Transcriptome-Wide Analysis of Stationary Phase Small ncRNAs in E. coli
Abstract
1. Introduction
2. Results
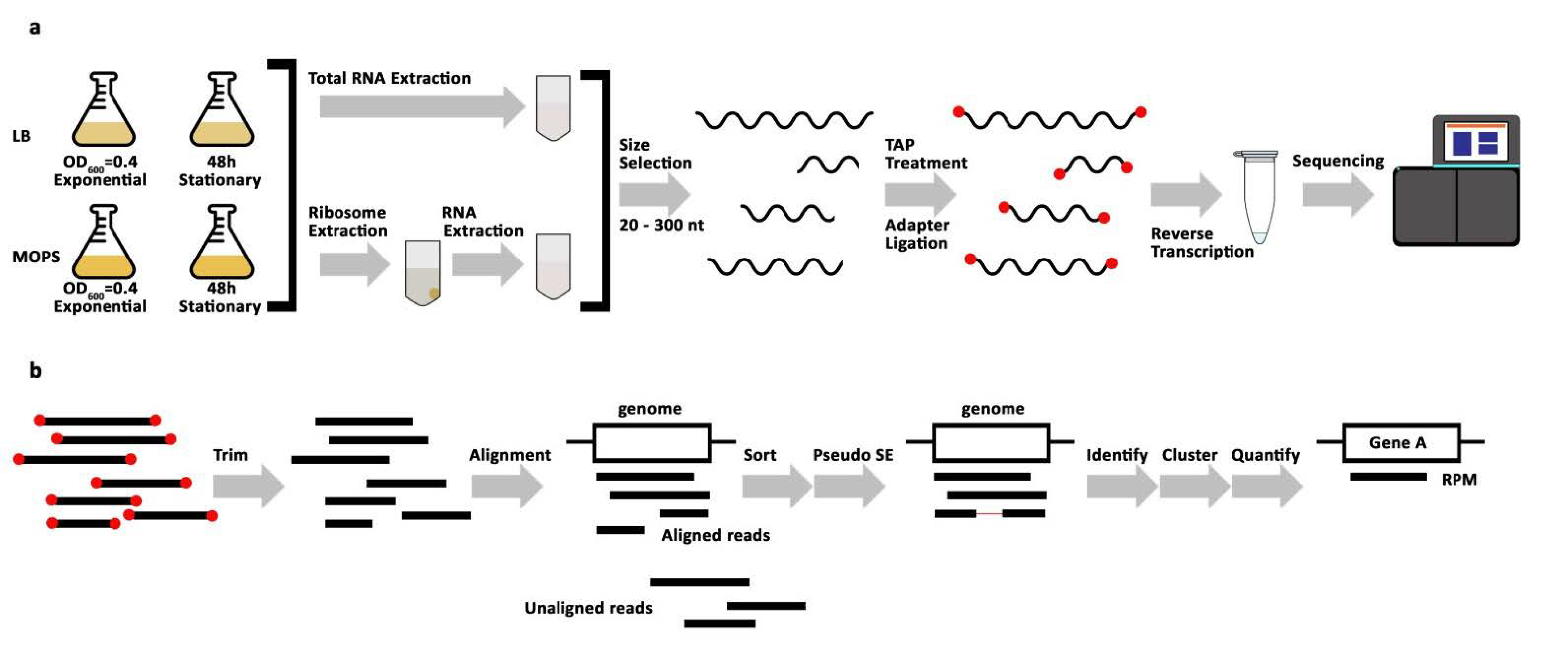
2.1. Library Preparation and Sample Collection
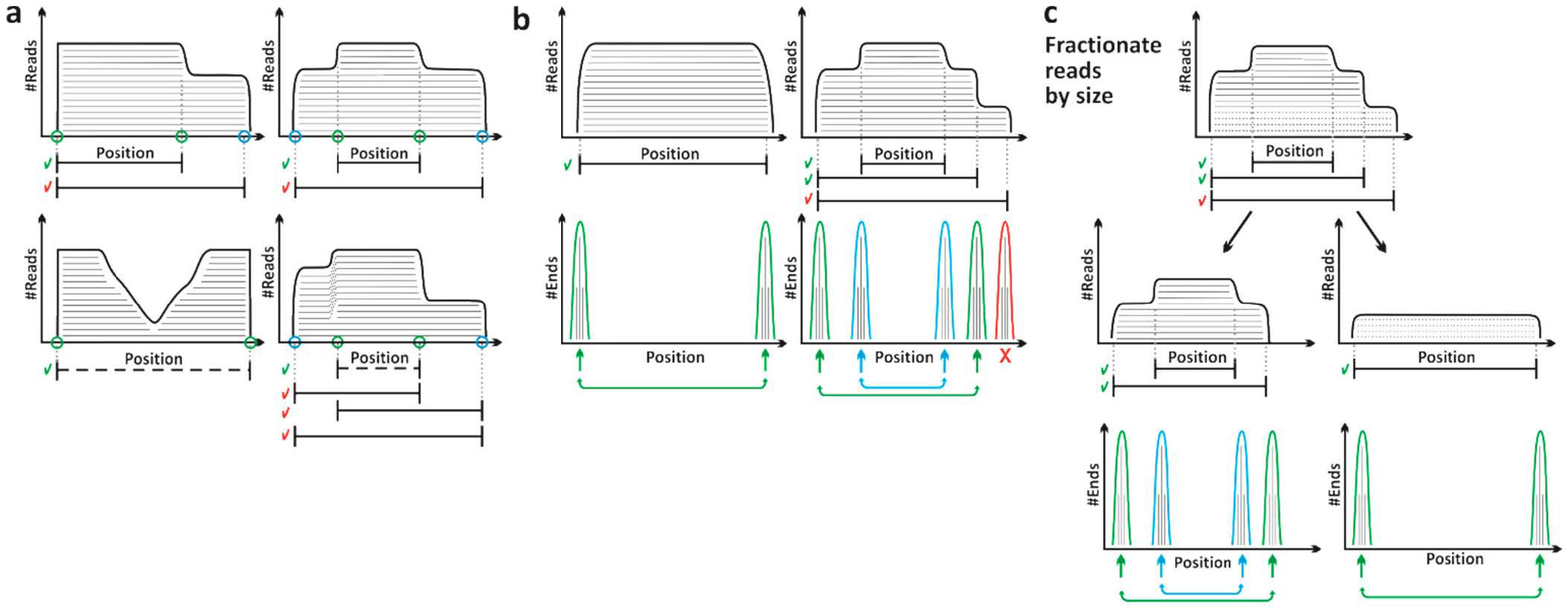
2.2. STARPA Workflow
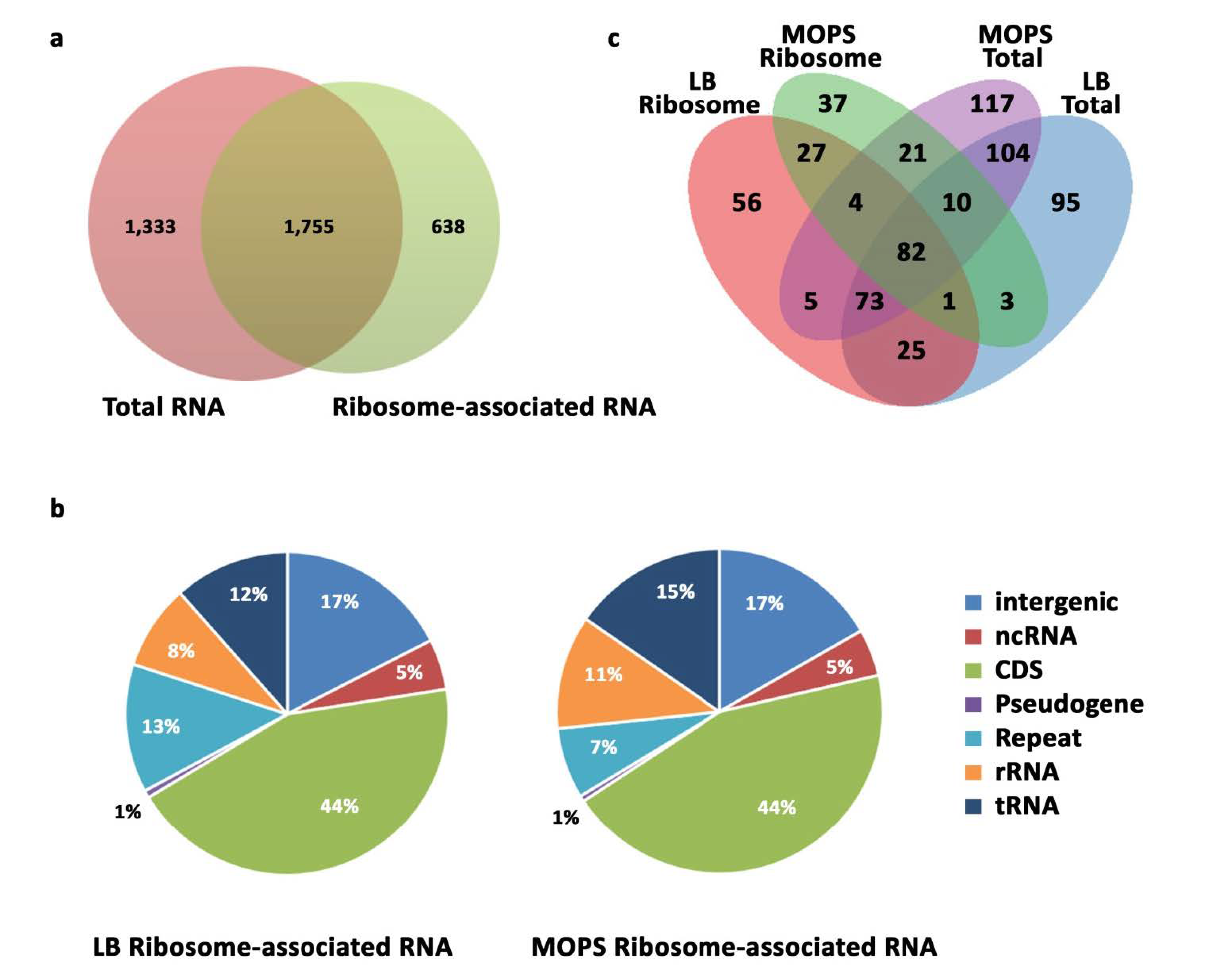
2.3. Overview of the Small E. coli Transcriptome
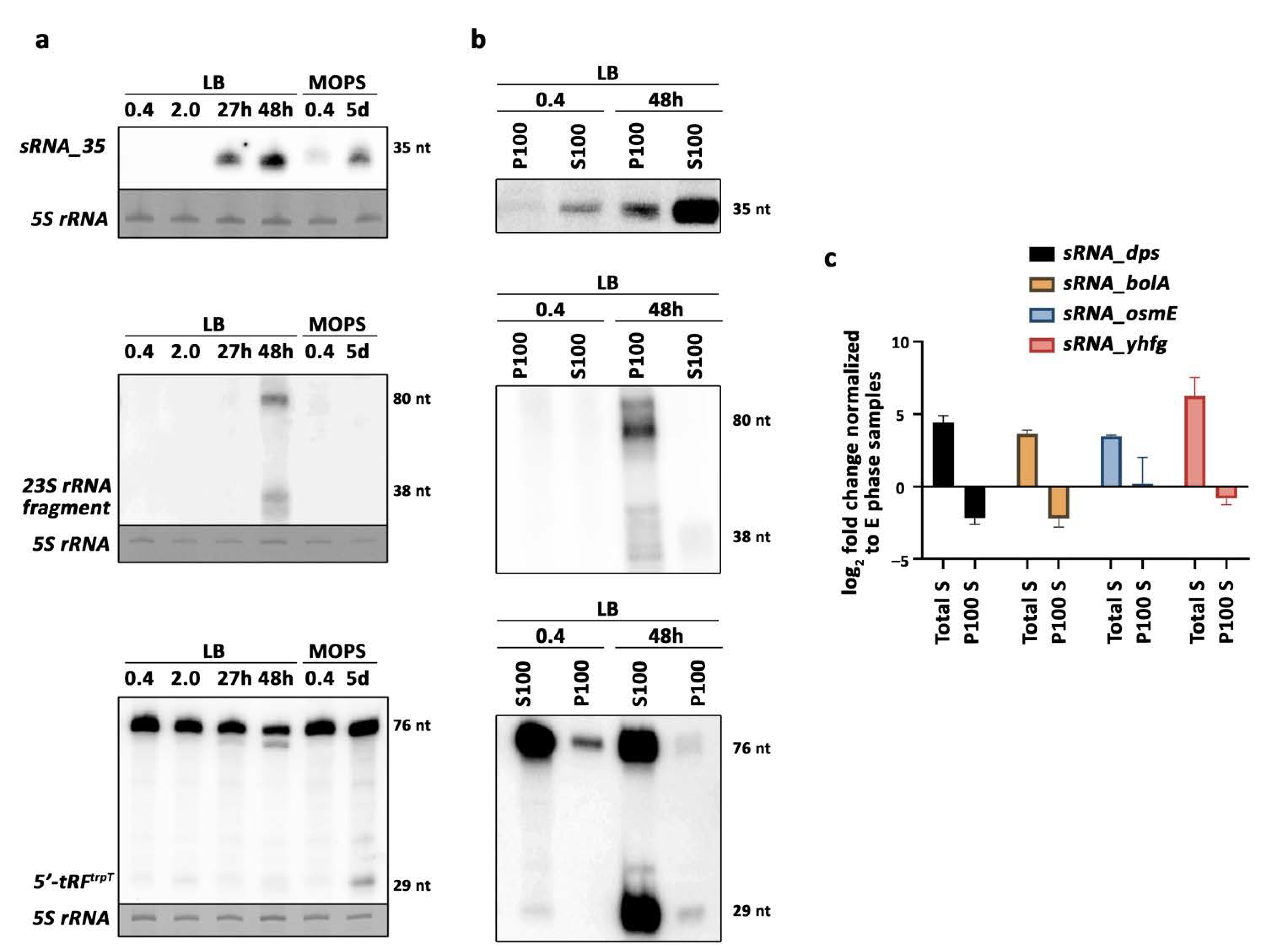
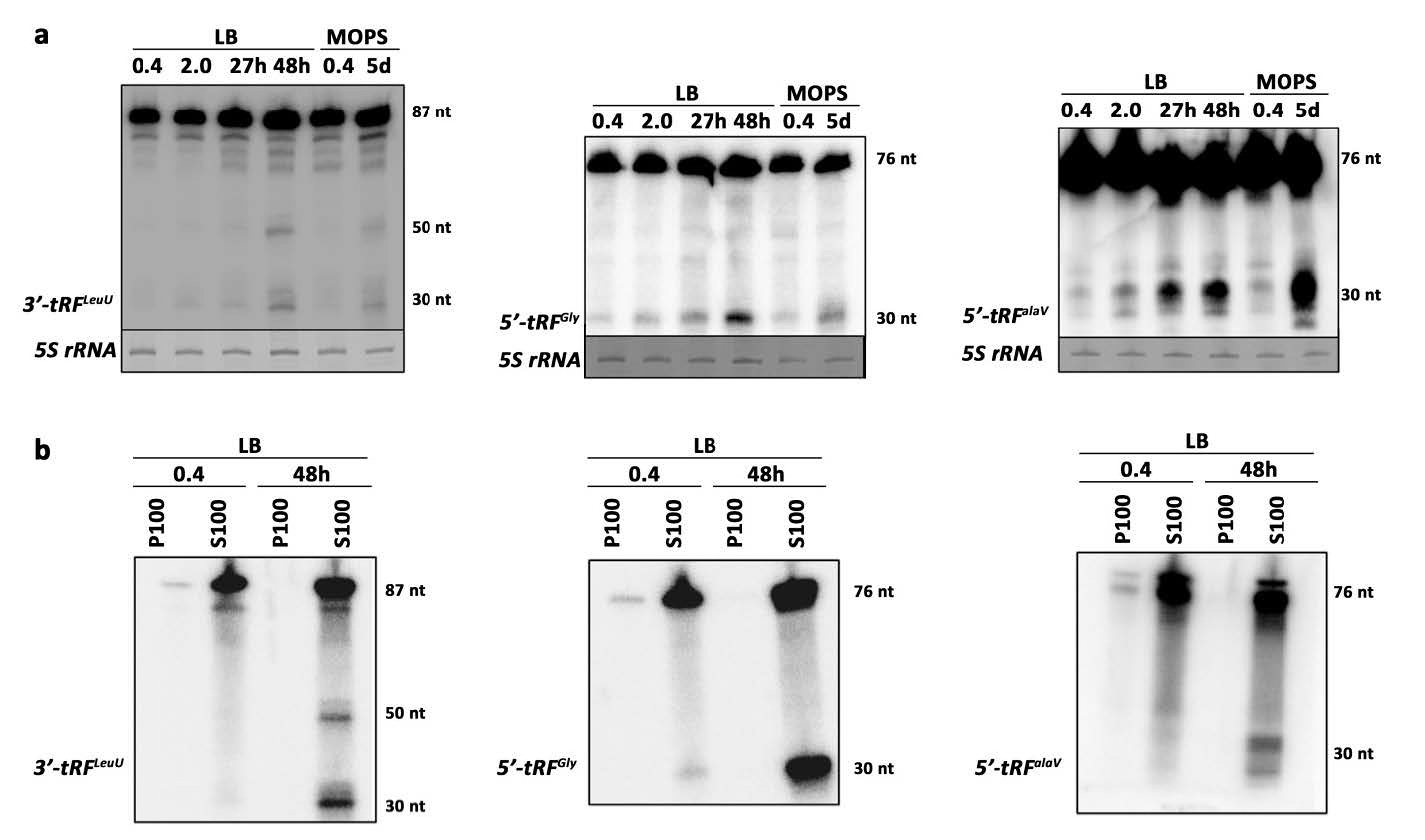
2.4. Validation of Selected Candidates
3. Discussion
3.1. sRNAs in Stationary Phase Biology
3.2. tRF Enrichment in Stationary Phase
4. Materials and Methods
4.1. Strains and Media
4.2. Cell Growth and Sample Collection
4.3. Construction of sRNA Expression Plasmids
4.4. Preparation of Crude Ribosomes
4.5. cDNA Library Preparation and Deep Sequencing
4.6. RNA Extraction and Northern Blot Analysis
4.7. Quantitative Real-Time RT-PCR (qPCR)
4.8. Bioinformatics Analysis
4.8.1. Trim—Cleaning of Reads
4.8.2. Align—Alignment to the Genome
4.8.3. Sam_sort—Sorting of Aligned Reads
4.8.4. Pseudo SE—SE to PE Conversion
4.8.5. Identify—Identification of Processing Products
4.8.6. Cluster—Clustering of Processing Products
Clustering by Overlap
Clustering by Sequence
4.8.7. Quantify—Quantification of Processing Products
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| cDNA | complementary DNA |
| CPM | counts per million |
| d | day |
| ETS | external transcribed spacer |
| FC | fold change |
| h | hour |
| ITS | internal transcribed spacer |
| mm | millimeter |
| mM | millimolar |
| ncRNA | noncoding RNA |
| nm | nanometers |
| nt | nucleotide |
| OD600 | optical density at wavelength 600nm |
| sRNA | small RNA |
| tRF | tRNA-derived RNA fragment |
References
- Roostalu, J.; Jõers, A.; Luidalepp, H.; Kaldalu, N.; Tenson, T. Cell division in Escherichia colicultures monitored at single cell resolution. BMC Microbiol. 2008, 8, 68. [Google Scholar] [CrossRef]
- Jaishankar, J.; Srivastava, P. Molecular Basis of Stationary Phase Survival and Applications. Front. Microbiol. 2017, 8, 2000. [Google Scholar] [CrossRef]
- Jõers, A.; Liske, E.; Tenson, T. Dividing subpopulation of Escherichia coli in stationary phase. Res. Microbiol. 2020, 171, 153–157. [Google Scholar] [CrossRef]
