Small-Molecule Inhibitors of the RNA M6A Demethylases FTO Potently Support the Survival of Dopamine Neurons
Abstract
1. Introduction
2. Results and Discussion
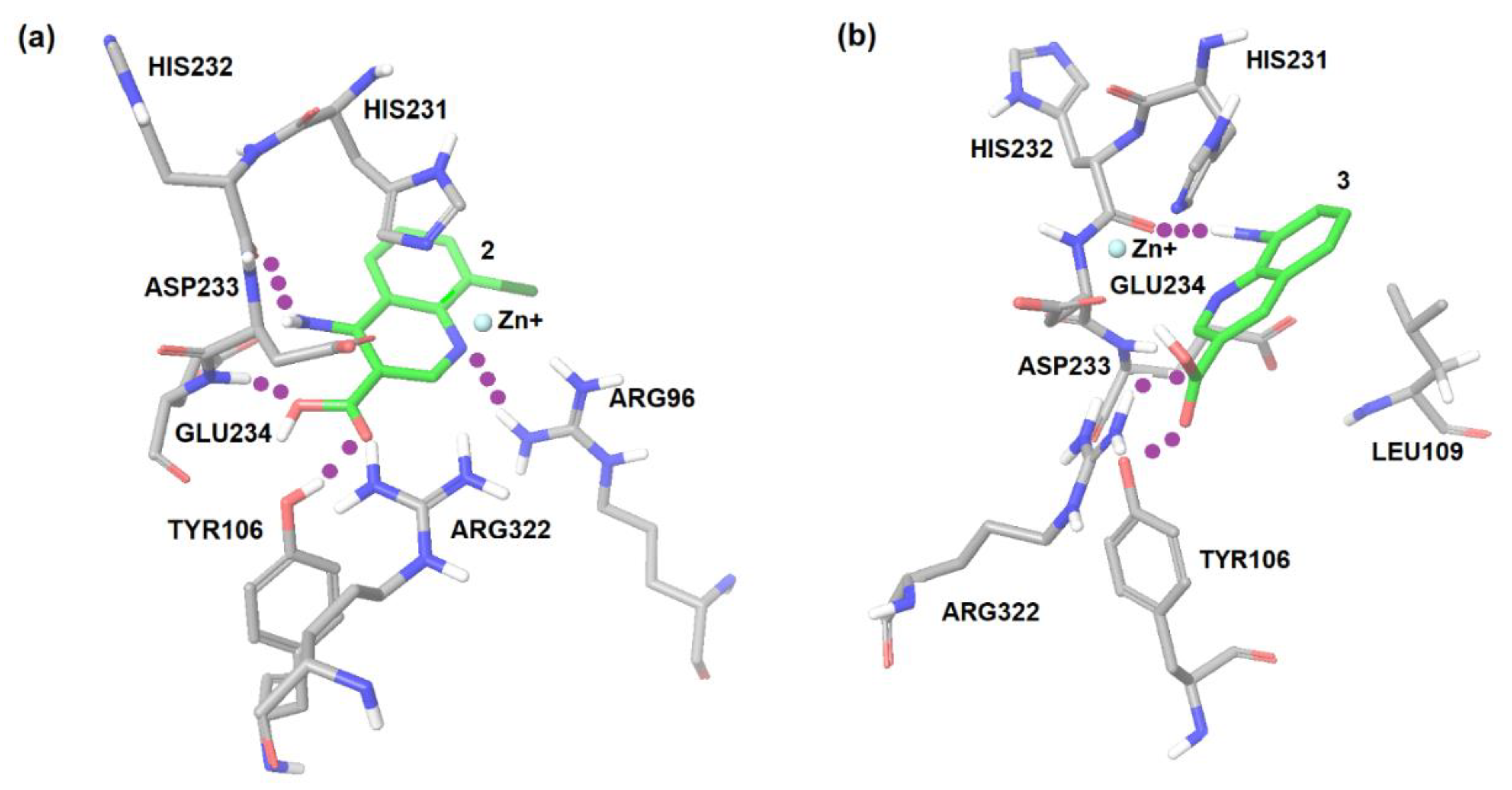
2.1. Computational Modeling of FTO Ligand Binding Site and Virtual Screening
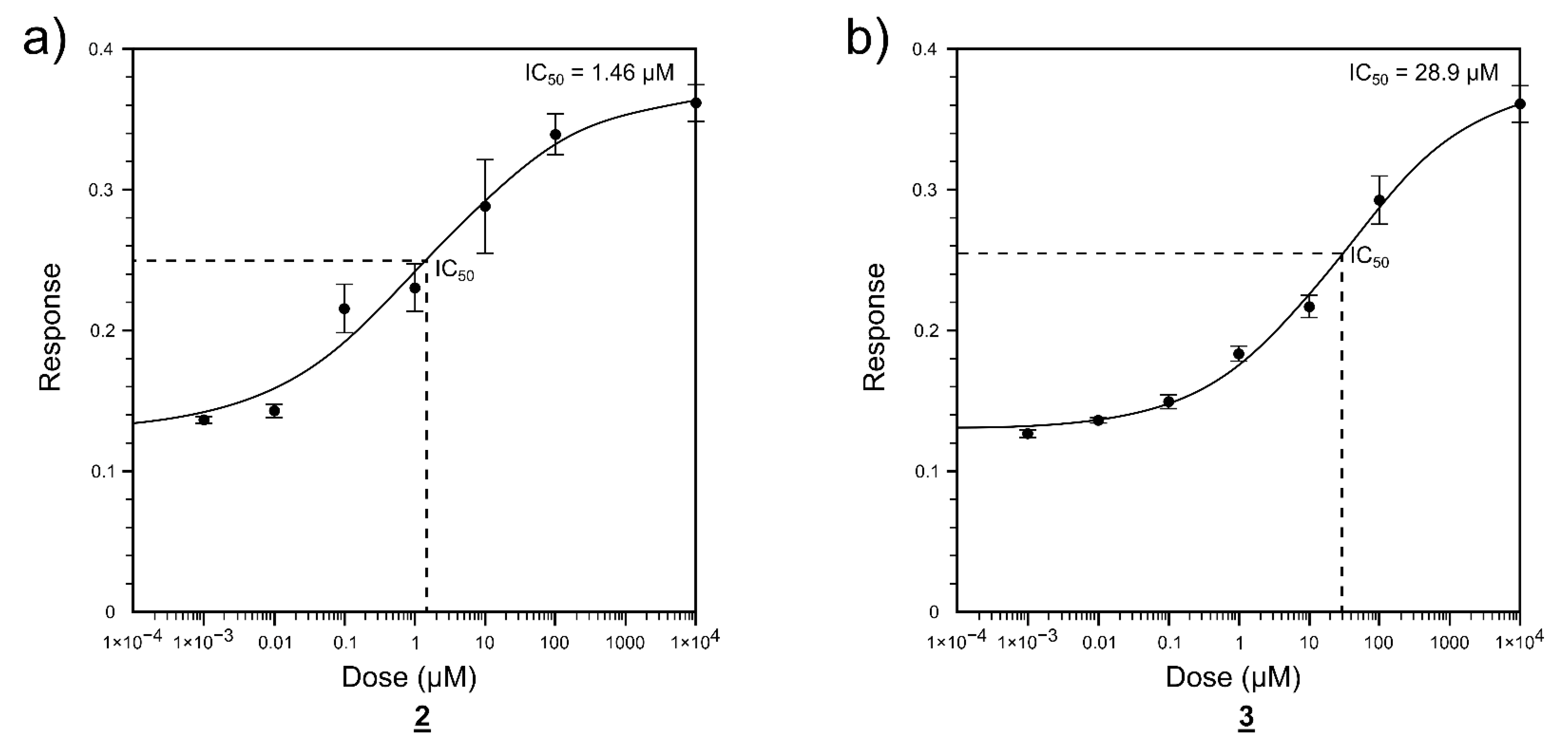
2.2. Enzyme Activity Inhibition
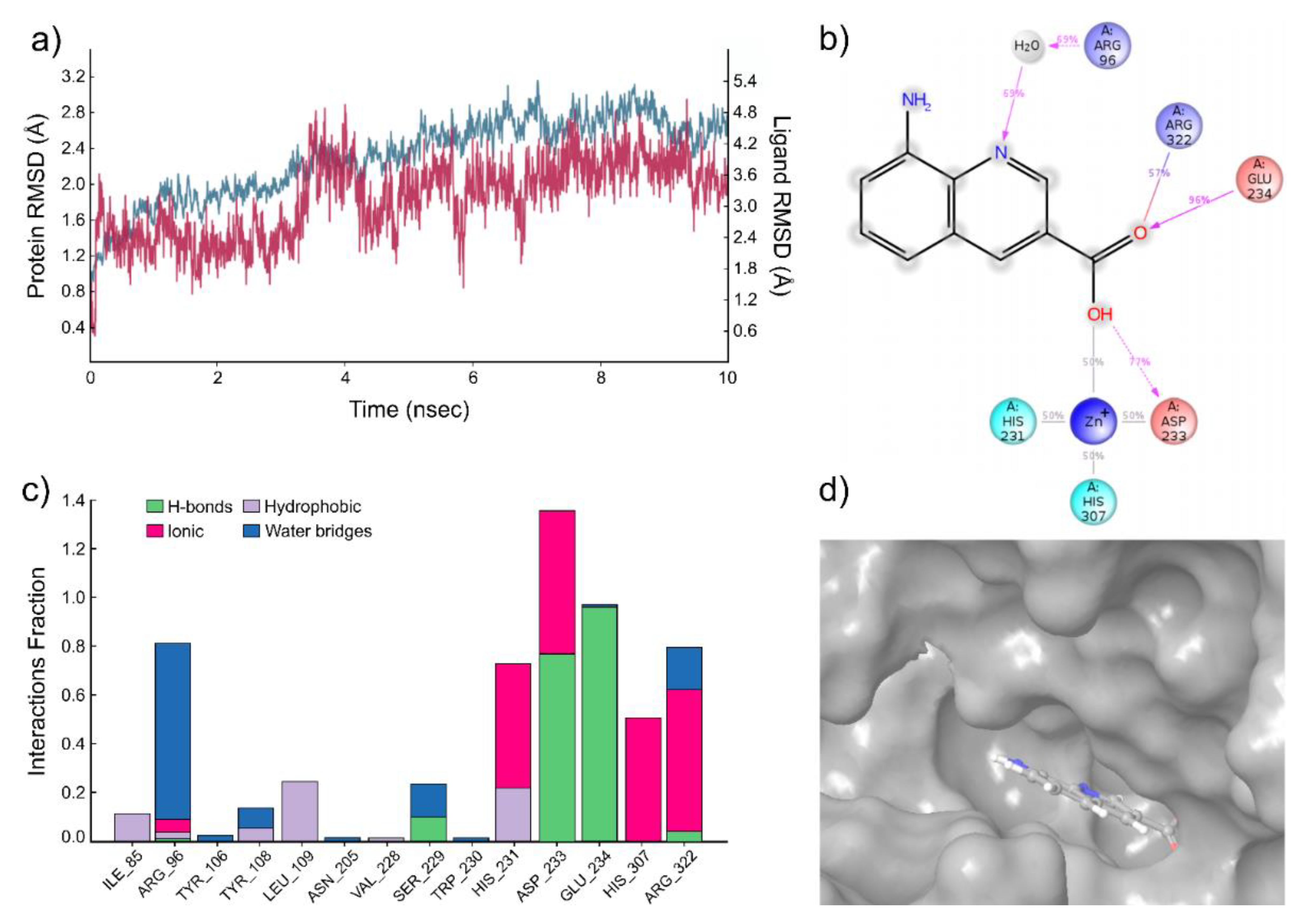
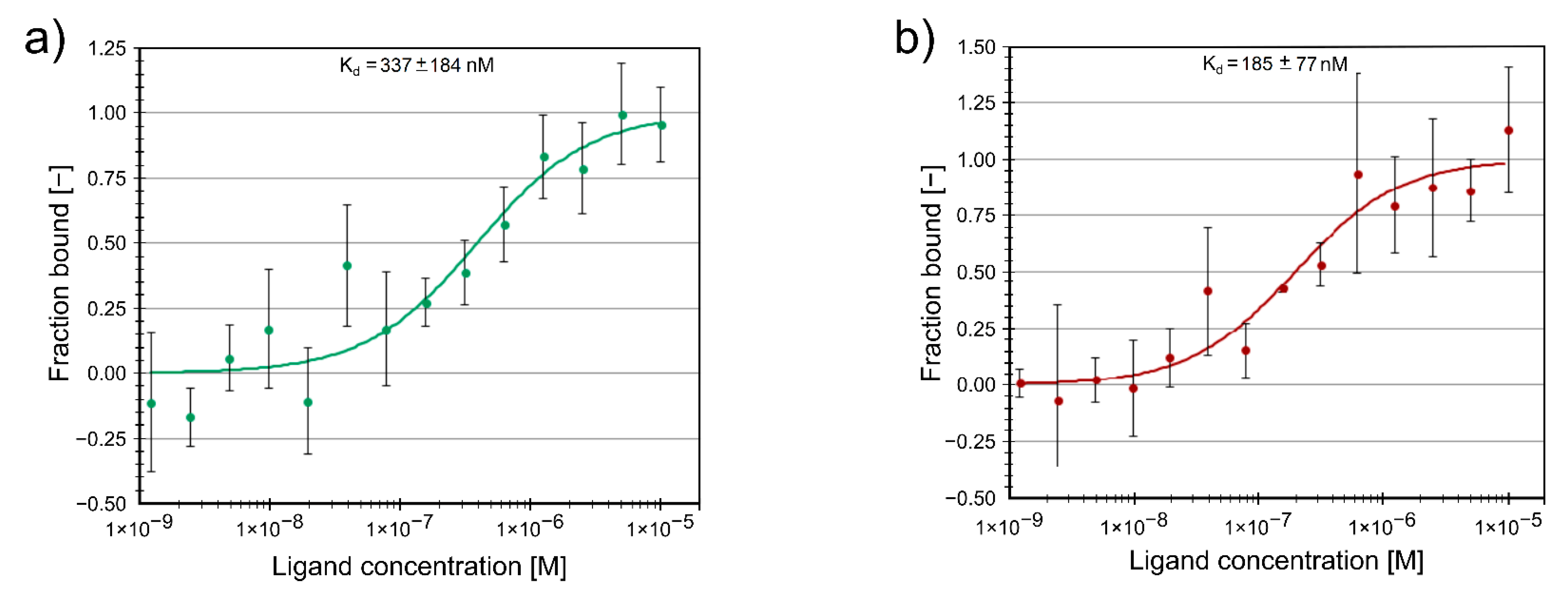
2.3. Protein Binding of Compounds
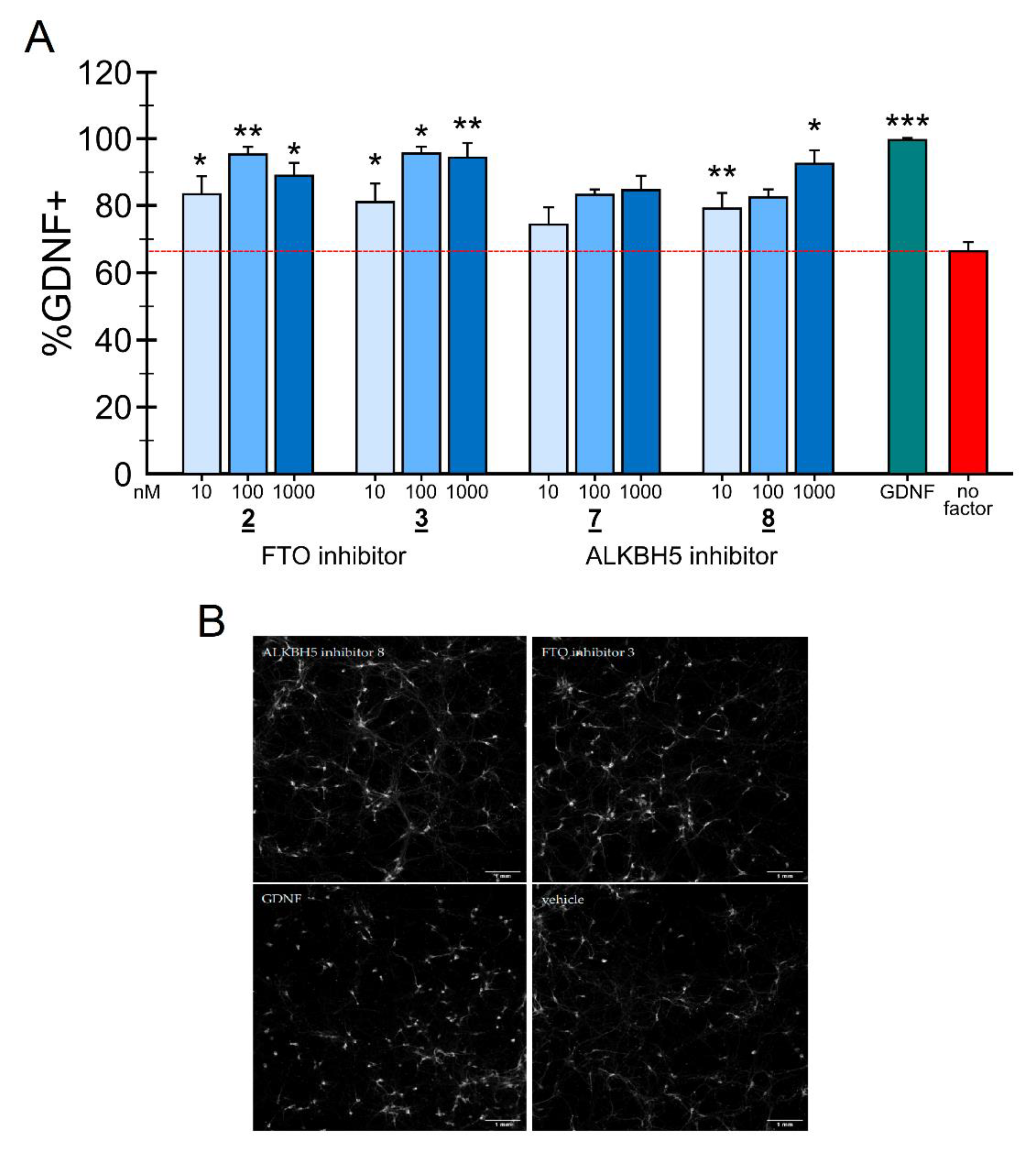
2.4. Neuronal Survival Experiments
2.5. Penetration of the FTO Inhibitors Through the Blood-Brain Barrier in Artificial In Vitro Model
3. Materials and Methods
3.1. Compounds
3.2. FTO Protein
3.3. Computational Modeling
3.4. Protein Binding Study Using Microscale Thermophoresis
3.5. Enzyme Inhibition Assay
3.6. Primary Cultures of Midbrain Dopamine Neurons and m6A Regulator Treatment
3.7. Artificial Blood–Brain Barrier Model
3.8. Quantification and Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Song, J.; Yi, C. Chemical Modifications to RNA: A New Layer of Gene Expression Regulation. ACS Chem. Biol. 2017, 12, 316–325. [Google Scholar] [CrossRef]
- Tzelepis, K.; Rausch, O.; Kouzarides, T. RNA-Modifying Enzymes and Their Function in a Chromatin Context. Nat. Struct. Mol. Biol. 2019, 26, 858–862. [Google Scholar] [CrossRef]
- Maity, A.; Das, B. N6-Methyladenosine Modification in MRNA: Machinery, Function and Implications for Health and Diseases. FEBS J. 2016, 283, 1607–1630. [Google Scholar] [CrossRef]
- Roundtree, I.A.; Evans, M.E.; Pan, T.; He, C. Dynamic RNA Modifications in Gene Expression Regulation. Cell 2017, 169, 1187–1200. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.A.; Kinzig, C.G.; DeGregorio, S.J.; Steitz, J.A. Methyltransferase-like Protein 16 Binds the 3′-Terminal Triple Helix of MALAT1 Long Noncoding RNA. Proc. Natl. Acad. Sci. USA 2016, 113, 14013–14018. [Google Scholar] [CrossRef]
