A Conserved Allosteric Site on Drug-Metabolizing CYPs: A Systematic Computational Assessment
Abstract
:1. Introduction
2. Results and Discussion
2.1. Simulation Techniques and Model Validation
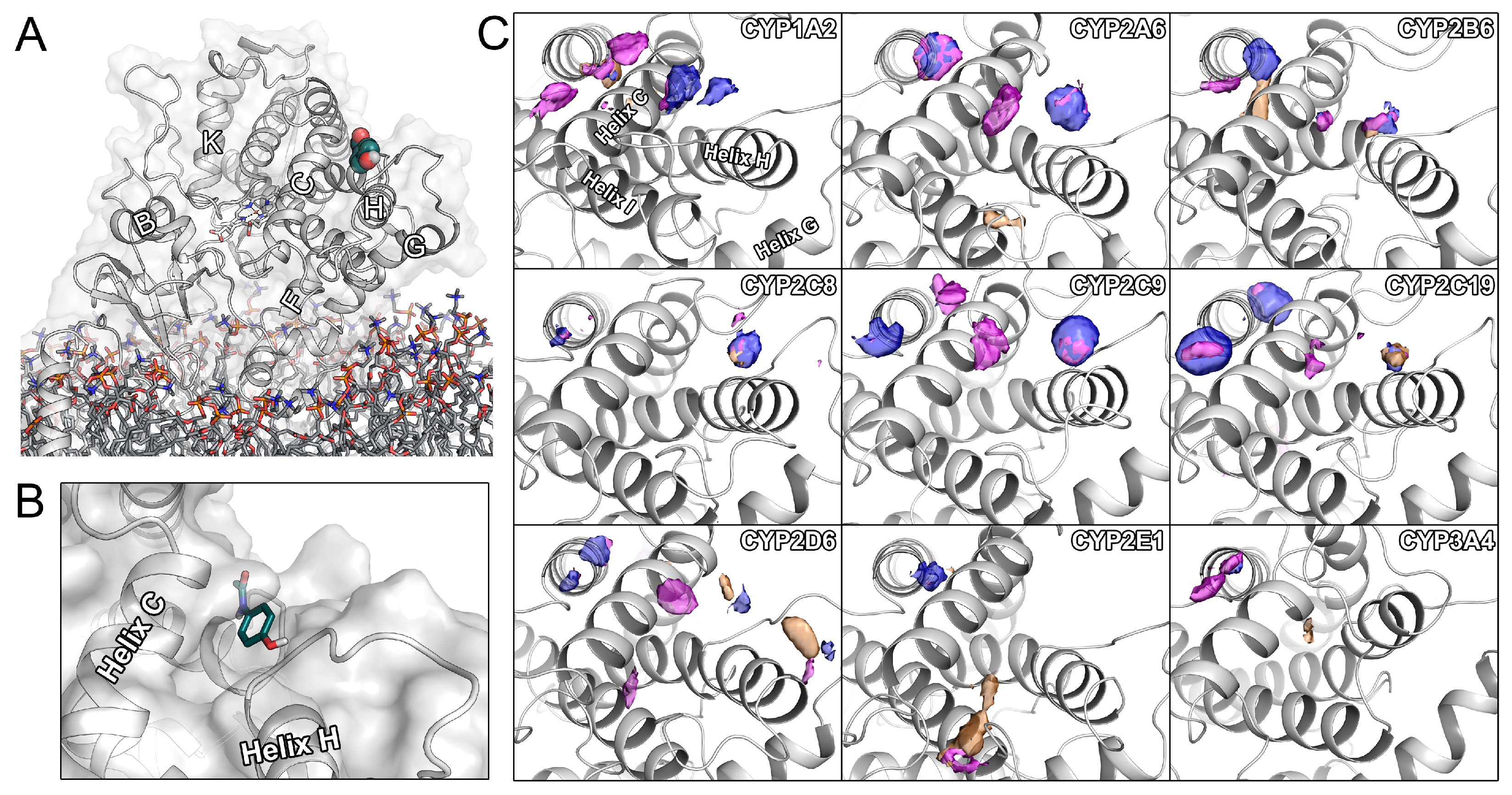
2.2. Small Molecules Regularly Associate with the H1 Site of Several CYPs
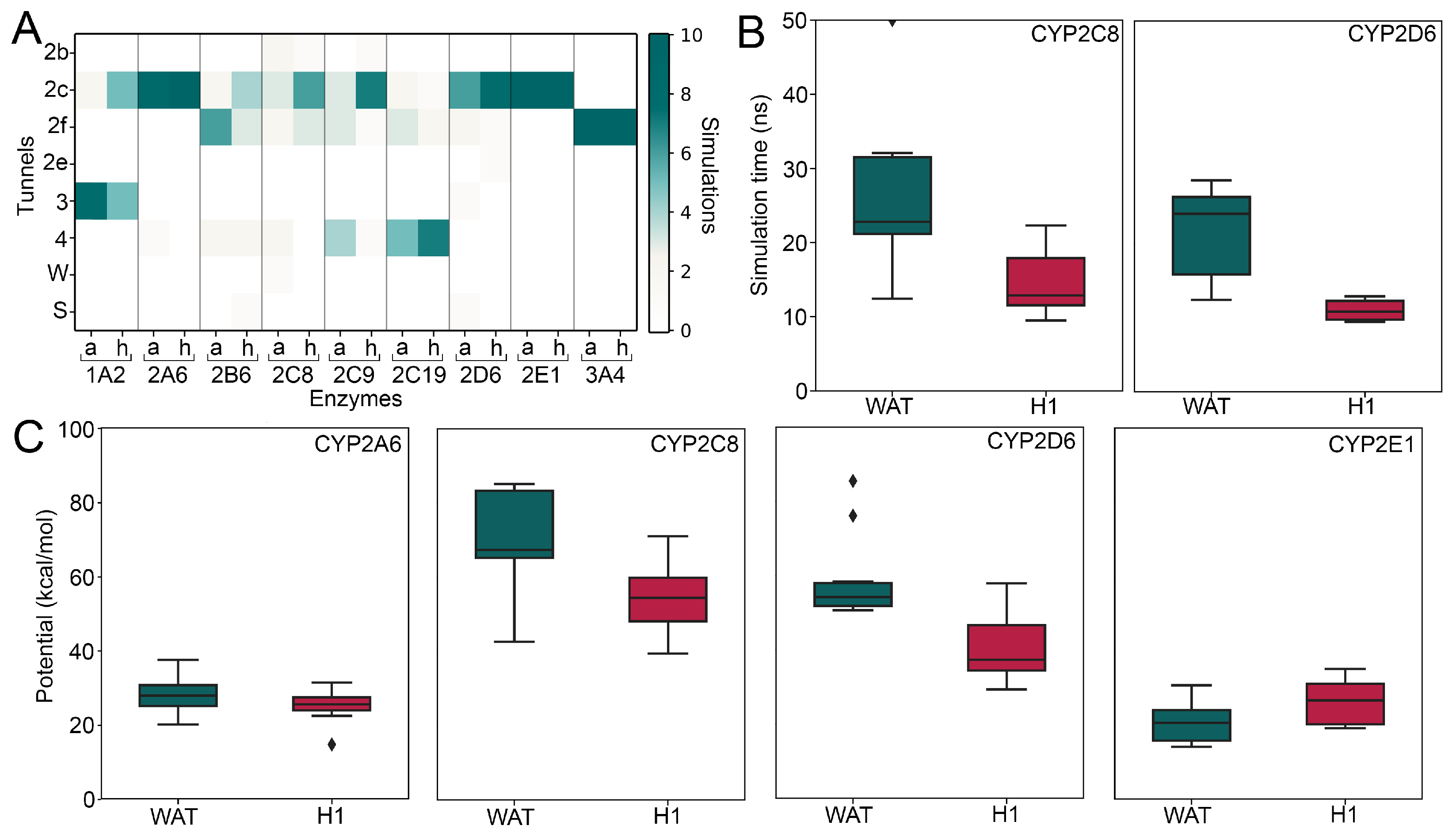
2.3. The Effect of Allosteric Ligands Bound to the H1 Site Is Isoform-Dependent
3. Conclusions
4. Materials and Methods
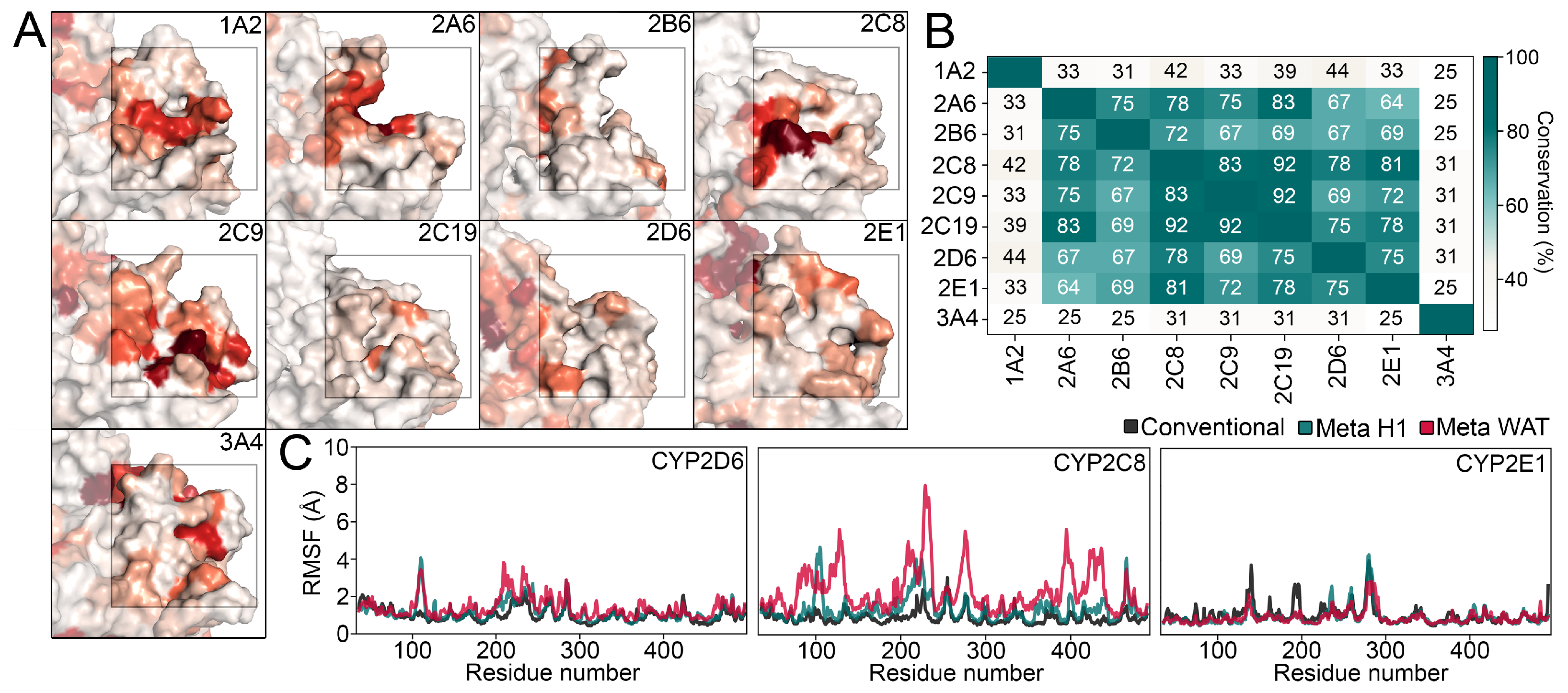
4.1. Bioinformatics Analysis
4.2. Model Building
4.3. MD Simulations
4.4. Evaluation of the MD Trajectories
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zanger, U.M.; Schwab, M. Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol. Ther. 2013, 138, 103–141. [Google Scholar] [CrossRef]
- Manikandan, P.; Nagini, S. Cytochrome P450 Structure, Function and Clinical Significance: A Review. Curr. Drug Targets 2018, 19, 38–54. [Google Scholar] [CrossRef] [PubMed]
- Laizure, S.C.; Herring, V.; Hu, Z.; Witbrodt, K.; Parker, R.B. The role of human carboxylesterases in drug metabolism: Have we overlooked their importance? Pharmacotherapy 2013, 33, 210–222. [Google Scholar] [CrossRef] [Green Version]
- Tyzack, J.D.; Kirchmair, J. Computational methods and tools to predict cytochrome P450 metabolism for drug discovery. Chem. Biol. Drug Des. 2019, 93, 377–386. [Google Scholar] [CrossRef]
- Van de Waterbeemd, H.; Gifford, E.; Arbor, A. ADMET in silico modelling: Towards prediction paradise? Nat. Rev. Drug Discov. 2003, 2, 192–204. [Google Scholar] [CrossRef] [PubMed]
- Kola, I.; Landis, J. Can the pharmaceutical industry reduce attrition rates? Nat. Rev. Drug Discov. 2004, 3, 711–715. [Google Scholar] [CrossRef] [PubMed]
- Fürst, M.J.L.J.; Fiorentini, F.; Fraaije, M.W. Beyond active site residues: Overall structural dynamics control catalysis in flavin-containing and heme-containing monooxygenases. Curr. Opin. Struct. Biol. 2019, 59, 29–37. [Google Scholar] [CrossRef]