- Levin, B.R.; Rozen, D.E. Non-inherited antibiotic resistance. Nat. Rev. Microbiol. 2006, 4, 556–562. [Google Scholar] [CrossRef]
- Lewis, K. Persister cells, dormancy and infectious disease. Nat. Rev. Microbiol. 2007, 5, 48–56. [Google Scholar] [CrossRef]
- Joers, A.; Tenson, T. Growth resumption from stationary phase reveals memory in Escherichia coli cultures. Sci. Rep. 2016, 6, 24055. [Google Scholar] [CrossRef] [PubMed]
- Gray, D.A.; Dugar, G.; Gamba, P.; Strahl, H.; Jonker, M.J.; Hamoen, L.W. Extreme slow growth as alternative strategy to survive deep starvation in bacteria. Nat. Commun. 2019, 10, 890. [Google Scholar] [CrossRef] [PubMed]
- Dong, T.; Joyce, C.; Schellhorn, H.E. The Role of RpoS in Bacterial Adaptation. In Bacterial Physiology: A Molecular Approach; El-Sharoud, W., Ed.; Springer: Berlin/Heidelberg, Germany, 2008; pp. 313–337. [Google Scholar]
- Tkachenko, A.G.; Kashevarova, N.M.; Karavaeva, E.A.; Shumkov, M.S. Putrescine controls the formation of Escherichia coli persister cells tolerant to aminoglycoside netilmicin. FEMS Microbiol. Lett. 2014, 361, 25–33. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tkachenko, A.G.; Kashevarova, N.M.; Tyuleneva, E.A.; Shumkov, M.S. Stationary-phase genes upregulated by polyamines are responsible for the formation of Escherichia coli persister cells tolerant to netilmicin. FEMS Microbiol. Lett. 2017, 364. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Wu, N.; Zhang, S.; Yuan, Y.; Zhang, W.; Zhang, Y. Variable Persister Gene Interactions with (p) ppGpp for Persister Formation in Escherichia coli. Front. Microbiol. 2017, 8, 1795. [Google Scholar] [CrossRef]
- Traxler, M.F.; Summers, S.M.; Nguyen, H.T.; Zacharia, V.M.; Hightower, G.A.; Smith, J.T.; Conway, T. The global, ppGpp-mediated stringent response to amino acid starvation in Escherichia coli. Mol. Microbiol. 2008, 68, 1128–1148. [Google Scholar] [CrossRef] [PubMed]
- Battesti, A.; Majdalani, N.; Gottesman, S. The RpoS-mediated general stress response in Escherichia coli. Annu. Rev. Microbiol. 2011, 65, 189–213. [Google Scholar] [CrossRef] [PubMed]
- Wagner, E.G.H.; Romby, P. Small RNAs in bacteria and archaea: Who they are, what they do, and how they do it. Adv. Genet. 2015, 90, 133–208. [Google Scholar] [CrossRef] [PubMed]