- Liu, J.; Yue, Y.; Han, D.; Wang, X.; Fu, Y.; Zhang, L.; Jia, G.; Yu, M.; Lu, Z.; Deng, X.; et al. A METTL3–METTL14 Complex Mediates Mammalian Nuclear RNA N 6 -Adenosine Methylation. Nat. Chem. Biol. 2014, 10, 93–95. [Google Scholar] [CrossRef] [PubMed]
- Meyer, K.D.; Jaffrey, S.R. Rethinking M6A Readers, Writers, and Erasers. Annu. Rev. Cell Dev. Biol. 2017, 33, 319–342. [Google Scholar] [CrossRef]
- Jia, G.; Fu, Y.; Zhao, X.; Dai, Q.; Zheng, G.; Yang, Y.; Yi, C.; Lindahl, T.; Pan, T.; Yang, Y.-G.; et al. N 6-Methyladenosine in Nuclear RNA Is a Major Substrate of the Obesity-Associated FTO. Nat. Chem. Biol. 2011, 7, 885–887. [Google Scholar] [CrossRef]
- Zheng, G.; Dahl, J.A.; Niu, Y.; Fedorcsak, P.; Huang, C.-M.; Li, C.J.; Vågbø, C.B.; Shi, Y.; Wang, W.-L.; Song, S.-H.; et al. ALKBH5 Is a Mammalian RNA Demethylase That Impacts RNA Metabolism and Mouse Fertility. Mol. Cell 2013, 49, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, J.; Gu, Q.; Ma, Y.; Yang, Y.; Zhu, J.; Zhang, Q. The Biological Function of M6A Demethylase ALKBH5 and Its Role in Human Disease. Cancer Cell Int. 2020, 20, 347. [Google Scholar] [CrossRef]
- Patil, D.P.; Pickering, B.F.; Jaffrey, S.R. Reading M6A in the Transcriptome: M6A-Binding Proteins. Trends Cell Biol. 2018, 28, 113–127. [Google Scholar] [CrossRef]
- Wojtas, M.N.; Pandey, R.R.; Mendel, M.; Homolka, D.; Sachidanandam, R.; Pillai, R.S. Regulation of M6A Transcripts by the 3ʹ→5ʹ RNA Helicase YTHDC2 Is Essential for a Successful Meiotic Program in the Mammalian Germline. Mol. Cell 2017, 68, 374–387.e12. [Google Scholar] [CrossRef]
- Angelova, M.T.; Dimitrova, D.G.; Dinges, N.; Lence, T.; Worpenberg, L.; Carré, C.; Roignant, J.-Y. The Emerging Field of Epitranscriptomics in Neurodevelopmental and Neuronal Disorders. Front. Bioeng. Biotechnol. 2018, 6. [Google Scholar] [CrossRef] [PubMed]
- Jung, Y.; Goldman, D. Role of RNA Modifications in Brain and Behavior. Genes Brain Behav. 2018, 17, e12444. [Google Scholar] [CrossRef]
