Abstract
In eukaryotic cells, the nucleus houses the genomic material of the cell. The physical properties of the nucleus and its ability to sense external mechanical cues are tightly linked to the regulation of cellular events, such as gene expression. Nuclear mechanics and morphology are altered in many diseases such as cancer and premature ageing syndromes. Therefore, it is important to understand how different components contribute to nuclear processes, organisation and mechanics, and how they are misregulated in disease. Although, over the years, studies have focused on the nuclear lamina—a mesh of intermediate filament proteins residing between the chromatin and the nuclear membrane—there is growing evidence that chromatin structure and factors that regulate chromatin organisation are essential contributors to the physical properties of the nucleus. Here, we review the main structural components that contribute to the mechanical properties of the nucleus, with particular emphasis on chromatin structure. We also provide an example of how nuclear stiffness can both impact and be affected by cellular processes such as DNA damage and repair.
1. Introduction
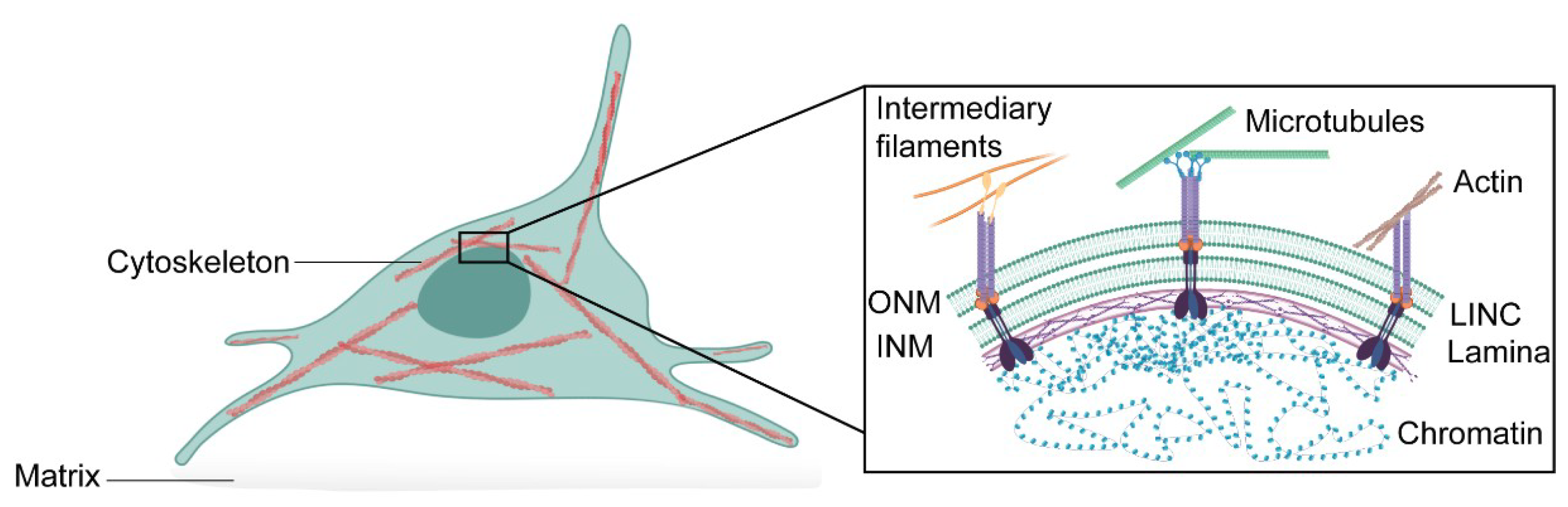
The nucleus houses the genetic information necessary for the activity and survival of the cell, but as we outline in this review, the nucleus is more than just a compartment to house DNA. Although the nucleus is the largest and stiffest cellular organelle, it is also a highly dynamic organelle that can sense the external environment and rapidly adapt [1,2,3]. The nuclear envelope comprises a double membrane—the outer nuclear membrane and the inner nuclear membrane—associated with various distinct transmembrane proteins, such as nuclear pore complexes and LEM (Lap2, emerin, and Man1)-domain proteins [4]. This is followed by an assembly of lamin filaments at the nuclear interior that provides structural stability to the organelle and tether chromatin to the nuclear envelope. From the outside, the nucleus is linked to the cytoskeleton through the Linker of Nucleoskleton and Cytoskeleton (LINC) complex, which also binds to the nuclear lamina [5]. This nuclear connectivity allows external signals to modulate nuclear functions, such as transcription [6,7] and DNA replication [8,9]. Moreover, it may allow communication in the opposite direction (Figure 1).
Figure 1.
Schematic representation of the interconnectivity between cytoskeleton, nuclear envelope and chromatin. The cytoskeleton is physically connected to the nuclear envelope consisting of the outer nuclear membrane (ONM) and the inner nuclear membrane (INM) through the LINC complex. The LINC complex is formed of trimers of SUN-domain proteins that bind different KASH-domain proteins at the nuclear membrane. LINC complexes can indirectly associate with intermediary filaments and microtubules through cyto-linker proteins or motor proteins, respectively, or directly interact with actin filaments. At the nuclear interior, the nuclear lamina tethers chromatin domains—lamina-associated domains—to the nuclear envelope. This allows effective mechanotransduction in the cell.
Within the nucleus, the DNA associates to histone cores to form nucleosomes, the building blocks of chromatin. Through epigenetic regulation, chromatin can be packaged into different conformations and higher-order structures, which determine the accessibility [10] of DNA to replication [11,12], transcription [13,14], and repair machinery [15,16]. Higher-order compact chromatin structures, known as heterochromatin, are largely inaccessible and are usually associated with genomic regions of low transcriptional activity at the nuclear periphery [17,18,19]. Meanwhile, more open conformations of chromatin, also known as euchromatin, are easily accessible and represent areas of active gene expression [20,21].
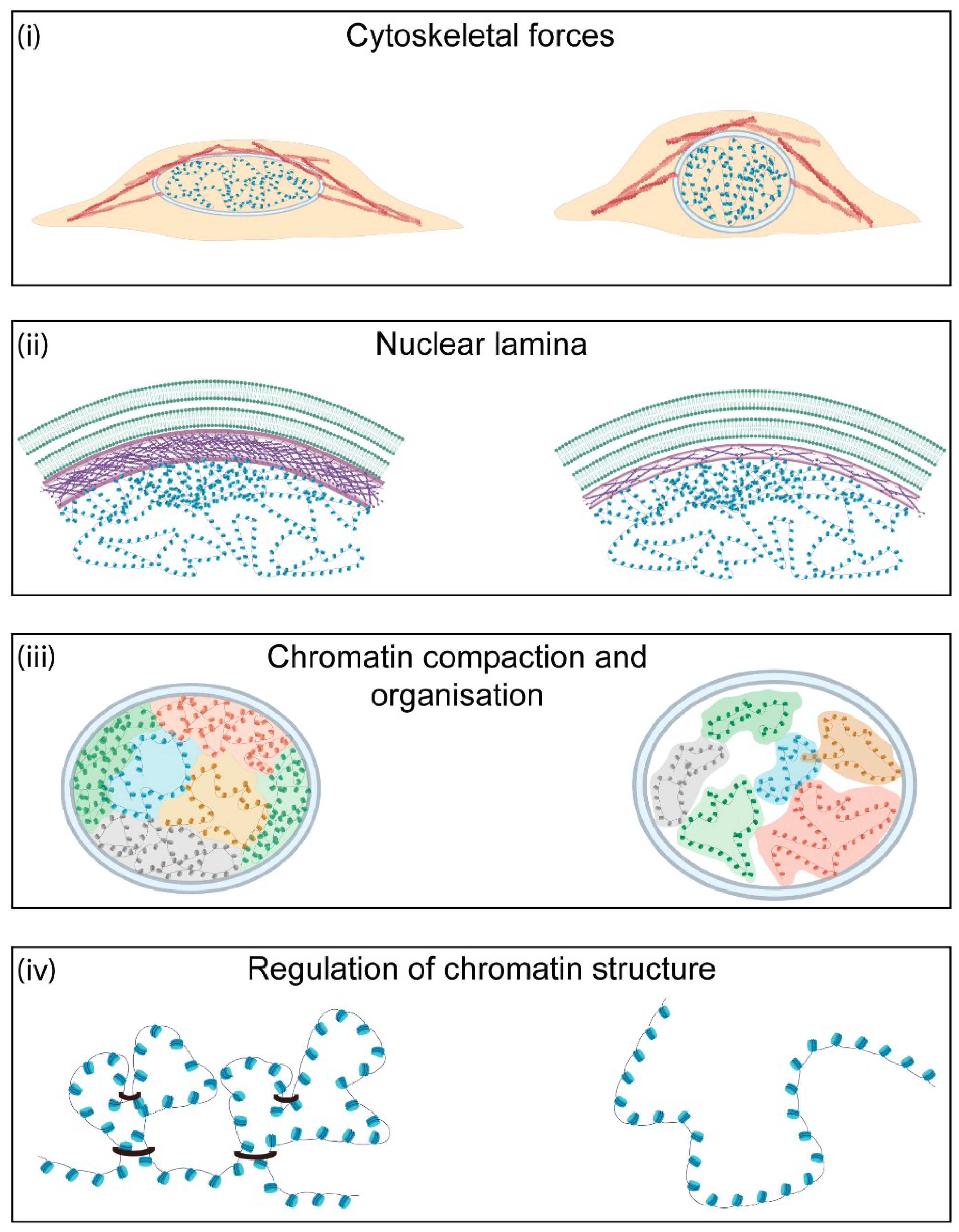
Variations to the biochemical components of the nucleus result in changes to the physical properties of the organelle and its morphology. The nuclear mechanical properties, comprising the viscoelastic behaviour and plasticity, are tightly linked to cellular function and vary between cell stages and types [22]. There are four major contributors to nuclear shape and the mechanical properties: the magnitude of cytoskeletal forces exerted on the organelle, the composition and thickness of the nuclear lamina, the level of chromatin compaction within the nucleus, and the activity of proteins that modulate DNA structure (Figure 2).
