Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential
Abstract
:1. Introduction
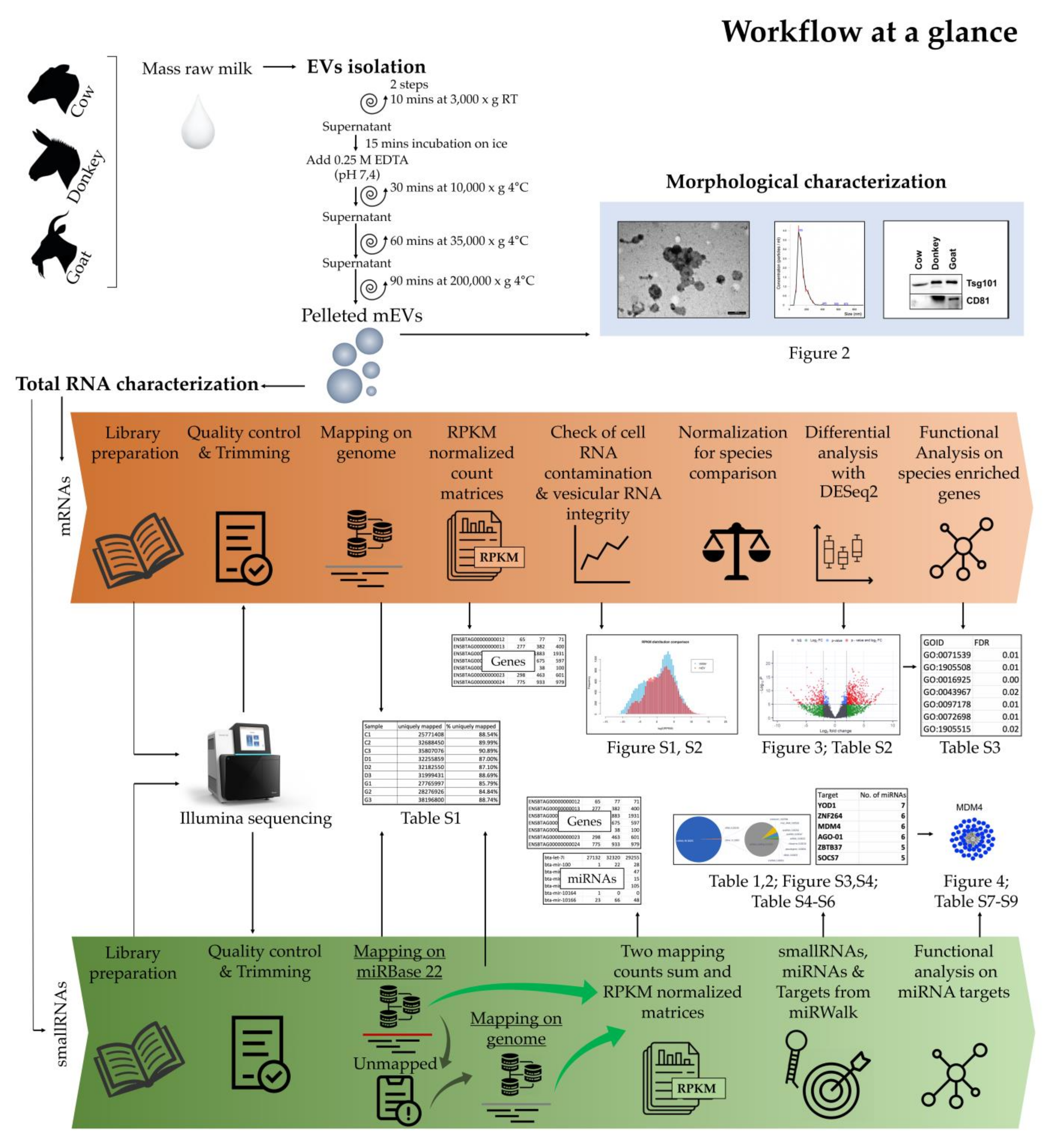
2. Results
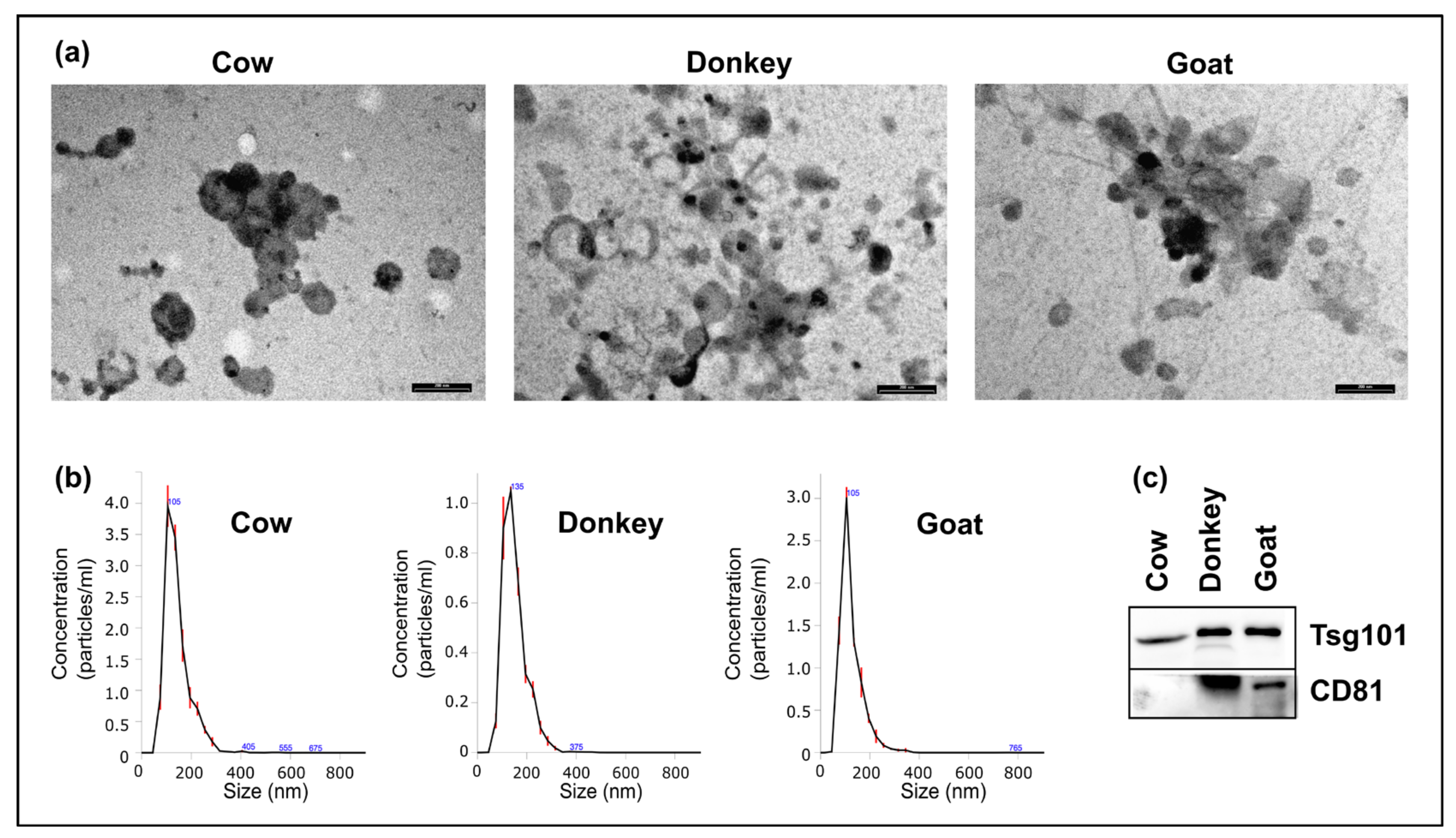
2.1. Morphologic Characterization of mEVs
2.2. Molecular Characterization of mEVs RNAs
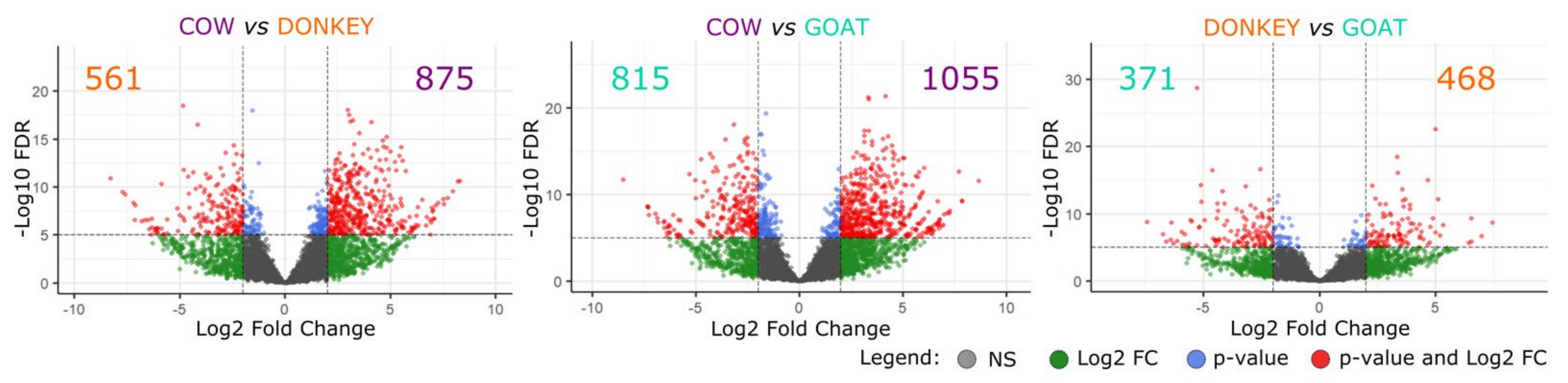
2.2.1. Messenger RNAs
