Ameliorating Fibrotic Phenotypes of Keloid Dermal Fibroblasts through an Epidermal Growth Factor-Mediated Extracellular Matrix Remodeling
Abstract
1. Introduction
2. Results
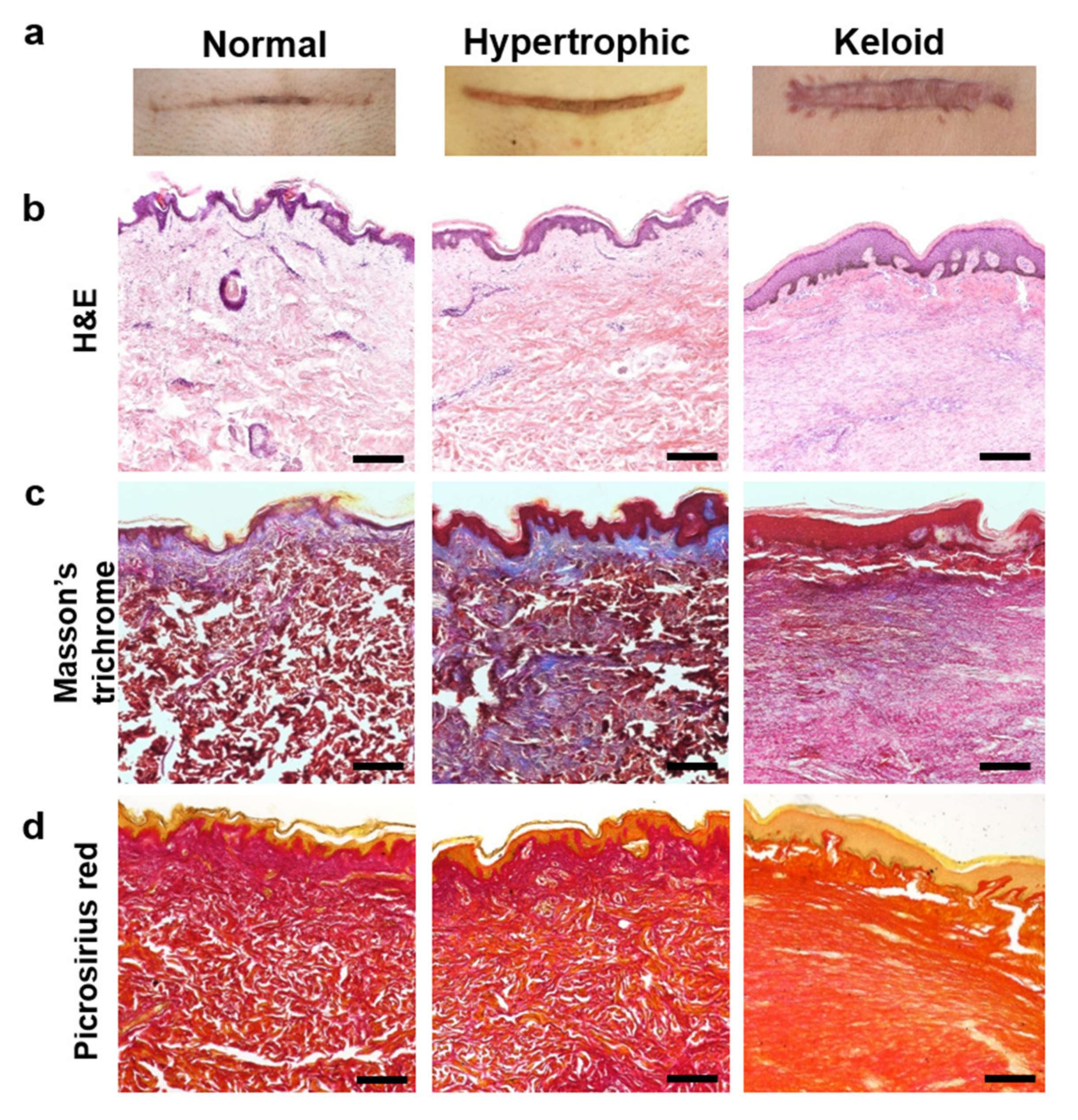
2.1. Histologic Evaluation of Normal, Hypertrophic, and Keloid (NHK) Scar Tissues
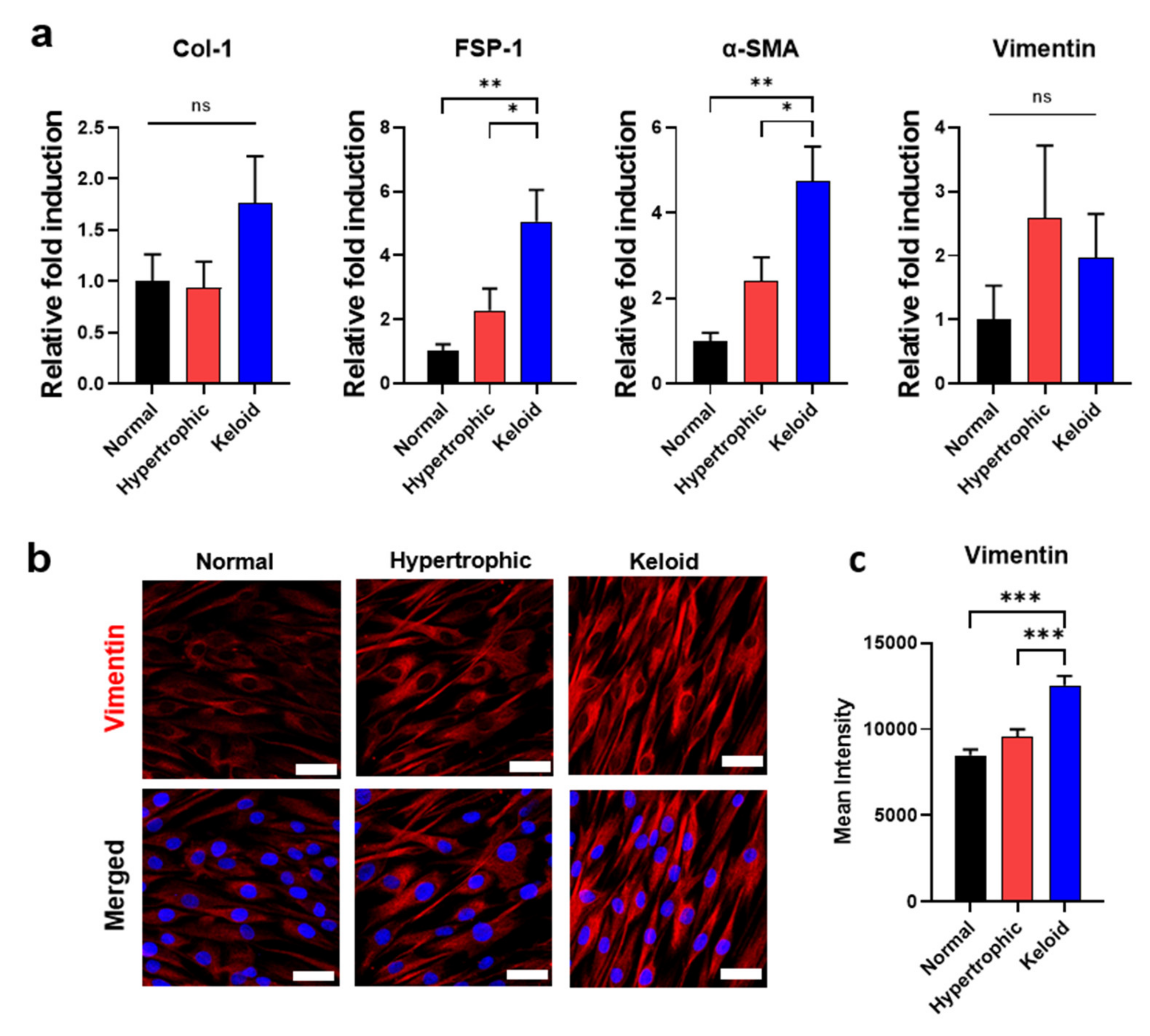
2.2. Isolation of Primary Dermal Fibroblasts from NHK Skin Tissues and Their Phenotypic Characterization
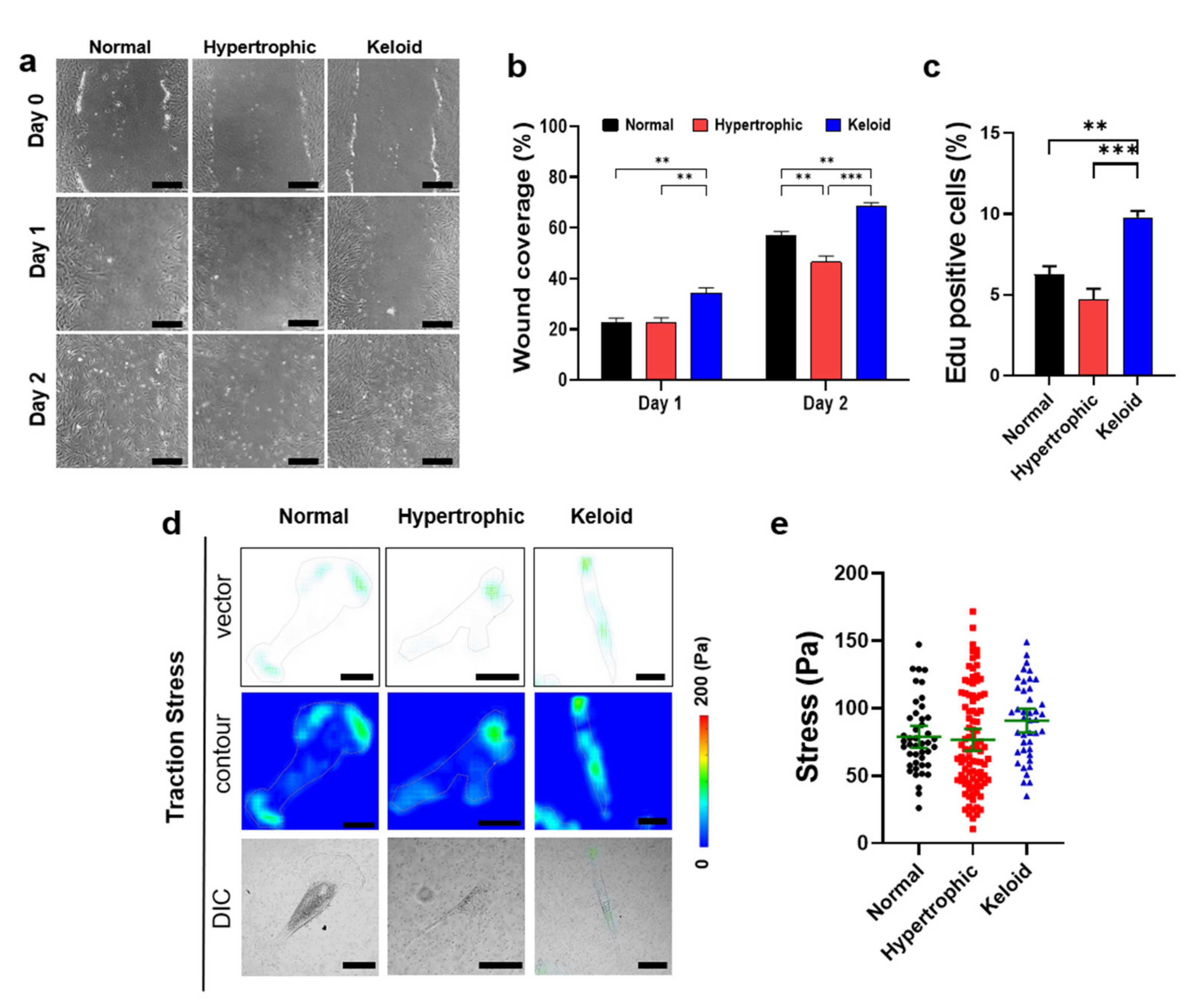
2.3. Cell–Matrix Interaction-Mediated Increases in the Cell Motility of Keloid Dermal Fibroblasts
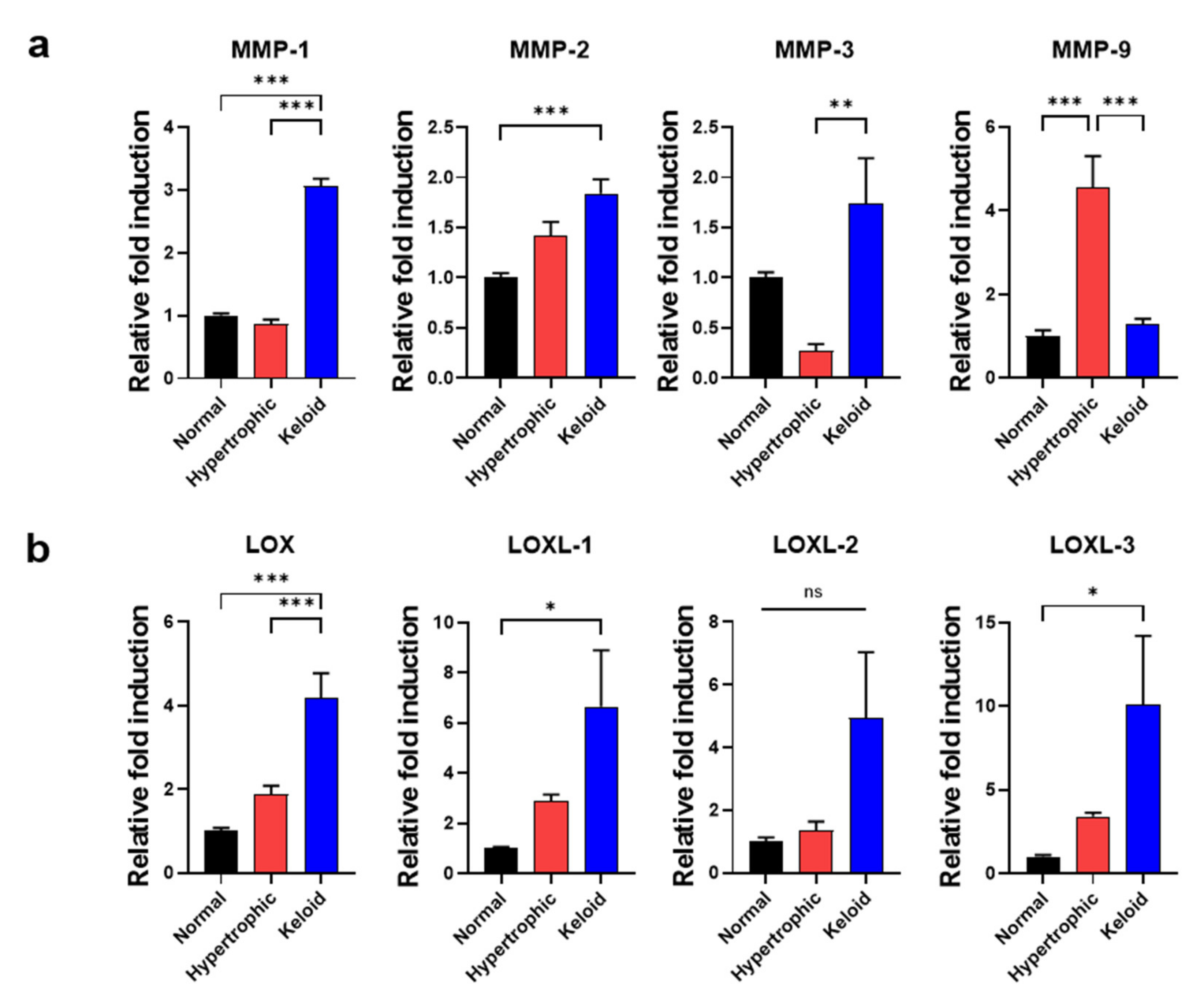
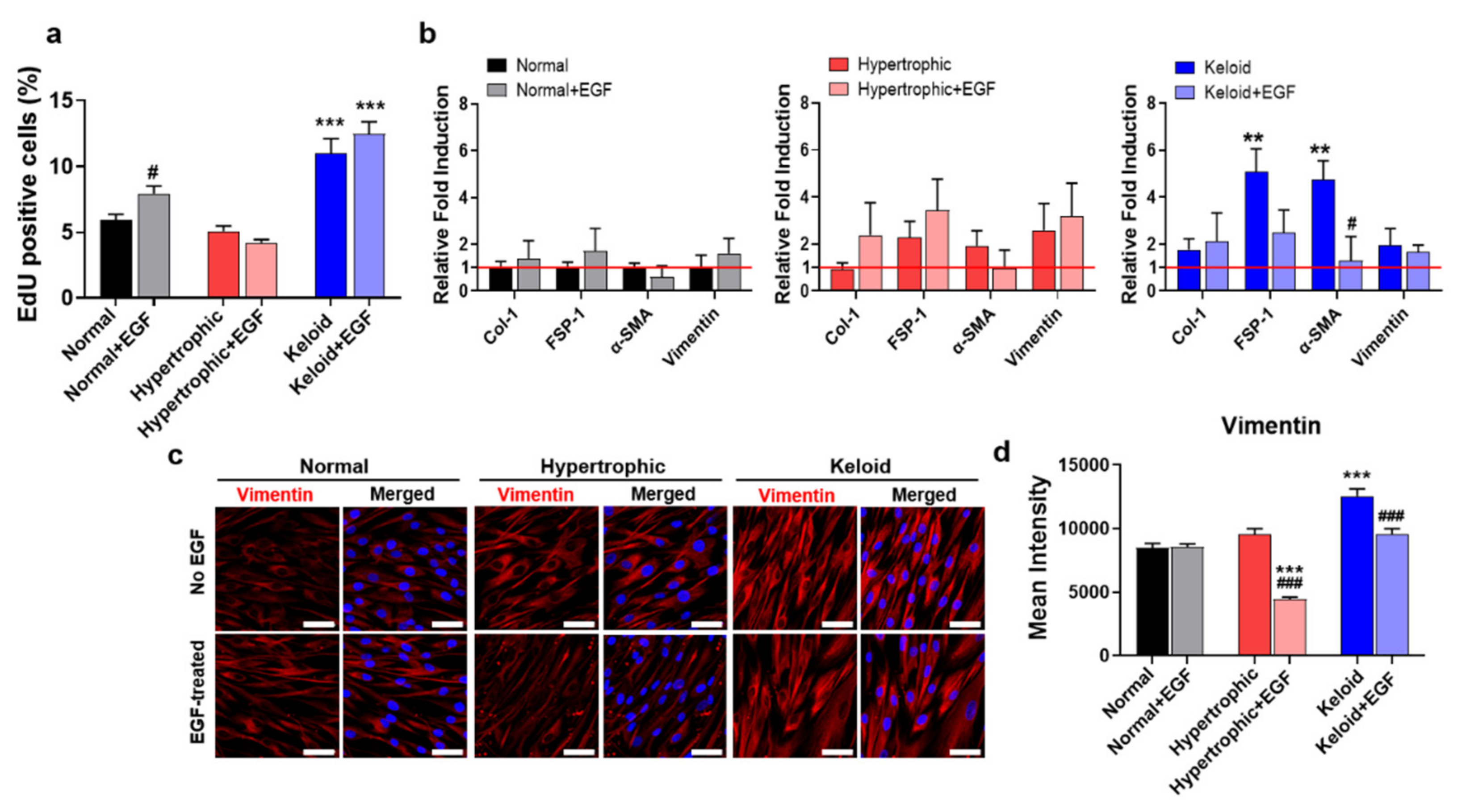
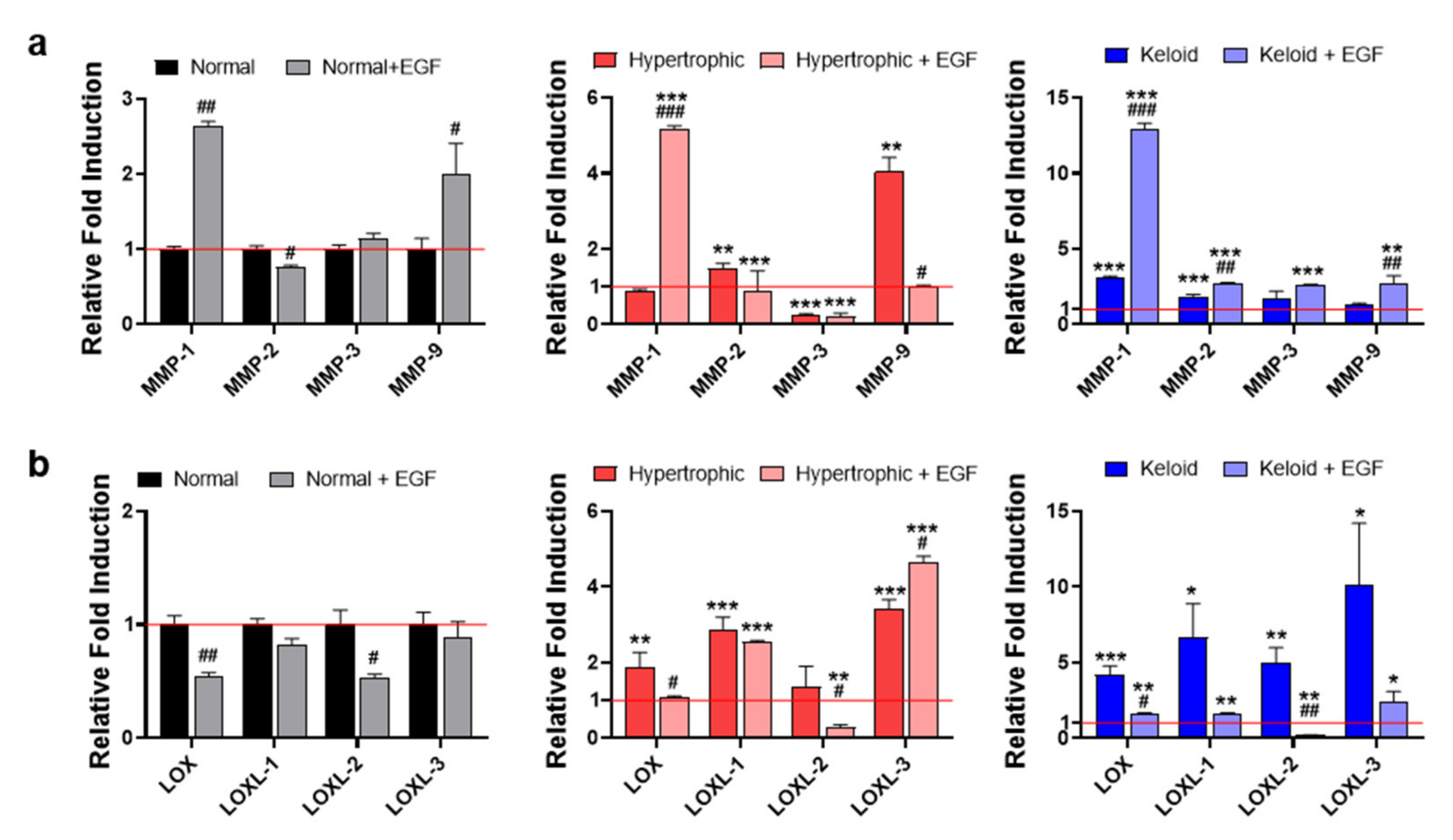
2.4. Exogenous EGF-Mediated Amelioration of Fibrotic Phenotypes of Keloid Dermal Fibroblasts through Modulation of ECM Remodeling
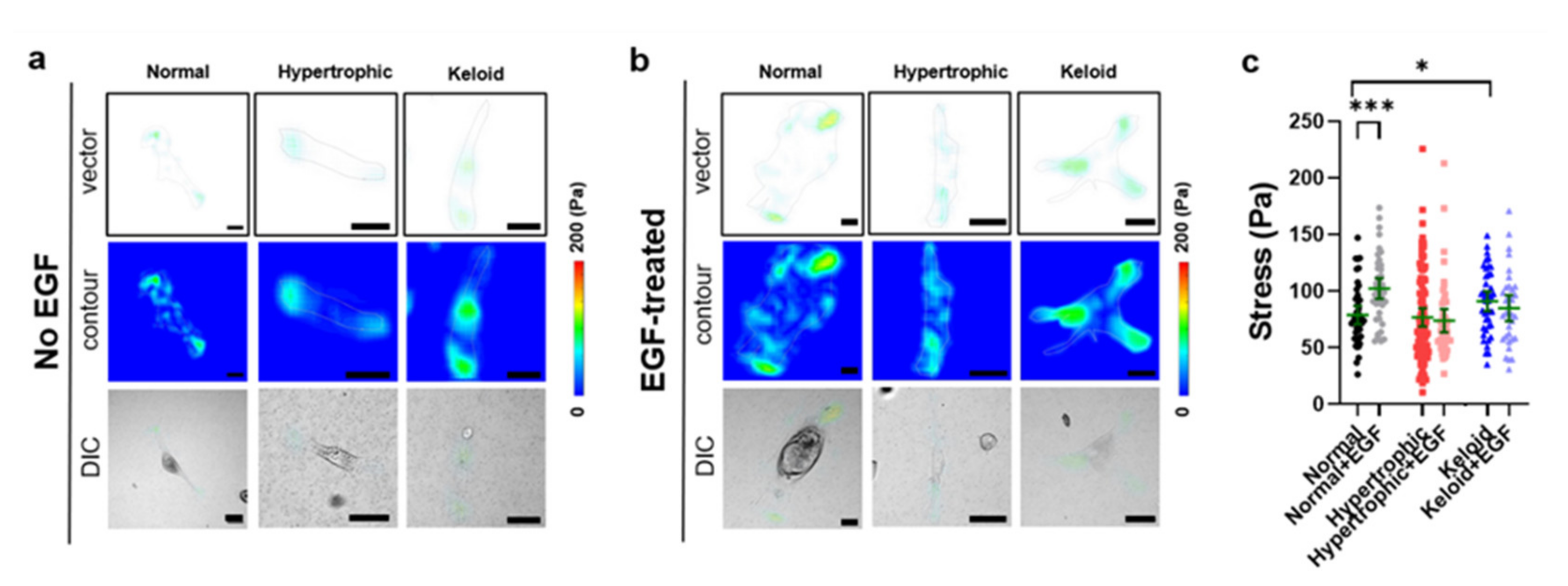
2.5. Exogenous EGF-Mediated Changes in Cell-Matrix Traction Force
3. Discussion
4. Materials and Methods
4.1. Patients and Sample Collection
4.2. Isolation of Dermal Fibroblasts and In Vitro Cell Culture
4.3. H&E and Masson’s Trichrome Staining
4.4. qPCR
4.5. Immunocytochemistry
4.6. Image Analysis
4.7. Cellular Motility
4.8. Cell Proliferation Assay
4.9. Preparation of PAA Gels
4.10. Cellular Traction Force Analysis
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AFM | atomic force microscopy |
| ANOVA | analysis of variance |
| α-SMA | alpha smooth muscle actin |
| BSA | bovine serum albumin |
| CI | confidence interval |
| Col | collagen |
| CTGF | connective tissue growth factor |
| DIC | differential interference contrast |
| DMEM | Dulbecco’s modified eagle’s medium |
| ECM | extracellular matrix |
| EDU | 5-ethynyl-2′-deoxyuridine |
| EGF | epidermal growth factor |
| FBS | fetal bovine serum |
| FEM | finite element method |
| FSP-1 | fibroblast specific protein 1 |
| GAPDH | glyceraldehyde-3-phosphate dehydrogenase |
| H&E | hematoxylin and eosin |
| IPF | idiopathic pulmonary fibrosis |
| LOX | lysyl oxidase |
| LOXL | lysyl oxidase-like |
| MMP | matrix metalloproteinase |
| PAA | polyacrylamide |
| PBS | phosphate buffered saline |
| PFA | paraformaldehyde |
| PIV | particle image velocimetry |
| SDS | sodium dodecyl sulfate |
| Sulfo-SANPAH | N-sulfosuccinimidyl-6-[4′-azido-2′-nitrophenylamino] hexanoate |
