A Structural Comparison of SARS-CoV-2 Main Protease and Animal Coronaviral Main Protease Reveals Species-Specific Ligand Binding and Dimerization Mechanism
Abstract
:1. Introduction
2. Results
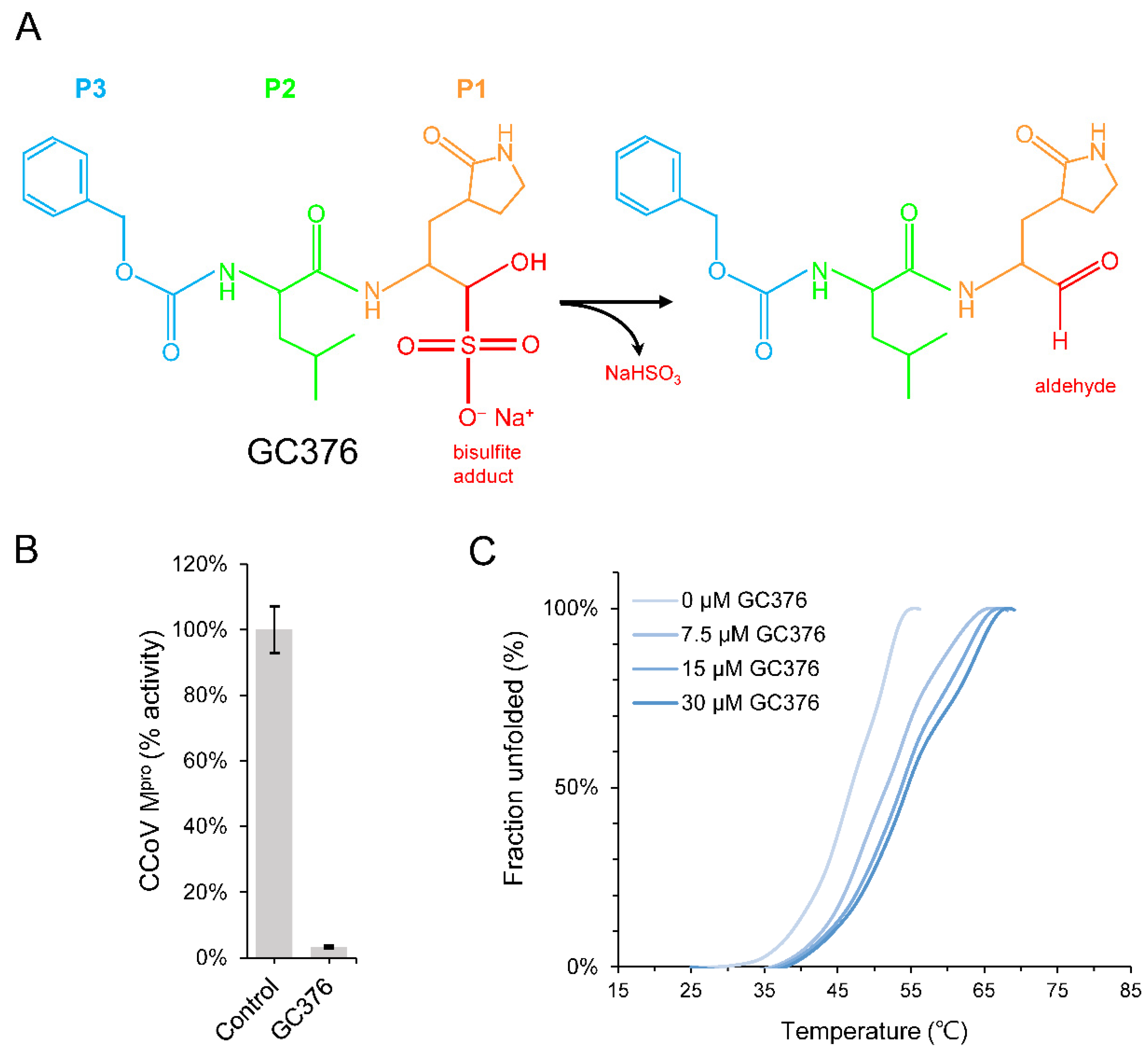
2.1. GC376 Is a Potential Lead Compound against CCoV Mpro
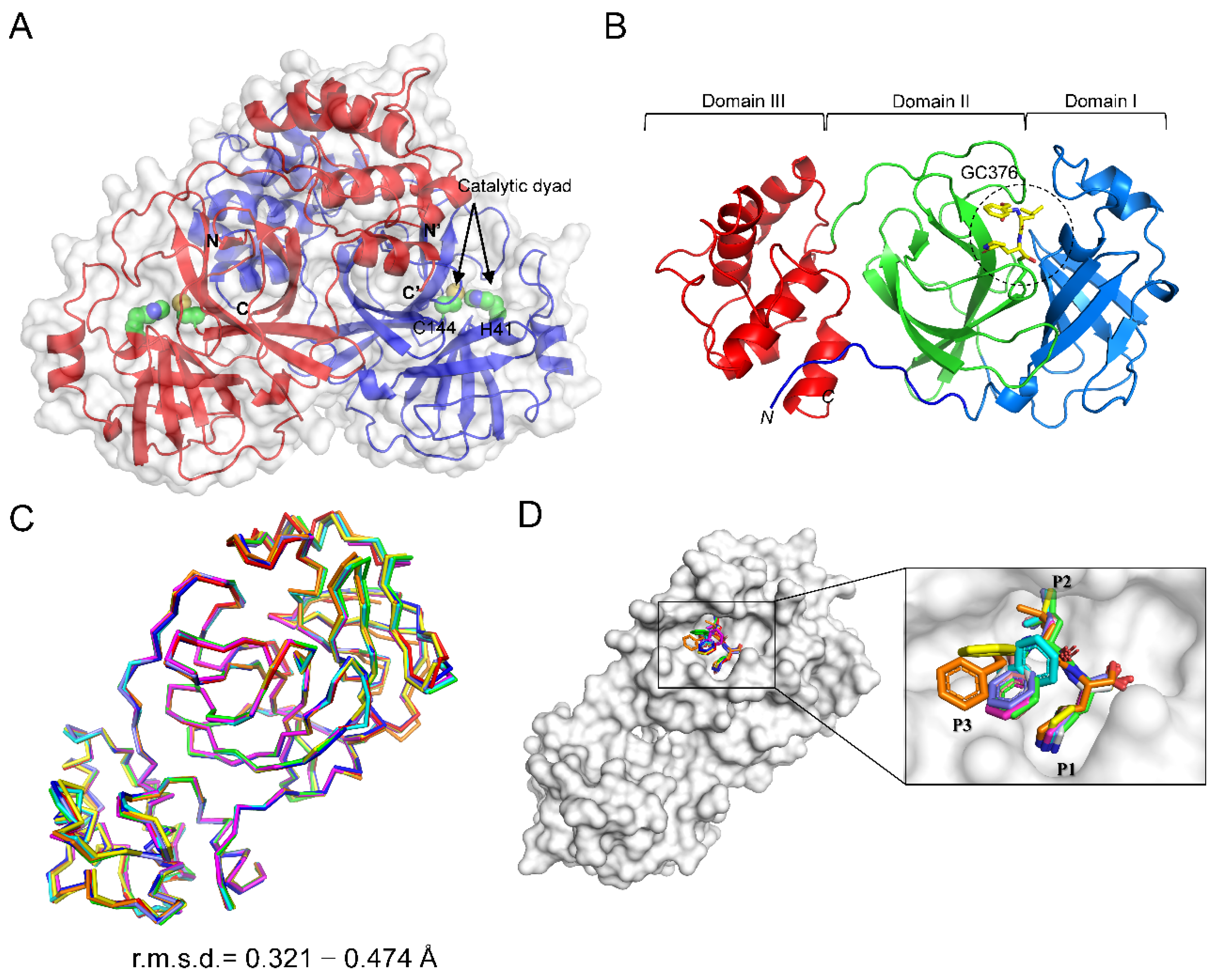
2.2. Overall Structure of CCoV Mpro in Complex with GC376
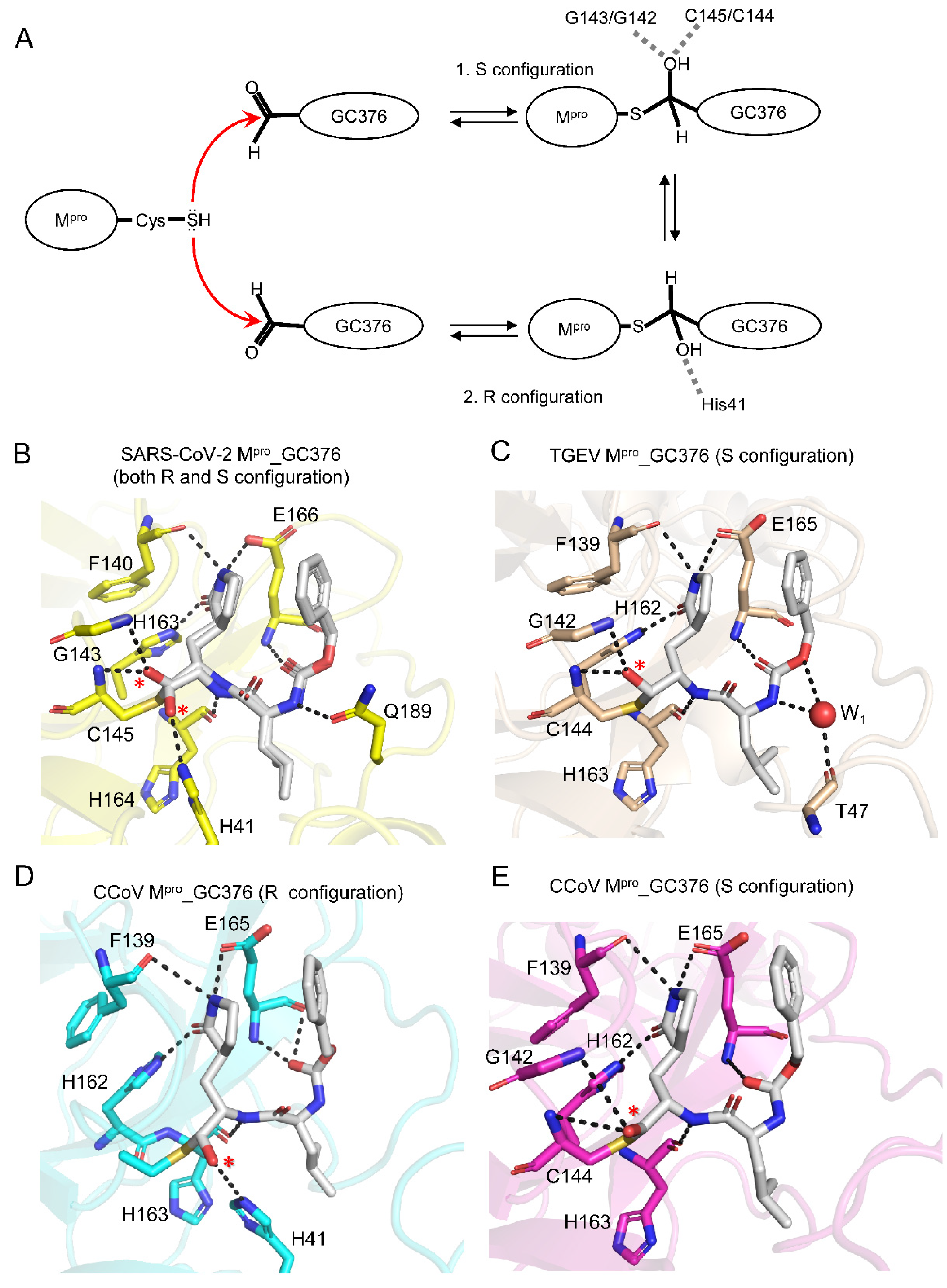
2.3. GC376 Covalently Linked to Catalytic Cys144 of CCoV Mpro, Forming an (R)- or (S)-Hemithioacetal
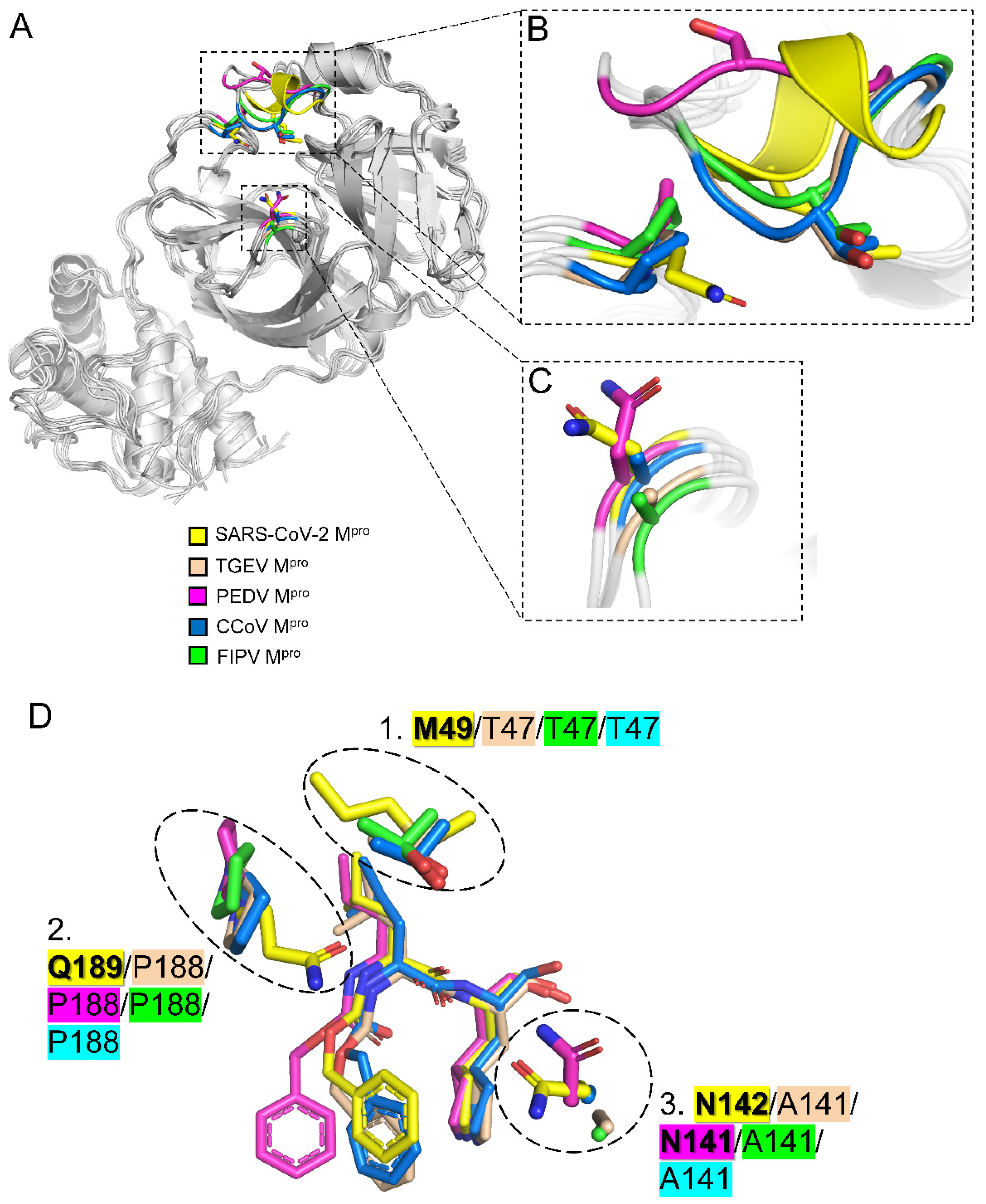
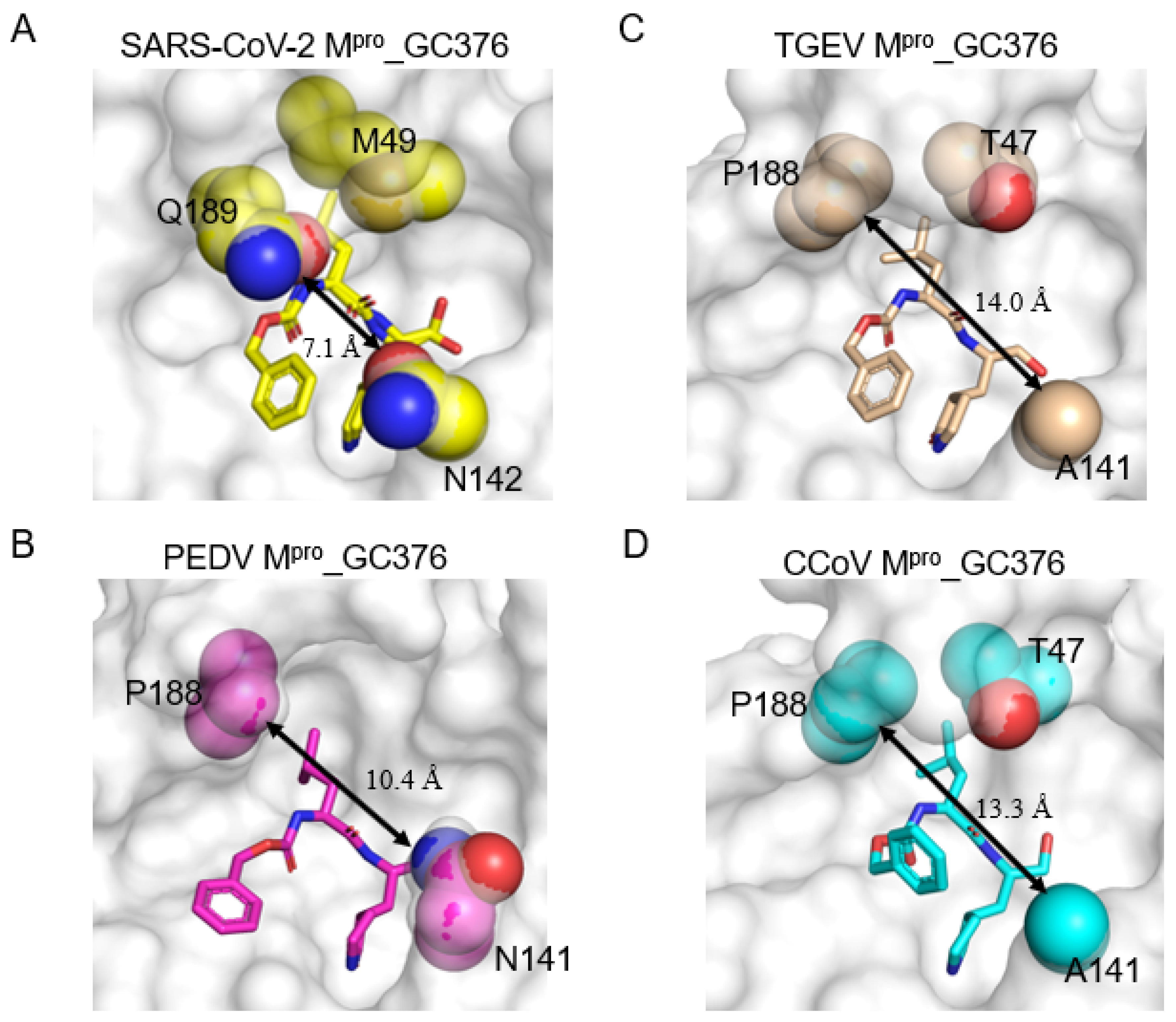
2.4. Three Conserved Structural Features Dictate the Substrate-Binding Pocket of CoV Mpros
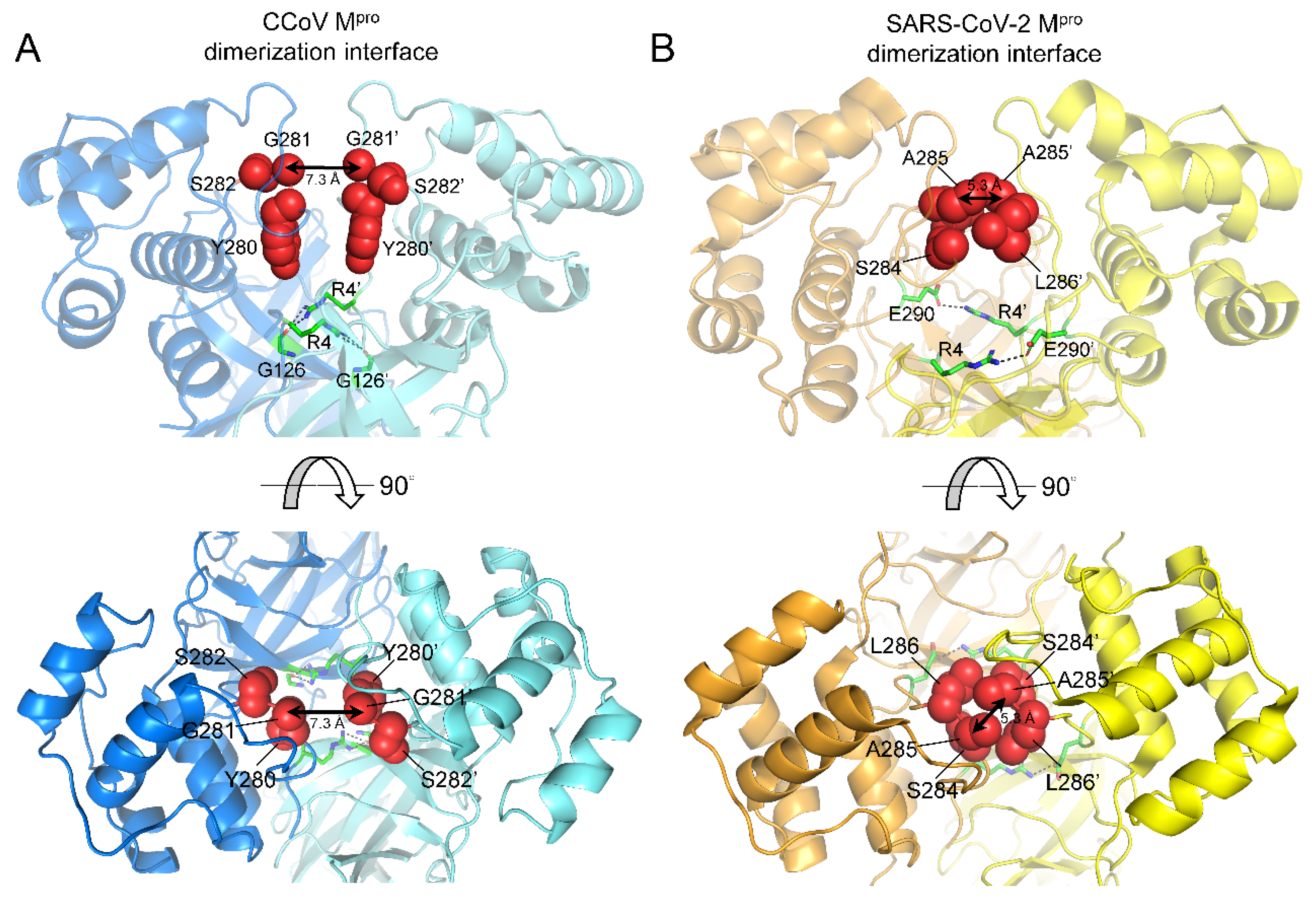
2.5. Species-Specific Dimerization of CoV Mpros
3. Discussion
4. Materials and Methods
4.1. Cloning, Expression, and Purification of CCoV Mpro
4.2. Fluorescence Resonance Energy Transfer (FRET)-Based Assay
4.3. Protein Thermal Shift Assay
4.4. Crystallization and Structural Determination of CCoV Mpro in Complex with GC376
4.5. Bioinformatic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wong, A.C.P.; Lau, S.K.P.; Woo, P.C.Y. Interspecies Jumping of Bat Coronaviruses. Viruses 2021, 13, 2188. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021, 19, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.-F.; Eaton, B.T. Bats, Civets and the Emergence of SARS. Curr. Top. Microbiol. Immunol. 2007, 315, 325–344. [Google Scholar] [PubMed]
