Microscopic Methods for Identification of Sulfate-Reducing Bacteria from Various Habitats
Abstract
1. Introduction
2. Ecological Characteristics of Sulfate-Reducing Bacteria
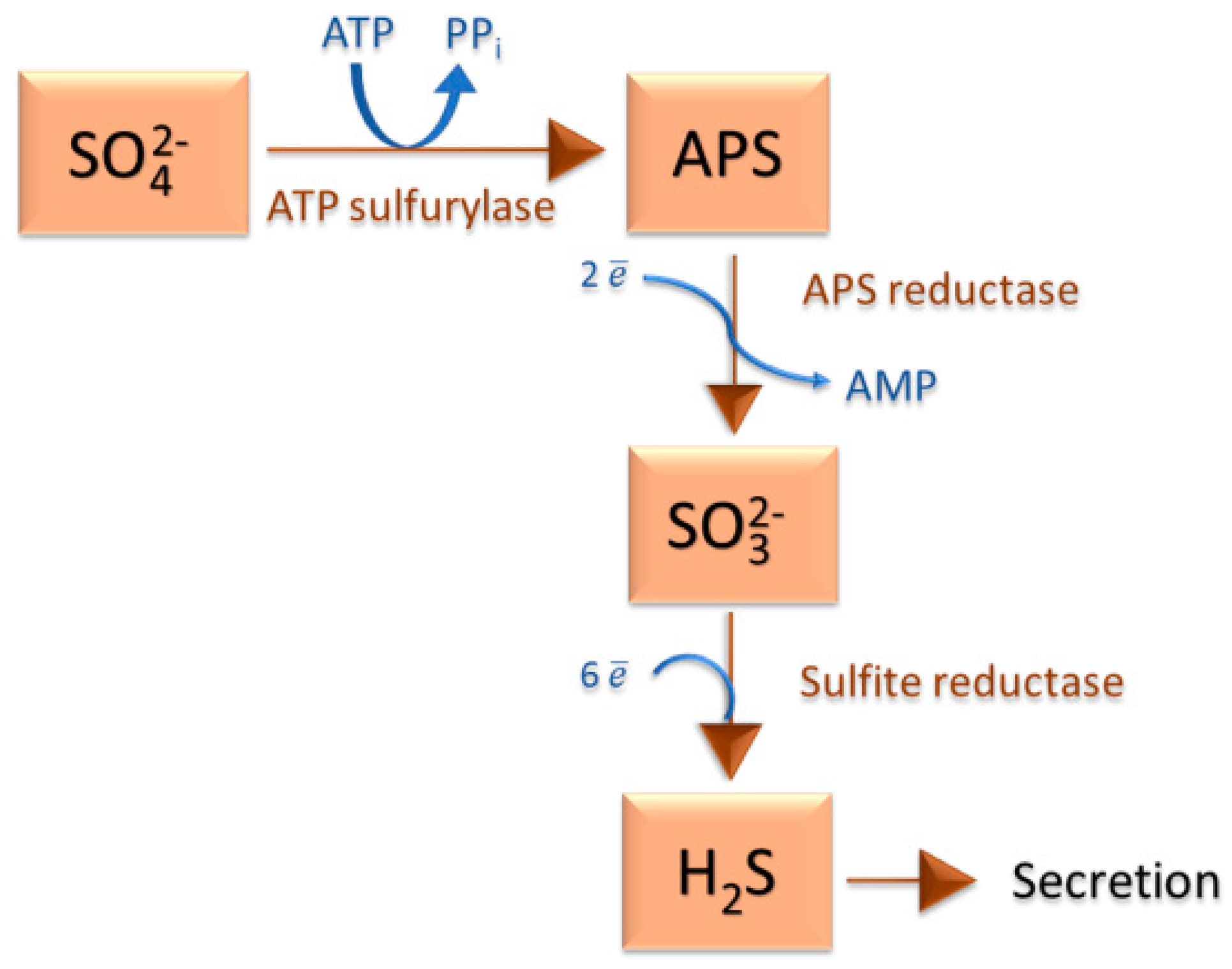
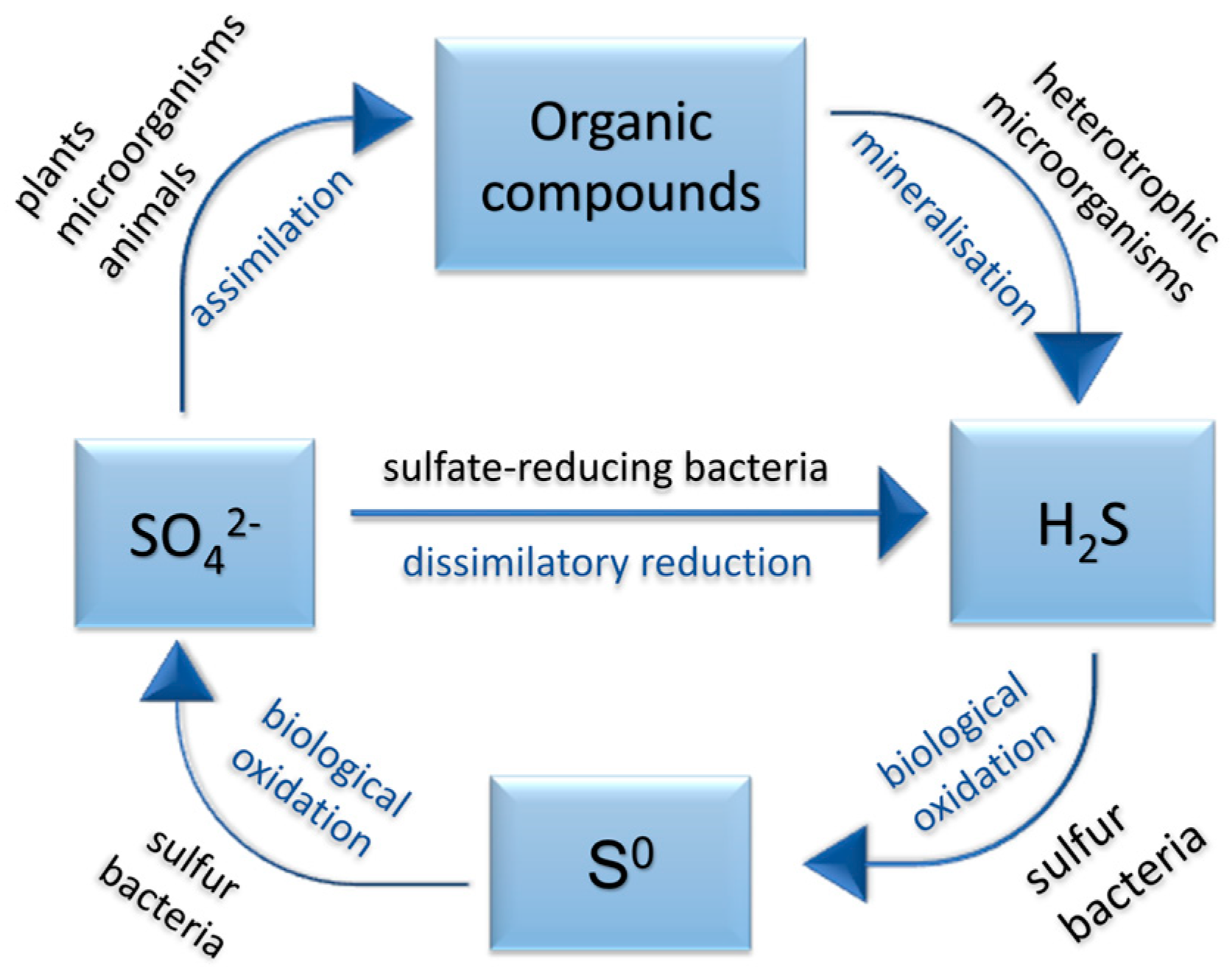
2.1. Sulfate-Reducing Bacteria and the Sulfur Cycle
2.2. Habitats of Sulfate-Reducing Bacteria
2.2.1. Water and Water Sediment
2.2.2. Soil and Mud
2.2.3. Oil Fields and Industrial Environment
2.2.4. Large Intestine of Humans and Animals
3. Morphological and Biochemical Characteristics of Selected SRB
4. Molecular Methods for Detection of Sulfate-Reducing Bacteria
4.1. Molecular Methods for Classification
4.2. Molecular Methods for Detection and Identification
4.3. Usage of Metagenomics
5. Microscopic Methods for Detection of Sulfate-Reducing Bacteria
5.1. Light Microscopy
5.2. Basic Light Microscopy Modifications
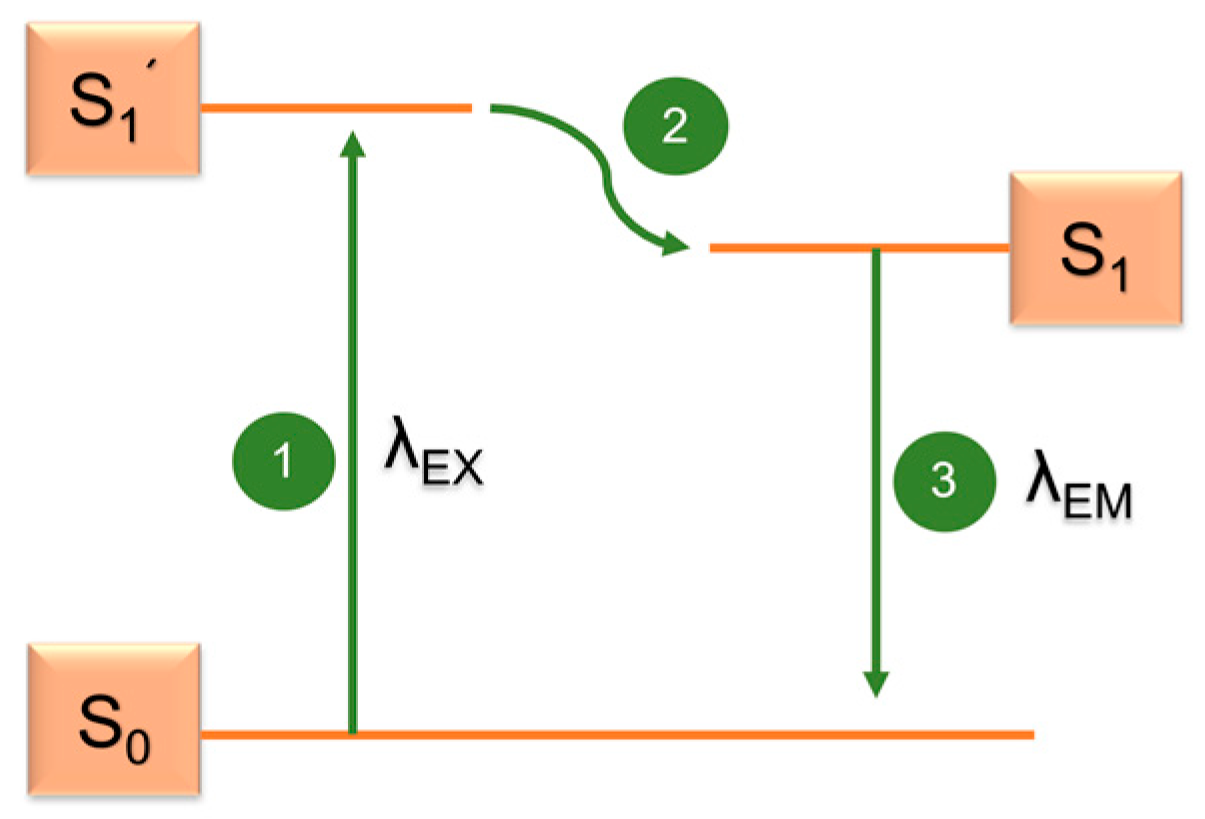
5.3. Fluorescence Microscopy
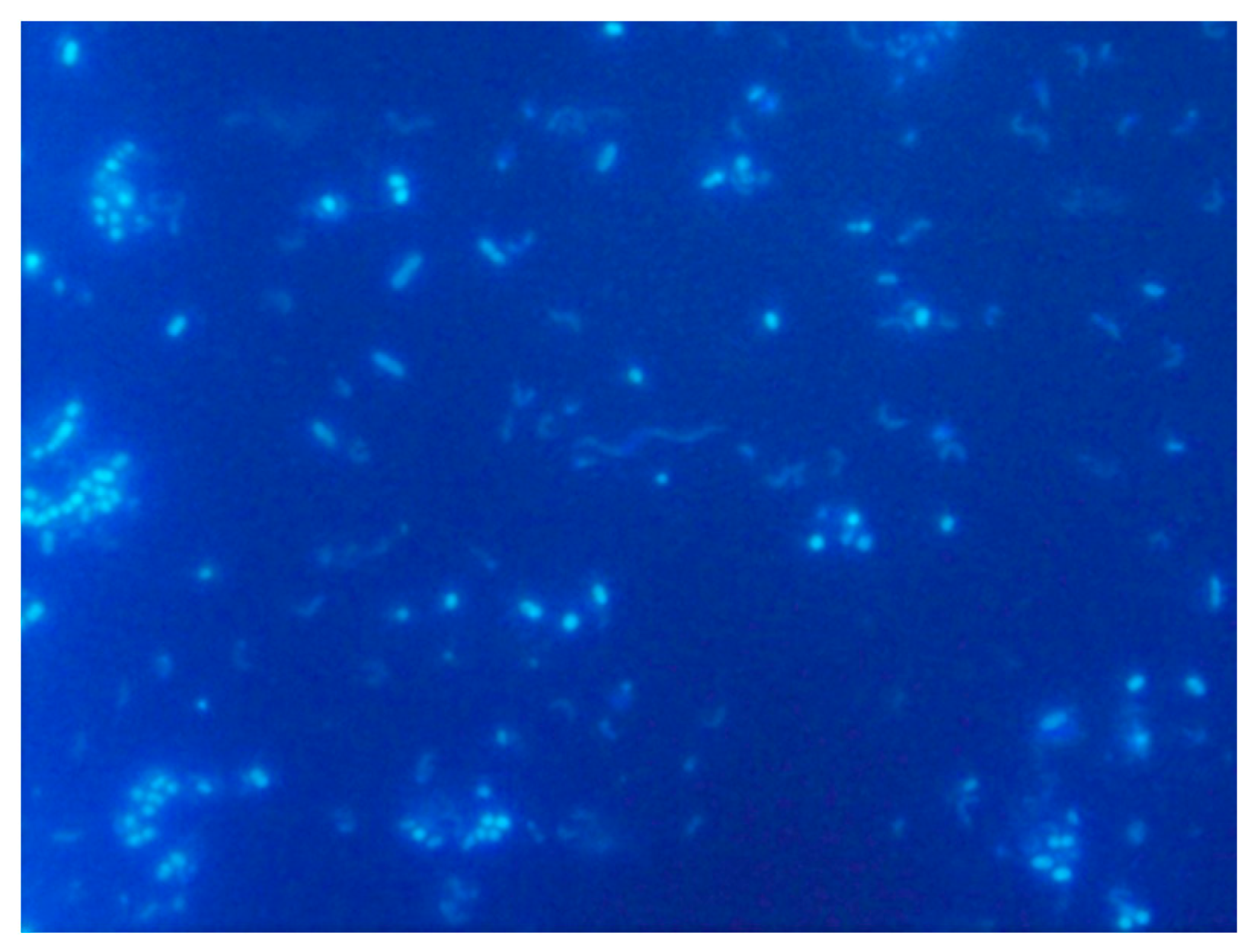
5.3.1. DAPI Fluorescent Dye
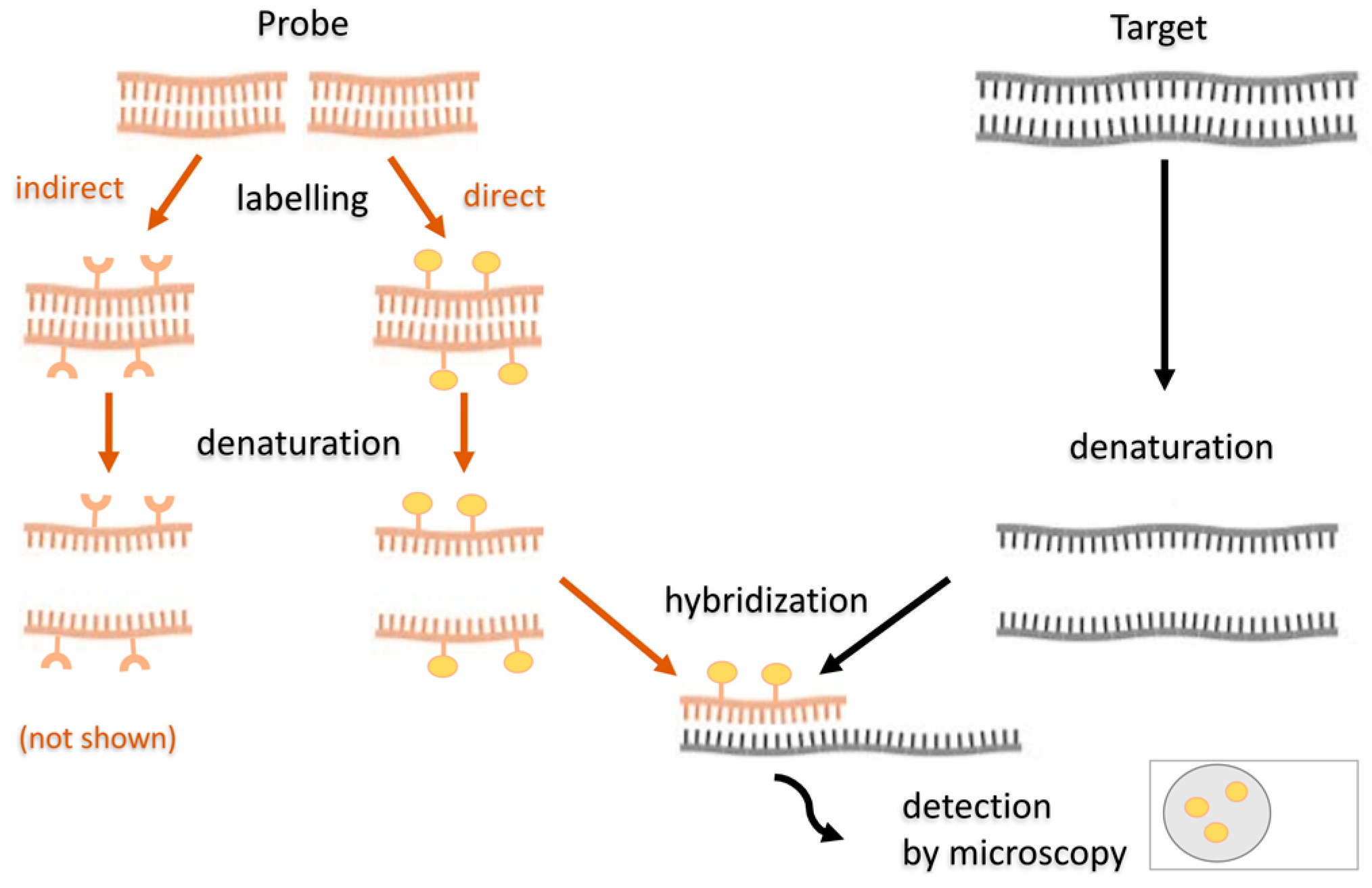
5.3.2. Fluorescence In Situ Hybridization
5.3.3. Fluorescence In Situ Hybridization in SRB Studies
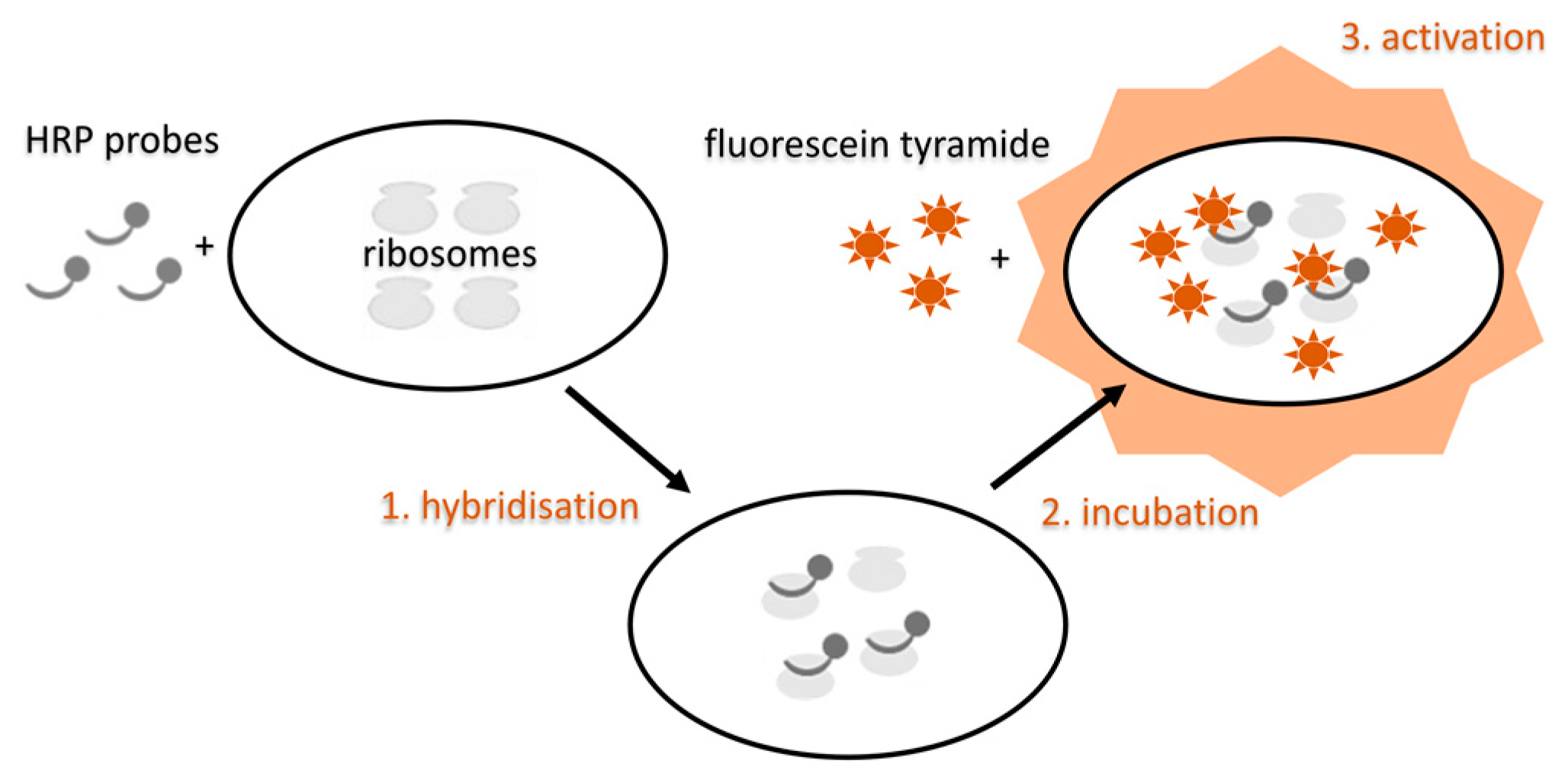
5.3.4. Modifications of the FISH Method
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Plugge, C.M.; Zhang, W.; Scholten, J.C.M.; Stams, A.J.M. Metabolic Flexibility of Sulfate-Reducing Bacteria. Front. Microbiol. 2002, 2. [Google Scholar] [CrossRef]
- Barton, L.L.; Fauque, G.D. Chapter 2 Biochemistry, Physiology and Biotechnology of Sulfate-Reducing Bacteria. In Advances in Applied Microbiology; Academic Press: Cambridge, MA, USA, 2009; Volume 68, pp. 41–98. [Google Scholar]
- Kushkevych, I.; Coufalová, M.; Vítězová, M.; Rittmann, S.K.-M.R. Sulfate-Reducing Bacteria of the Oral Cavity and Their Relation with Periodontitis—Recent Advances. JCM 2020, 9, 2347. [Google Scholar] [CrossRef]
- Kushkevych, I. Isolation and Purification of Sulfate-Reducing Bacteria. In Microorganisms; Blumenberg, M., Shaaban, M., Elgaml, A., Eds.; IntechOpen: London, UK, 2020; ISBN 978-1-83880-187-8. [Google Scholar]
- Gibson, G.R. Physiology and Ecology of the Sulphate-Reducing Bacteria. J. Appl. Bacteriol. 1990, 69, 769–797. [Google Scholar] [CrossRef]
- Dalsgaard, T.; Bak, F. Nitrate Reduction in a Sulfate-Reducing Bacterium, Desulfovibrio Desulfuricans, Isolated from Rice Paddy Soil: Sulfide Inhibition, Kinetics, and Regulation. Appl. Environ. Microbiol. 1994, 60, 291–297. [Google Scholar] [CrossRef]
- Kushkevych, I.V. Kinetic Properties of Pyruvate Ferredoxin Oxidoreductase of Intestinal Sulfate-Reducing Bacteria Desulfovibrio Piger Vib-7 and Desulfomicrobium Sp. Rod-9. Pol. J. Microbiol. 2015, 64, 107–114. [Google Scholar] [CrossRef]
- Kushkevych, I.V. Activity and Kinetic Properties of Phosphotransacetylase from Intestinal Sulfate-Reducing Bacteria. Acta Biochim. Pol. 2015, 62, 103–108. [Google Scholar] [CrossRef]
- Kushkevych, I.; Fafula, R.; Parák, T.; Bartoš, M. Activity of Na+/K+-Activated Mg2+-Dependent ATP-Hydrolase in the Cell-Free Extracts of the Sulfate-Reducing Bacteria Desulfovibrio Piger Vib-7 and Desulfomicrobium Sp. Rod-9. Acta Vet. Brno. 2015, 84, 3–12. [Google Scholar] [CrossRef]
- Fauque, G. Ecology of Sulfate-Reducing Bacteria. In The Sulfate-Reducing Bacteria; Springer: New York, NY, USA, 1995; pp. 217–241. ISBN 978-1-4899-1584-9. [Google Scholar]
- Postgate, J.R. The Sulphate-Reducing Bacteria, 2nd ed.; Cambridge University Press: Cambridge, UK, 1984; ISBN 978-0-521-25791-3. [Google Scholar]
- Sen, A.M.; Johnson, B. Acidophilic sulphate-reducing bacteria: Candidates for bioremediation of acid mine drainage. In Process Metallurgy; Amils, R., Ballester, A., Eds.; Elsevier: Amsterdam, The Netherlands, 1999; Volume 9, pp. 709–718. [Google Scholar]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Toxicity of Hydrogen Sulfide toward Sulfate-Reducing Bacteria Desulfovibrio Piger Vib-7. Arch. Microbiol. 2019, 201, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Kollar, P.; Vítězová, M.; Drago, L. Hydrogen Sulfide as a Toxic Product in the Small–Large Intestine Axis and Its Role in IBD Development. JCM 2019, 8, 1054. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Kotrsová, V.; Dordević, D.; Buňková, L.; Vítězová, M.; Amedei, A. Hydrogen Sulfide Effects on the Survival of Lactobacilli with Emphasis on the Development of Inflammatory Bowel Diseases. Biomolecules 2019, 9, 752. [Google Scholar] [CrossRef] [PubMed]