- Noack, F.; Calegari, F. Epitranscriptomics: A New Regulatory Mechanism of Brain Development and Function. Front. Neurosci. 2018, 12. [Google Scholar] [CrossRef] [PubMed]
- Widagdo, J.; Anggono, V. The M6A-Epitranscriptomic Signature in Neurobiology: From Neurodevelopment to Brain Plasticity. J. Neurochem. 2018, 147, 137–152. [Google Scholar] [CrossRef]
- Du, K.; Zhang, L.; Lee, T.; Sun, T. M6A RNA Methylation Controls Neural Development and Is Involved in Human Diseases. Mol. Neurobiol. 2019, 56, 1596–1606. [Google Scholar] [CrossRef] [PubMed]
- Livneh, I.; Moshitch-Moshkovitz, S.; Amariglio, N.; Rechavi, G.; Dominissini, D. The m 6 A Epitranscriptome: Transcriptome Plasticity in Brain Development and Function. Nat. Rev. Neurosci. 2020, 21, 36–51. [Google Scholar] [CrossRef] [PubMed]
- Yoon, K.-J.; Koo, B.-K.; Im, S.-K.; Jeong, H.-W.; Ghim, J.; Kwon, M.; Moon, J.-S.; Miyata, T.; Kong, Y.-Y. Mind Bomb 1-Expressing Intermediate Progenitors Generate Notch Signaling to Maintain Radial Glial Cells. Neuron 2008, 58, 519–531. [Google Scholar] [CrossRef] [PubMed]
- Weng, Y.-L.; Wang, X.; An, R.; Cassin, J.; Vissers, C.; Liu, Y.; Liu, Y.; Xu, T.; Wang, X.; Wong, S.Z.H.; et al. Epitranscriptomic M6A Regulation of Axon Regeneration in the Adult Mammalian Nervous System. Neuron 2018, 97, 313–325.e6. [Google Scholar] [CrossRef]
- Du, T.; Rao, S.; Wu, L.; Ye, N.; Liu, Z.; Hu, H.; Xiu, J.; Shen, Y.; Xu, Q. An Association Study of the M6A Genes with Major Depressive Disorder in Chinese Han Population. J. Affect. Disord. 2015, 183, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Zhang, X.; Weng, Y.-L.; Lu, Z.; Liu, Y.; Lu, Z.; Li, J.; Hao, P.; Zhang, Y.; Zhang, F.; et al. M6A Facilitates Hippocampus-Dependent Learning and Memory through Ythdf1. Nature 2018, 563, 249–253. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zang, L.; Zhang, F.; Chen, J.; Shen, H.; Shu, L.; Liang, F.; Feng, C.; Chen, D.; Tao, H.; et al. Fat Mass and Obesity-Associated (FTO) Protein Regulates Adult Neurogenesis. Hum. Mol. Genet. 2017, 26, 2398–2411. [Google Scholar] [CrossRef]