- Sharif-Yakan, A.; Kanj, S.S. Emergence of MERS-CoV in the Middle East: Origins, transmission, treatment, and perspectives. PLoS Pathog. 2014, 10, e1004457. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Latif, A.A.; Mukaratirwa, S. Zoonotic origins and animal hosts of coronaviruses causing human disease pandemics: A review. Onderstepoort J. Vet. Res. 2020, 87, e1–e9. [Google Scholar] [CrossRef]
- Murphy, F.A.; Fauquet, C.M.; Bishop, D.H.; Ghabrial, S.A.; Jarvis, A.W.; Martelli, G.P.; Mayo, M.A.; Summers, M.D. (Eds.) Virus Taxonomy: Classification and Nomenclature of Viruses; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2012; Volume 10. [Google Scholar]
- Sun, R.-Q.; Cai, R.-J.; Chen, Y.-Q.; Liang, P.-S.; Chen, D.-K.; Song, C.-X. Outbreak of Porcine Epidemic Diarrhea in Suckling Piglets, China. Emerg. Infect. Dis. 2012, 18, 161–163. [Google Scholar] [CrossRef]
- Decaro, N.; Buonavoglia, C. An update on canine coronaviruses: Viral evolution and pathobiology. Vet. Microbiol. 2008, 132, 221–234. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Li, K.S.M.; Tsang, A.K.L.; Lam, C.S.F.; Ahmed, S.; Chen, H.; Chan, K.-H.; Woo, P.C.Y.; Yuen, K.-Y. Genetic Characterization of Betacoronavirus Lineage C Viruses in Bats Reveals Marked Sequence Divergence in the Spike Protein of Pipistrellus Bat Coronavirus HKU5 in Japanese Pipistrelle: Implications for the Origin of the Novel Middle East Respiratory Syndrome Coronavirus. J. Virol. 2013, 87, 8638–8650. [Google Scholar]
- Malik, Y.A. Properties of Coronavirus and SARS-CoV-2. Malays. J. Pathol. 2020, 42, 3–11. [Google Scholar]
- De Wit, J.J.S.; Cook, J.K.A. Spotlight on avian coronaviruses. Avian Pathol. 2020, 49, 313–316. [Google Scholar] [CrossRef]