- Waters, L.S.; Storz, G. Regulatory RNAs in bacteria. Cell 2009, 136, 615–628. [Google Scholar] [CrossRef]
- Fröhlich, K.S.; Gottesman, S. Small Regulatory RNAs in the Enterobacterial Response to Envelope Damage and Oxidative Stress. In Regulating with RNA in Bacteria and Archaea; John Wiley & Sons: Hoboken, NJ, USA, 2018. [Google Scholar]
- Raghavan, R.; Groisman, E.A.; Ochman, H. Genome-wide detection of novel regulatory RNAs in E. coli. Genome Res. 2011, 21, 1487–1497. [Google Scholar] [CrossRef] [PubMed]
- Shinhara, A.; Matsui, M.; Hiraoka, K.; Nomura, W.; Hirano, R.; Nakahigashi, K.; Tomita, M.; Mori, H.; Kanai, A. Deep sequencing reveals as-yet-undiscovered small RNAs in Escherichia coli. BMC Genom. 2011, 12, 428. [Google Scholar] [CrossRef]
- Miyakoshi, M.; Matera, G.; Maki, K.; Sone, Y.; Vogel, J. Functional expansion of a TCA cycle operon mRNA by a 3′ end-derived small RNA. Nucleic Acids Res. 2019, 47, 2075–2088. [Google Scholar] [CrossRef]
- Feng, L.; Rutherford, S.T.; Papenfort, K.; Bagert, J.D.; van Kessel, J.C.; Tirrell, D.A.; Wingreen, N.S.; Bassler, B.L. A qrr noncoding RNA deploys four different regulatory mechanisms to optimize quorum-sensing dynamics. Cell 2015, 160, 228–240. [Google Scholar] [CrossRef]
- Borgmann, J.; Schakermann, S.; Bandow, J.E.; Narberhaus, F. A Small Regulatory RNA Controls Cell Wall Biosynthesis and Antibiotic Resistance. MBio 2018, 9. [Google Scholar] [CrossRef]
- Chao, Y.; Vogel, J. A 3’ UTR-Derived Small RNA Provides the Regulatory Noncoding Arm of the Inner Membrane Stress Response. Mol. Cell 2016, 61, 352–363. [Google Scholar] [CrossRef] [PubMed]
- Westermann, A.J.; Forstner, K.U.; Amman, F.; Barquist, L.; Chao, Y.; Schulte, L.N.; Muller, L.; Reinhardt, R.; Stadler, P.F.; Vogel, J. Dual RNA-seq unveils noncoding RNA functions in host-pathogen interactions. Nature 2016, 529, 496–501. [Google Scholar] [CrossRef]
- Zhang, S.; Liu, S.; Wu, N.; Yuan, Y.; Zhang, W.; Zhang, Y. Small Non-coding RNA RyhB Mediates Persistence to Multiple Antibiotics and Stresses in Uropathogenic Escherichia coli by Reducing Cellular Metabolism. Front. Microbiol. 2018, 9, 136. [Google Scholar] [CrossRef] [PubMed]