| TFM | traction force microscopy |
| TGF-β1 | transforming growth factor beta 1 |
| TBS | tris-buffered saline |
| VSS | Vancouver scar scale |
References
- Lee, W.J.; Lee, J.H.; Ahn, H.M.; Song, S.Y.; Kim, Y.O.; Lew, D.H.; Yun, C.O. Heat Shock Protein 90 Inhibitor Decreases Collagen Synthesis of Keloid Fibroblasts and Attenuates the Extracellular Matrix on the Keloid Spheroid Model. Plast. Reconstr. Surg. 2015, 136, 328e–337e. [Google Scholar] [CrossRef]
- Shin, J.U.; Lee, W.J.; Tran, T.N.; Jung, I.; Lee, J.H. Hsp70 Knockdown by siRNA Decreased Collagen Production in Keloid Fibroblasts. Yonsei Med. J. 2015, 56, 1619–1626. [Google Scholar] [CrossRef]
- Bran, G.M.; Goessler, U.R.; Hormann, K.; Riedel, F.; Sadick, H. Keloids: Current concepts of pathogenesis (review). Int. J. Mol. Med. 2009, 24, 283–293. [Google Scholar] [CrossRef]
- Gauglitz, G.G.; Korting, H.C.; Pavicic, T.; Ruzicka, T.; Jeschke, M.G. Hypertrophic scarring and keloids: Pathomechanisms and current and emerging treatment strategies. Mol. Med. 2011, 17, 113–125. [Google Scholar] [CrossRef]
- Ehrlich, H.P.; Desmouliere, A.; Diegelmann, R.F.; Cohen, I.K.; Compton, C.C.; Garner, W.L.; Kapanci, Y.; Gabbiani, G. Morphological and immunochemical differences between keloid and hypertrophic scar. Am. J. Pathol. 1994, 145, 105–113. [Google Scholar] [PubMed]
- Niessen, F.B.; Spauwen, P.H.; Schalkwijk, J.; Kon, M. On the nature of hypertrophic scars and keloids: A review. Plast. Reconstr. Surg. 1999, 104, 1435–1458. [Google Scholar] [CrossRef]
- Atiyeh, B.S.; Costagliola, M.; Hayek, S.N. Keloid or hypertrophic scar: The controversy: Review of the literature. Ann. Plast. Surg. 2005, 54, 676–680. [Google Scholar] [CrossRef]
- Al-Attar, A.; Mess, S.; Thomassen, J.M.; Kauffman, C.L.; Davison, S.P. Keloid pathogenesis and treatment. Plast. Reconstr. Surg. 2006, 117, 286–300. [Google Scholar] [CrossRef]
- Piera-Velazquez, S.; Jimenez, S.A. Molecular mechanisms of endothelial to mesenchymal cell transition (EndoMT) in experimentally induced fibrotic diseases. Fibrogenesis Tissue Repair 2012, 5, S7. [Google Scholar] [CrossRef] [PubMed]
- Pakyari, M.; Farrokhi, A.; Maharlooei, M.K.; Ghahary, A. Critical Role of Transforming Growth Factor Beta in Different Phases of Wound Healing. Adv. Wound Care 2013, 2, 215–224. [Google Scholar] [CrossRef]
