.; Trautwein, C.; et al. Reduced expression of fibroblast growth factor receptor 2IIIb in hepatocellular carcinoma induces a more aggressive growth. Am. J. Pathol. 2010, 176, 1433–1442. [Google Scholar] [CrossRef] [PubMed]
- Gartside, M.G.; Chen, H.; Ibrahimi, O.A.; Byron, S.A.; Curtis, A.V.; Wellens, C.L.; Bengston, A.; Yudt, L.M.; Eliseenkova, A.V.; Ma, J.; et al. Loss-of-function fibroblast growth factor receptor-2 mutations in melanoma. Mol. Cancer Res. 2009, 7, 41–54. [Google Scholar] [CrossRef] [PubMed]
- Babina, I.S.; Turner, N.C. Advances and challenges in targeting FGFR signalling in cancer. Nat. Rev. Cancer 2017, 17, 318–332. [Google Scholar] [CrossRef] [PubMed]
- Dutt, A.; Salvesen, H.B.; Chen, T.H.; Ramos, A.H.; Onofrio, R.C.; Hatton, C.; Nicoletti, R.; Winckler, W.; Grewal, R.; Hanna, M.; et al. Drug-sensitive FGFR2 mutations in endometrial carcinoma. Proc. Natl. Acad. Sci. USA 2008, 105, 8713–8717. [Google Scholar] [CrossRef]
- Greenman, C.; Stephens, P.; Smith, R.; Dalgliesh, G.L.; Hunter, C.; Bignell, G.; Davies, H.; Teague, J.; Butler, A.; Stevens, C.; et al. Patterns of somatic mutation in human cancer genomes. Nature 2007, 446, 153–158. [Google Scholar] [CrossRef]
- Su, N.; Jin, M.; Chen, L. Role of FGF/FGFR signaling in skeletal development and homeostasis: Learning from mouse models. Bone Res. 2014, 2, 14003. [Google Scholar] [CrossRef]
- Mansukhani, A.; Bellosta, P.; Sahni, M.; Basilico, C. Signaling by fibroblast growth factors (FGF) and fibroblast growth factor receptor 2 (FGFR2)-activating mutations blocks mineralization and induces apoptosis in osteoblasts. J. Cell Biol. 2000, 149, 1297–1308. [Google Scholar] [CrossRef]
- Wakahara, T.; Kusu, N.; Yamauchi, H.; Kimura, I.; Konishi, M.; Miyake, A.; Itoh, N. Fibin, a novel secreted lateral plate mesoderm signal, is essential for pectoral fin bud initiation in zebrafish. Dev. Biol. 2007, 303, 527–535. [Google Scholar] [CrossRef]
- Lakner, J.; Seyer, C.; Hermsdorf, T.; Schoneberg, T. Characterization of the expression, promoter activity and molecular architecture of fibin. BMC Biochem. 2011, 12, 26. [Google Scholar] [CrossRef]
- Rhim, J.S.; Putman, D.L.; Arnstein, P.; Huebner, R.J.; McAllister, R.M. Characterization of human cells transformed in vitro by N-methyl-N′-nitro-N-nitrosoguanidine. Int. J. Cancer 1977, 19, 505–510. [Google Scholar] [CrossRef]
- Flores, R.J.; Li, Y.; Yu, A.; Shen, J.; Rao, P.H.; Lau, S.S.; Vannucci, M.; Lau, C.C.; Man, T.K. A systems biology approach reveals common metastatic pathways in osteosarcoma. BMC Syst. Biol. 2012, 6, 50. [Google Scholar] [CrossRef]
- Fogh, J.; Fogh, J.M.; Orfeo, T. One hundred and twenty-seven cultured human tumor cell lines producing tumors in nude mice. J. Natl. Cancer Inst. 1977, 59, 221–226. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hadley, D.; Liu, R.; Glessner, J.; Grant, S.F.; Hakonarson, H.; Bucan, M. PennCNV: An integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 2007, 17, 1665–1674. [Google Scholar] [CrossRef]
- Van Loo, P.; Nordgard, S.H.; Lingjærde, O.C.; Russnes, H.G.; Rye, I.H.; Sun, W.; Weigman, V.J.; Marynen, P.; Zetterberg, A.; Naume, B. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. USA 2010, 107, 16910–16915. [Google Scholar] [CrossRef]
- Mermel, C.H.; Schumacher, S.E.; Hill, B.; Meyerson, M.L.; Beroukhim, R.; Getz, G. GISTIC2. 0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 2011, 12, R41. [Google Scholar] [CrossRef]
- Meyer, L.R.; Zweig, A.S.; Hinrichs, A.S.; Karolchik, D.; Kuhn, R.M.; Wong, M.; Sloan, C.A.; Rosenbloom, K.R.; Roe, G.; Rhead, B. The UCSC Genome Browser database: Extensions and updates 2013. Nucleic Acids Res. 2012, 41, D64–D69. [Google Scholar] [CrossRef]
- Gaujoux, R.; Seoighe, C. A flexible R package for nonnegative matrix factorization. BMC Bioinform. 2010, 11, 367. [Google Scholar] [CrossRef]
- Cox, D.R. Regression models and life-tables. J. R. Stat. Soc. Ser. B 1972, 34, 187–202. [Google Scholar] [CrossRef]
- Kaplan, E.L.; Meier, P. Nonparametric estimation from incomplete observations. J. Am. Stat. Assoc. 1958, 53, 457–481. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Society. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing, Vienna, Austria. Available online: http://www.R-project.org (accessed on 19 May 2013).