- Kim, Y.; Liu, H.; Galasiti Kankanamalage, A.C.; Weerasekara, S.; Hua, D.H.; Groutas, W.C.; Chang, K.O.; Pedersen, N.C. Reversal of the Progression of Fatal Coronavirus Infection in Cats by a Broad-Spectrum Coronavirus Protease Inhibitor. PLoS Pathog. 2016, 12, e1005531. [Google Scholar]
- Pedersen, N.C.; Kim, Y.; Liu, H.; Galasiti Kankanamalage, A.C.; Eckstrand, C.; Groutas, W.C.; Bannasch, M.; Meadows, J.M.; Chang, K.O. Efficacy of a 3C-like protease inhibitor in treating various forms of acquired feline infectious peritonitis. J. Feline Med. Surg. 2018, 20, 378–392. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Lovell, S.; Tiew, K.-C.; Mandadapu, S.R.; Alliston, K.R.; Battaile, K.P.; Groutas, W.C.; Chang, K.-O. Broad-Spectrum Antivirals against 3C or 3C-Like Proteases of Picornaviruses, Noroviruses, and Coronaviruses. J. Virol. 2012, 86, 11754–11762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ye, G.; Wang, X.; Tong, X.; Shi, Y.; Fu, Z.F.; Peng, G. Structural Basis for Inhibiting Porcine Epidemic Diarrhea Virus Replication with the 3C-Like Protease Inhibitor GC376. Viruses 2020, 12, 240. [Google Scholar] [CrossRef] [Green Version]
- Vuong, W.; Khan, M.B.; Fischer, C.; Arutyunova, E.; Lamer, T.; Shields, J.; Saffran, H.A.; McKay, R.T.; van Belkum, M.J.; Joyce, M.A.; et al. Feline coronavirus drug inhibits the main protease of SARS-CoV-2 and blocks virus replication. Nat. Commun. 2020, 11, 4282. [Google Scholar] [CrossRef]
- Kankanamalage, A.C.G.; Kim, Y.; Damalanka, V.C.; Rathnayake, A.D.; Fehr, A.; Mehzabeen, N.; Battaile, K.; Lovell, S.; Lushington, G.; Perlman, S.; et al. Structure-guided design of potent and permeable inhibitors of MERS coronavirus 3CL protease that utilize a piperidine moiety as a novel design element. Eur. J. Med. Chem. 2018, 150, 334–346. [Google Scholar] [CrossRef]