- Hess, M.E.; Hess, S.; Meyer, K.D.; Verhagen, L.A.W.; Koch, L.; Brönneke, H.S.; Dietrich, M.O.; Jordan, S.D.; Saletore, Y.; Elemento, O.; et al. The Fat Mass and Obesity Associated Gene (Fto) Regulates Activity of the Dopaminergic Midbrain Circuitry. Nat. Neurosci. 2013, 16, 1042–1048. [Google Scholar] [CrossRef]
- Chen, X.; Yu, C.; Guo, M.; Zheng, X.; Ali, S.; Huang, H.; Zhang, L.; Wang, S.; Huang, Y.; Qie, S.; et al. Down-Regulation of M6A MRNA Methylation Is Involved in Dopaminergic Neuronal Death. ACS Chem. Neurosci. 2019, 10, 2355–2363. [Google Scholar] [CrossRef]
- Chen, B.; Ye, F.; Yu, L.; Jia, G.; Huang, X.; Zhang, X.; Peng, S.; Chen, K.; Wang, M.; Gong, S.; et al. Development of Cell-Active N6-Methyladenosine RNA Demethylase FTO Inhibitor. J. Am. Chem. Soc. 2012, 134, 17963–17971. [Google Scholar] [CrossRef] [PubMed]
- Aik, W.; Demetriades, M.; Hamdan, M.K.K.; Bagg, E.A.L.; Yeoh, K.K.; Lejeune, C.; Zhang, Z.; McDonough, M.A.; Schofield, C.J. Structural Basis for Inhibition of the Fat Mass and Obesity Associated Protein (FTO). J. Med. Chem. 2013, 56, 3680–3688. [Google Scholar] [CrossRef]
- Zheng, G.; Cox, T.; Tribbey, L.; Wang, G.Z.; Iacoban, P.; Booher, M.E.; Gabriel, G.J.; Zhou, L.; Bae, N.; Rowles, J.; et al. Synthesis of a FTO Inhibitor with Anticonvulsant Activity. ACS Chem. Neurosci. 2014, 5, 658–665. [Google Scholar] [CrossRef]
- Huang, Y.; Yan, J.; Li, Q.; Li, J.; Gong, S.; Zhou, H.; Gan, J.; Jiang, H.; Jia, G.-F.; Luo, C.; et al. Meclofenamic Acid Selectively Inhibits FTO Demethylation of M6A over ALKBH5. Nucleic Acids Res. 2015, 43, 373–384. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, B.S.; Roundtree, I.A.; Lu, Z.; Han, D.; Ma, H.; Weng, X.; Chen, K.; Shi, H.; He, C. N6-Methyladenosine Modulates Messenger RNA Translation Efficiency. Cell 2015, 161, 1388–1399. [Google Scholar] [CrossRef]