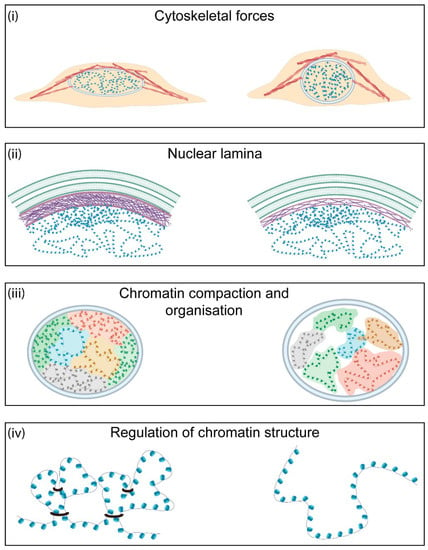
Figure 2.
Major contributors to nuclear morphology and mechanics. There are four major contributors to nuclear mechanics in the cell. (i) Cytoskeletal forces determine nuclear shape and morphology. Increased actin polymerisation leads to higher nuclear tension. (ii) The nuclear lamina is one of the major contributors to nuclear stiffness. This meshwork of intermediary filaments at the nuclear periphery is important for nuclear stability and chromatin organisation. Higher levels of lamin A/C or a thicker nuclear lamina lead to increased nuclear stiffness. (iii) Chromatin behaves as a crosslinked polymer gel. As a result, changes in chromatin organisation and levels of heterochromatin and euchromatin directly affect nuclear shape and mechanics. (iv) Regulation of chromatin architecture is dependent on the activity of several factors, such as cohesins, that allow crosslinking of chromatin and the formation of higher-order structures. The activity of these proteins can affect local and global chromatin conformation and, hence, is important for nuclear mechanics.
Altered nuclear morphology and mechanics are usually accompanied by changes in gene expression and cell function. Changes in the shape and size of the nucleus have been reported for different diseases, and in some cases, this can also be used to help diagnosis. For example, abnormally shaped nuclei can be found in cardiomyopathies, progeria and in cancer. In particular, nuclei of cervical cancer cells present herniations or blebbing, and this constitutes part of the Pap smear test diagnosis [23]; in breast cancer, nuclear pleomorphisms (altered nuclear morphology) are used for tumour grading and correlates with patient outcome [24]. It is therefore essential to understand how these changes in nuclear morphology arise, how they reflect altered mechanical properties of the nucleus and how this affects overall cellular function, mechanosensing and force transduction.
Here, we pay special attention to the newly emerging data on the importance of chromatin dynamics and the regulation of its spatial organisation. We also discuss some new technological approaches in mechanobiology and in the study of chromatin architecture. Finally, we will discuss how nuclear mechanics can influence cellular processes such as DNA damage.
2. Contributing Factors to Nuclear Mechanics
2.1. Cytoskeletal Forces in Nuclear Mechanics
Mechanotransduction refers to the process by which cells respond to external mechanical cues through the activation of biochemical pathways, changes in structure, and activation or repression of specific genes. This is a key mechanism for sensing and adapting to the extracellular microenvironment. The nucleus is the largest and the most mechanically prominent organelle in the cell, and so it would be expected to play a dominant role in cellular mechanics. It is, therefore, not surprising that it has been receiving increasing attention in the field of cell mechanics over the last decade.
Extracellular forces propagate into the nucleus through the LINC complex, located on the nuclear envelope. The LINC complex physically connects the nucleus to the cytoskeleton and, therefore, to cellular adhesions that can sense the mechanical microenvironment. Disruption of the LINC complex results in defective force transduction from the cytoplasm to the nucleus and is shown to disrupt the expression of mechanosensitive genes [25,26]. This occurs, at least in part, because cytoskeletal forces can directly affect the localisation and nuclear import of mechanosensitive transcription regulators. One functional example is the Yes-associated protein (YAP) and its transcriptional coactivator with PDZ-binding motif (TAZ). The mechanical regulation of YAP/TAZ requires cytoskeletal integrity and a functional LINC complex [27]. As a result, translocation of YAP/TAZ to the nucleus and consequent activation of YAP-dependent genes varies according to extracellular matrix rigidity [28,29], cytoskeletal formation [30], LINC complex integrity [31], and nuclear stiffness [27,32].
Cytoskeletal forces also contribute to the regulation of nuclear movement, shape, orientation and morphology. In extreme cases, the accumulation of aberrant actin fibres around the nucleus can induce actin-dependent nuclear deformation through increased nuclear tension and lead to blebbing, herniation and even rupture of the nuclear envelope [33]. Equally, chemical or genetic perturbation of the cytoskeleton results in deficient force transmission to the nucleus.
The tissue microenvironment is an important factor in cytoskeletal formation and structure, and hence it is a key determinant of the mechanical forces transmitted to the nucleus. This is particularly important when studying cancer biology and therapeutic approaches to disease. We are beginning to have insight into the mechanical consequences of different microenvironments and how they affect nuclear processes such as DNA repair, transcription and chromatin regulation.
However, studying the effects of cytoskeletal forces on nuclear processes comes with specific challenges. Many studies use drugs to disrupt cytoskeletal forces, and it is unclear how these may directly affect nuclear events, such as transcription or chromatin organisation. Latrunculin B, for example, disrupts cytoskeletal formation through inhibition of actin polymerisation, but it also disrupts nuclear actin and myosin functions, directly affecting transcription levels and RNA Polymerase II (RNAPII) spatial organisation [34]. Methods such as micropatterning [35,36] or the use of gels as substrates for cells [37,38,39] are becoming increasingly common to change the stresses exerted upon the cytoskeleton. Although these do not allow complete disruption of the cytoskeleton, they can mimic different tissue microenvironments and permit the fine-tuning of cytoskeletal forces. The data are both easier to interpret and, in some cases, can be a good alternative to chemical disruption of the cytoskeleton.
Accumulating evidence shows that actomyosin contractility around the nucleus has not only important repercussions to the deformability of the organelle, and therefore to cell migration and nuclear integrity, but it can also directly affect lamina structure and chromatin organisation. For example, using micropatterned substrates, Makhija and colleagues observed that cells lacking long stress fibres had reduced levels of lamin A/C expression, resulting in softer, more deformable nuclei. Interestingly, these cells also displayed increased chromatin and telomere dynamics, suggesting a direct relationship between geometric cell constraints and genome organisation [40]. Similarly, in rod photoreceptor cells, actomyosin deformation of the nucleus results in chromocenter clustering (condensed centromere heterochromatin regions) and inverted chromatin architecture, with euchromatin regions being redistributed to the nuclear periphery [41].
Although the molecular mechanisms that link cytoskeletal forces to chromatin regulation are still largely unexplored, spatial redistribution and misregulation of nuclear transport of transcription factors and chromatin regulators, such as transcription coactivator MKL1 [42] or histone deacetylase HDAC3 [43], have been observed and could account, at least in part, for chromatin architectural changes.
A challenge often encountered in the study of nuclear mechanics—especially when performing measurements on adhered cells—is the masking effect that the stiff cytoskeletal fibres around the nucleus have on measurements. Indeed, small changes in nuclear mechanics can be imperceptible if measured under a fully-formed cytoskeleton. Because of this, many studies resort to the use of isolated nuclei, removing the cytoskeletal impact entirely. This has proven useful when comparing nuclei across different cell types [44] to study how the expression of nuclear envelope components affects nuclear stiffness [45], or how the nucleus itself—and independently of cytosolic intervention—adapts and responds to external forces [46]. The clear trade-off is the loss of physiological environment and understanding how nuclei in live-cells respond to matrix stiffness, drug treatments, radiation or other challenges. To overcome these limitations, dos Santos et al. have recently performed mechanical measurements, using atomic force microscopy (AFM), in live cells, but at initially adhered stages. In this case, cells were allowed sufficient time to attach to their surface but, with their cytoskeleton not yet fully formed, the nucleus then occupies most of the cell volume and is the most important contributor to cell mechanics. This has allowed the detection of mechanical changes that would have otherwise not been observed [47].
Overall, there is a complex interplay of interactions between the nucleus and cytoskeletal components that contribute to the mechanics of the organelle, and much work is needed to understand how changes in cytoskeletal forces directly affect nuclear organisation and nuclear processes.
2.2. The Nuclear Lamina
The nuclear lamina is located between the INM and the chromatin. This is a dense, complex meshwork of proteins with a thickness up to 100 nm. It provides major structural support to the nucleus as well as support for a variety of nuclear functions, such as transcription, replication, DNA repair, and genome organisation. Lamina proteins fall into two separate classes, A-type and B type—lamin A and C, which are splice-isoforms of the LMNA gene—belong to the former and lamin B1 and B2, encoded by LMNB1 and LMNB2, respectively, belong to the latter [48]. Lamins belong to a class of proteins called intermediate filaments, which contain rod domains that are critical to the formation of the meshwork [49].
Post-translational modifications of lamin proteins allow the regulation of this peripheral meshwork. One important post-translational modification is the farnesylation of both lamin A and B at their C-terminal domains, which is thought to be important for the localisation and retention of these proteins at the nuclear envelope. Whilst lamin B is permanently farnesylated, lamin A is further processed by proteolytic cleavage, which includes removal of this group [48,50,51].
Whilst B-type lamins are expressed throughout development and in all nucleated cells, the levels of type-A lamins are reduced or not present at early embryonic stages. Expression of lamin A onset is highly varied in different tissues during development and, in some cases, such as for stem cells or cells of the hemopoietic system, lamin A is never expressed [52].
The nuclear lamina is crucial for maintaining nuclear envelope integrity. Depletion or mutation of lamin components leads to severe nuclear instability, morphological defects in the nucleus and gives rise to disease, as in the case of laminopathies. For instance, in mice, loss of either lamin B1 or B2 leads to neuronal defects and perinatal death [53,54]. Similarly, in humans, mutations in lamin A are associated with premature ageing and muscular malfunction, as observed in Hutchinson–Gilford progeria, muscular dystrophy, and cardiomyopathies [55].
The morphological defects observed in the nucleus of cells with lamina defects are also indicative of the important role of these proteins in nuclear mechanics. This is not surprising, as expression levels of lamin proteins, but in particular of lamin A/C, scale with nuclear stiffness. Depletion of lamin A/C makes the nucleus softer and more deformable, whilst expression of a shorter and permanently farnesylated mutant version, progerin, confers higher stiffness to the nucleus. In both cases, the expression of mechano-responsive genes is severely disrupted [22,56]. Importantly, in these cells, nuclear processes such as chromatin structure regulation, DNA replication, DNA repair and gene expression are also invariably misregulated.