- Ma, C.; Sacco, M.D.; Hurst, B.; Townsend, J.A.; Hu, Y.; Szeto, T.; Zhang, X.; Tarbet, B.; Marty, M.T.; Chen, Y.; et al. Boceprevir, GC-376, and calpain inhibitors II, XII inhibit SARS-CoV-2 viral replication by targeting the viral main protease. Cell Res. 2020, 30, 678–692. [Google Scholar] [CrossRef]
- Wang, Y.C.; Yang, W.H.; Yang, C.S.; Hou, M.H.; Tsai, C.L.; Chou, Y.Z.; Hung, M.C.; Chen, Y. Structural basis of SARS-CoV-2 main protease inhibition by a broad-spectrum anti-coronaviral drug. Am. J. Cancer Res. 2020, 10, 2535–2545. [Google Scholar]
- Decaro, N.; Balboni, A.; Bertolotti, L.; Martino, P.A.; Mazzei, M.; Mira, F.; Pagnini, U. SARS-CoV-2 Infection in Dogs and Cats: Facts and Speculations. Front. Vet. Sci. 2021, 8, 619207. [Google Scholar] [CrossRef]
- Sit, T.H.C.; Brackman, C.J.; Ip, S.M.; Tam, K.W.S.; Law, P.Y.T.; To, E.M.W.; Yu, V.Y.T.; Sims, L.D.; Tsang, D.N.C.; Chu, D.K.W.; et al. Infection of dogs with SARS-CoV-2. Nature 2020, 586, 776–778. [Google Scholar] [CrossRef]
- Sanchez-Morales, L.; Sanchez-Vizcaino, J.M.; Perez-Sancho, M.; Domínguez, L.; Barroso-Arévalo, S. The Omicron (B.1.1.529) SARS-CoV-2 variant of concern also affects companion animals. bioRxiv 2022. [Google Scholar] [CrossRef]
- Vlasova, A.N.; Diaz, A.; Damtie, D.; Xiu, L.; Toh, T.-H.; Lee, J.S.-Y.; Saif, L.J.; Gray, G.C. Novel Canine Coronavirus Isolated from a Hospitalized Patient with Pneumonia in East Malaysia. Clin. Infect. Dis. 2022, 74, 446–454. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Wei, Z.; Song, J. Dissection study on the severe acute respiratory syndrome 3C-like protease reveals the critical role of the extra domain in dimerization of the enzyme: Defining the extra domain as a new target for design of highly specific protease inhibitors. J. Biol. Chem. 2004, 279, 24765–24773. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Günther, S.; Reinke, P.Y.A.; Fernández-García, Y.; Lieske, J.; Lane, T.J.; Ginn, H.M.; Koua, F.; Ehrt, C.; Ewert, W.; Oberthuer, D.; et al. X-ray screening identifies active site and allosteric inhibitors of SARS-CoV-2 main protease. Science 2021, 372, 642–646. [Google Scholar] [CrossRef]
- Zhang, L.; Lin, D.; Sun, X.; Curth, U.; Drosten, C.; Sauerhering, L.; Becker, S.; Rox, K.; Hilgenfeld, R. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 2020, 368, 409–412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, L.; Shi, J.; Mu, Y.; Song, J. Dynamically-driven enhancement of the catalytic machinery of the SARS 3C-like protease by the S284-T285-I286/A mutations on the extra domain. PLoS ONE 2014, 9, e101941. [Google Scholar] [CrossRef]
- Yang, H.; Xie, W.; Xue, X.; Yang, K.; Ma, J.; Liang, W.; Zhao, Q.; Zhou, Z.; Pei, D.; Ziebuhr, J.; et al. Design of wide-spectrum inhibitors targeting coronavirus main proteases. PLoS Biol. 2005, 3, e324. [Google Scholar]
- Kneller, D.W.; Phillips, G.; O’Neill, H.M.; Jedrzejczak, R.; Stols, L.; Langan, P.; Joachimiak, A.; Coates, L.; Kovalevsky, A. Structural plasticity of SARS-CoV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography. Nat. Commun. 2020, 11, 3202. [Google Scholar] [CrossRef]