- Massé, E.; Gottesman, S. A Small RNA Regulates the Expression of Genes Involved in Iron Metabolism in Escherichia coli. Proc. Natl. Acad. Sci. USA 2002, 99, 4620–4625. [Google Scholar] [CrossRef]
- Smirnov, A.; Forstner, K.U.; Holmqvist, E.; Otto, A.; Gunster, R.; Becher, D.; Reinhardt, R.; Vogel, J. Grad-seq guides the discovery of ProQ as a major small RNA-binding protein. Proc. Natl. Acad. Sci. USA 2016, 113, 11591–11596. [Google Scholar] [CrossRef] [PubMed]
- Müller, P.; Gimpel, M.; Wildenhain, T.; Brantl, S. A new role for CsrA: Promotion of complex formation between an sRNA and its mRNA target in Bacillus subtilis. RNA Biol. 2019, 16, 972–987. [Google Scholar] [CrossRef]
- Vogel, J.; Luisi, B.F. Hfq and its constellation of RNA. Nat. Rev. Microbiol. 2011, 9, 578–589. [Google Scholar] [CrossRef] [PubMed]
- Melamed, S.; Peer, A.; Faigenbaum-Romm, R.; Gatt, Y.E.; Reiss, N.; Bar, A.; Altuvia, Y.; Argaman, L.; Margalit, H. Global Mapping of Small RNA-Target Interactions in Bacteria. Mol. Cell 2016, 63, 884–897. [Google Scholar] [CrossRef]
- Iosub, I.A.; van Nues, R.W.; McKellar, S.W.; Nieken, K.J.; Marchioretto, M.; Sy, B.; Tree, J.J.; Viero, G.; Granneman, S. Hfq CLASH uncovers sRNA-target interaction networks linked to nutrient availability adaptation. Elife 2020, 9, e54655. [Google Scholar] [CrossRef]
- Holmqvist, E.; Li, L.; Bischler, T.; Barquist, L.; Vogel, J. Global Maps of ProQ Binding In Vivo Reveal Target Recognition via RNA Structure and Stability Control at mRNA 3’ Ends. Mol. Cell 2018, 70, 971–982 e976. [Google Scholar] [CrossRef]
- Pircher, A.; Gebetsberger, J.; Polacek, N. Ribosome-associated ncRNAs: An emerging class of translation regulators. RNA Biol. 2014, 11, 1335–1339. [Google Scholar] [CrossRef] [PubMed]
- Gonskikh, Y.; Gerstl, M.; Kos, M.; Borth, N.; Schosserer, M.; Grillari, J.; Polacek, N. Modulation of mammalian translation by a ribosome-associated tRNA half. RNA Biol. 2020, 17, 1125–1136. [Google Scholar] [CrossRef] [PubMed]
- Fricker, R.; Brogli, R.; Luidalepp, H.; Wyss, L.; Fasnacht, M.; Joss, O.; Zywicki, M.; Helm, M.; Schneider, A.; Cristodero, M.; et al. A tRNA half modulates translation as stress response in Trypanosoma brucei. Nat. Commun. 2019, 10, 118. [Google Scholar] [CrossRef] [PubMed]