However, the nuclear envelope is not mechanically isolated. Instead, it is physically connected to the cytoskeleton on one side, through the LINC complex, and to the chromatin, through lamina-associated domains (LADs) on the other. This means that mechanical changes to one of these components have large structural implications for the others. For example, Swift et al. described how matrix and cytoskeletal stiffness could directly influence lamin A expression and turnover [57]. In stiffer matrices, phosphorylation of lamin A, which promotes disassembly, is reduced, and this increases total amounts of lamin A at the nuclear envelope, which in turn increases nuclear stiffness [57,58]. Similarly, progerin-expressing cells, such as those from Hutchinson–Gilford progeria patients, display increased Filamentous-actin (F-actin) polymerisation and cytoskeletal stiffness, as well as reduced levels of heterochromatin [59]. In these cells, destabilisation of microtubules to reduce cytoskeletal tension may be a promising therapeutic approach. It has been shown to restore nuclear morphology and alleviate premature ageing in progeria in mice [60]. Conversely, a separate study also described how inhibition of histone demethylation, which directly leads to increased heterochromatin levels, also rescues morphological defects in progerin-expressing cells [61].
Chromatin function is highly dependent on its conformation. This includes correct tethering to LADs at the nuclear periphery. Loss of lamin A function, for example, leads to higher chromatin dynamics and more diffuse genomic loci, representative of higher levels of decondensed chromatin. Complete lamin loss results in detachment of LADs and disruption of global 3D chromatin-chromatin interactions [62]. Similarly, lamin B1 also has an important role in maintaining chromatin structure and distribution, especially at the nuclear periphery [63]. As a result, lamin regulation and chromatin structure are tightly linked as key components of nuclear mechanics. Future insights into how changes to nuclear envelope components affect the mechanical properties of the nucleus will be especially important in the study of ageing and cancer, where lamina mutations are often found.
2.3. Chromatin Is a Key Component of Nuclear Mechanics
Chromatin organisation is highly regulated through epigenetic histone modifications that determine local and global levels of DNA compaction. The degree of chromatin condensation and the nuclear content of hetero versus euchromatin affects not only nuclear size and morphology but also determines DNA accessibility to transcription machinery and all forms of DNA processing. In conventional nuclei, the highly compacted heterochromatin is spatially segregated from active, decondensed euchromatin, with the former usually occupying regions in the nuclear periphery and the latter in the nuclear core. Developments in genome-wide chromosome conformation capture (Hi-C) have shown that in addition to LAD formation, chromatin can also associate with itself to form sub-compartments called topologically associated domains (TADs) [64,65].
We now know that chromatin is also a major contributor to nuclear stiffness and morphology. Historically, research has focused on the more clear-cut structural contributions of lamins, in particular, lamin A. Early experiments using micropipetting showed that the lamina dominates the mechanics of swollen nuclei, whilst chromatin is the main contributor to the mechanics of shrunken nuclei. Although this suggested two different types of mechanical contribution in the nucleus, the chromatin is often regarded as a minor viscous component that flows upon applied force [22,66,67]. This view of chromatin as a secondary mechanical component, less important than the lamina, has started to change, and some recent studies have now shed light on how chromatin architecture can affect nuclear stiffness. A recent report by Strickfaden et al. highlights this by showing that self-associated condensed chromatin behaves as a solid or elastic gel instead of a liquid. This indicates that the intrinsic mechanical properties of chromatin have an impact on overall nuclear mechanics and its response to external force stimuli [68].
In line with this, experiments by Chalut et al., using an optical stretching technique, show how nuclear deformability is directly related to the degree of chromatin condensation [69]. Furthermore, experiments using MNase, for chromatin digestion also show that its structure determines nuclear morphology and governs nuclear responses to short-extension strains (<30%) [70]. We now know that changes in nuclear mechanics and shape, including the occurrence of nuclear blebbing, can occur independently of lamin perturbation due to changes in the levels in euchromatin and heterochromatin. Overexpression of nucleosome binding protein HMGN5 leads to chromatin decondensation, increased nuclear area, morphological aberrations to the nuclear envelope and nuclear softening. In mice, this leads to premature death as a result of cardiac defects [71]. Similarly, treatment with deacetylase inhibitor trichostatin A, which alters histone modifications and leads to chromatin decondensation, also results in nuclear softening and blebbing [47,61,72].
Trying to resolve individual mechanical contributions from the lamina and chromatin is a major challenge for the field of nuclear mechanics. Their physical linkage means that force measurements provide a composite value for the contribution of both components. New methodologies in force measurements could prove particularly useful. For instance, a combined AFM-side-view light-sheet fluorescence microscope developed by Nelsen et al. allowed the 3D (x/z cross-section) visualisation of live-cells whilst performing force measurements with high spatio-temporal resolution [73]. In fact, Hobson et al. recently used this approach to propose a two-regime nucleus, where lamina and chromatin respond differently to volume deformation and nuclear-area stretching [73]. Using 3D fluorescent imaging of the nucleus combined with mechanical measurements could be a powerful tool not only to probe the mechanical behaviour of different nuclear components but also to look at force transduction from the cytoskeleton, or how nuclear processes, such as DNA repair, affect cell mechanics.
Whilst mechanical differences between the lamina and chromatin are readily intuitive, differences within the chromatin itself at the nuclear interior are less so. Chromatin architecture is not homogeneous; its organisation is highly regulated, with different domains occupying different regions. This means that the nuclear interior is mechanically non-uniform. Using AFM microrheology on isolated nuclei, Lherbette et al. were able to detect mechanical variations within the nucleus, representative of a largely inhomogeneous chromatin interior. The authors observed two different mechanical regions within the nucleus, independently of the nuclear lamina—a more viscous periphery and a stiffer and predominantly elastic nuclear core [44]. A new methodology that allows a more in-depth study of chromatin environments could be important to understand how changes to chromatin organisation influence the mechanical behaviour of the nucleus.
As chromatin becomes more prominent in the field of nuclear mechanics, it becomes obvious that processes and proteins that regulate DNA structure and conformation have a large impact on the mechanical properties of the nucleus. As will be discussed later, DNA damage, occurring either through ionising radiation or genotoxic agents, such as chemotherapy drugs, causes large genomic alterations, including changes to chromatin architecture and transcription levels [74]. This can result in global relaxation of the chromatin and mechanical softening of the nuclear envelope [47]. Our knowledge of how these processes affect nuclear mechanics is still limited but may be of crucial importance.
2.4. Chromatin Conformation and Crosslinking in Nuclear Mechanics
In the nucleus, chromatin associates with a large variety of DNA-binding proteins that regulate its structure and spatial organisation. Examples of this are the previously mentioned lamins that allow crosslinking of the chromatin to the nuclear periphery. This is achieved through interactions with lamina-associated proteins and is important in the regulation of global chromatin structure. Chromatin conformations and mobility are irregular throughout the nucleus and largely determined by its level of crosslinking.
Using liquid chromatin Hi-C, Belaghzal and colleagues recently showed that chromatin loci dynamics and association are largely determined by chromatin-associated factors, such as cohesins and lamins. The authors found that chromatin behaves as a crosslinked polymer gel that, even upon digestion (within 10–25 kb), maintains its structural and mechanical connections. Furthermore, chromatin digestion at this scale did not affect nuclear stiffness, measured by micropipetting. Interestingly, after extensive chromatin digestion (<6 kb), loss of chromosome compartmentalisation was achieved, together with loss of chromatin-associated cohesins, which resulted in a 75% decrease in nuclear stiffness [75]. This indicates that compartmental segregation of chromosomes and nuclear mechanics are highly dependent on the crosslinking capabilities of proteins that modulate DNA structure.
Cohesins are highly conserved protein complexes that can loop chromatin through their ring domain to create bundles that restrict chromatin movement [76,77]. Together with the chromatin insulator CTCF, cohesin function is the driver behind TAD formation and hence crucial for chromatin 3D organisation [78,79]. CTCF/cohesin anchoring of chromatin has been shown to be important at different genomic length-scales, allowing the formation of long-range Topologically Associating Domain (TAD) compartments (megabase-sized), as well as intermediate (100 kb–1 Mb) and small-range (<100 kb) sub-compartments [80,81]. Heterochromatin protein 1 (HP1) is another protein that is known for its role in chromatin organisation. HP1 binds to H3K9me3-rich areas, which represent constitutive heterochromatin regions [82], and is capable of bridging nucleosomes [83]. This crosslinking effect of HP1 is thought to stabilise compacted chromatin states and to be essential in heterochromatin phase separation [84,85] through the formation of membrane-less condensates. A recent report by Strom et al. also showed that HP1 chromatin crosslinking capabilities are important for nuclear shape maintenance, and its degradation leads to decreased chromatin stiffness and nuclear rigidity [86].
Chromatin crosslinking also occurs at active chromatin regions. For example, the transcriptional coactivator BRD4 can create condensates at active super-enhancer regions [87]. Similarly, during transcription, RNAPII forms large molecular clusters with transcription factors. These networks have not only been associated with phase separation events but also with the formation of transient chromatin bridges [88]. Interestingly, whilst active chromatin regions are usually associated with open and dynamic conformations, recent reports challenge this view by showing that the bridging effect of RNAPII and associated transcription machinery can increase chromatin constraints, reduce chromatin mobility and possibly affect local stiffness [89,90].
The arising field of nuclear actin and myosins also provides an interesting new perspective in chromatin mechanics. Actin and myosins, in the cytoplasm, are usually associated with structural and mechanical roles. Whilst actin is one of the most abundant cytoplasmic proteins; its nuclear levels are comparatively low and tightly regulated—actin is actively transported in and out of the nucleus by importin-9 and exportin-6, respectively. Due to the low concentrations of actin and the highly dynamic and transient nature of nuclear actin filaments, visualisation of these structures is particularly challenging, and much debate still surrounds them.
There is, however, growing evidence that nuclear F-actin assembly plays important roles in transcription [91], mitosis [92], DNA replication [93], DNA repair [94] and chromatin regulation [95]. Nuclear F-actin filament formation has been observed following DNA repair response (DDR) activation, caused by telomere uncapping, treatment with genotoxic drugs or UV radiation. F-actin is recruited to sites of damage, and the formation of polymeric actin structures appear to be necessary for repair factors to cluster at DNA lesions [96,97,98,99]. These observations followed early studies showing the actin-dependent intranuclear movement of genomic loci.