- Bzówka, M.; Mitusińska, K.; Raczyńska, A.; Samol, A.; Tuszyński, J.A.; Góra, A. Structural and Evolutionary Analysis Indicate That the SARS-CoV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design. Int. J. Mol. Sci. 2020, 21, 3099. [Google Scholar] [CrossRef]
- Kneller, D.W.; Galanie, S.; Phillips, G.; O’Neill, H.M.; Coates, L.; Kovalevsky, A. Malleability of the SARS-CoV-2 3CL M(pro) Active-Site Cavity Facilitates Binding of Clinical Antivirals. Structure 2020, 28, 1313–1320.e3. [Google Scholar] [CrossRef]
- Ferreira, G.M.; Kronenberger, T.; Tonduru, A.K.; Hirata, R.; Hirata, M.H.; Poso, A. SARS-COV-2 M(pro) conformational changes induced by covalently bound ligands. J. Biomol. Struct. Dyn. 2021, 39, 1–11. [Google Scholar]
- Boivin, S.; Kozak, S.; Meijers, R. Optimization of protein purification and characterization using Thermofluor screens. Protein Expr. Purif. 2013, 91, 192–206. [Google Scholar] [CrossRef] [PubMed]
- Otwinowski, Z.; Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997, 276, 307–326. [Google Scholar] [PubMed]
- Vagin, A.; Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 22–25. [Google Scholar] [CrossRef]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.W.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 2011, 67, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Kantardjieff, K.A.; Rupp, B. Matthews coefficient probabilities: Improved estimates for unit cell contents of proteins, DNA, and protein-nucleic acid complex crystals. Protein Sci. 2003, 12, 1865–1871. [Google Scholar] [CrossRef]
- Vagin, A.A.; Steiner, R.A.; Lebedev, A.A.; Potterton, L.; McNicholas, S.; Long, F.; Murshudov, G.N. REFMAC5 dictionary: Organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, 60, 2184–2195. [Google Scholar] [CrossRef] [Green Version]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef] [Green Version]