- Langendijk, P.S.; Hanssen, J.T.; Van der Hoeven, J.S. Sulfate-Reducing Bacteria in Association with Human Periodontitis. J. Clin. Periodontol. 2000, 27, 943–950. [Google Scholar] [CrossRef] [PubMed]
- Barton, L.L.; Ritz, N.L.; Fauque, G.D.; Lin, H.C. Sulfur Cycling and the Intestinal Microbiome. Dig. Dis. Sci. 2017, 62, 2241–2257. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Kollar, P.; Ferreira, A.L.; Palma, D.; Duarte, A.; Lopes, M.M.; Bartos, M.; Pauk, K.; Imramovsky, A.; Jampilek, J. Antimicrobial Effect of Salicylamide Derivatives against Intestinal Sulfate-Reducing Bacteria. J. Appl. Biomed. 2016, 14, 125–130. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kollar, P.; Suchy, P.; Parak, T.; Pauk, K.; Imramovsky, A. Activity of Selected Salicylamides against Intestinal Sulfate-Reducing Bacteria. Neuro Endocrinol. Lett. 2015, 36 (Suppl. 1), 106–113. [Google Scholar]
- Kushkevych, I.; Vítězová, M.; Kos, J.; Kollár, P.; Jampílek, J. Effect of Selected 8-Hydroxyquinoline-2-Carboxanilides on Viability and Sulfate Metabolism of Desulfovibrio Piger. J. Appl. Biomed. 2018, 16, 241–246. [Google Scholar] [CrossRef]
- Rabus, R.; Dworkin, M.; Falkow, S.; Rosenberg, E.; Schleifer, K.-H.; Stackebrandt, E.; Hansen, T.A.; Widdel, F. Dissimilatory Sulfate- and Sulfur-Reducing Prokaryotes. In The Prokaryotes; Springer: Berlin, Germany, 2006; pp. 659–768. ISBN 978-0-387-25492-0. [Google Scholar]
- The Sulfate-Reducing Bacteria: Contemporary Perspectives; Odom, J.M., Singleton, R., Eds.; Brock/Springer Series in Contemporary Bioscience; Springer: New York, NY, USA, 1993; ISBN 978-1-4613-9265-1. [Google Scholar]
- Widdel, F.; Bak, F. Gram-Negative Mesophilic Sulfate-Reducing Bacteria. In The Prokaryotes: A Handbook on the Biology of Bacteria: Ecophysiology, Isolation, Identification, Applications; Balows, A., Trüper, H.G., Dworkin, M., Harder, W., Schleifer, K.-H., Eds.; Springer: New York, NY, USA, 1992; pp. 3352–3378. ISBN 978-1-4757-2191-1. [Google Scholar]
- Kushkevych, I.; Cejnar, J.; Treml, J.; Dordević, D.; Kollar, P.; Vítězová, M. Recent Advances in Metabolic Pathways of Sulfate Reduction in Intestinal Bacteria. Cells 2020, 9, 698. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Kováč, J.; Kaucká, P.; Jesionek, W.; Bartoš, M.; Barton, L. A New Combination of Substrates: Biogas Production and Diversity of the Methanogenic Microorganisms. Open Life Sci. 2018, 13, 119–128. [Google Scholar] [CrossRef]
- Černý, M.; Vítězová, M.; Vítěz, T.; Bartoš, M.; Kushkevych, I. Variation in the Distribution of Hydrogen Producers from the Clostridiales Order in Biogas Reactors Depending on Different Input Substrates. Energies 2018, 11, 3270. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Vítěz, T.; Bartoš, M. Production of Biogas: Relationship between Methanogenic and Sulfate-Reducing Microorganisms. Open Life Sci. 2017, 12, 82–91. [Google Scholar] [CrossRef]
- Postgate, J. Sulphate Reduction by Bacteria. Annu. Rev. Microbiol. 1959, 13, 505–520. [Google Scholar] [CrossRef]
- Tang, K.; Baskaran, V.; Nemati, M. Bacteria of the Sulphur Cycle: An Overview of Microbiology, Biokinetics and Their Role in Petroleum and Mining Industries. Biochem. Eng. J. 2009, 44, 73–94. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M.; Rittmann, S.K.-M.R. Environmental Impact of Sulfate-Reducing Bacteria, Their Role in Intestinal Bowel Diseases, and Possible Control by Bacteriophages. Appl. Sci. 2021, 11, 735. [Google Scholar] [CrossRef]
- Abdulina, D.; Kováč, J.; Iutynska, G.; Kushkevych, I. ATP Sulfurylase Activity of Sulfate-Reducing Bacteria from Various Ecotopes. 3 Biotech 2020, 10, 55. [Google Scholar] [CrossRef]
- Cypionka, H. Oxygen Respiration by Desulfovibrio Species. Annu. Rev. Microbiol. 2000, 54, 827–848. [Google Scholar] [CrossRef]