To visualise nuclear F-actin polymers, Baarlink and co-workers developed a phalloidin-based assay and used dSTORM super-resolution imaging. The authors observed that these structures are necessary for reshaping nuclei following mitosis. Depletion of nuclear actin filaments, either by overexpression of exportin-6 or a polymerisation-deficient mutant of nuclear actin (NLS-actinR62D), prevented the usual chromatin decondensation and nuclear expansion that occurs following cell division. In addition, this also led to impaired transcription and, in the case of mouse embryos, impaired development [92].
Budding yeast experiments support the idea of a role for nuclear F-actin in chromatin organisation. In this case, inhibition of actin polymerisation was shown to reduce telomere and chromosome dynamics, and, interestingly, it also reduced the efficiency of DNA repair by homologous recombination (HR) [100].
It seems very likely that mechanical changes to the nucleus would occur upon the formation of nuclear actin filaments. In the cytoplasm, the extent of actin fibre formation scales with cellular stiffness. However, it is still unclear how nuclear actin filament formation affects nuclear mechanics and nuclear mechanosensing. Nuclear actin may induce direct changes to local mechanics or alternatively through altered chromatin compaction and dynamics.
In the cytoplasm, the role of actin filaments is tightly linked to the function of myosins. These molecular motors use ATP hydrolysis to generate force and movement, and thus, are essential in several cellular processes such as cell migration [101], intracellular transport [102,103] and membrane regulation [104]. Similar to F-actin, these proteins are also present in the nucleus. However, there are still many unanswered questions regarding their nuclear functions.
One example is Myosin VI (MVI). MVI associates with RNAPII and is important in transcription regulation [105,106,107]. However, the specific molecular roles of MVI in the nucleus and how they might be connected to nuclear actin filament formation have only recently started to come to light. A report by Zorca et al. showed that MVI is necessary for chromatin reorganisation at the early stages of transcription. In this study, the authors found that inhibition of MVI or actin prevented allelic pairing and gene repositioning after transcription stimulation [108]. This suggests MVI has the ability to reorganise and move chromatin across long nuclear distances and also to crosslink chromatin to allow gene proximity during transcription. More recent work by Große-Berkenbusch et al. allowed single-molecule tracking of MVI along actin filaments in the nucleus of HeLa cells [109]. This movement was observed across several micrometres, resembling the cytoplasmic motility of this molecular motor. The authors also reported ATPase-depend movement of MVI on chromatin in vitro and described how this could have an important function in chromatin organisation by regulating long-range chromatin movement. In agreement with this, using STORM imaging, Hari-Gupta et al. have shown that MVI regulates the spatial organisation of RNA Polymerase II (RNAPII) during transcription initiation. Both disruption of nuclear actin polymerisation and MVI force-sensing—through the introduction of an MVI spring mutant that changes its response to force—abrogate RNAPII distribution and have severe consequences for gene expression using RNA-Seq [34]. The role of MVI in regulating RNAPII clustering also suggests a role for this molecular motor in RNAPII-dependent chromatin bridging and droplet formation [89,90], thus leading to local changes to chromatin mechanics.
Another myosin, nuclear myosin I (NMI), also appears to be involved in a plethora of nuclear roles, including chromatin movement and DNA repair. NMI is a component of the chromatin remodelling complex B-WICH, which allows chromosomal rearrangement for RNAPI-related transcription [110,111]. Recently, KO experiments of NMI in mouse embryonic fibroblasts led to increased levels of heterochromatin and lower levels of active chromatin markers such as H3K9ac [112]. Interestingly, another study showed that with cisplatin damage, NMI is recruited to the chromatin and facilitates chromosome territory relocation in an ATM and γH2AX-dependent manner [113].
Work connecting the activity of NMI, actin and the nuclear lamina also reinforce the idea that these proteins might have more extensive structural and mechanical roles in the nucleus than previously thought [114,115,116]. As these proteins are well-known for their role in force transduction in the cytoplasm, they may have a similar role in the nucleus [117,118]. Future studies are necessary to shed light on the response of nuclear F-actin and nuclear myosins to external mechanical cues.
3. The Relationship between DNA Damage and Nuclear Mechanics
Having described the contributing factors to nuclear mechanics, we will now exemplify how this can impact cellular processes using DNA damage. DNA damage events continuously occur throughout the life cycle of a cell. Following DNA damage, the cell activates the DNA damage response (DDR) to ensure appropriate repair of lesions and their survival. Failure within these mechanisms leads to apoptosis or drives genomic instability leading to diseases such as cancer. Both DNA damage and the DDR lead to large nuclear reorganisation and activation of different biochemical pathways that may result in changes to the mechanical properties of the nucleus.
DNA damage can arise endogenously as a result of replication errors [119,120], topoisomerase activity [121] or reactive oxygen species (ROS) [122]. Conversely, external insults to the cell, such as ionising radiation or chemotherapy drugs, also cause significant DNA damage and genomic instability. Types of damage range from base-mismatches, adducts and crosslinks introduced into the DNA helix, single-strand DNA breaks or, the more deleterious and toxic, double-strand breaks. In order to allow efficient repair and prevent the propagation of mutations to daughter cells, once a lesion is detected, DDR is activated, cell cycle progression is halted, and transcriptional activity becomes markedly reduced. This is accompanied by changes to the 3D organisation of chromatin.
Two key DDR components in mammalian cells are the Ataxia Telangiectasia-mutated (ATM) and the ATM Rad3-related protein (ATR) kinases. These early responders are recruited to sites of damage to initiate DNA damage checkpoints and are responsible for the phosphorylation of additional DNA repair factors that recruit and regulate repair machinery [123]. Histone modifications also have a major role in DDR. An example is histone H2AX, which following double-strand break detection is phosphorylated (γH2AX) close to the lesion site. This creates repair centres, known as repair foci, which act as signalling hubs for the rapid recruitment of repair factors.
DNA damage and repair are not independent of the chromatin architecture. Two very early studies highlighted this relationship by showing that more accessible, open structures of chromatin are more susceptible to nuclease digestion [124,125]. We now know that local chromatin dynamics change following DNA damage and how this may be important for DDR [126]. For example, Hauer et al. describe how induction of a single double-strand break in yeast increases damaged chromatin flexibility and motility as a result of chromatin decondensation and histone eviction [127]. This was dependent on the activity of ATP-dependent chromatin remodelling complex INO80. Similarly, work in both yeast and mammalian cells show the recruitment of different remodelling complexes, such Swi/Snf family members, and changes to overall chromatin structure during DDR [128,129,130,131,132,133]. Chromatin relaxation is now considered to be an important part of DDR that improves repair efficiency.
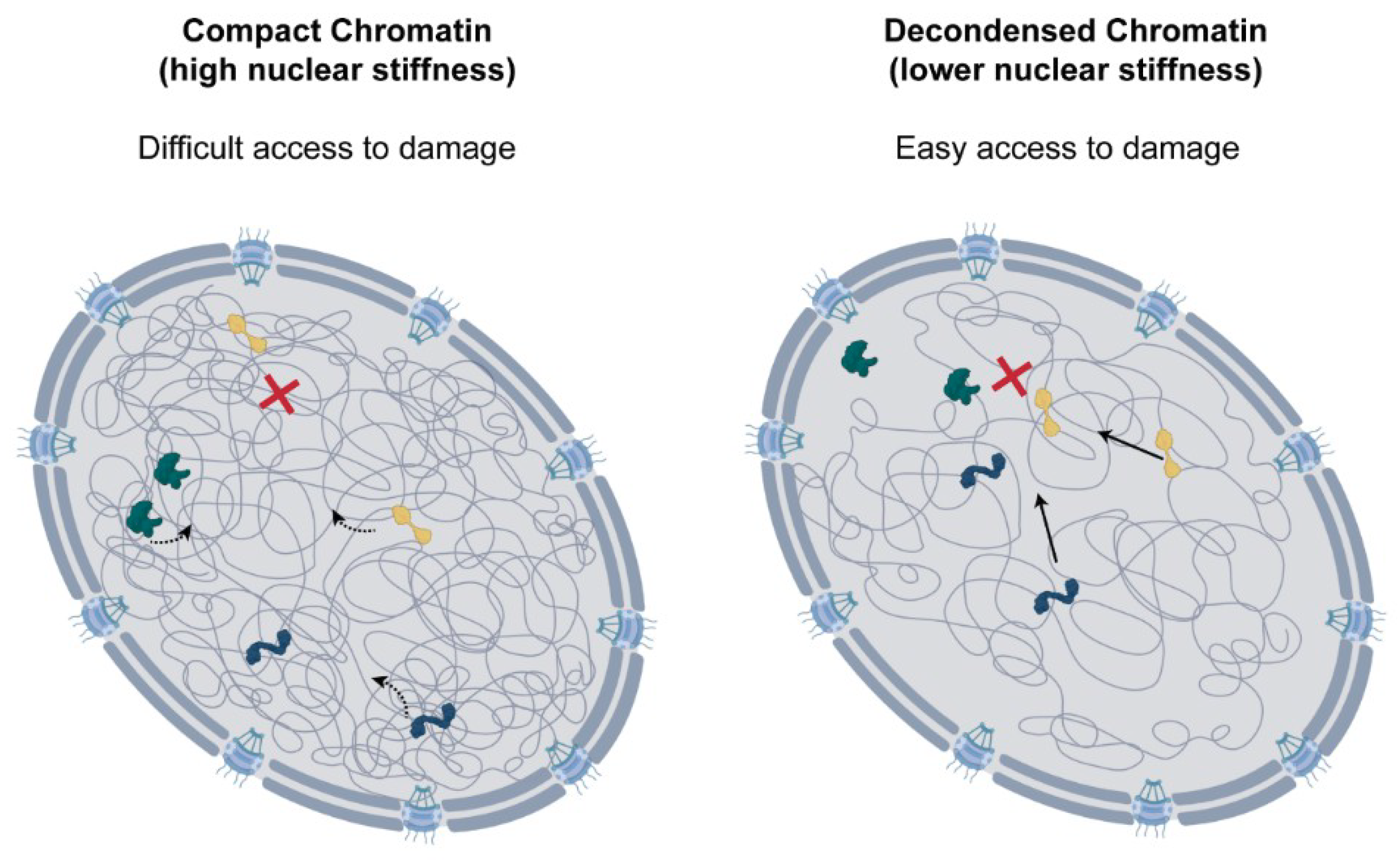
The chromatin context and nuclear positioning of the DNA lesion are also crucial determinants for DDR pathway selection [134,135,136]. Lesions occurring in active transcription regions of euchromatin, associated with histone mark H3K36me3, are preferentially repaired by the error-free HR [137,138]. For these regions, the major determinant in pathway selection appears to be chromatin accessibility and not transcriptional activity [139,140,141]. Paradoxically, DNA breaks in heterochromatin, which packages inactive transcription sites, can also preferentially undergo HR [142,143,144,145]. However, for this to occur, DNA lesions need to be relocated to more permissive environments. Chiolo et al. described that, in Drosophila, following a double-strand break, the heterochromatin domain where the break occurs expands, and the lesion is relocated to the periphery or to the exterior of this domain [146]. This movement of DNA lesions to euchromatin regions allows access to repair machinery and enables HR [146,147]. Similar dynamics of double-strand breaks have also been observed in yeast [148,149,150]. However, this is not always the case, and there is a large variability between the repair of lesions in different heterochromatin environments. For DNA breaks at LADs, HR is restricted to favour error-prone non-homologous end-joining, and, in this case, lesions do not migrate to euchromatin regions [139]. Similarly, differences between repair in centromeric and pericentric heterochromatin have also been reported, suggesting other factors might also influence repair strategies [151]. Although chromatin relaxation and reorganisation following DNA damage has been well documented, very little is known about how this impacts nuclear structure and mechanics. To shed light on this question, dos Santos et al. recently used AFM to probe the mechanical properties of the nucleus in cisplatin-treated cells. This work showed that DNA damage leads to significant softening of the nucleus, and this was the direct result of global chromatin decondensation [47]. Reduction in nuclear stiffness correlated with the levels of damage and was ATM-dependent, suggesting that DDR activation is necessary. In line with previous work, the chromatin decondensation observed led to more rapid molecular diffusion in the nucleus, which could lead to higher accessibility of repair factors and improve repair efficiency (Figure 3). Interestingly, a recent report also revealed a role for the ATR kinase in nuclear mechanics. Kidiyoor et al. showed that ATR ensures coupling of the nuclear envelope to the cytoskeleton, and its depletion leads to changes in chromatin organisation, nuclear softening and reduction in nuclear circularity [152].