- Huang, Y.; Su, R.; Sheng, Y.; Dong, L.; Dong, Z.; Xu, H.; Ni, T.; Zhang, Z.S.; Zhang, T.; Li, C.; et al. Small-Molecule Targeting of Oncogenic FTO Demethylase in Acute Myeloid Leukemia. Cancer Cell 2019, 35, 677–691.e10. [Google Scholar] [CrossRef] [PubMed]
- Karelson, M.; Selberg, S. Method of Suppressing Cancer by RNA M6A Demethylase AlkBH5 Inhibitors. WO2020207550, 15 October 2020. [Google Scholar]
- Irwin, J.J.; Shoichet, B.K. ZINC—A Free Database of Commercially Available Compounds for Virtual Screening. J. Chem. Inf. Model. 2005, 45, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Bento, A.P.; Gaulton, A.; Hersey, A.; Bellis, L.J.; Chambers, J.; Davies, M.; Krüger, F.A.; Light, Y.; Mak, L.; McGlinchey, S.; et al. The ChEMBL Bioactivity Database: An Update. Nucleic Acids Res. 2014, 42, D1083–D1090. [Google Scholar] [CrossRef]
- Zhang, X.; Wei, L.-H.; Wang, Y.; Xiao, Y.; Liu, J.; Zhang, W.; Yan, N.; Amu, G.; Tang, X.; Zhang, L.; et al. Structural Insights into FTO’s Catalytic Mechanism for the Demethylation of Multiple RNA Substrates. Proc. Natl. Acad. Sci. USA 2019, 116, 2919–2924. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.-Y.; Jokitalo, E.; Sun, Y.-F.; Mehlen, P.; Lindholm, D.; Saarma, M.; Arumäe, U. GDNF-Deprived Sympathetic Neurons Die via a Novel Nonmitochondrial Pathway. J. Cell Biol. 2003, 163, 987–997. [Google Scholar] [CrossRef] [PubMed]
- Lindholm, P.; Voutilainen, M.H.; Laurén, J.; Peränen, J.; Leppänen, V.-M.; Andressoo, J.-O.; Lindahl, M.; Janhunen, S.; Kalkkinen, N.; Timmusk, T.; et al. Novel Neurotrophic Factor CDNF Protects and Rescues Midbrain Dopamine Neurons in Vivo. Nature 2007, 448, 73–77. [Google Scholar] [CrossRef]
- Yu, L.; Saarma, M.; Arumäe, U. Death Receptors and Caspases but Not Mitochondria Are Activated in the GDNF- or BDNF-Deprived Dopaminergic Neurons. J. Neurosci. 2008, 28, 7467–7475. [Google Scholar] [CrossRef]