- Fischer, A.; Smieško, M. Spontaneous Ligand Access Events to Membrane-Bound Cytochrome P450 2D6 Sampled at Atomic Resolution. Sci. Rep. 2019, 9, 16411. [Google Scholar] [CrossRef] [PubMed]
- Urban, P.; Lautier, T.; Pompon, D.; Truan, G. Ligand Access Channels in Cytochrome P450 Enzymes: A Review. Int. J. Mol. Sci. 2018, 19, 1617. [Google Scholar] [CrossRef] [Green Version]
- Berka, K.; Hendrychová, T.; Anzenbacher, P.; Otyepka, M. Membrane position of ibuprofen agrees with suggested access path entrance to cytochrome P450 2C9 active site. J. Phys. Chem. A 2011, 115, 11248–11255. [Google Scholar] [CrossRef] [PubMed]
- Berka, K.; Paloncýová, M.; Anzenbacher, P.; Otyepka, M. Behavior of human cytochromes P450 on lipid membranes. J. Phys. Chem. B 2013, 117, 11556–11564. [Google Scholar] [CrossRef] [PubMed]
- Gora, A.; Brezovsky, J.; Damborsky, J. Gates of enzymes. Chem. Rev. 2013, 113, 5871–5923. [Google Scholar] [CrossRef]
- Fischer, A.; Frehner, G.; Lill, M.A.; Smieško, M. Conformational Changes of Thyroid Receptors in Response to Antagonists. J. Chem. Inf. Model. 2021, 61, 1010–1019. [Google Scholar] [CrossRef]
- Follmer, A.H.; Mahomed, M.; Goodin, D.B.; Poulos, T.L. Substrate-Dependent Allosteric Regulation in Cytochrome P450cam (CYP101A1). J. Am. Chem. Soc. 2018, 140, 16222–16228. [Google Scholar] [CrossRef]
- Cojocaru, V.; Winn, P.J.; Wade, R.C. The ins and outs of cytochrome P450s. Biochim. Biophys. Acta. 2007, 1770, 390–401. [Google Scholar] [CrossRef] [PubMed]
- Verkhivker, G.M.; Agajanian, S.; Hu, G.; Tao, P. Allosteric Regulation at the Crossroads of New Technologies: Multiscale Modeling, Networks, and Machine Learning. Front. Mol. Biosci. 2020, 7, 136. [Google Scholar] [CrossRef] [PubMed]