- Liebschner, D.; Afonine, P.V.; Baker, M.L.; Bunkóczi, G.; Chen, V.B.; Croll, T.; Hintze, B.; Hung, L.W.; Jain, S.; McCoy, A.; et al. Macromolecular structure determination using X-rays, neutrons and electrons: Recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 2019, 75, 861–877. [Google Scholar] [CrossRef] [Green Version]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.-T.; Von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| GC376 Bound CCoV Mpro | |
|---|---|
| PDB Code | 7XJW |
| Data collection | |
| Diffraction source | TPS 05A, 3 GeV TPS, NSRRC |
| Wavelength (Å) | 0.99984 |
| Detector | MX300-HS |
| Crystal-detector distance (mm) | 300 |
| Space group | C2 |
| a, b, c (Å) | 156.975, 125.749, 160.418 |
| α, β, γ (°) | 90, 97.467, 90 |
| Resolution range (Å) | 30.0–2.75 (2.85–2.75) |
| Total no. of reflections | 295,387 (28,408) |
| No. of unique reflections | 79,392 (7891) |
| Completeness (%) | 99.3 (99.7) |
| Multiplicity | 3.7 (3.6) |
| 〈I/σ(I)〉 | 20.39 (3.39) |
| Rmerge | 0.062 (0.386) |
| Rp.i.m. | 0.036 (0.234) |
| CC1/2 | (0.914) |
| Refinement | |
| Resolution range (Å) | 27.83–2.75 (2.82–2.75) |
| Final Rwork (%) | 21.0 (27.2) |
| Final Rfree (%) | 25.7 (28.0) |
| No. of non-H atoms | 18,564 |
| No. of atoms | |
| Protein | 18,224 |
| Ligand | 232 |
| Water | 108 |
| B factors (Å2) | 54.4 |
| Protein | 54.6 |
| Ligand | 48.6 |
| Water | 31.9 |
| R.m.s. deviations | |
| Bonds (Å) | 0.015 |
| Angles (°) | 1.72 |
| Ramachandran plot | |
| Most favoured (%) | 92.44 |