- Shih, B.; Bayat, A. Genetics of keloid scarring. Arch. Derm. Res. 2010, 302, 319–339. [Google Scholar] [CrossRef] [PubMed]
- Xue, M.; Jackson, C.J. Extracellular Matrix Reorganization During Wound Healing and Its Impact on Abnormal Scarring. Adv. Wound Care 2015, 4, 119–136. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Kwan, J.Y.Y.; Yip, K.; Liu, P.P.; Liu, F.F. Targeting metabolic dysregulation for fibrosis therapy. Nat. Rev. Drug Discov. 2020, 19, 57–75. [Google Scholar] [CrossRef]
- Aumiller, V.; Strobel, B.; Romeike, M.; Schuler, M.; Stierstorfer, B.E.; Kreuz, S. Comparative analysis of lysyl oxidase (like) family members in pulmonary fibrosis. Sci. Rep. 2017, 7, 149. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Ren, J.; Loo, W.T.Y.; Hao, L.; Wang, M. Lysyl oxidases expression and histopathological changes of the diabetic rat nephron. Mol. Med. Rep. 2018, 17, 2431–2441. [Google Scholar] [CrossRef]
- Erasmus, M.; Samodien, E.; Lecour, S.; Cour, M.; Lorenzo, O.; Dludla, P.; Pheiffer, C.; Johnson, R. Linking LOXL2 to Cardiac Interstitial Fibrosis. Int. J. Mol. Sci. 2020, 21, 5913. [Google Scholar] [CrossRef]
- Szauter, K.M.; Cao, T.; Boyd, C.D.; Csiszar, K. Lysyl oxidase in development, aging and pathologies of the skin. Pathol. Biol. 2005, 53, 448–456. [Google Scholar] [CrossRef]
- Chadli, L.; Sotthewes, B.; Li, K.; Andersen, S.N.; Cahir-McFarland, E.; Cheung, M.; Cullen, P.; Dorjee, A.; de Vries-Bouwstra, J.K.; Huizinga, T.W.J.; et al. Identification of regulators of the myofibroblast phenotype of primary dermal fibroblasts from early diffuse systemic sclerosis patients. Sci. Rep. 2019, 9, 4521. [Google Scholar] [CrossRef]
- Barrientos, S.; Stojadinovic, O.; Golinko, M.S.; Brem, H.; Tomic-Canic, M. Growth factors and cytokines in wound healing. Wound Repair Regen. 2008, 16, 585–601. [Google Scholar] [CrossRef]
- Hu, Y.; Fu, J.; Liu, X.; Xue, X. ERK1/2 Signaling Pathway Activated by EGF Promotes Proliferation, Transdifferentiation, and Migration of Cultured Primary Newborn Rat Lung Fibroblasts. Biomed. Res. Int. 2020, 2020, 7176169. [Google Scholar] [CrossRef]
- Mimura, Y.; Ihn, H.; Jinnin, M.; Asano, Y.; Yamane, K.; Tamaki, K. Epidermal growth factor affects the synthesis and degradation of type I collagen in cultured human dermal fibroblasts. Matrix Biol. 2006, 25, 202–212. [Google Scholar] [CrossRef]