- Lisi, G.P.; Loria, J.P. Allostery in enzyme catalysis. Curr. Opin. Struct. Biol. 2017, 47, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Ghanakota, P.; Carlson, H.A. Driving Structure-Based Drug Discovery through Cosolvent Molecular Dynamics. J. Med. Chem. 2016, 59, 10383–10399. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Vivo, M.; Masetti, M.; Bottegoni, G.; Cavalli, A. Role of Molecular Dynamics and Related Methods in Drug Discovery. J. Med. Chem. 2016, 59, 4035–4061. [Google Scholar] [CrossRef]
- Hospital, A.; Goñi, J.R.; Orozco, M.; Gelpi, J. Molecular dynamics simulations: Advances and applications. Adv. Appl. Bioinform. Chem. 2015, 8, 37–47. [Google Scholar] [CrossRef] [Green Version]
- Fischer, A.; Don, C.G.; Smieško, M. Molecular Dynamics Simulations Reveal Structural Differences among Allelic Variants of Membrane-Anchored Cytochrome P450 2D6. J. Chem. Inf. Model. 2018, 58, 1962–1975. [Google Scholar] [CrossRef]
- Ohta, Y.; Kawato, S.; Tagashira, H.; Takemori, S.; Kominami, S. Dynamic Structures of Adrenocortical Cytochrome P-450 in Proteoliposomes and Microsomes: Protein Rotation Study. Biochemistry 1992, 31, 12680–12687. [Google Scholar] [CrossRef] [PubMed]
- Bayburt, T.H.; Sligar, S.G. Single-molecule height measurements on microsomal cytochrome P450 in nanometer-scale phospholipid bilayer disks. Proc. Natl. Acad. Sci. USA 2002, 99, 6725–6730. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, Y.T.; Wilson, R.F.; Rupniewski, I.; Goodin, D.B. P450cam visits an open conformation in the absence of substrate. Biochemistry 2010, 49, 3412–3419. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haritha, P.; Shanmugavadivu, P.; Dhamodharan, S. A Comprehensive Review on Protein Sequence Analysis Techniques. Int. J. Comput. Sci. Eng. 2018, 6, 1433–1442. [Google Scholar] [CrossRef]
- Paloncýova, M.; Navrátilova, V.; Berka, K.; Laio, A.; Otyepka, M. Role of Enzyme Flexibility in Ligand Access and Egress to Active Site: Bias-Exchange Metadynamics Study of 1,3,7-Trimethyluric Acid in Cytochrome P450 3A4. J. Chem. Theory Comput. 2016, 12, 2101–2109. [Google Scholar] [CrossRef]