| # | Process Networks | In Data | Total | p-Value | FDR | Network Objects from Active Data |
|---|---|---|---|---|---|---|
| 1 | Cell adhesion_Integrin priming | 9 | 110 | 1.05 × 10−5 | 6.71 × 10−4 | ACTA2, G-protein alpha-i family, PIB4, SDF-1, SOS, Actin, Collagen III, PLC-beta, SOS1 |
| 2 | Cell adhesion_Leucocyte chemotaxis | 12 | 205 | 1.09 × 10−5 | 6.71 × 10−4 | G-protein alpha-i family, VCAM1, PIB4, Galpha(i)-specific EDG GPCRs, CCL2, CCL13, SDF-1, CXCL13, Actin, LPA3 receptor, PLC-beta, Galpha(q)-specific EDG GPCRs |
| 3 | Development_Regulation of angiogenesis | 11 | 223 | 1.25 × 10−4 | 4.66 × 10−3 | FAP48, G-protein alpha-i1, Angiopoietin 1, Ephrin-A receptors, G-protein alpha-i family, IL-6, CCL2, PGAR, N-cadherin, SOS, PLC-beta |
| 4 | Development_Blood vessel morphogenesis | 11 | 228 | 1.52 × 10−4 | 4.66 × 10−3 | G-protein alpha-i1, Angiopoietin 1, G-protein alpha-i family, VCAM1, ErbB4, Galpha(i)-specific EDG GPCRs, PGAR, SDF-1, PLGF, SOS, HGF receptor (Met) |
| 5 | Development_Ossification and bone remodeling | 8 | 157 | 8.88 × 10−4 | 2.18 × 10−2 | AEBP1, Frizzled, SFRP4, OSF-2, DMP1, MEPE, Osteomodulin, Bone sialoprotein |
| 6 | Cell adhesion_Cadherins | 8 | 180 | 2.13 × 10−3 | 3.53 × 10−2 | Frizzled, SFRP4, DKK1, N-cadherin, PTPR-zeta, WIF1, Actin, HGF receptor (Met) |
| 7 | Development_EMT_Regulation of epithelial-to-mesenchymal transition | 9 | 225 | 2.33 × 10−3 | 3.53 × 10−2 | HGF, ACTA2, Frizzled, G-protein alpha-i family, N-cadherin, SOS, Actin, HGF receptor (Met), Collagen III |
| 8 | Development_Skeletal muscle development | 7 | 144 | 2.46 × 10−3 | 3.53 × 10−2 | ACTA2, ER81, Actin muscle, ITGA11, ACTG2, Actin, HGF receptor (Met) |
| 9 | Inflammation_Protein C signaling | 6 | 108 | 2.59 × 10−3 | 3.53 × 10−2 | G-protein alpha-i family, PIB4, Galpha(i)-specific EDG GPCRs, IL-6, Actin, PLC-beta |
| 10 | Inflammation_Histamine signaling | 8 | 213 | 5.95 × 10−3 | 6.28 × 10−2 | Kappa chain (Ig light chain), G-protein alpha-i family, VCAM1, PIB4, IL-6, CCL2, Actin, PLC-beta |
| HuEx Discovery Set (n = 88) | |||||||||
| Probeset ID | Associated Gene(s) | Event-free Survival Model | Overall Survival Model | ||||||
| Hazard ratio † | p-value †,‡ | Corrected p-value * | Full-model p-value | Hazard ratio † | p-value †,‡ | Corrected p-value * | Full-model p-value | ||
| 3310041 | FGFR2 | 0.718 | 1.377 × 10−2 | 1.316 × 10−1 | 5.840 × 10−5 | 0.561 | 5.440 × 10−4 | 3.878 × 10−2 | 8.294 × 10−6 |
| 3324447 | FIBIN | 0.698 | 6.198 × 10−3 | 1.042 × 10−1 | 2.256 × 10−5 | 0.559 | 5.170 × 10−4 | 3.878 × 10−2 | 6.542 × 10−6 |
| 3074857 | PTN///DGKI | 0.729 | 3.402 × 10−3 | 8.746 × 10−2 | 1.018 × 10−5 | 0.618 | 6.211 × 10−4 | 3.878 × 10−2 | 3.849 × 10−6 |