- Okabe, S. Ecophysiology of sulphate-reducing bacteria in environmental biofilms. In Sulphate-Reducing Bacteria: Environmental and Engineered Systems; Barton, L.L., Hamilton, W.A., Eds.; Cambridge University Press: Cambridge, UK, 2007; pp. 359–382. ISBN 978-0-521-85485-6. [Google Scholar]
- Okabe, S.; Itoh, T.; Satoh, H.; Watanabe, Y. Analyses of Spatial Distributions of Sulfate-Reducing Bacteria and Their Activity in Aerobic Wastewater Biofilms. Appl. Environ. Microbiol. 1999, 65, 5107–5116. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Nielsen, J.; Okabe, S.; Watanabe, Y.; Nielsen, P. Phylogenetic Identification and Substrate Uptake Patterns of Sulfate-Reducing Bacteria Inhabiting an Oxic-Anoxic Sewer Biofilm Determined by Combining Microautoradiography and Fluorescent In Situ Hybridization. Appl. Environ. Microbiol. 2002, 68, 356–364. [Google Scholar] [CrossRef] [PubMed]
- Ollivier, B.; Hatchikian, C.E.; Prensier, G.; Guezennec, J.; Garcia, J.-L. Desulfohalobium Retbaense Gen. Nov., Sp. Nov., a Halophilic Sulfate-Reducing Bacterium from Sediments of a Hypersaline Lake in Senegal. IJSB 1991, 41, 74–81. [Google Scholar]
- Kushkevych, I.; Leščanová, O.; Dordević, D.; Jančíková, S.; Hošek, J.; Vítězová, M.; Buňková, L.; Drago, L. The Sulfate-Reducing Microbial Communities and Meta-Analysis of Their Occurrence during Diseases of Small–Large Intestine Axis. JCM 2019, 8, 1656. [Google Scholar] [CrossRef] [PubMed]
- Beech, I.B.; Sunner, J. Sulphate-reducing bacteria and their role in corrosion of ferrous materials. In Sulphate-Reducing Bacteria–Environmental and Engineered Systems; Cambridge University Press: Cambridge, UK, 2007; pp. 459–482. ISBN 978-0-511-29028-2. [Google Scholar]
- McInerney, M.J.; Bryant, M.P. Anaerobic Degradation of Lactate by Syntrophic Associations of Methanosarcina Barkeri and Desulfovibrio Species and Effect of H(2) on Acetate Degradation. Appl. Environ. Microbiol. 1981, 41, 346–354. [Google Scholar] [CrossRef]
- Ouattara, A.; Jacq, V. Characterization of Sulfate-Reducing Bacteria Isolated from Senegal Ricefields. FEMS Microbiol. Ecol. 1992, 10, 217–228. [Google Scholar] [CrossRef]
- Kushkevych, I.; Abdulina, D.; Kováč, J.; Dordević, D.; Vítězová, M.; Iutynska, G.; Rittmann, S.K.-M.R. Adenosine-5′-Phosphosulfate- and Sulfite Reductases Activities of Sulfate-Reducing Bacteria from Various Environments. Biomolecules 2020, 10, 921. [Google Scholar] [CrossRef]
- Hamilton, W.A.; Lee, W. Biocorrosion. In Sulfate-Reducing Bacteria; Barton, L.L., Ed.; Biotechnology Handbooks; Springer: Boston, MA, USA, 1995; pp. 243–264. ISBN 978-1-4899-1582-5. [Google Scholar]
- Rabus, R.; Fukui, M.; Wilkes, H.; Widdle, F. Degradative Capacities and 16S RRNA-Targeted Whole-Cell Hybridization of Sulfate-Reducing Bacteria in an Anaerobic Enrichment Culture Utilizing Alkylbenzenes from Crude Oil. Appl. Environ. Microbiol. 1996, 62, 3605–3613. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Vítězová, M. Possible Synergy Effect of Hydrogen Sulfide and Acetate Produced by Sulfate-Reducing Bacteria on Inflammatory Bowel Disease Development. J. Adv. Res. 2020, S2090123220300564. [Google Scholar] [CrossRef]
- Kushkevych, I.; Dordević, D.; Kollár, P. Analysis of Physiological Parameters of Desulfovibrio Strains from Individuals with Colitis. Open Life Sci. 2019, 13, 481–488. [Google Scholar] [CrossRef]
- Dordević, D.; Jančíková, S.; Vítězová, M.; Kushkevych, I. Hydrogen Sulfide Toxicity in the Gut Environment: Meta-Analysis of Sulfate-Reducing and Lactic Acid Bacteria in Inflammatory Processes. J. Adv. Res. 2020, S2090123220300527. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Dordević, D.; Vítězová, M.; Kollár, P. Cross-Correlation Analysis of the Desulfovibrio Growth Parameters of Intestinal Species Isolated from People with Colitis. Biologia 2018, 73, 1137–1143. [Google Scholar] [CrossRef]
- Kushkevych, I.; Castro Sangrador, J.; Dordević, D.; Rozehnalová, M.; Černý, M.; Fafula, R.; Vítězová, M.; Rittmann, S.K.-M.R. Evaluation of Physiological Parameters of Intestinal Sulfate-Reducing Bacteria Isolated from Patients Suffering from IBD and Healthy People. JCM 2020, 9, 1920. [Google Scholar] [CrossRef] [PubMed]