- Chovancova, E.; Pavelka, A.; Benes, P.; Strnad, O.; Brezovsky, J.; Kozlikova, B.; Gora, A.; Sustr, V.; Klvana, M.; Medek, P.; et al. CAVER 3.0: A Tool for the Analysis of Transport Pathways in Dynamic Protein Structures. PLoS Comput. Biol. 2012, 8, e1002708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sündermann, A.; Oostenbrink, C. Molecular dynamics simulations give insight into the conformational change, complex formation, and electron transfer pathway for cytochrome P450 reductase. Protein Sci. 2013, 22, 1183–1195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lobanov, M.Y.; Bogatyreva, N.S.; Galzitskaya, O.V. Radius of gyration as an indicator of protein structure compactness. Mol. Biol. 2008, 42, 623–628. [Google Scholar] [CrossRef]
- Zwier, M.C.; Chong, L.T. Reaching biological timescales with all-atom molecular dynamics simulations. Curr. Opin. Pharmacol. 2010, 10, 745–752. [Google Scholar] [CrossRef] [PubMed]
- Vécsey-Semjén, B.; Lesieur, C.; Möllby, R.; Van Der Goot, F.G. Conformational changes due to membrane binding and channel formation by staphylococcal α-toxin. J. Biol. Chem. 1997, 272, 5709–5717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sui, S.F. Membrane-induced conformational change of proteins. Adv. Colloid Interface Sci. 2000, 85, 257–267. [Google Scholar] [CrossRef]
- Trendelkamp-Schroer, B.; Noé, F. Efficient Estimation of Rare-Event Kinetics. Phys. Rev. X 2016, 6, 11009. [Google Scholar] [CrossRef] [Green Version]
- Laio, A.; Gervasio, F.L. Metadynamics: A method to simulate rare events and reconstruct the free energy in biophysics, chemistry and material science. Rep. Prog. Phys. 2008, 71, 126601. [Google Scholar] [CrossRef]
- Hamelberg, D.; Mongan, J.; McCammon, J.A. Accelerated molecular dynamics: A promising and efficient simulation method for biomolecules. J. Chem. Phys. 2004, 120, 11919–11929. [Google Scholar] [CrossRef] [Green Version]