- Hur, S.S.; del Alamo, J.C.; Park, J.S.; Li, Y.S.; Nguyen, H.A.; Teng, D.; Wang, K.C.; Flores, L.; Alonso-Latorre, B.; Lasheras, J.C.; et al. Roles of cell confluency and fluid shear in 3-dimensional intracellular forces in endothelial cells. Proc. Natl. Acad. Sci. USA 2012, 109, 11110–11115. [Google Scholar] [CrossRef] [PubMed]
- Hur, S.S.; Jeong, J.H.; Ban, M.J.; Park, J.H.; Yoon, J.K.; Hwang, Y. Traction force microscopy for understanding cellular mechanotransduction. BMB Rep. 2020, 53, 74–81. [Google Scholar] [CrossRef] [PubMed]
- Hur, S.S.; Zhao, Y.; Li, Y.S.; Botvinick, E.; Chien, S. Live Cells Exert 3-Dimensional Traction Forces on Their Substrata. Cell Mol. Bioeng. 2009, 2, 425–436. [Google Scholar] [CrossRef] [PubMed]
- Limandjaja, G.C.; Niessen, F.B.; Scheper, R.J.; Gibbs, S. The Keloid Disorder: Heterogeneity, Histopathology, Mechanisms and Models. Front. Cell Dev. Biol. 2020, 8, 360. [Google Scholar] [CrossRef] [PubMed]
- Park, T.H.; Park, J.H.; Tirgan, M.H.; Halim, A.S.; Chang, C.H. Clinical implications of single- versus multiple-site keloid disorder: A retrospective study in an Asian population. Ann. Plast. Surg. 2015, 74, 248–251. [Google Scholar] [CrossRef]
- Ryu, Y.H.; Lee, Y.J.; Kim, K.J.; Lee, S.J.; Han, Y.N.; Rhie, J.W. Epidermal Growth Factor (EGF)-Like Repeats and Discoidin I-Like Domains 3 (EDIL3): A Potential New Therapeutic Tool for the Treatment of Keloid Scars. Tissue Eng. Regen. Med. 2017, 14, 267–277. [Google Scholar] [CrossRef]
- Kim, S.W.; Kim, K.J.; Rhie, J.W.; Ahn, S.T. Effects of adipose-derived stem cells on keloid fibroblasts based on paracrine function. Tissue Eng. Regen. Med. 2015, 12, 435–441. [Google Scholar] [CrossRef]
- Rinkevich, Y.; Walmsley, G.G.; Hu, M.S.; Maan, Z.N.; Newman, A.M.; Drukker, M.; Januszyk, M.; Krampitz, G.W.; Gurtner, G.C.; Lorenz, H.P.; et al. Skin fibrosis. Identification and isolation of a dermal lineage with intrinsic fibrogenic potential. Science 2015, 348, aaa2151. [Google Scholar] [CrossRef]
- Nakasaki, M.; Hwang, Y.; Xie, Y.; Kataria, S.; Gund, R.; Hajam, E.Y.; Samuel, R.; George, R.; Danda, D.; Paul, M.J.; et al. The matrix protein Fibulin-5 is at the interface of tissue stiffness and inflammation in fibrosis. Nat. Commun. 2015, 6, 8574. [Google Scholar] [CrossRef]