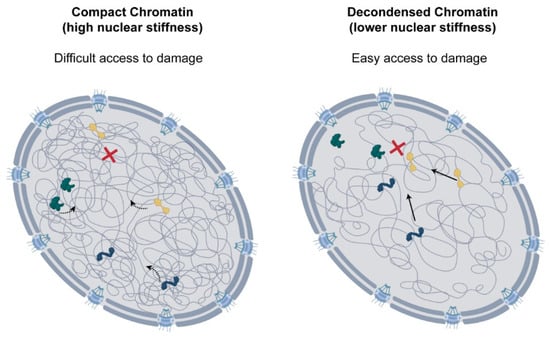
Figure 3.
Role of chromatin decondensation following DNA damage. Chromatin decondensation leads to higher nuclear diffusion, which could be important for rapid access of repair factors to DNA lesions. This could support the efficient repair of DNA damage.
The consequences of DDR-induced nuclear softening on mechanotransduction are still unclear. However, it is likely that mechanical changes to the organelle during repair will have repercussions for cell migration, force transduction to the cytoplasm and transcription regulation.
The mechanical properties of the cell and the nucleus prior to the damage, are also known to influence the extent of DNA damage that the cell suffers, as well as the outcome and efficiency of repair. Increased cytoskeletal stiffness can lead to higher levels of DNA damage in the cell [153], and this could be particularly important during confined migration. When cells migrate, especially through spaces smaller than their cross-sectional area, substantial compression forces are exerted on the nucleus. The nuclear deformation caused by these forces can in itself cause DNA damage, but it can also lead to rupture of the nuclear envelope, which, by exposing the nuclear contents to cytoplasmic factors, can cause severe genomic instability [154,155]. Furthermore, during migration, the deformation of the nuclear envelope, as well as its rupture, can cause local loss of repair factors and other mobile proteins and have severe implications to the efficiency of DDR pathways [156].
Both the cytoskeletal forces exerted on the nucleus, and nuclear stiffness, are important in determining nuclear deformability and the propensity of the nuclear envelope for rupture. As a result, the mechanical properties of the nucleus are tightly linked to the levels of DNA damage caused by migration and the outcome of DDR. Softer nuclei, with reduced levels of lamin A/C, require less pressure for nuclear envelope rupture to occur during migration and, therefore, display increased levels of DNA damage [154,156,157]. Similarly, in experiments using trichostatin A, nuclear softening caused by chromatin decompaction increased the likelihood of nuclear envelope rupture after shear stress application via syringe passes [61].
Although this work indicates that nuclear stiffening can have a protective role for the cell, cells expressing progerin, which increases nuclear stiffness, display higher levels of DNA damage; the mechanisms through which this occurs are not entirely clear, especially as progerin-expressing cells appear to require higher forces for nuclear envelope rupture [158]. However, this reinforces the idea of a link between altered nuclear mechanics and the incidence of DNA damage in the cell. Interestingly, cells treated with cisplatin display different levels of DNA damage, depending on the stiffness of their substrate. Using polyacrylamide gels of different stiffness, dos Santos et al. showed that on softer surfaces, where the nuclear tension is lower due to reduced cytoskeletal forces, cells have lower levels of DNA damage marker γH2AX following cisplatin treatment. This is also true for cells treated with blebbistatin prior to DNA damage induction [47]. The reasons for this are not yet clear, and much work is still needed in order to understand how the mechanical state of the nucleus relates to the propensity of the cells for DNA damage. Furthermore, how the pre-existing mechanical state of the nucleus might influence the DDR pathway choice is also not clear. As we discussed, changes in cytoskeletal forces and lamina composition affect nuclear stiffness and are also likely to affect chromatin organisation. Similarly, dysregulation of chromatin compaction due to, for example, mutations in chromatin remodelling complexes are often observed in cancer, but there are no data on how this may affect overall nuclear mechanics. It is perhaps possible that, as nuclear mechanics is reflective of the chromatin environment, nuclear stiffness might influence DDR pathway choice and, as a result, genomic stability. Understanding this relationship is important, particularly in therapeutics, since the mechanical properties of the cells and of the nucleus are often altered in disease, and a mechanically compromised nucleus could mean a different outcome in terms of drug efficacy.
4. Conclusions
The physical properties of the nucleus, including its shape and viscoelastic behaviour, are a result of the combined contribution of cytoskeletal forces, nuclear envelope composition, chromatin structure and its regulation by DNA-binding factors.
Cytoskeletal tension exerted on the nucleus directly affects nuclear stability and can alter nuclear envelope components, such as lamin A expression and turnover. Originally, the nuclear lamina was seen as the major contributor to nuclear rigidity and structure. Its composition and thickness directly correlate with nuclear stiffness and affect nuclear processes, including transcription and the spatial organisation of chromatin. Recently, chromatin has emerged as a key contributor to nuclear mechanics. We now know that chromatin behaves as a polymer gel that responds to mechanical stimuli and contributes to overall nuclear stiffness. Chromatin architecture is tightly regulated in cells but is often disrupted in diseases, such as cancer. This is usually accompanied by changes in nuclear morphology and genomic instability.
Cellular processes such as the DNA damage response and repair are known to cause changes in chromatin organisation, and this can alter the mechanical properties of the nucleus. Although changes to chromatin structure that occur during DDR lead to more efficient repair, which may drive chemotherapeutic resistance, it is still unclear how the resulting mechanical changes to the nucleus affect cellular mechanosensing, migration, and cell survival.
Overall, understanding how the mechanical properties of the nucleus affect cellular processes will lead to a better understanding of disease and ageing, along with the development of targeted therapeutic strategies.
Author Contributions
All authors have read and agreed to the published version of the manuscript.