- Borrelli, K.W.; Vitalis, A.; Alcantara, R.; Guallar, V. PELE: Protein energy landscape exploration. A novel Monte Carlo based technique. J. Chem. Theory Comput. 2005, 1, 1304–1311. [Google Scholar] [CrossRef]
- Bortolato, A.; Deflorian, F.; Weiss, D.R.; Mason, J.S. Decoding the Role of Water Dynamics in Ligand–Protein Unbinding: CRF1R as a Test Case. J. Chem. Inf. Model. 2015, 55, 1857–1866. [Google Scholar] [CrossRef]
- Sun, H.; Li, Y.; Shen, M.; Li, D.; Kang, Y.; Hou, T. Characterizing Drug–Target Residence Time with Metadynamics: How To Achieve Dissociation Rate Efficiently without Losing Accuracy against Time-Consuming Approaches. J. Chem. Inf. Model. 2017, 57, 1895–1906. [Google Scholar] [CrossRef]
- Bateman, A.; Martin, M.J.; O’Donovan, C.; Magrane, M.; Alpi, E.; Antunes, R.; Bely, B.; Bingley, M.; Bonilla, C.; Britto, R.; et al. UniProt: The universal protein knowledgebase. Nucleic Acids Res. 2017, 45, D158–D169. [Google Scholar] [CrossRef] [Green Version]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [Green Version]
- Okonechnikov, K.; Golosova, O.; Fursov, M.; Varlamov, A.; Vaskin, Y.; Efremov, I.; German Grehov, O.G.; Kandrov, D.; Rasputin, K.; Syabro, M.; et al. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fischer, A.; Smieško, M. Allosteric binding sites on nuclear receptors: Focus on drug efficacy and selectivity. Int. J. Mol. Sci. 2020, 21, 534. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrödinger LCC. Maestro Small-Molecule Drug Discovery Suite 2019-3; Schrödinger, LCC: New York, NY, USA, 2019. [Google Scholar]
- Madhavi Sastry, G.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput.-Aided Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Rendic, S. Summary of information on human CYP enzymes: Human P450 metabolism data. Drug Metab. Rev. 2002, 34, 83–448. [Google Scholar] [CrossRef]
- Lomize, M.A.; Pogozheva, I.D.; Joo, H.; Mosberg, H.I.; Lomize, A.L. OPM database and PPM web server: Resources for positioning of proteins in membranes. Nucleic Acids Res. 2012, 40, 370–376. [Google Scholar] [CrossRef] [PubMed]