- Mendez, M.G.; Kojima, S.I.; Goldman, R.D. Vimentin induces changes in cell shape, motility, and adhesion during the epithelial to mesenchymal transition. FASEB J. 2010, 24, 1838–1851. [Google Scholar] [CrossRef]
- Hahn, J.M.; McFarland, K.L.; Combs, K.A.; Supp, D.M. Partial epithelial-mesenchymal transition in keloid scars: Regulation of keloid keratinocyte gene expression by transforming growth factor-beta 1. Burn. Trauma 2016, 4, 30. [Google Scholar] [CrossRef]
- Strutz, F.; Okada, H.; Lo, C.W.; Danoff, T.; Carone, R.L.; Tomaszewski, J.E.; Neilson, E.G. Identification and characterization of a fibroblast marker: FSP1. J. Cell. Biol. 1995, 130, 393–405. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.P.; Phan, T.T.; Lim, I.J.; Cao, X. Stat3 contributes to keloid pathogenesis via promoting collagen production, cell proliferation and migration. Oncogene 2006, 25, 5416–5425. [Google Scholar] [CrossRef]
- Achterberg, V.F.; Buscemi, L.; Diekmann, H.; Smith-Clerc, J.; Schwengler, H.; Meister, J.J.; Wenck, H.; Gallinat, S.; Hinz, B. The nano-scale mechanical properties of the extracellular matrix regulate dermal fibroblast function. J. Investig. Derm. 2014, 134, 1862–1872. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Kim, S.Y.; Mun, S.K.; Rhee, S.; Kim, B.J. Epidermal growth factor improves the migration and contractility of aged fibroblasts cultured on 3D collagen matrices. Int. J. Mol. Med. 2015, 35, 1017–1025. [Google Scholar] [CrossRef]
- Satish, L.; Babu, M.; Tran, K.T.; Hebda, P.A.; Wells, A. Keloid fibroblast responsiveness to epidermal growth factor and activation of downstream intracellular signaling pathways. Wound Repair Regen. 2004, 12, 183–192. [Google Scholar] [CrossRef]
- Limandjaja, G.C.; Belien, J.M.; Scheper, R.J.; Niessen, F.B.; Gibbs, S. Hypertrophic and keloid scars fail to progress from the CD34(-)/alpha-smooth muscle actin (alpha-SMA)(+) immature scar phenotype and show gradient differences in alpha-SMA and p16 expression. Br. J. Derm. 2020, 182, 974–986. [Google Scholar] [CrossRef] [PubMed]
- Schultz, G.S.; Davidson, J.M.; Kirsner, R.S.; Bornstein, P.; Herman, I.M. Dynamic reciprocity in the wound microenvironment. Wound Repair Regen. 2011, 19, 134–148. [Google Scholar] [CrossRef]