- Pircher, A.; Bakowska-Zywicka, K.; Schneider, L.; Zywicki, M.; Polacek, N. An mRNA-derived noncoding RNA targets and regulates the ribosome. Mol. Cell 2014, 54, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Gebetsberger, J.; Wyss, L.; Mleczko, A.M.; Reuther, J.; Polacek, N. A tRNA-derived fragment competes with mRNA for ribosome binding and regulates translation during stress. RNA Biol. 2017, 14, 1364–1373. [Google Scholar] [CrossRef] [PubMed]
- Wyss, L.; Waser, M.; Gebetsberger, J.; Zywicki, M.; Polacek, N. mRNA-specific translation regulation by a ribosome-associated ncRNA in Haloferax volcanii. Sci. Rep. 2018, 8, 12502. [Google Scholar] [CrossRef]
- Hoogstrate, Y.; Jenster, G.; Martens-Uzunova, E.S. FlaiMapper: Computational annotation of small ncRNA-derived fragments using RNA-seq high-throughput data. Bioinformatics 2015, 31, 665–673. [Google Scholar] [CrossRef]
- Nitzan, M.; Rehani, R.; Margalit, H. Integration of Bacterial Small RNAs in Regulatory Networks. Annu. Rev. Biophys. 2017, 46, 131–148. [Google Scholar] [CrossRef]
- Zywicki, M.; Bakowska-Zywicka, K.; Polacek, N. Revealing stable processing products from ribosome-associated small RNAs by deep-sequencing data analysis. Nucleic Acids Res. 2012, 40, 4013–4024. [Google Scholar] [CrossRef]
- Janssen, B.D.; Hayes, C.S. The tmRNA ribosome-rescue system. Adv. Protein Chem. Struct. Biol. 2012, 86, 151–191. [Google Scholar] [CrossRef]
- Luidalepp, H.; Berger, S.; Joss, O.; Tenson, T.; Polacek, N. Ribosome Shut-Down by 16S rRNA Fragmentation in Stationary-Phase Escherichia coli. J. Mol. Biol. 2016, 428, 2237–2247. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pinel-Marie, M.L.; Brielle, R.; Riffaud, C.; Germain-Amiot, N.; Polacek, N.; Felden, B. RNA antitoxin SprF1 binds ribosomes to attenuate translation and promote persister cell formation in Staphylococcus aureus. Nat. Microbiol. 2021, 6, 209–220. [Google Scholar] [CrossRef] [PubMed]
- Wassarman, K.M.; Storz, G. 6S RNA regulates E. coli RNA polymerase activity. Cell 2000, 101, 613–623. [Google Scholar] [CrossRef]