- Jasmin, M.; Ahn, E.H.; Voutilainen, M.H.; Fombonne, J.; Guix, C.; Viljakainen, T.; Kang, S.S.; Yu, L.; Saarma, M.; Mehlen, P.; et al. Netrin-1 and Its Receptor DCC Modulate Survival and Death of Dopamine Neurons and Parkinson’s Disease Features. EMBO J. 2021, 40, e105537. [Google Scholar] [CrossRef]
- Sidorova, Y.A.; Saarma, M. Can Growth Factors Cure Parkinson’s Disease? Trends Pharmacol. Sci. 2020, 41, 909–922. [Google Scholar] [CrossRef]
- Mauer, J.; Luo, X.; Blanjoie, A.; Jiao, X.; Grozhik, A.V.; Patil, D.P.; Linder, B.; Pickering, B.F.; Vasseur, J.-J.; Chen, Q.; et al. Reversible Methylation of m 6 A m in the 5′ Cap Controls MRNA Stability. Nature 2017, 541, 371–375. [Google Scholar] [CrossRef]
- Mauer, J.; Sindelar, M.; Despic, V.; Guez, T.; Hawley, B.R.; Vasseur, J.-J.; Rentmeister, A.; Gross, S.S.; Pellizzoni, L.; Debart, F.; et al. FTO Controls Reversible m 6 Am RNA Methylation during SnRNA Biogenesis. Nat. Chem. Biol. 2019, 15, 340–347. [Google Scholar] [CrossRef]
- Wei, C.-M.; Gershowitz, A.; Moss, B. N 6, O 2 ′-Dimethyladenosine a Novel Methylated Ribonucleoside next to the 5′ Terminal of Animal Cell and Virus MRNAs. Nature 1975, 257, 251–253. [Google Scholar] [CrossRef]
- Gerken, T.; Girard, C.A.; Tung, Y.-C.L.; Webby, C.J.; Saudek, V.; Hewitson, K.S.; Yeo, G.S.H.; McDonough, M.A.; Cunliffe, S.; McNeill, L.A.; et al. The Obesity-Associated FTO Gene Encodes a 2-Oxoglutarate–Dependent Nucleic Acid Demethylase. Science 2007, 318, 1469–1472. [Google Scholar] [CrossRef]
- Airaksinen, M.S.; Saarma, M. The GDNF Family: Signalling, Biological Functions and Therapeutic Value. Nat. Rev. Neurosci. 2002, 3, 383–394. [Google Scholar] [CrossRef] [PubMed]