- Gibson, G.R.; Macfarlane, G.T.; Cummings, J.H. Occurrence of Sulphate-Reducing Bacteria in Human Faeces and the Relationship of Dissimilatory Sulphate Reduction to Methanogenesis in the Large Gut. J. Appl. Bacteriol. 1988, 65, 103–111. [Google Scholar] [CrossRef]
- Vítězová, M.; Kohoutová, A.; Vítěz, T.; Hanišáková, N.; Kushkevych, I. Methanogenic Microorganisms in Industrial Wastewater Anaerobic Treatment. Processes 2020, 8, 1546. [Google Scholar] [CrossRef]
- Christl, S.U.; Gibson, G.R.; Cummings, J.H. Role of Dietary Sulphate in the Regulation of Methanogenesis in the Human Large Intestine. Gut 1992, 33, 1234–1238. [Google Scholar] [CrossRef]
- Florin, T.H.J.; Neale, G.; Goretski, S.; Cummings, J.H. The Sulfate Content of Foods and Beverages. J. Food Compos. Anal. 1993, 6, 140–151. [Google Scholar] [CrossRef]
- Lozniewski, A.; Maurer, P.; Schuhmacher, H.; Carlier, J.P.; Mory, F. First Isolation of Desulfovibrio Species as Part of a Polymicrobial Infection from a Brain Abscess. Eur. J. Clin. Microbiol. Infect. Dis 1999, 18, 602–603. [Google Scholar] [CrossRef]
- Loubinoux, J.; Bronowicki, J.-P.; Pereira, I.A.C.; Mougenel, J.-L.; Faou, A.E. Sulfate-Reducing Bacteria in Human Feces and Their Association with Inflammatory Bowel Diseases. FEMS Microbiol. Ecol. 2002, 40, 107–112. [Google Scholar] [CrossRef]
- Ichiishi, S.; Tanaka, K.; Nakao, K.; Izumi, K.; Mikamo, H.; Watanabe, K. First Isolation of Desulfovibrio from the Human Vaginal Flora. Anaerobe 2010, 16, 229–233. [Google Scholar] [CrossRef] [PubMed]
- Kushkevych, I.; Abdulina, D.; Dordević, D.; Rozehnalová, M.; Vítězová, M.; Černý, M.; Svoboda, P.; Rittmann, S.K.-M.R. Basic Bioelement Contents in Anaerobic Intestinal Sulfate-Reducing Bacteria. Appl. Sci. 2021, 11, 1152. [Google Scholar] [CrossRef]
- Kushkevych, I.; Vítězová, M.; Fedrová, P.; Vochyanová, Z.; Paráková, L.; Hošek, J. Kinetic Properties of Growth of Intestinal Sulphate-Reducing Bacteria Isolated from Healthy Mice and Mice with Ulcerative Colitis. Acta Vet. Brno 2017, 86, 405–411. [Google Scholar] [CrossRef]
- Cummings, J.H.; Macfarlane, G.T.; Macfarlane, S. Intestinal Bacteria and Ulcerative Colitis. Curr. Issues Intest. Microbiol. 2003, 4, 9–20. [Google Scholar] [PubMed]
- Dzierzewicz, Z.; Cwalina, B.; Szczerba, J.; Wȩglarz, L.; Wilczok, T. Diversity of Ribosomal RNA Operon in Strains of Desulfovibrio desulfuricans. In Focus on DNA fingerprinting research; Nova Biomedical Books: New York, NY, USA, 2006; pp. 61–98. ISBN 978-1-59454-953-3. [Google Scholar]
- Petroleum Microbiology; Ollivier, B., Magot, M., Eds.; ASM Press: Washington, DC, USA, 2005; ISBN 978-1-68367-196-1. [Google Scholar]
- Holt, J.G.; Krieg, N.R.; Sneath, P.H. (Eds.) Bergey’s Manual of Determinative Bacteriology, 9th ed.; Lippincott, Williams & Wilkins: Baltimore, MD, USA, 1994; ISBN 10-0-683-00603-7. [Google Scholar]
- Whitman, W.B. Bergey’s Manual of Systematics of Archaea and Bacteria, 1st ed.; Wiley: Hoboken, NJ, USA, 2015; ISBN 978-1-118-96060-8. [Google Scholar]
- Kováč, J.; Kushkevych, I. New Modification of Cultivation Medium for Isolation and Growth of Intestinal Sulfate-Reducing Bacteria. In Proceedings of the MendelNet 2017, Mendel Univ Brno, Fac AgriSciences, Brno, Czech Republic, 6 November 2019; Volume 2017, pp. 702–707. [Google Scholar]
- Kuever, J.; Rainey, F.; Widdel, F. Genus Desulfovibrio. In Bergey’s Manual of Systematic Bacteriology; Springer: New York, NY, USA, 2005; pp. 926–930. ISBN 0-387-24145-0. [Google Scholar]
- Loubinoux, J.; Valente, F.M.A.; Pereira, I.A.C.; Costa, A.; Grimont, P.A.D.; Le Faou, A.E. Reclassification of the Only Species of the Genus Desulfomonas, Desulfomonas Pigra, as Desulfovibrio Piger Comb. Nov. Int. J. Syst. Evol. Microbiol. 2002, 52, 1305–1308. [Google Scholar] [CrossRef]
- Kuever, J.; Rainey, F.; Widdel, F. Genus Desulfobacter. In Bergey’s Manual of Systematic Bacteriology; Springer: New York, NY, USA, 2005; pp. 961–964. ISBN 0-387-24145-0. [Google Scholar]
- Kuever, J.; Rainey, F.; Widdel, F. Genus Desulfobulbus. In Bergey’s Manual of Systematic Bacteriology; Springer: New York, NY, USA, 2005; pp. 988–992. ISBN 0-387-24145-0. [Google Scholar]
- Kuever, J.; Rainey, F.A. Genus Desulfotomaculum. In Bergey’s Manual of Systematics of Archaea and Bacteria; American Cancer Society: Atlanta, GA, USA, 2015; pp. 1–12. ISBN 978-1-118-96060-8. [Google Scholar]