- Steuten, B.; Hoch, P.G.; Damm, K.; Schneider, S.; Köhler, K.; Wagner, R.; Hartmann, R.K. Regulation of transcription by 6S RNAs: Insights from the Escherichia coli and Bacillus subtilis model systems. RNA Biol. 2014, 11, 508–521. [Google Scholar] [CrossRef]
- Han, K.; Kim, K.S.; Bak, G.; Park, H.; Lee, Y. Recognition and discrimination of target mRNAs by Sib RNAs, a cis-encoded sRNA family. Nucleic Acids Res. 2010, 38, 5851–5866. [Google Scholar] [CrossRef] [PubMed]
- Jahanshahi, S.; Li, Y. An Effective Method for Quantifying RNA Expression of IbsC-SibC, a Type I Toxin-Antitoxin System in Escherichia coli. Chembiochem 2020, 21, 3120–3130. [Google Scholar] [CrossRef]
- Rasmussen, A.A.; Johansen, J.; Nielsen, J.S.; Overgaard, M.; Kallipolitis, B.; Valentin-Hansen, P. A conserved small RNA promotes silencing of the outer membrane protein YbfM. Mol. Microbiol. 2009, 72, 566–577. [Google Scholar] [CrossRef] [PubMed]
- Mandin, P.; Gottesman, S. A genetic approach for finding small RNAs regulators of genes of interest identifies RybC as regulating the DpiA/DpiB two-component system. Mol. Microbiol. 2009, 72, 551–565. [Google Scholar] [CrossRef] [PubMed]
- Weichart, D.; Lange, R.; Henneberg, N.; Hengge-Aronis, R. Identification and characterization of stationary phase inducible genes in Escherichia coli. Mol. Microbiol. 1993, 10, 407–420. [Google Scholar] [CrossRef] [PubMed]
- Polacek, N.; Ivanov, P. The regulatory world of tRNA fragments beyond canonical tRNA biology. RNA Biol. 2020, 17, 1057–1059. [Google Scholar] [CrossRef] [PubMed]
- Cristodero, M.; Polacek, N. The multifaceted regulatory potential of tRNA-derived fragments. Non-Coding RNA Investig. 2017, 170, 61–71.e11. [Google Scholar] [CrossRef]
- Goodarzi, H.; Liu, X.; Nguyen, H.C.; Zhang, S.; Fish, L.; Tavazoie, S.F. Endogenous tRNA-Derived Fragments Suppress Breast Cancer Progression via YBX1 Displacement. Cell 2015, 161, 790–802. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.K.; Fuchs, G.; Wang, S.; Wei, W.; Zhang, Y.; Park, H.; Roy-Chaudhuri, B.; Li, P.; Xu, J.; Chu, K.; et al. A transfer-RNA-derived small RNA regulates ribosome biogenesis. Nature 2017, 552, 57–62. [Google Scholar] [CrossRef]