Funding
UKRI-MRC (MR/M020606/1) to CPT.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Maniotis, A.J.; Chen, C.S.; Ingber, D.E. Demonstration of mechanical connections between integrins, cytoskeletal filaments, and nucleoplasm that stabilize nuclear structure. Proc. Natl. Acad. Sci. USA 1997, 94, 849. [Google Scholar] [CrossRef]
- Heo, S.-J.; Cosgrove, B.D.; Dai, E.N.; Mauck, R.L. Mechano-adaptation of the stem cell nucleus. Nucleus 2018, 9, 9–19. [Google Scholar] [CrossRef]
- Kirby, T.J.; Lammerding, J. Emerging views of the nucleus as a cellular mechanosensor. Nat. Cell Biol. 2018, 20, 373–381. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.-J.; Iwamoto, M.; Hiraoka, Y.; Haraguchi, T. Function of nuclear membrane proteins in shaping the nuclear envelope integrity during closed mitosis. J. Biochem. 2017, 161, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Crisp, M. Coupling of the nucleus and cytoplasm: Role of the LINC complex. J. Cell Biol. 2005, 172, 41–53. [Google Scholar] [CrossRef]
- Mammoto, A.; Mammoto, T.; Ingber, D.E. Mechanosensitive mechanisms in transcriptional regulation. J. Cell Sci. 2012, 125, 3061–3073. [Google Scholar] [CrossRef]
- Alam, S.G. The mammalian LINC complex regulates genome transcriptional responses to substrate rigidity. Sci. Rep. 2016, 6, 38063. [Google Scholar] [CrossRef]
- Leno, G.H. Regulation of DNA replication by the nuclear envelope. Semin. Cell Biol. 1992, 3, 237–243. [Google Scholar] [CrossRef]
- Wang, S. Mechanotransduction via the LINC complex regulates DNA replication in myonuclei. J. Cell Biol. 2018, 217, 2005–2018. [Google Scholar] [CrossRef] [PubMed]
- Klemm, S.L.; Shipony, Z.; Greenleaf, W.J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 2019, 20, 207–220. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.T.; Masukata, H. Regulation of DNA replication by chromatin structures: Accessibility and recruitment. Chromosoma 2011, 120, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Bellush, J.M.; Whitehouse, I. DNA replication through a chromatin environment. Philos. Trans. R. Soc. B Biol. Sci. 2017, 372, 20160287. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, K. Chromatin Accessibility Impacts Transcriptional Reprogramming in Oocytes. Cell Rep. 2018, 24, 304–311. [Google Scholar] [CrossRef]
- Dong, X.; Weng, Z. The correlation between histone modifications and gene expression. Epigenomics 2013, 5, 113–116. [Google Scholar] [CrossRef]
- House, N.C.M.; Koch, M.R.; Freudenreich, C.H. Chromatin modifications and DNA repair: Beyond double-strand breaks. Front. Genet. 2014, 5. [Google Scholar] [CrossRef]
- Hauer, M.H.; Gasser, S.M. Chromatin and nucleosome dynamics in DNA damage and repair. Genes Dev. 2017, 31, 2204–2221. [Google Scholar] [CrossRef] [PubMed]
- Saksouk, N.; Simboeck, E.; Déjardin, J. Constitutive heterochromatin formation and transcription in mammals. Epigenet. Chromatin 2015, 8, 3. [Google Scholar] [CrossRef] [PubMed]
- Pickersgill, H. Characterization of the Drosophila melanogaster genome at the nuclear lamina. Nat. Genet. 2006, 38, 1005–1014. [Google Scholar] [CrossRef]
- Van Steensel, B.; Belmont, A.S. Lamina-Associated Domains: Links with Chromosome Architecture, Heterochromatin, and Gene Repression. Cell 2017, 169, 780–791. [Google Scholar] [CrossRef]
- Solovei, I.; Thanisch, K.; Feodorova, Y. How to rule the nucleus: Divide et impera. Curr. Opin. Cell Biol. 2016, 40, 47–59. [Google Scholar] [CrossRef]
- Stevens, T.J. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 2017, 544, 59–64. [Google Scholar] [CrossRef]
- Pajerowski, J.D.; Dahl, K.N.; Zhong, F.L.; Sammak, P.J.; Discher, D.E. Physical plasticity of the nucleus in stem cell differentiation. Proc. Natl. Acad. Sci. USA 2007, 104, 15619. [Google Scholar] [CrossRef]
- Papanicolaou, G.N.; Traut, H.F. The Diagnostic Value of Vaginal Smears in Carcinoma of the Uterus. Am. J. Obstet. Gynecol. 1941, 42, 193–206. [Google Scholar] [CrossRef]
- Dunne, B.; Going, J.J. Scoring nuclear pleomorphism in breast cancer. Histopathology 2001, 39, 259–265. [Google Scholar] [CrossRef] [PubMed]
- Lombardi, M.L. The interaction between nesprins and sun proteins at the nuclear envelope is critical for force transmission between the nucleus and cytoskeleton. J. Biol. Chem. 2011, 286, 26743–26753. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, I. Targeted Ablation of Nesprin 1 and Nesprin 2 from Murine Myocardium Results in Cardiomyopathy, Altered Nuclear Morphology and Inhibition of the Biomechanical Gene Response. PLOS Genet. 2014, 10, e1004114. [Google Scholar] [CrossRef] [PubMed]
- Elosegui-Artola, A. Force Triggers YAP Nuclear Entry by Regulating Transport across Nuclear Pores. Cell 2017, 171, 1397–1410. [Google Scholar] [CrossRef]
- Calvo, F. Mechanotransduction and YAP-dependent matrix remodelling is required for the generation and maintenance of cancer-associated fibroblasts. Nat. Cell Biol. 2013, 15, 637–646. [Google Scholar] [CrossRef]
- Chaudhuri, O. Hydrogels with tunable stress relaxation regulate stem cell fate and activity. Nat. Mater. 2016, 15, 326–334. [Google Scholar] [CrossRef]
- Das, A.; Fischer, R.S.; Pan, D.; Waterman, C.M. YAP Nuclear Localization in the Absence of Cell-Cell Contact Is Mediated by a Filamentous Actin-dependent, Myosin II- and Phospho-YAP-independent Pathway during Extracellular Matrix Mechanosensing. J. Biol. Chem. 2016, 291, 6096–6110. [Google Scholar] [CrossRef]
- Driscoll, T.P.; Cosgrove, B.D.; Heo, S.J.; Shurden, Z.E.; Mauck, R.L. Cytoskeletal to Nuclear Strain Transfer Regulates YAP Signaling in Mesenchymal Stem Cells. Biophys. J. 2015, 108, 2783–2793. [Google Scholar] [CrossRef] [PubMed]
- Koushki, N. Lamin A redistribution mediated by nuclear deformation determines dynamic localization of YAP. bioRxiv 2020. [Google Scholar] [CrossRef]
- Wesolowska, N. Actin assembly ruptures the nuclear envelope by prying the lamina away from nuclear pores and nuclear membranes in starfish oocytes. eLife 2020, 9, e49774. [Google Scholar] [CrossRef] [PubMed]
- Hari-Gupta, Y. Nuclear myosin VI regulates the spatial organization of mammalian transcription initiation. bioRxiv 2020. [Google Scholar] [CrossRef]
- Théry, M. Micropatterning as a tool to decipher cell morphogenesis and functions. J. Cell Sci. 2010, 123, 4201. [Google Scholar] [CrossRef] [PubMed]
- Bautista, M.; Fernandez, A.; Pinaud, F. A Micropatterning Strategy to Study Nuclear Mechanotransduction in Cells. Micromachines 2019, 10, 810. [Google Scholar] [CrossRef]
- Charrier, E.E.; Pogoda, K.; Wells, R.G.; Janmey, P.A. Control of cell morphology and differentiation by substrates with independently tunable elasticity and viscous dissipation. Nat. Commun. 2018, 9, 449. [Google Scholar] [CrossRef]
- Vertelov, G. Rigidity of silicone substrates controls cell spreading and stem cell differentiation. Sci. Rep. 2016, 6, 33411. [Google Scholar] [CrossRef]
- Rehfeldt, F. Hyaluronic acid matrices show matrix stiffness in 2D and 3D dictates cytoskeletal order and myosin-II phosphorylation within stem cells. Integr. Biol. 2012, 4, 422–430. [Google Scholar] [CrossRef] [PubMed]
- Makhija, E.; Jokhun, D.S.; Shivashankar, G.V. Nuclear deformability and telomere dynamics are regulated by cell geometric constraints. Proc. Natl. Acad. Sci. USA 2016, 113, E32. [Google Scholar] [CrossRef]
- Seirin-Lee, S. Role of dynamic nuclear deformation on genomic architecture reorganization. PLoS Comput. Biol. 2019, 15, e1007289. [Google Scholar] [CrossRef]
- Hu, X. MKL1-actin pathway restricts chromatin accessibility and prevents mature pluripotency activation. Nat. Commun. 2019, 10, 1695. [Google Scholar] [CrossRef]
- Jain, N.; Iyer, K.V.; Kumar, A.; Shivashankar, G.V. Cell geometric constraints induce modular gene-expression patterns via redistribution of HDAC3 regulated by actomyosin contractility. Proc. Natl. Acad. Sci. USA 2013, 201300801. [Google Scholar] [CrossRef]
- Lherbette, M. Atomic Force Microscopy micro-rheology reveals large structural inhomogeneities in single cell-nuclei. Sci. Rep. 2017, 7, 8116. [Google Scholar] [CrossRef] [PubMed]
- Ferrera, D. Lamin B1 overexpression increases nuclear rigidity in autosomal dominant leukodystrophy fibroblasts. FASEB J. 2014, 28, 3906–3918. [Google Scholar] [CrossRef] [PubMed]
- Guilluy, C. Isolated nuclei adapt to force and reveal a mechanotransduction pathway in the nucleus. Nat. Cell Biol. 2014, 16, 376–381. [Google Scholar] [CrossRef]
- Dos Santos, Á. DNA damage alters nuclear mechanics through chromatin reorganization. Nucleic Acids Res. 2021, 49, 340–353. [Google Scholar] [CrossRef] [PubMed]
- Dittmer, T.A.; Misteli, T. The lamin protein family. Genome Biol. 2011, 12, 222. [Google Scholar] [CrossRef]
- Aebi, U.; Cohn, J.; Buhle, L.; Gerace, L. The nuclear lamina is a meshwork of intermediate-type filaments. Nature 1986, 323, 560–564. [Google Scholar] [CrossRef] [PubMed]
- Adam, S.A.; Butin-Israeli, V.; Cleland, M.M.; Shimi, T.; Goldman, R.D. Disruption of lamin B1 and lamin B2 processing and localization by farnesyltransferase inhibitors. Nucleus 2013, 4, 142–150. [Google Scholar] [CrossRef] [PubMed]
- Sinensky, M. The processing pathway of prelamin A. J. Cell Sci. 1994, 107, 61. [Google Scholar]