- Sharak Genthner, B.R.; Devereux, R. Desulfomicrobium. In Bergey’s Manual of Systematics of Archaea and Bacteria; American Cancer Society: Atlanta, GA, USA, 2015; pp. 1–9. ISBN 978-1-118-96060-8. [Google Scholar]
- Thevenieau, F.; Fardeau, M.-L.; Ollivier, B.; Joulian, C.; Baena, S. Desulfomicrobium Thermophilum Sp. Nov., a Novel Thermophilic Sulphate-Reducing Bacterium Isolated from a Terrestrial Hot Spring in Colombia. Extremophiles 2007, 11, 295–303. [Google Scholar] [CrossRef] [PubMed]
- Langendijk, P.S.; Kulik, E.M.; Sandmeier, H.; Meyer, J.; van der Hoeven, J.S. Isolation of Desulfomicrobium Orale Sp. Nov. and Desulfovibrio Strain NY682, Oral Sulfate-Reducing Bacteria Involved in Human Periodontal Disease. Int. J. Syst. Evol. Microbiol. 2001, 51, 1035–1044. [Google Scholar] [CrossRef]
- Sharak Gerthner, B.R.; Devereux, R. Genus Desulfomicrobium. In Bergey’s Manual of Systematic Bacteriology; Springer: New York, NY, USA, 2005; pp. 944–948. ISBN 0-387-24145-0. [Google Scholar]
- Chun, J.; Oren, A.; Ventosa, A.; Christensen, H.; Arahal, D.R.; da Costa, M.S.; Rooney, A.P.; Yi, H.; Xu, X.-W.; De Meyer, S.; et al. Proposed Minimal Standards for the Use of Genome Data for the Taxonomy of Prokaryotes. Int. J. Syst. Evol. Microbiol. 2018, 68, 461–466. [Google Scholar] [CrossRef]
- Cho, J.C.; Tiedje, J.M. Bacterial Species Determination from DNA-DNA Hybridization by Using Genome Fragments and DNA Microarrays. Appl. Environ. Microbiol. 2001, 67, 3677–3682. [Google Scholar] [CrossRef] [PubMed]
- Auch, A.F.; von Jan, M.; Klenk, H.-P.; Göker, M. Digital DNA-DNA Hybridization for Microbial Species Delineation by Means of Genome-to-Genome Sequence Comparison. Stand. Genomic Sci. 2010, 2, 117–134. [Google Scholar] [CrossRef] [PubMed]
- Stackebrandt, E.; Frederiksen, W.; Garrity, G.M.; Grimont, P.A.D.; Kämpfer, P.; Maiden, M.C.J.; Nesme, X.; Rosselló-Mora, R.; Swings, J.; Trüper, H.G.; et al. Report of the Ad Hoc Committee for the Re-Evaluation of the Species Definition in Bacteriology. Int. J. Syst. Evol. Microbiol. 2002, 52, 1043–1047. [Google Scholar] [CrossRef]
- Goris, J.; Konstantinidis, K.T.; Klappenbach, J.A.; Coenye, T.; Vandamme, P.; Tiedje, J.M. DNA–DNA Hybridization Values and Their Relationship to Whole-Genome Sequence Similarities. Int. J. Syst. Evol. Microbiol. 2007, 57, 81–91. [Google Scholar] [CrossRef] [PubMed]
- Felis, G.; Dellaglio, F. Taxonomy of Lactobacilli and Bifidobacteria. Curr. Issues Intest. Microbiol. 2007, 8, 44–61. [Google Scholar]
- Woese, C.R. Bacterial Evolution. Microbiol. Rev. 1987, 51, 221–271. [Google Scholar] [CrossRef] [PubMed]
- Stackebrandt, E.; Stahl, D.A.; Devereux, R. Taxonomic Relationships. In Sulfate-Reducing Bacteria; Barton, L.L., Ed.; Biotechnology Handbooks; Springer: Boston, MA, USA, 1995; pp. 49–87. ISBN 978-1-4899-1582-5. [Google Scholar]
- Castro, H.F.; Williams, N.H.; Ogram, A. Phylogeny of Sulfate-Reducing Bacteria(1). FEMS Microbiol. Ecol. 2000, 31, 1–9. [Google Scholar] [CrossRef]
- Daly, K.; Sharp, R.J.; McCarthy, A.J. Development of Oligonucleotide Probes and PCR Primers for Detecting Phylogenetic Subgroups of Sulfate-Reducing Bacteria. Microbiology 2000, 146 Pt 7, 1693–1705. [Google Scholar] [CrossRef]
- Wegmann, U.; Nueno Palop, C.; Mayer, M.J.; Crost, E.; Narbad, A. Complete Genome Sequence of Desulfovibrio Piger FI11049. Genome Announc. 2017, 5. [Google Scholar] [CrossRef]
- Kováč, J.; Vítězová, M.; Kushkevych, I. Metabolic Activity of Sulfate-Reducing Bacteria from Rodents with Colitis. Open Med. 2018, 13, 344–349. [Google Scholar] [CrossRef]
- Kushkevych, I.; Kos, J.; Kollar, P.; Kralova, K.; Jampilek, J. Activity of Ring-Substituted 8-Hydroxyquinoline-2-Carboxanilides against Intestinal Sulfate-Reducing Bacteria Desulfovibrio Piger. Med. Chem. Res. 2018, 27, 278–284. [Google Scholar] [CrossRef]
- Kushkevych, I. Identification of sulfate-reducing bacteria strains of human large intestine. Studia Biol. 2013, 7, 115–132. [Google Scholar] [CrossRef]
- Pepper, I.L.; Gerba, C.P.; Brendecke, J.W. Environmental Microbiology: A Laboratory Manual; Academic Press: San Diego, CA, USA, 1995; ISBN 978-0-12-550655-7. [Google Scholar]