- Oberbauer, V.; Schaefer, M.R. tRNA-Derived Small RNAs: Biogenesis, Modification, Function and Potential Impact on Human Disease Development. Genes 2018, 9, 607. [Google Scholar] [CrossRef] [PubMed]
- Lalaouna, D.; Carrier, M.C.; Semsey, S.; Brouard, J.S.; Wang, J.; Wade, J.T.; Masse, E. A 3’ external transcribed spacer in a tRNA transcript acts as a sponge for small RNAs to prevent transcriptional noise. Mol. Cell 2015, 58, 393–405. [Google Scholar] [CrossRef] [PubMed]
- Melamed, S.; Adams, P.P.; Zhang, A.; Zhang, H.; Storz, G. RNA-RNA Interactomes of ProQ and Hfq Reveal Overlapping and Competing Roles. Mol. Cell 2020, 77, 411–425.e417. [Google Scholar] [CrossRef]
- Blattner, F.R.; Plunkett, G.; Bloch, C.A.; Perna, N.T.; Burland, V.; Riley, M.; Collado-Vides, J.; Glasner, J.D.; Rode, C.K.; Mayhew, G.F.; et al. The Complete Genome Sequence of Escherichia coli K-12. Science 1997, 277, 1453–1462. [Google Scholar] [CrossRef]
- Vimberg, V.; Tats, A.; Remm, M.; Tenson, T. Translation initiation region sequence preferences in Escherichia coli. BMC Mol. Biol. 2007, 8, 100. [Google Scholar] [CrossRef] [PubMed]
- Luidalepp, H.; Joers, A.; Kaldalu, N.; Tenson, T. Age of inoculum strongly influences persister frequency and can mask effects of mutations implicated in altered persistence. J. Bacteriol. 2011, 193, 3598–3605. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.S.; Krupa, R.A.; Zhang, F.; Hajimorad, M.; Holtz, W.J.; Prasad, N.; Lee, S.K.; Keasling, J.D. BglBrick vectors and datasheets: A synthetic biology platform for gene expression. J. Biol. Eng. 2011, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, K. Chapter seventeen-MEGAWHOP Cloning: A Method of Creating Random Mutagenesis Libraries via Megaprimer PCR of Whole Plasmids. In Methods in Enzymology; Voigt, C., Ed.; Academic Press: Cambridge, MA, USA, 2011; Volume 498, pp. 399–406. [Google Scholar]
- Qi, D.; Scholthof, K.B. A one-step PCR-based method for rapid and efficient site-directed fragment deletion, insertion, and substitution mutagenesis. J. Virol. Methods 2008, 149, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Gebetsberger, J.; Zywicki, M.; Kunzi, A.; Polacek, N. tRNA-derived fragments target the ribosome and function as regulatory non-coding RNA in Haloferax volcanii. Archaea 2012, 2012, 260909. [Google Scholar] [CrossRef] [PubMed]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2ˆ (-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinforma. Biomath. 