- Rober, R.A.; Sauter, H.; Weber, K.; Osborn, M. Cells of the cellular immune and hemopoietic system of the mouse lack lamins A/C: Distinction versus other somatic cells. J. Cell Sci. 1990, 95, 587. [Google Scholar]
- Chen, N.Y. An absence of lamin B1 in migrating neurons causes nuclear membrane ruptures and cell death. Proc. Natl. Acad. Sci. USA 2019, 116, 25870. [Google Scholar] [CrossRef]
- Coffinier, C. Deficiencies in lamin B1 and lamin B2 cause neurodevelopmental defects and distinct nuclear shape abnormalities in neurons. Mol. Biol. Cell 2011, 22, 4683–4693. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, K.H.; Kennedy, B.K. When lamins go bad: Nuclear structure and disease. Cell 2013, 152, 1365–1375. [Google Scholar] [CrossRef]
- Lammerding, J. Lamin A/C deficiency causes defective nuclear mechanics and mechanotransduction. J. Clin. Investig. 2004, 113, 370–378. [Google Scholar] [CrossRef]
- Swift, J. Nuclear Lamin-A Scales with Tissue Stiffness and Enhances Matrix-Directed Differentiation. Science 2013, 341, 1240104. [Google Scholar] [CrossRef]
- Buxboim, A. Matrix Elasticity Regulates Lamin-A,C Phosphorylation and Turnover with Feedback to Actomyosin. Curr. Biol. 2014, 24, 1909–1917. [Google Scholar] [CrossRef] [PubMed]
- Mu, X. Cytoskeleton stiffness regulates cellular senescence and innate immune response in Hutchinson–Gilford Progeria Syndrome. Aging Cell 2020, 19, e13152. [Google Scholar] [CrossRef]
- Larrieu, D.; Britton, S.; Demir, M.; Rodriguez, R.; Jackson, S.P. Chemical inhibition of NAT10 corrects defects of laminopathic cells. Science 2014, 344, 527–532. [Google Scholar] [CrossRef]
- Stephens, A.D. Chromatin histone modifications and rigidity affect nuclear morphology independent of lamins. Mol. Biol. Cell 2018, 29, 220–233. [Google Scholar] [CrossRef]
- Zheng, X. Lamins Organize the Global Three-Dimensional Genome from the Nuclear Periphery. Mol. Cell 2018, 71, 802–815.e807. [Google Scholar] [CrossRef]
- Chang, L. Nuclear peripheral chromatin-lamin B1 interaction is required for global integrity of chromatin architecture and dynamics in human cells. Protein Cell 2020. [Google Scholar] [CrossRef]
- Nora, E.P. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 2012, 485, 381–385. [Google Scholar] [CrossRef]
- Dixon, J.R. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 2012, 485, 376–380. [Google Scholar] [CrossRef]
- Dahl, K.N.; Engler, A.J.; Pajerowski, J.D.; Discher, D.E. Power-Law Rheology of Isolated Nuclei with Deformation Mapping of Nuclear Substructures. Biophys. J. 2005, 89, 2855–2864. [Google Scholar] [CrossRef]
- Mazumder, A.; Roopa, T.; Basu, A.; Mahadevan, L.; Shivashankar, G.V. Dynamics of Chromatin Decondensation Reveals the Structural Integrity of a Mechanically Prestressed Nucleus. Biophys. J. 2008, 95, 3028–3035. [Google Scholar] [CrossRef]
- Strickfaden, H. Condensed Chromatin Behaves like a Solid on the Mesoscale In Vitro and in Living Cells. Cell 2020, 183, 1772–1784.e1713. [Google Scholar] [CrossRef] [PubMed]
- Chalut, K.J. Chromatin Decondensation and Nuclear Softening Accompany Nanog Downregulation in Embryonic Stem Cells. Biophys. J. 2012, 103, 2060–2070. [Google Scholar] [CrossRef]
- Stephens, A.D.; Banigan, E.J.; Adam, S.A.; Goldman, R.D.; Marko, J.F. Chromatin and lamin A determine two different mechanical response regimes of the cell nucleus. Mol. Biol. Cell 2017, 28, 1984–1996. [Google Scholar] [CrossRef]
- Furusawa, T. Chromatin decompaction by the nucleosomal binding protein HMGN5 impairs nuclear sturdiness. Nat. Commun. 2015, 6, 6138. [Google Scholar] [CrossRef] [PubMed]
- Shimamoto, Y.; Tamura, S.; Masumoto, H.; Maeshima, K. Nucleosome–nucleosome interactions via histone tails and linker DNA regulate nuclear rigidity. Mol. Biol. Cell 2017, 28, 1580–1589. [Google Scholar] [CrossRef]
- Nelsen, E. Combined Atomic Force Microscope and Volumetric Light Sheet System for Correlative Force and Fluorescence Mechanobiology Studies. Sci. Rep. 2020, 10, 8133. [Google Scholar] [CrossRef] [PubMed]
- Dinant, C.; Houtsmuller, A.B.; Vermeulen, W. Chromatin structure and DNA damage repair. Epigenet. Chromatin 2008, 1, 9. [Google Scholar] [CrossRef]
- Belaghzal, H. Liquid chromatin Hi-C characterizes compartment-dependent chromatin interaction dynamics. Nat. Genet. 2021. [Google Scholar] [CrossRef]
- Ashwin, S.S.; Nozaki, T.; Maeshima, K.; Sasai, M. Organization of fast and slow chromatin revealed by single-nucleosome dynamics. Proc. Natl. Acad. Sci. USA 2019, 116, 19939. [Google Scholar] [CrossRef]
- Nozaki, T. Dynamic Organization of Chromatin Domains Revealed by Super-Resolution Live-Cell Imaging. Mol. Cell 2017, 67, 282–293.e287. [Google Scholar] [CrossRef]
- Wendt, K.S. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 2008, 451, 796–801. [Google Scholar] [CrossRef]
- Vietri Rudan, M.; Hadjur, S. Genetic Tailors: CTCF and Cohesin Shape the Genome During Evolution. Trends Genet. 2015, 31, 651–660. [Google Scholar] [CrossRef]
- Phillips-Cremins, J.E. Architectural Protein Subclasses Shape 3D Organization of Genomes during Lineage Commitment. Cell 2013, 153, 1281–1295. [Google Scholar] [CrossRef]
- Sofueva, S. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 2013, 32, 3119–3129. [Google Scholar] [CrossRef] [PubMed]
- Bannister, A.J. Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 2001, 410, 120–124. [Google Scholar] [CrossRef]
- Canzio, D. Chromodomain-mediated oligomerization of HP1 suggests a nucleosome-bridging mechanism for heterochromatin assembly. Mol. Cell 2011, 41, 67–81. [Google Scholar] [CrossRef]
- Strom, A.R. Phase separation drives heterochromatin domain formation. Nature 2017, 547, 241–245. [Google Scholar] [CrossRef]
- Larson, A.G. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 2017, 547, 236–240. [Google Scholar] [CrossRef]
- Strom, A.R. HP1α is a chromatin crosslinker that controls nuclear and mitotic chromosome mechanics. bioRxiv 2020. [Google Scholar] [CrossRef]
- Sabari, B.R. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361, eaar3958. [Google Scholar] [CrossRef] [PubMed]
- Cisse, I.I. Real-Time Dynamics of RNA Polymerase II Clustering in Live Human Cells. Science 2013, 341, 664. [Google Scholar] [CrossRef]
- Nagashima, R. Single nucleosome imaging reveals loose genome chromatin networks via active RNA polymerase II. J. Cell Biol. 2019, 218, 1511–1530. [Google Scholar] [CrossRef]
- Ashwin, S.S.; Maeshima, K.; Sasai, M. Heterogeneous fluid-like movements of chromatin and their implications to transcription. Biophys. Rev. 2020, 12, 461–468. [Google Scholar] [CrossRef]
- Wei, M. Nuclear actin regulates inducible transcription by enhancing RNA polymerase II clustering. Sci. Adv. 2020, 6, eaay6515. [Google Scholar] [CrossRef]
- Baarlink, C. A transient pool of nuclear F-actin at mitotic exit controls chromatin organization. Nat. Cell Biol. 2017, 19, 1389–1399. [Google Scholar] [CrossRef]
- Lamm, N. Nuclear F-actin counteracts nuclear deformation and promotes fork repair during replication stress. Nat. Cell Biol. 2020, 22, 1460–1470. [Google Scholar] [CrossRef] [PubMed]
- Caridi, C.P.; Plessner, M.; Grosse, R.; Chiolo, I. Nuclear actin filaments in DNA repair dynamics. Nat. Cell Biol. 2019, 21, 1068–1077. [Google Scholar] [CrossRef]
- Kapoor, P.; Shen, X. Mechanisms of nuclear actin in chromatin-remodeling complexes. Trends Cell Biol. 2014, 24, 238–246. [Google Scholar] [CrossRef]
- Belin, B.J.; Lee, T.; Mullins, R.D. DNA damage induces nuclear actin filament assembly by Formin-2 and Spire-1/2 that promotes efficient DNA repair. eLife 2015, 4, e07735. [Google Scholar] [CrossRef] [PubMed]
- Andrin, C. A requirement for polymerized actin in DNA double-strand break repair. Nucleus 2012, 3, 384–395. [Google Scholar] [CrossRef]
- Belin, B.J.; Cimini, B.A.; Blackburn, E.H.; Mullins, R.D. Visualization of actin filaments and monomers in somatic cell nuclei. Mol. Biol. Cell 2013, 24, 982–994. [Google Scholar] [CrossRef] [PubMed]
- Schrank, B.R. Nuclear ARP2/3 drives DNA break clustering for homology-directed repair. Nature 2018, 559, 61–66. [Google Scholar] [CrossRef]
- Spichal, M. Evidence for a dual role of actin in regulating chromosome organization and dynamics in yeast. J. Cell Sci. 2016, 129, 681–692. [Google Scholar] [CrossRef]
- Vicente-Manzanares, M.; Ma, X.; Adelstein, R.S.; Horwitz, A.R. Non-muscle myosin II takes centre stage in cell adhesion and migration. Nat. Rev. Mol. Cell Biol. 2009, 10, 778–790. [Google Scholar] [CrossRef]
- Fili, N.; Toseland, C.P. Unconventional Myosins: How Regulation Meets Function. Int. J. Mol. Sci. 2019, 21, 67. [Google Scholar] [CrossRef] [PubMed]
- Titus, M.A. Myosin-Driven Intracellular Transport. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Nambiar, R.; McConnell, R.E.; Tyska, M.J. Control of cell membrane tension by myosin-I. Proc. Natl. Acad. Sci. USA 2009, 106, 11972. [Google Scholar] [CrossRef]
- Vreugde, S. Nuclear myosin VI enhances RNA polymerase II-dependent transcription. Mol. Cell 2006, 23, 749–755. [Google Scholar] [CrossRef] [PubMed]
- Fili, N. NDP52 activates nuclear myosin VI to enhance RNA polymerase II transcription. Nat. Commun. 2017, 8, 1871. [Google Scholar] [CrossRef]
- Fili, N. Competition between two high- and low-affinity protein-binding sites in myosin VI controls its cellular function. J. Biol. Chem. 2020, 295, 337–347. [Google Scholar] [CrossRef]