- McKleroy, W.; Lee, T.H.; Atabai, K. Always cleave up your mess: Targeting collagen degradation to treat tissue fibrosis. Am. J. Physiol. Lung Cell Mol. Physiol. 2013, 304, L709–L721. [Google Scholar] [CrossRef]
- Tanriverdi-Akhisaroglu, S.; Menderes, A.; Fau-Oktay, G.; Oktay, G. Matrix metalloproteinase-2 and -9 activities in human keloids, hypertrophic and atrophic scars: A pilot study. Cell Biochem. Funct. 2009, 27, 81–87. [Google Scholar] [CrossRef]
- Fujiwara, M.; Muragaki, Y.; Ooshima, A. Keloid-derived fibroblasts show increased secretion of factors involved in collagen turnover and depend on matrix metalloproteinase for migration. Br. J. Derm. 2005, 153, 295–300. [Google Scholar] [CrossRef]
- Caley, M.P.; Martins, V.L.; O’Toole, E.A. Metalloproteinases and Wound Healing. Adv. Wound Care 2015, 4, 225–234. [Google Scholar] [CrossRef] [PubMed]
- Simon, F.; Bergeron, D.; Larochelle, S.; Lopez-Valle, C.A.; Genest, H.; Armour, A.; Moulin, V.J. Enhanced secretion of TIMP-1 by human hypertrophic scar keratinocytes could contribute to fibrosis. Burns 2012, 38, 421–427. [Google Scholar] [CrossRef] [PubMed]
- Iimuro, Y.; Nishio, T.; Morimoto, T.; Nitta, T.; Stefanovic, B.; Choi, S.K.; Brenner, D.A.; Yamaoka, Y. Delivery of matrix metalloproteinase-1 attenuates established liver fibrosis in the rat. Gastroenterology 2003, 124, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Foronjy, R.F.; Sun, J.; Lemaitre, V.; D’Armiento, J.M. Transgenic expression of matrix metalloproteinase-1 inhibits myocardial fibrosis and prevents the transition to heart failure in a pressure overload mouse model. Hypertens. Res. 2008, 31, 725–735. [Google Scholar] [CrossRef]
- Kaar, J.L.; Li, Y.; Blair, H.C.; Asche, G.; Koepsel, R.R.; Huard, J.; Russell, A.J. Matrix metalloproteinase-1 treatment of muscle fibrosis. Acta Biomater. 2008, 4, 1411–1420. [Google Scholar] [CrossRef]
- Shiraha, H.; Gupta, K.; Drabik, K.; Wells, A. Aging fibroblasts present reduced epidermal growth factor (EGF) responsiveness due to preferential loss of EGF receptors. J. Biol. Chem. 2000, 275, 19343–19351. [Google Scholar] [CrossRef]
- Huebsch, N.; Arany, P.R.; Mao, A.S.; Shvartsman, D.; Ali, O.A.; Bencherif, S.A.; Rivera-Feliciano, J.; Mooney, D.J. Harnessing traction-mediated manipulation of the cell/matrix interface to control stem-cell fate. Nat. Mater. 2010, 9, 518–526. [Google Scholar] [CrossRef]