| Allowed (%) | 7.22 |
| Outliers (%) | 0.34 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ho, C.-Y.; Yu, J.-X.; Wang, Y.-C.; Lin, Y.-C.; Chiu, Y.-F.; Gao, J.-Y.; Lai, S.-J.; Chen, M.-J.; Huang, W.-C.; Tien, N.; et al. A Structural Comparison of SARS-CoV-2 Main Protease and Animal Coronaviral Main Protease Reveals Species-Specific Ligand Binding and Dimerization Mechanism. Int. J. Mol. Sci. 2022, 23, 5669. https://doi.org/10.3390/ijms23105669
Ho C-Y, Yu J-X, Wang Y-C, Lin Y-C, Chiu Y-F, Gao J-Y, Lai S-J, Chen M-J, Huang W-C, Tien N, et al. A Structural Comparison of SARS-CoV-2 Main Protease and Animal Coronaviral Main Protease Reveals Species-Specific Ligand Binding and Dimerization Mechanism. International Journal of Molecular Sciences. 2022; 23(10):5669. https://doi.org/10.3390/ijms23105669
Chicago/Turabian StyleHo, Chien-Yi, Jia-Xin Yu, Yu-Chuan Wang, Yu-Chuan Lin, Yi-Fang Chiu, Jing-Yan Gao, Shu-Jung Lai, Ming-Jen Chen, Wei-Chien Huang, Ni Tien, and et al. 2022. "A Structural Comparison of SARS-CoV-2 Main Protease and Animal Coronaviral Main Protease Reveals Species-Specific Ligand Binding and Dimerization Mechanism" International Journal of Molecular Sciences 23, no. 10: 5669. https://doi.org/10.3390/ijms23105669
APA StyleHo, C.-Y., Yu, J.-X., Wang, Y.-C., Lin, Y.-C., Chiu, Y.-F., Gao, J.-Y., Lai, S.-J., Chen, M.-J., Huang, W.-C., Tien, N., & Chen, Y. (2022). A Structural Comparison of SARS-CoV-2 Main Protease and Animal Coronaviral Main Protease Reveals Species-Specific Ligand Binding and Dimerization Mechanism. International Journal of Molecular Sciences, 23(10), 5669. https://doi.org/10.3390/ijms23105669