| 3074857 | PTN///DGKI | 0.729 | 3.402 × 10−3 | 8.746 × 10−2 | 1.018 × 10−5 | 0.618 | 6.211 × 10−4 | 3.878 × 10−2 | 3.849 × 10−6 |
| U133 Validation Set (n = 60) | |||||||||
| Probeset ID | Associated Gene | Event-free Survival Model | Overall Survival Model | ||||||
| Hazard ratio † | p-value †,‡ | Full model p-value | Hazard ratio † | p-value †,‡ | Full model p-value | ||||
| 211399_at | FGFR2 | 0.026 | 1.825 × 10−3 | 2.749 × 10−5 | 0.042 | 5.347 × 10−3 | 4.658 × 10−6 | ||
| 231001_at | FIBIN | 0.602 | 1.129 × 10−2 | 1.216 × 10−4 | 0.595 | 2.149 × 10−2 | 1.262 × 10−5 | ||
| 208408_at | PTN | 8.914 | 4.325 × 10−2 | 3.864 × 10−4 | 24.039 | 1.163 × 10−2 | 1.985 × 10−5 | ||
| 206806_at | DGKI | 0.755 | 3.288 × 10−1 | 1.197 × 10−3 | 0.502 | 4.080 × 10−2 | 1.741 × 10−5 | ||
| Cytoband | Location(Mbs) | Width(Mbs) | Residual q Value | Frequency | High Frequency ‘ | Key Genes | |
|---|---|---|---|---|---|---|---|
| 17p11.2 | chr17:18.123–18.237 | 0.114 | 0 | 44.9 | 30.6 | TOP3A, FLI1 | * |
| 8q24.21 | chr8:128.357–128.772 | 0.415 | 0 | 46.9 | 27.2 | MYC | * |
| 20p13 | chr20:1.52–1.529 | 0.009 | 0 | 49 | 20.4 | ||
| 15q26.3 | chr15:99.366–99.408 | 0.043 | 0 | 42.2 | 20.4 | IGF1R | * |
| 1q21.3 | chr1:149.996–151.21 | 1.214 | 0.001 | 48.3 | 19 | * | |
| 13q34 | chr13:105.817–114.882 | 9.065 | 0.116 | 44.2 | 19 | ||
| 19p13.2 | chr19:12.686–13.498 | 0.812 | 0 | 43.5 | 18.4 | * | |
| 6p21.1 | chr6:43.323–44.511 | 1.187 | 0 | 40.1 | 17.7 | * | |
| 19q12 | chr19:30.082–30.306 | 0.224 | 0 | 40.1 | 17 | CCNE1 | * |
| 8p11.1 | chr8:41.441–50.441 | 9 | 0.033 | 50.3 | 16.3 | ||
| 17p13.1 | chr17:7.305–7.329 | 0.024 | 0.001 | 58.5 | |||
| 19q12 | chr19:28.283–30.098 | 1.814 | 0 | 58.5 | |||
| 13q14.2 | chr13:48.834–49.065 | 0.231 | 0 | 56.5 | RB1 | * | |
| 17p13.1 | chr17:10.372–10.532 | 0.16 | 0 | 56.5 | |||
| 3q13.31 | chr3:116.162–118.625 | 2.463 | 0 | 55.1 | LSAMP1, LSAMP-AS1 | * | |
| 8q24.3 | chr8:146.066–146.28 | 0.214 | 0 | 55.1 | |||
| 17p13.1 | chr17:7.611–7.763 | 0.152 | 0 | 55.1 | |||
| 4q35.2 | chr4:190.883–191.154 | 0.271 | 0 | 53.7 | |||
| 16q24.3 | chr16:89.995–90.355 | 0.36 | 0 | 53.1 | |||
| 20q13.33 | chr20:62.735–62.89 | 0.155 | 0 | 53.1 |
| Cytoband | Change | Genes | Full Model | Including Initial Metastasis | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| OS | EFS | OS | EFS | |||||||
| HR | p-Value | HR | p-Value | HR | p-Value | HR | p-Value | |||
| 15q26.3 | Amp | IGF1R | 1.110 | 9.00 × 10−3 | 1.108 | 3.40 × 10−2 | 1.116 | 1.30 × 10−2 | 1.093 | 2.25 × 10−1 |
| 8q24.21 | Amp | MYC, POU5F1B, LOC727677 | 1.170 | 1.10 × 10−2 | 1.211 | 4.00 × 10−3 | ||||
| All Samples | HuEx Samples | U133 Samples | Copy Number Only Samples | ||||||
| # | % | # | % | # | % | # | % | ||
| Total | 214 | 100 | 103 | 100 | 64 | 100 | 47 | 100 | |