- Madhavi Sastry, G.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and Ligand Preparation: Parameters, Protocols, and Influence on Virtual Screening Enrichments. J. Comput. Aided Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Stephens, P.J.; Devlin, F.J.; Chabalowski, C.F.; Frisch, M.J. Ab Initio Calculation of Vibrational Absorption and Circular Dichroism Spectra Using Density Functional Force Fields. J. Phys. Chem. 1994, 98, 11623–11627. [Google Scholar] [CrossRef]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing (SC ’06), Tampa, FL, USA, 11–17 November 2006; p. 43. [Google Scholar]
- Banks, J.L.; Beard, H.S.; Cao, Y.; Cho, A.E.; Damm, W.; Farid, R.; Felts, A.K.; Halgren, T.A.; Mainz, D.T.; Maple, J.R.; et al. Integrated Modeling Program, Applied Chemical Theory (IMPACT). J. Comput. Chem. 2005, 26, 1752–1780. [Google Scholar] [CrossRef]
- Toukmaji, A.Y.; Board, J.A. Ewald Summation Techniques in Perspective: A Survey. Comput. Phys. Commun. 1996, 95, 73–92. [Google Scholar] [CrossRef]
- Zielkiewicz, J. Structural Properties of Water: Comparison of the SPC, SPCE, TIP4P, and TIP5P Models of Water. J. Chem. Phys. 2005, 123, 104501. [Google Scholar] [CrossRef]
- McQuin, C.; Goodman, A.; Chernyshev, V.; Kamentsky, L.; Cimini, B.A.; Karhohs, K.W.; Doan, M.; Ding, L.; Rafelski, S.M.; Thirstrup, D.; et al. CellProfiler 3.0: Next-Generation Image Processing for Biology. PLoS Biol. 2018, 16, e2005970. [Google Scholar] [CrossRef]
- Mahato, A.K.; Kopra, J.; Renko, J.-M.; Visnapuu, T.; Korhonen, I.; Pulkkinen, N.; Bespalov, M.M.; Domanskyi, A.; Ronken, E.; Piepponen, T.P.; et al. Glial Cell Line–Derived Neurotrophic Factor Receptor Rearranged During Transfection Agonist Supports Dopamine Neurons In Vitro and Enhances Dopamine Release In Vivo. Mov. Disord. 2020, 35, 245–255. [Google Scholar] [CrossRef]
- Joncour, V.L.; Karaman, S.; Laakkonen, P.M. Predicting In Vivo Payloads Delivery Using a Blood-Brain Tumor-Barrier in a Dish. J. Vis. Exp. 2019, e59384. [Google Scholar] [CrossRef] [PubMed]
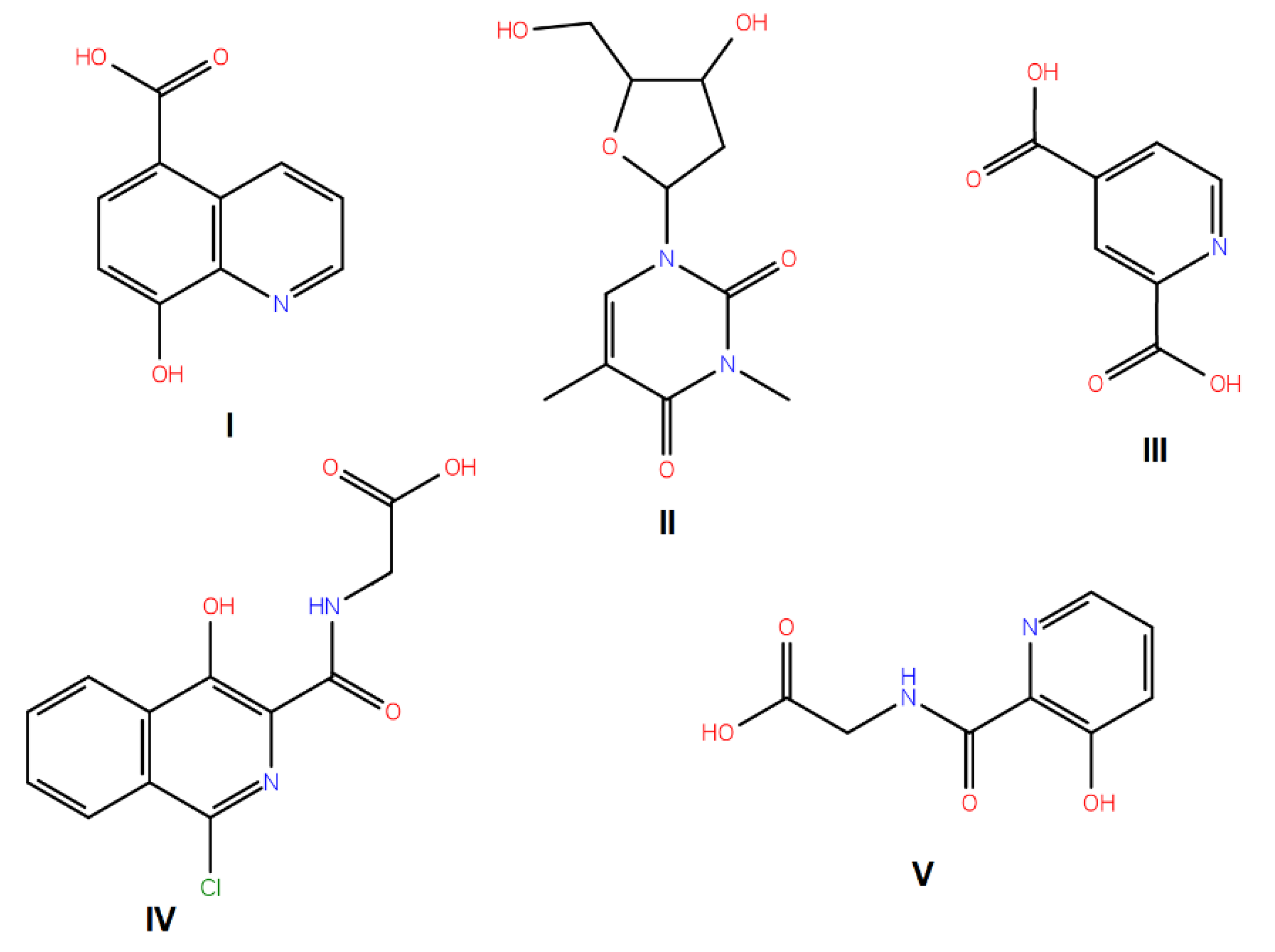