- Chang, I.S.; Ballard, J.D.; Krumholz, L.R. Evidence for Chimeric Sequences Formed during Random Arbitrarily Primed PCR. J. Microbiol. Methods 2003, 54, 427–431. [Google Scholar] [CrossRef]
- Versalovic, J.; Koeuth, T.; Lupski, J.R. Distribution of Repetitive DNA Sequences in Eubacteria and Application to Fingerprinting of Bacterial Genomes. Nucleic Acids Res. 1991, 19, 6823–6831. [Google Scholar] [CrossRef] [PubMed]
- Dzierżewicz, Z.; Szczerba, J.; Wȩglarz, L.; Świa̧tkowska, L.; Jasinska, D.; Wilczok, T. Intraspecies Variability of Desulfovibrio Desulfuricans Strains Determined by the Genetic Profiles. FEMS Microbiol. Lett. 2003, 219, 69–74. [Google Scholar] [CrossRef]
- Devereux, R.; Kane, M.D.; Winfrey, J.; Stahl, D.A. Genus- and Group-Specific Hybridization Probes for Determinative and Environmental Studies of Sulfate-Reducing Bacteria. Syst. Appl. Microbiol. 1992, 15, 601–609. [Google Scholar] [CrossRef]
- Loubinoux, J.; Mory, F.; Pereira, I.A.C.; Faou, A.E.L. Bacteremia Caused by a Strain of Desulfovibrio Related to the Provisionally Named Desulfovibrio Fairfieldensis. J. Clin. Microbiol. 2000, 38, 1707. [Google Scholar] [CrossRef]
- McDougall, R.; Robson, J.; Paterson, D.; Tee, W. Bacteremia Caused by a Recently Described Novel Desulfovibrio Species. J. Clin. Microbiol. 1997, 35, 1805–1808. [Google Scholar] [CrossRef] [PubMed]
- Tee, W.; Dyall-Smith, M.; Woods, W.; Eisen, D. Probable New Species of Desulfovibrio Isolated from a Pyogenic Liver Abscess. J. Clin. Microbiol. 1996, 34, 1760–1764. [Google Scholar] [CrossRef]
- Geets, J.; Borremans, B.; Diels, L.; Springael, D.; Vangronsveld, J.; van der Lelie, D.; Vanbroekhoven, K. DsrB Gene-Based DGGE for Community and Diversity Surveys of Sulfate-Reducing Bacteria. J. Microbiol. Methods 2006, 66, 194–205. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Jiménez, J.R.; Kerkhof, L.J. Phylogeography of Sulfate-Reducing Bacteria among Disturbed Sediments, Disclosed by Analysis of the Dissimilatory Sulfite Reductase Genes (DsrAB). AEM 2005, 71, 1004–1011. [Google Scholar] [CrossRef]
- Tuffin, M.; Anderson, D.; Heath, C.; Cowan, D.A. Metagenomic Gene Discovery: How Far Have We Moved into Novel Sequence Space? Biotechnol. J. 2009, 4, 1671–1683. [Google Scholar] [CrossRef] [PubMed]
- Pinnell, L.J.; Turner, J.W. Shotgun Metagenomics Reveals the Benthic Microbial Community Response to Plastic and Bioplastic in a Coastal Marine Environment. Front. Microbiol. 2019, 10, 1252. [Google Scholar] [CrossRef] [PubMed]
- Dyksma, S.; Pjevac, P.; Ovanesov, K.; Mussmann, M. Evidence for H 2 Consumption by Uncultured Desulfobacterales in Coastal Sediments: H2 -Consuming Sulfate Reducers in Coastal Sediments. Environ. Microbiol. 2018, 20, 450–461. [Google Scholar] [CrossRef]
- Li, J.; Jia, P.; Wang, X.; Feng, S.; Yang, T.; Fang, Z.; Liu, J.; Liao, B.; Shu, W.; Liang, J.-L. Metagenomic Insights into Sulfate-Reducing Bacteria in a Revegetated Acidic Mine Wasteland. Res. Sq. 2020, 1–21. [Google Scholar] [CrossRef]
- Ran, S.; Mu, C.; Zhu, W. Diversity and Community Pattern of Sulfate-Reducing Bacteria in Piglet Gut. J. Anim. Sci. Biotechnol. 2019, 10, 40. [Google Scholar] [CrossRef]
- Kocherginskaya, S.A.; Aminov, R.I.; White, B.A. Analysis of the Rumen Bacterial Diversity under Two Different Diet Conditions Using Denaturing Gradient Gel Electrophoresis, Random Sequencing, and Statistical Ecology Approaches. Anaerobe 2001, 7, 119–134. [Google Scholar] [CrossRef]
- Regensbogenova, M.; Pristas, P.; Javorsky, P.; Moon-van der Staay, S.Y.; van der Staay, G.W.M.; Hackstein, J.H.P.; Newbold, C.J.; McEwan, N.R. Assessment of Ciliates in the Sheep Rumen by DGGE. Lett. Appl. Microbiol. 2004, 39, 144–147. [Google Scholar] [CrossRef]
- Galbraith, E.A.; Antonopoulos, D.A.; White, B.A. Suppressive Subtractive Hybridization as a Tool for Identifying Genetic Diversity in an Environmental Metagenome: The Rumen as a Model. Environ. Microbiol. 2004, 6, 928–937. [Google Scholar] [CrossRef]
- Ferrer, M.; Golyshina, O.V.; Chernikova, T.N.; Khachane, A.N.; Reyes-Duarte, D.; Santos, V.A.P.M.D.; Strompl, C.; Elborough, K.; Jarvis, G.; Neef, A.; et al. Novel Hydrolase Diversity Retrieved from a Metagenome Library of Bovine Rumen Microflora: Enzymatic Diversity from Bovine Rumen Metagenome. Environ. Microbiol. 2005, 7, 1996–2010. [Google Scholar] [CrossRef] [PubMed]