2013, 3, 71–85. [Google Scholar]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. EdgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J. 2011, 17, 3. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Flyvbjerg, H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics 2015, 31, 3476–3482. [Google Scholar] [CrossRef] [PubMed]
- Mohanty, B.K.; Kushner, S.R. The majority of Escherichia coli mRNAs undergo post-transcriptional modification in exponentially growing cells. Nucleic Acids Res. 2006, 34, 5695–5704. [Google Scholar] [CrossRef] [PubMed]
- Fu, H.; Feng, J.; Liu, Q.; Sun, F.; Tie, Y.; Zhu, J.; Xing, R.; Sun, Z.; Zheng, X. Stress induces tRNA cleavage by angiogenin in mammalian cells. FEBS Lett. 2009, 583, 437–442. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]

| Description | LB Total | MOPS Total | LB Ribosome | MOPS Ribosome |
|---|---|---|---|---|
| Downregulated | 1024 | 1098 | 323 | 299 |
| Not significant | 1671 | 1574 | 1797 | 1909 |
| Upregulated | 393 | 416 | 273 | 185 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raad, N.; Luidalepp, H.; Fasnacht, M.; Polacek, N. Transcriptome-Wide Analysis of Stationary Phase Small ncRNAs in E. coli. Int. J. Mol. Sci. 2021, 22, 1703. https://doi.org/10.3390/ijms22041703
Raad N, Luidalepp H, Fasnacht M, Polacek N. Transcriptome-Wide Analysis of Stationary Phase Small ncRNAs in E. coli. International Journal of Molecular Sciences. 2021; 22(4):1703. https://doi.org/10.3390/ijms22041703
Chicago/Turabian StyleRaad, Nicole, Hannes Luidalepp, Michel Fasnacht, and Norbert Polacek. 2021. "Transcriptome-Wide Analysis of Stationary Phase Small ncRNAs in E. coli" International Journal of Molecular Sciences 22, no. 4: 1703. https://doi.org/10.3390/ijms22041703
APA StyleRaad, N., Luidalepp, H., Fasnacht, M., & Polacek, N. (2021). Transcriptome-Wide Analysis of Stationary Phase Small ncRNAs in E. coli. International Journal of Molecular Sciences, 22(4), 1703. https://doi.org/10.3390/ijms22041703