| No. | Compound Structure | ΔG (kcal/mol) | DE |
|---|---|---|---|
| 1 | 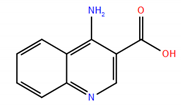 | −7.37 | 0.53 |
| 2 | 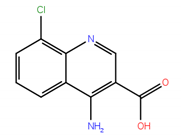 | −7.70 | 0.51 |
| 3 | 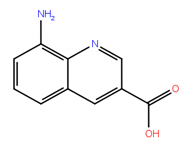 | −7.03 | 0.50 |
| 4 | 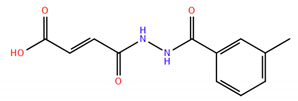 | −8.78 | 0.49 |
| 5 | 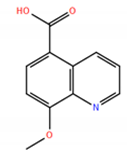 | −7.17 | 0.48 |
| 6 | 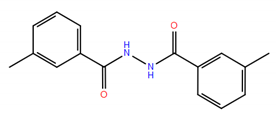 | −9.45 | 0.47 |
| 7 * | 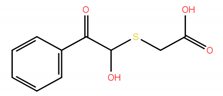 | −6.53 | 0.44 |
| 8 * | 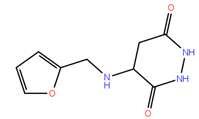 | −4.78 | 0.32 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Selberg, S.; Yu, L.-Y.; Bondarenko, O.; Kankuri, E.; Seli, N.; Kovaleva, V.; Herodes, K.; Saarma, M.; Karelson, M. Small-Molecule Inhibitors of the RNA M6A Demethylases FTO Potently Support the Survival of Dopamine Neurons. Int. J. Mol. Sci. 2021, 22, 4537. https://doi.org/10.3390/ijms22094537
Selberg S, Yu L-Y, Bondarenko O, Kankuri E, Seli N, Kovaleva V, Herodes K, Saarma M, Karelson M. Small-Molecule Inhibitors of the RNA M6A Demethylases FTO Potently Support the Survival of Dopamine Neurons. International Journal of Molecular Sciences. 2021; 22(9):4537. https://doi.org/10.3390/ijms22094537
Chicago/Turabian StyleSelberg, Simona, Li-Ying Yu, Olesja Bondarenko, Esko Kankuri, Neinar Seli, Vera Kovaleva, Koit Herodes, Mart Saarma, and Mati Karelson. 2021. "Small-Molecule Inhibitors of the RNA M6A Demethylases FTO Potently Support the Survival of Dopamine Neurons" International Journal of Molecular Sciences 22, no. 9: 4537. https://doi.org/10.3390/ijms22094537
APA StyleSelberg, S., Yu, L.-Y., Bondarenko, O., Kankuri, E., Seli, N., Kovaleva, V., Herodes, K., Saarma, M., & Karelson, M. (2021). Small-Molecule Inhibitors of the RNA M6A Demethylases FTO Potently Support the Survival of Dopamine Neurons. International Journal of Molecular Sciences, 22(9), 4537. https://doi.org/10.3390/ijms22094537