- Fichtel, K.; Mathes, F.; Könneke, M.; Cypionka, H.; Engelen, B. Isolation of Sulfate-Reducing Bacteria from Sediments Above the Deep-Subseafloor Aquifer. Front. Microbiol. 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Pluta, M. Nomarski’s DIC Microscopy: A Review; Pluta, M., Szyjer, M., Eds.; Materials Science, Engineering: Warsaw, Poland, 1994; pp. 10–25. [Google Scholar] [CrossRef]
- Fan, X.; Healy, J.J.; O’Dwyer, K.; Hennelly, B.M. Label-Free Color Staining of Quantitative Phase Images of Biological Cells by Simulated Rheinberg Illumination. Appl. Opt. 2019, 58, 3104. [Google Scholar] [CrossRef]
- Lichtman, J.W.; Conchello, J.-A. Fluorescence Microscopy. Nat. Methods 2005, 2, 910–919. [Google Scholar] [CrossRef]
- Life Technologies. The Molecular Probes Handbook: A Guide to Fluorescent Probes and Labeling Technologies, 11th ed.; Johnson, I., Ed.; Life Technologies: Carlsbad, CA, USA, 2010; ISBN 978-0-9829279-1-5. [Google Scholar]
- Gough, H.L.; Stahl, D.A. Optimization of Direct Cell Counting in Sediment. J. Microbiol. Methods 2003, 52, 39–46. [Google Scholar] [CrossRef]
- Amalfitano, S.; Fazi, S. Recovery and Quantification of Bacterial Cells Associated with Streambed Sediments. J. Microbiol. Methods 2008, 75, 237–243. [Google Scholar] [CrossRef]
- Zhu, X.; Al-Moniee, M.A. 22—Molecular microbiology techniques. In Trends in Oil and Gas Corrosion Research and Technologies; El-Sherik, A.M., Ed.; Woodhead Publishing Series in Energy; Woodhead Publishing: Boston, MA, USA, 2017; pp. 513–536. ISBN 978-0-08-101105-8. [Google Scholar]
- Araujo, J.C.; Mortara, R.; Campos, J.R.; Vazoller, R.F. Development and Analysis of Anaerobic Biofilms onto Hydrophobic and Hydrophilic Surfaces. Environ. Technol. 2004, 25, 809–817. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyaa, J. Low Cost Wastewater Bioremediation Technology: Innovative Treatment of Sulphate and Metal.-Rich Wastewater, 1st ed.; Elsevier: Cambridge, CA, USA, 2017; ISBN 978-0-12-812510-6. [Google Scholar]
- Speicher, M.; Carter, N. The New Cytogenetics: Blurring the Boundaries with Molecular Biology. Nat. Rev. Genet. 2005. [Google Scholar] [CrossRef]
- Amann, R.; Moraru, C. Two Decades of Fluorescence in Situ Hybridization in Systematic and Applied Microbiology. Syst. Appl. Microbiol. 2012, 35, 483–484. [Google Scholar] [CrossRef]
- Amann, R.; Fuchs, B.M.; Behrens, S. The Identification of Microorganisms by Fluorescence in Situ Hybridisation. Curr. Opin. Biotechnol. 2001, 12, 231–236. [Google Scholar] [CrossRef]
- Boetius, A.; Ravenschlag, K.; Schubert, C.J.; Rickert, D.; Widdel, F.; Gieseke, A.; Amann, R.; Jørgensen, B.B.; Witte, U.; Pfannkuche, O. A Marine Microbial Consortium Apparently Mediating Anaerobic Oxidation of Methane. Nature 2000, 407, 623–626. [Google Scholar] [CrossRef]
- Probst, A.J.; Holman, H.-Y.N.; DeSantis, T.Z.; Andersen, G.L.; Birarda, G.; Bechtel, H.A.; Piceno, Y.M.; Sonnleitner, M.; Venkateswaran, K.; Moissl-Eichinger, C. Tackling the Minority: Sulfate-Reducing Bacteria in an Archaea-Dominated Subsurface Biofilm. ISME J. 2013, 7, 635–651. [Google Scholar] [CrossRef] [PubMed]
- Zwirglmaier, K. Fluorescence in Situ Hybridisation (FISH)--the next Generation. FEMS Microbiol. Lett. 2005, 246, 151–158. [Google Scholar] [CrossRef]
- Schmidt, H.; Eickhorst, T.; Mußmann, M. Gold-FISH: A New Approach for the in Situ Detection of Single Microbial Cells Combining Fluorescence and Scanning Electron Microscopy. Syst. Appl. Microbiol. 2012, 35, 518–525. [Google Scholar] [CrossRef]
- Lee, N.; Nielsen, P.H.; Andreasen, K.H.; Juretschko, S.; Nielsen, J.L.; Schleifer, K.-H.; Wagner, M. Combination of Fluorescent In Situ Hybridization and Microautoradiography—A New Tool for Structure-Function Analyses in Microbial Ecology. Appl. Environ. Microbiol. 1999, 65, 1289–1297. [Google Scholar] [CrossRef] [PubMed]
- Okabe, S.; Kindaichi, T.; Ito, T. MAR-FISH—An Ecophysiological Approach to Link Phylogenetic Affiliation and In Situ Metabolic Activity of Microorganisms at a Single-Cell Resolution. Microbes Environ. 2004, 19, 83–98. [Google Scholar] [CrossRef]