- Bowers, K.; Chow, E.; Xu, H.; Dror, R.; Eastwood, M.; Gregersen, B.; Klepeis, J.; Kolossvary, I.; Moraes, M.; Sacerdoti, F.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proceedings of the SC ‘06: Proceedings of the 2006 ACM/IEEE Conference on Supercomputing, Tampa, FL, USA, 11–17 November 2006; p. 43. [Google Scholar] [CrossRef] [Green Version]
- Shaw, D.E.; Grossman, J.P.; Bank, J.A.; Batson, B.; Butts, J.A.; Chao, J.C.; Deneroff, M.M.; Dror, R.O.; Even, A.; Fenton, C.H.; et al. Anton 2: Raising the Bar for Performance and Programmability in a Special-Purpose Molecular Dynamics Supercomputer. In Proceedings of the SC ‘14: Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis, New Orleans, LA, USA, 16–21 November 2014; pp. 41–53. [Google Scholar] [CrossRef]
- Halgren, T.A.; Murphy, R.B.; Friesner, R.A.; Beard, H.S.; Frye, L.L.; Pollard, W.T.; Banks, J.L. Glide: A New Approach for Rapid, Accurate Docking and Scoring. 2. Enrichment Factors in Database Screening. J. Med. Chem. 2004, 47, 1750–1759. [Google Scholar] [CrossRef] [PubMed]
- Ducassou, L.; Dhers, L.; Jonasson, G.; Pietrancosta, N.; Boucher, J.L.; Mansuy, D.; André, F. Membrane-bound human orphan cytochrome P450 2U1: Sequence singularities, construction of a full 3D model, and substrate docking. Biochimie 2017, 140, 166–175. [Google Scholar] [CrossRef] [PubMed]
- Kozlikova, B.; Sebestova, E.; Sustr, V.; Brezovsky, J.; Strnad, O.; Daniel, L.; Bednar, D.; Pavelka, A.; Manak, M.; Bezdeka, M.; et al. CAVER Analyst 1.0: Graphic tool for interactive visualization and analysis of tunnels and channels in protein structures. Bioinformatics 2014, 30, 2684–2685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trapl, D.; Spiwok, V. Analysis of the Results of Metadynamics Simulations by metadynminer and metadynminer3d. arXiv 2020, arXiv:2009.02241. [Google Scholar]