| Gender | Male | 122 | 57 | 55 | 53 | 36 | 56 | 31 | 66 |
| Female | 92 | 43 | 48 | 47 | 28 | 44 | 16 | 34 | |
| Age at Diagnosis | <12 | 151 | 71 | 70 | 68 | 46 | 72 | 35 | 74 |
| >12 | 63 | 29 | 33 | 32 | 18 | 28 | 12 | 26 | |
| Location | Leg/Foot | 183 | 86 | 90 | 87 | 58 | 91 | 35 | 74 |
| Arm/Hand | 17 | 8 | 10 | 10 | 2 | 3 | 5 | 11 | |
| Other | 11 | 5 | 1 | 1 | 4 | 6 | 6 | 13 | |
| No Data | 3 | 1 | 2 | 2 | 0 | 0 | 1 | 2 | |
| SNP Data | Yes | 147 | 69 | 93 | 90 | 7 | 11 | 47 | 100 |
| No | 67 | 31 | 10 | 10 | 57 | 89 | 0 | 0 | |
| Event | Occurred | 87 | 41 | 38 | 37 | 31 | 48 | 18 | 38 |
| Censored | 100 | 47 | 50 | 49 | 29 | 45 | 21 | 45 | |
| No Data | 27 | 13 | 15 | 15 | 4 | 6 | 8 | 17 | |
| Death | Occurred | 68 | 32 | 27 | 26 | 28 | 44 | 13 | 28 |
| Censored | 119 | 56 | 61 | 59 | 32 | 50 | 26 | 55 | |
| No Data | 27 | 13 | 15 | 15 | 4 | 6 | 8 | 17 | |
| Metastasis at Diagnosis | No | 170 | 79 | 81 | 79 | 54 | 84 | 35 | 74 |
| Yes | 44 | 21 | 22 | 21 | 10 | 16 | 12 | 26 | |
| Mean | SD | Mean | SD | Mean | SD | Mean | SD | ||
| Age at Diagnosis | Years | 13.89 | 3.78 | 13.36 | 3.6 | 14.4 | 3.65 | 14.37 | 4.25 |
| Follow-up of Survivors | Years | 6.47 | 2.77 | 6.83 | 2.78 | 6.26 | 2.9 | 5.87 | 2.53 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Taylor, A.M.; Sun, J.M.; Yu, A.; Voicu, H.; Shen, J.; Barkauskas, D.A.; Triche, T.J.; Gastier-Foster, J.M.; Man, T.-K.; Lau, C.C. Integrated DNA Copy Number and Expression Profiling Identifies IGF1R as a Prognostic Biomarker in Pediatric Osteosarcoma. Int. J. Mol. Sci. 2022, 23, 8036. https://doi.org/10.3390/ijms23148036
Taylor AM, Sun JM, Yu A, Voicu H, Shen J, Barkauskas DA, Triche TJ, Gastier-Foster JM, Man T-K, Lau CC. Integrated DNA Copy Number and Expression Profiling Identifies IGF1R as a Prognostic Biomarker in Pediatric Osteosarcoma. International Journal of Molecular Sciences. 2022; 23(14):8036. https://doi.org/10.3390/ijms23148036
Chicago/Turabian StyleTaylor, Aaron M., Jiayi M. Sun, Alexander Yu, Horatiu Voicu, Jianhe Shen, Donald A. Barkauskas, Timothy J. Triche, Julie M. Gastier-Foster, Tsz-Kwong Man, and Ching C. Lau. 2022. "Integrated DNA Copy Number and Expression Profiling Identifies IGF1R as a Prognostic Biomarker in Pediatric Osteosarcoma" International Journal of Molecular Sciences 23, no. 14: 8036. https://doi.org/10.3390/ijms23148036
APA StyleTaylor, A. M., Sun, J. M., Yu, A., Voicu, H., Shen, J., Barkauskas, D. A., Triche, T. J., Gastier-Foster, J. M., Man, T.-K., & Lau, C. C. (2022). Integrated DNA Copy Number and Expression Profiling Identifies IGF1R as a Prognostic Biomarker in Pediatric Osteosarcoma. International Journal of Molecular Sciences, 23(14), 8036. https://doi.org/10.3390/ijms23148036