- Ayala, R.; Zhang, C.; Yang, D.; Hwang, Y.; Aung, A.; Shroff, S.S.; Arce, F.T.; Lal, R.; Arya, G.; Varghese, S. Engineering the cell-material interface for controlling stem cell adhesion, migration, and differentiation. Biomaterials 2011, 32, 3700–3711. [Google Scholar] [CrossRef]
- Harn, H.I.; Wang, Y.K.; Hsu, C.K.; Ho, Y.T.; Huang, Y.W.; Chiu, W.T.; Lin, H.H.; Cheng, C.M.; Tang, M.J. Mechanical coupling of cytoskeletal elasticity and force generation is crucial for understanding the migrating nature of keloid fibroblasts. Exp. Derm. 2015, 24, 579–584. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Hwang, Y.; Suk, S.; Lin, S.; Tierney, M.; Du, B.; Seo, T.; Mitchell, A.; Sacco, A.; Varghese, S. Directed in vitro myogenesis of human embryonic stem cells and their in vivo engraftment. PLoS ONE 2013, 8, e72023. [Google Scholar] [CrossRef]
- Hwang, Y.; Seo, T.; Hariri, S.; Choi, C.; Varghese, S. Matrix Topographical Cue-Mediated Myogenic Differentiation of Human Embryonic Stem Cell Derivatives. Polymers 2017, 9, 580. [Google Scholar] [CrossRef]
- Liang, C.C.; Park, A.Y.; Guan, J.L. In vitro scratch assay: A convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2007, 2, 329–333. [Google Scholar] [CrossRef]
- Tse, J.R.; Engler, A.J. Preparation of Hydrogel Substrates with Tunable Mechanical Properties. Curr. Protoc. Cell Biol. 2010, 47, 10.16.1–10.16.16. [Google Scholar] [CrossRef]
| Variable | Normal Skin Tissue | Hypertrophic Scar | Keloid Scar | Comparison (* p) |
|---|---|---|---|---|
| (n = 6) | (n = 7) | (n = 7) | ||
| Age (years) | 47 | 31 | 34 | 0.0805 |
| Gender | ||||
| Male | 6 | 5 | 7 | |
| Female | 0 | 2 | 0 | |
| Location of keloid | ||||
| Ear | 0 | 3 | 1 | |
| Extremity | 0 | 0 | 2 | |
| Trunk | 6 | 4 | 4 | |
| Ethnicity | Asian | Asian | Asian | |
| Vancouver Scar Scale | ||||
| Vascularity | 0 | 0.71 ± 1.11 | 2.29 ± 0.76 | 0.017 |
| Pigmentation | 0 | 1.14 ± 1.07 | 1.57 ± 0.98 | 0.535 |
| Pliability | 0 | 2.43 ± 0.079 | 4.14 ± 1.21 | 0.017 |
| Height | 0 | 2.00 ± 0.82 | 2.16 ± 0.69 | 0.805 |
| Total | 0 | 6.29 ± 1.704 | 10.00 ± 2.08 | 0.007 |
| Gene | Primer Sequence (5′ to 3′) |
|---|---|
| GAPDH | F-CAC TCC ACC TTT GAC GC |
| R-GGT CCA GGG GTC TTA CTC C | |
| Col-1 | F-CAA GAC AGT GAT TGA ATA CAA AAC CA |
| R-GGT CCA GGG GTC TTA CTC C | |