- Zorca, C.E. Myosin VI regulates gene pairing and transcriptional pause release in T cells. Proc. Natl. Acad. Sci. USA 2015, 112, E1587. [Google Scholar] [CrossRef]
- Große-Berkenbusch, A. Myosin VI moves on nuclear actin filaments and supports long-range chromatin rearrangements. bioRxiv 2020. [Google Scholar] [CrossRef]
- Percipalle, P. The chromatin remodelling complex WSTF–SNF2h interacts with nuclear myosin 1 and has a role in RNA polymerase I transcription. EMBO Rep. 2006, 7, 525–530. [Google Scholar] [CrossRef] [PubMed]
- Sarshad, A. Nuclear Myosin 1c Facilitates the Chromatin Modifications Required to Activate rRNA Gene Transcription and Cell Cycle Progression. PLOS Genet. 2013, 9, e1003397. [Google Scholar] [CrossRef]
- Venit, T. Nuclear myosin 1 activates p21 gene transcription in response to DNA damage through a chromatin-based mechanism. Commun. Biol. 2020, 3, 115. [Google Scholar] [CrossRef]
- Kulashreshtha, M.; Mehta, I.S.; Kumar, P.; Rao, B.J. Chromosome territory relocation during DNA repair requires nuclear myosin 1 recruitment to chromatin mediated by ϒ-H2AX signaling. Nucleic Acids Res. 2016, 44, 8272–8291. [Google Scholar] [CrossRef]
- Simon, D.N.; Zastrow, M.S.; Wilson, K.L. Direct actin binding to A- and B-type lamin tails and actin filament bundling by the lamin A tail. Nucleus 2010, 1, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Mehta, I.S.; Elcock, L.S.; Amira, M.; Kill, I.R.; Bridger, J.M. Nuclear motors and nuclear structures containing A-type lamins and emerin: Is there a functional link? Biochem. Soc. Trans. 2008, 36, 1384–1388. [Google Scholar] [CrossRef]
- Holaska, J.M.; Wilson, K.L. An emerin “proteome”: Purification of distinct emerin-containing complexes from HeLa cells suggests molecular basis for diverse roles including gene regulation, mRNA splicing, signaling, mechanosensing, and nuclear architecture. Biochemistry 2007, 46, 8897–8908. [Google Scholar] [CrossRef]
- Cook, A.W.; Gough, R.E.; Toseland, C.P. Nuclear myosins—Roles for molecular transporters and anchors. J. Cell Sci. 2020, 133. [Google Scholar] [CrossRef]
- Cook, A.W.; Toseland, C.P. The roles of nuclear myosin in the DNA damage response. J. Biochem. 2020. [Google Scholar] [CrossRef]
- Viguera, E.; Canceill, D.; Ehrlich, S.D. Replication slippage involves DNA polymerase pausing and dissociation. EMBO J. 2001, 20, 2587–2595. [Google Scholar] [CrossRef]
- Loeb, L.A.; Monnat, R.J. DNA polymerases and human disease. Nat. Rev. Genet. 2008, 9, 594–604. [Google Scholar] [CrossRef]
- Deweese, J.E.; Osheroff, N. The DNA cleavage reaction of topoisomerase II: Wolf in sheep’s clothing. Nucleic Acids Res. 2009, 37, 738–748. [Google Scholar] [CrossRef]
- Cadet, J.; Wagner, J.R. DNA base damage by reactive oxygen species, oxidizing agents, and UV radiation. Cold Spring Harb. Perspect. Biol. 2013, 5, a012559. [Google Scholar] [CrossRef] [PubMed]
- Maréchal, A.; Zou, L. DNA damage sensing by the ATM and ATR kinases. Cold Spring Harb. Perspect. Biol. 2013, 5. [Google Scholar] [CrossRef]
- Smerdon, M.J.; Lieberman, M.W. Nucleosome rearrangement in human chromatin during UV-induced DNA- reapir synthesis. Proc. Natl. Acad. Sci. USA 1978, 75, 4238. [Google Scholar] [CrossRef]
- Cleaver, J.E. Nucleosome structure controls rates of excision repair in DNA of human cells. Nature 1977, 270, 451–453. [Google Scholar] [CrossRef]
- Adam, S.; Dabin, J.; Polo, S.E. Chromatin plasticity in response to DNA damage: The shape of things to come. DNA Repair 2015, 32, 120–126. [Google Scholar] [CrossRef]
- Hauer, M.H. Histone degradation in response to DNA damage enhances chromatin dynamics and recombination rates. Nat. Struct. Mol. Biol. 2017, 24, 99–107. [Google Scholar] [CrossRef]
- Lou, J. Phasor histone FLIM-FRET microscopy quantifies spatiotemporal rearrangement of chromatin architecture during the DNA damage response. Proc. Natl. Acad. Sci. USA 2019, 116, 7323. [Google Scholar] [CrossRef]
- Kruhlak, M.J. Changes in chromatin structure and mobility in living cells at sites of DNA double-strand breaks. J. Cell Biol. 2006, 172, 823–834. [Google Scholar] [CrossRef] [PubMed]
- Murga, M. Global chromatin compaction limits the strength of the DNA damage response. J. Cell Biol. 2007, 178, 1101–1108. [Google Scholar] [CrossRef]
- Lu, H.; Saha, J.; Beckmann, P.J.; Hendrickson, E.A.; Davis, A.J. DNA-PKcs promotes chromatin decondensation to facilitate initiation of the DNA damage response. Nucleic Acids Res. 2019, 47, 9467–9479. [Google Scholar] [CrossRef] [PubMed]
- Strickfaden, H. Poly(ADP-ribosyl)ation-dependent Transient Chromatin Decondensation and Histone Displacement following Laser Microirradiation. J. Biol. Chem. 2016, 291, 1789–1802. [Google Scholar] [CrossRef]
- Ziv, Y. Chromatin relaxation in response to DNA double-strand breaks is modulated by a novel ATM- and KAP-1 dependent pathway. Nat. Cell Biol. 2006, 8, 870–876. [Google Scholar] [CrossRef]
- Clouaire, T. Comprehensive Mapping of Histone Modifications at DNA Double-Strand Breaks Deciphers Repair Pathway Chromatin Signatures. Mol. Cell 2018, 72, 250–262.e256. [Google Scholar] [CrossRef]
- Kalousi, A.; Soutoglou, E. Nuclear compartmentalization of DNA repair. Curr. Opin. Genet. Dev. 2016, 37, 148–157. [Google Scholar] [CrossRef]
- Scully, R.; Panday, A.; Elango, R.; Willis, N.A. DNA double-strand break repair-pathway choice in somatic mammalian cells. Nat. Rev. Mol. Cell Biol. 2019, 20, 698–714. [Google Scholar] [CrossRef]
- Aymard, F. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nat. Struct. Mol. Biol. 2014, 21, 366–374. [Google Scholar] [CrossRef]
- Pfister, S.X. SETD2-Dependent Histone H3K36 Trimethylation Is Required for Homologous Recombination Repair and Genome Stability. Cell Rep. 2014, 7, 2006–2018. [Google Scholar] [CrossRef] [PubMed]
- Lemaître, C. Nuclear position dictates DNA repair pathway choice. Genes Dev. 2014, 28, 2450–2463. [Google Scholar] [CrossRef]
- Jha, D.K.; Strahl, B.D. An RNA polymerase II-coupled function for histone H3K36 methylation in checkpoint activation and DSB repair. Nat. Commun. 2014, 5, 3965. [Google Scholar] [CrossRef]
- Pai, C.-C. A histone H3K36 chromatin switch coordinates DNA double-strand break repair pathway choice. Nat. Commun. 2014, 5, 4091. [Google Scholar] [CrossRef]
- Sun, Y. Histone H3 methylation links DNA damage detection to activation of the tumour suppressor Tip60. Nat. Cell Biol. 2009, 11, 1376–1382. [Google Scholar] [CrossRef]
- Baldeyron, C.; Soria, G.; Roche, D.; Cook, A.J.; Almouzni, G. HP1alpha recruitment to DNA damage by p150CAF-1 promotes homologous recombination repair. J. Cell Biol. 2011, 193, 81–95. [Google Scholar] [CrossRef]
- Lee, Y.H.; Kuo, C.Y.; Stark, J.M.; Shih, H.M.; Ann, D.K. HP1 promotes tumor suppressor BRCA1 functions during the DNA damage response. Nucleic Acids Res. 2013, 41, 5784–5798. [Google Scholar] [CrossRef]
- Alagoz, M. SETDB1, HP1 and SUV39 promote repositioning of 53BP1 to extend resection during homologous recombination in G2 cells. Nucleic Acids Res. 2015, 43, 7931–7944. [Google Scholar] [CrossRef]
- Chiolo, I. Double-Strand Breaks in Heterochromatin Move Outside of a Dynamic HP1a Domain to Complete Recombinational Repair. Cell 2011, 144, 732–744. [Google Scholar] [CrossRef]
- Ryu, T. Heterochromatic breaks move to the nuclear periphery to continue recombinational repair. Nat. Cell Biol. 2015, 17, 1401–1411. [Google Scholar] [CrossRef]
- Oza, P.; Jaspersen, S.L.; Miele, A.; Dekker, J.; Peterson, C.L. Mechanisms that regulate localization of a DNA double-strand break to the nuclear periphery. Genes Dev. 2009, 23, 912–927. [Google Scholar] [CrossRef]
- Horigome, C. SWR1 and INO80 Chromatin Remodelers Contribute to DNA Double-Strand Break Perinuclear Anchorage Site Choice. Mol. Cell 2014, 55, 626–639. [Google Scholar] [CrossRef]
- Khadaroo, B. The DNA damage response at eroded telomeres and tethering to the nuclear pore complex. Nat. Cell Biol. 2009, 11, 980–987. [Google Scholar] [CrossRef]
- Tsouroula, K. Temporal and Spatial Uncoupling of DNA Double Strand Break Repair Pathways within Mammalian Heterochromatin. Mol. Cell 2016, 63, 293–305. [Google Scholar] [CrossRef]
- Kidiyoor, G.R. ATR is essential for preservation of cell mechanics and nuclear integrity during interstitial migration. Nat. Commun. 2020, 11, 4828. [Google Scholar] [CrossRef] [PubMed]
- Hatch, E.M.; Hetzer, M.W. Nuclear envelope rupture is induced by actin-based nucleus confinement. J. Cell Biol. 2016, 215, 27–36. [Google Scholar] [CrossRef]
- Denais, C.M. Nuclear envelope rupture and repair during cancer cell migration. Science 2016, 352, 353–358. [Google Scholar] [CrossRef]
- Raab, M. ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death. Science 2016, 352, 359. [Google Scholar] [CrossRef]
- Irianto, J. Nuclear constriction segregates mobile nuclear proteins away from chromatin. Mol. Biol. Cell 2016, 27, 4011–4020. [Google Scholar] [CrossRef] [PubMed]
- Chen, N.Y. Fibroblasts lacking nuclear lamins do not have nuclear blebs or protrusions but nevertheless have frequent nuclear membrane ruptures. Proc. Natl. Acad. Sci. USA 2018, 115, 10100. [Google Scholar] [CrossRef]
- Dahl, K.N. Distinct structural and mechanical properties of the nuclear lamina in Hutchinson–Gilford progeria syndrome. Proc. Natl. Acad. Sci. USA 2006, 103, 10271. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).