- Alonso, C. Tips and Tricks for High Quality MAR-FISH Preparations: Focus on Bacterioplankton Analysis. Syst. Appl. Microbiol. 2012, 35, 503–512. [Google Scholar] [CrossRef] [PubMed]
- Sekar, R.; Pernthaler, A.; Pernthaler, J.; Warnecke, F.; Posch, T.; Amann, R. An Improved Protocol for Quantification of Freshwater Actinobacteria by Fluorescence In Situ Hybridization. AEM 2003, 69, 2928–2935. [Google Scholar] [CrossRef] [PubMed]
- Pernthaler, A.; Pernthaler, J. Fluorescence in Situ Hybridization for the Identification of Environmental Microbes. Methods Mol. Biol. 2007, 353, 153–164. [Google Scholar] [CrossRef]
- FISH Protocols. Available online: https://www.arb-silva.de/fileadmin/graphics_fish/CARD_FISH.jpg (accessed on 13 March 2019).
| Species of Desulfovibrio | Source | Habitat |
|---|---|---|
| D. africanus | NCIB 8401 | well water |
| D. alcoholovorans | DSM 5433 | fermenter |
| D. carbinolicus | DSM 3852 | wastewater treatment plant |
| D. desulfuricans | DSM 642 | soil |
| D. piger | ATCC 29098 | human feces |
| D. gigas | NCIB 9332 | pond water |
| D. longus | DSM 6739 | oil field |
| D. sulfodismutans | DSM 3696 | freshwater mud |
| D. termitidis | DSM 5308 | intestines of termites |
| Species | Source | Habitat |
|---|---|---|
| Desulfobulbus marinus | DSM 2058 | marine mud |
| Desulfomicrobium apsheronum | AUCCM 1105 | oil field |
| Desulfobotulus sapovorans | DSM 2055 | freshwater mud |
| Desulfohalobium retbaense | DSM 5692 | hypersaline lake sediments |
| Food | Sulfate Content (µmol g−1) |
|---|---|
| Dried apples | 49 |
| Dried apricots | 30 |
| Commercial dark wheat bread | 15 |
| Commercial light wheat bread | 13 |
| Soy flour | 12 |
| Sausage | 10 |
| Almonds, hazelnuts | 9 |
| Brussels sprouts, broccoli | 9 |
| Cabbage (red, white) | 8 |
| Commercial rye bread | 8 |
| Jams, marmalades | 7 |
| Muesli | 6 |
| Group | Genus | G+C % | Desulfoviridin | Cytochromes | Acetate Oxidation |
|---|---|---|---|---|---|
| Gram-positive sporulating | |||||
| 1 | Desulfotomaculum | 48–52 | – | b, c | complete/partial |
| Gram-negative mesophilic | |||||
| 2 | Desulfobulbus | 59–60 | – | b, c, c3 | partial |
| 3 | Desulfobacterium | 41–52 | – | b, c | complete |
| 4 | Desulfobacter | 44–46 | – | no data | complete |
| 5 | Desulfococcus | 46–57 | +/– | b, c | complete |
| Desulfosarcina | 51 | – | b, c | complete | |
| 6 | Desulfovibrio | 49–66 | +/– | c3, b, c | partial |
| Desulfomicrobium | 52–67 | – | b, c | partial | |
| Thermophilic bacterial | |||||
| Thermodesulfobacterium | 30–38 | – | c3, c | partial | |
| Thermophilic archaeal | |||||
| Archaeoglobus | 41–46 | – | no data | partial | |
| Fluorophore | λEX | λEM |
|---|---|---|
| Fluorescein isothiocyanate (FITC) | 494 nm | 518 nm |
| Tetramethylrhodamine-isothiocyanate (TRITC) | 555 nm | 580 nm |
| Hoechst 33342 and 33258 | 352 nm | 461 nm |
| Propidium iodide (PI) | 536 nm | 617 nm |
| Ethidium bromide | 518 nm | 605 nm |
| DAPI (4′,6-diamidino-2-phenylindole) | 358 nm | 461 nm |
| Cy3 | 550 nm | 570 nm |
| Cy5 | 649 nm | 670 nm |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kushkevych, I.; Hýžová, B.; Vítězová, M.; Rittmann, S.K.-M.R. Microscopic Methods for Identification of Sulfate-Reducing Bacteria from Various Habitats. Int. J. Mol. Sci. 2021, 22, 4007. https://doi.org/10.3390/ijms22084007
Kushkevych I, Hýžová B, Vítězová M, Rittmann SK-MR. Microscopic Methods for Identification of Sulfate-Reducing Bacteria from Various Habitats. International Journal of Molecular Sciences. 2021; 22(8):4007. https://doi.org/10.3390/ijms22084007
Chicago/Turabian StyleKushkevych, Ivan, Blanka Hýžová, Monika Vítězová, and Simon K.-M. R. Rittmann. 2021. "Microscopic Methods for Identification of Sulfate-Reducing Bacteria from Various Habitats" International Journal of Molecular Sciences 22, no. 8: 4007. https://doi.org/10.3390/ijms22084007
APA StyleKushkevych, I., Hýžová, B., Vítězová, M., & Rittmann, S. K.-M. R. (2021). Microscopic Methods for Identification of Sulfate-Reducing Bacteria from Various Habitats. International Journal of Molecular Sciences, 22(8), 4007. https://doi.org/10.3390/ijms22084007