| FSP-1 | F-TCT TTC TTG GTT TGA TCC TGA CT |
| R-AGT TCT GAC TTG TTG AGC TTG A | |
| α-SMA | F-AAG CAC AGA GCA AAA GAG GAA T |
| R-ATG TCG TCC CAG TTG GTG AT | |
| Vimentin | F-AAT CCA AGT TTG CTG ACC TCT CTG A |
| R-ACT GCA CCT GTC TCC GGT ACT C | |
| MMP-1 | F-GGG GCT TTG ATG TAC CCT AGC |
| R-TGT CAC ACG CTT TTG GGG TTT | |
| MMP-2 | F-GAT ACC CCT TTG ACG GTA AGG A |
| R-CCT TCT CCC AAG GTC CAT AGC | |
| MMP-3 | F-CTG GAC TCC GAC ACT CTG GA |
| R-CAG GAA AGG TTC TGA AGT GAC C | |
| MMP-9 | F-GGG ACG CAG ACA TCG TCA TC |
| R-TCG TCA TCG TCG AAA TGG GC | |
| LOX | F-TTC CAG TAC GGT CTC CCA GA |
| R-TGG CCA GAC AGT TTT CCT CC | |
| LOXL-1 | F-GAG GCC ACC GAC TAC GAT GT |
| R-CTG TGG TAA TGC TGG TGG CAG | |
| LOXL-2 | F-GTA CAA GCC AGA GCA ACC CC |
| R-CCT GTG CAC TGG ATC TCG TT | |
| LOXL-3 | F-AAG CAA CAA CAG TCG AAG CC |
| R-TCC AGA GCA GCG AAC TTC AC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, H.; Anggradita, L.D.; Lee, S.-J.; Hur, S.S.; Bae, J.; Hwang, N.S.-Y.; Nam, S.M.; Hwang, Y. Ameliorating Fibrotic Phenotypes of Keloid Dermal Fibroblasts through an Epidermal Growth Factor-Mediated Extracellular Matrix Remodeling. Int. J. Mol. Sci. 2021, 22, 2198. https://doi.org/10.3390/ijms22042198
Kim H, Anggradita LD, Lee S-J, Hur SS, Bae J, Hwang NS-Y, Nam SM, Hwang Y. Ameliorating Fibrotic Phenotypes of Keloid Dermal Fibroblasts through an Epidermal Growth Factor-Mediated Extracellular Matrix Remodeling. International Journal of Molecular Sciences. 2021; 22(4):2198. https://doi.org/10.3390/ijms22042198
Chicago/Turabian StyleKim, Hyunbum, Laurensia Danis Anggradita, Sun-Jae Lee, Sung Sik Hur, Joonsuk Bae, Nathaniel Suk-Yeon Hwang, Seung Min Nam, and Yongsung Hwang. 2021. "Ameliorating Fibrotic Phenotypes of Keloid Dermal Fibroblasts through an Epidermal Growth Factor-Mediated Extracellular Matrix Remodeling" International Journal of Molecular Sciences 22, no. 4: 2198. https://doi.org/10.3390/ijms22042198
APA StyleKim, H., Anggradita, L. D., Lee, S.-J., Hur, S. S., Bae, J., Hwang, N. S.-Y., Nam, S. M., & Hwang, Y. (2021). Ameliorating Fibrotic Phenotypes of Keloid Dermal Fibroblasts through an Epidermal Growth Factor-Mediated Extracellular Matrix Remodeling. International Journal of Molecular Sciences, 22(4), 2198. https://doi.org/10.3390/ijms22042198