| Type | Membrane | Duration (ns) | Replicas a | LigOrtho b | LigAllo b |
|---|---|---|---|---|---|
| Cosolvent | no | 60 | 10 | no | no |
| Association | no | 500 | 1 | no | yes c |
| Equilibration | yes | 300 | 1 | yes | no |
| Sampling | yes | 1005 | 3 | no | yes |
| Metadynamics | yes | 50 | 10 | yes | no |
| Metadynamics | yes | 50 | 10 | yes | yes |
| Enzyme | Outcome T a | Outcome Pmax a | Significance | 2b | rB b 2c | 2f |
|---|---|---|---|---|---|---|
| CYP1A2 | H1 (0.386) | H1 (0.400) | no | = | + | = |
| CYP2A6 | H1 (0.237) | H1 (0.179) | no | + | = | = |
| CYP2B6 | H1 (0.204) | H1 (0.522) | no | + | n/a c | + |
| CYP2C8 | H1 (0.010) | H1 (0.025) | yes | – | – | + |
| CYP2C9 | WAT (0.675) | WAT (0.559) | no | = | – | + |
| CYP2C19 | H1 (0.454) | H1 (0.862) | no | – | + | + |
| CYP2D6 | H1 (0.001) | H1 (0.001) | yes | – | + | – |
| CYP2E1 | WAT (0.232) | WAT (0.085) | yes | + | – | = |
| CYP3A4 | H1 (0.571) | WAT (0.713) | no | n/a c | = | – |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fischer, A.; Smieško, M. A Conserved Allosteric Site on Drug-Metabolizing CYPs: A Systematic Computational Assessment. Int. J. Mol. Sci. 2021, 22, 13215. https://doi.org/10.3390/ijms222413215
Fischer A, Smieško M. A Conserved Allosteric Site on Drug-Metabolizing CYPs: A Systematic Computational Assessment. International Journal of Molecular Sciences. 2021; 22(24):13215. https://doi.org/10.3390/ijms222413215
Chicago/Turabian StyleFischer, André, and Martin Smieško. 2021. "A Conserved Allosteric Site on Drug-Metabolizing CYPs: A Systematic Computational Assessment" International Journal of Molecular Sciences 22, no. 24: 13215. https://doi.org/10.3390/ijms222413215
APA StyleFischer, A., & Smieško, M. (2021). A Conserved Allosteric Site on Drug-Metabolizing CYPs: A Systematic Computational Assessment. International Journal of Molecular Sciences, 22(24), 13215. https://doi.org/10.3390/ijms222413215