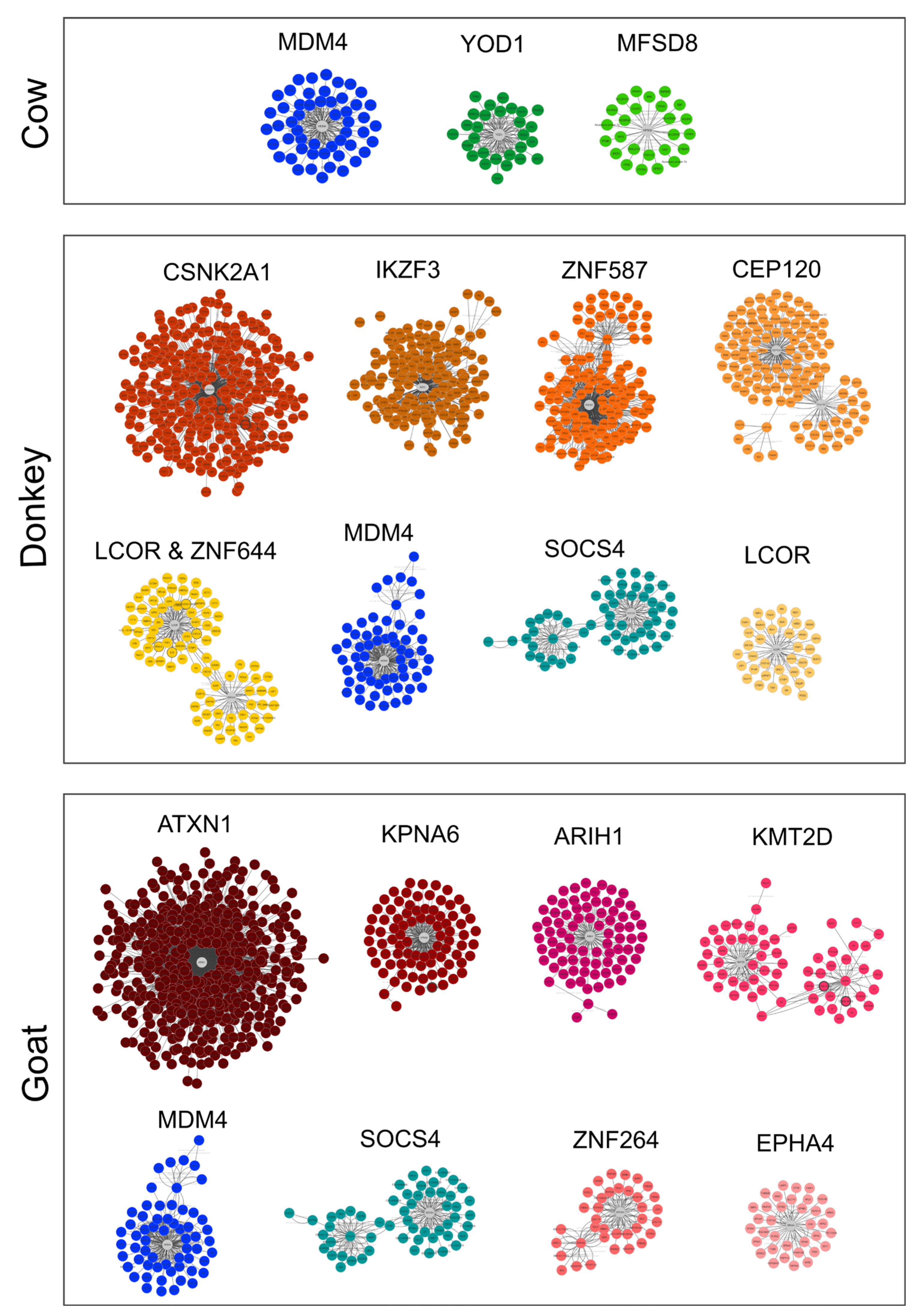
2.2.2. Small RNAs
3. Discussion
4. Materials and Methods
4.1. Milk Collection
4.2. EVs Isolation
4.3. Milk EVs Characterization
4.3.1. Western Blotting
4.3.2. Transmission Electron Microscopy (TEM) and Nanoparticle Tracking Assay (NTA)
4.4. RNA Extraction and Library Preparation
4.5. Bioinformatic Analysis
4.5.1. From Sequencing to Datasets
4.5.2. Differential Gene Expression Analysis between the Three Species
4.5.3. miRNA Targets Retrieving and Functional Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Verduci, E.; Giannì, M.L.; Vizzari, G.; Vizzuso, S.; Cerasani, J.; Mosca, F.; Zuccotti, G.V. The Triad Mother-Breast Milk-Infant as Predictor of Future Health: A Narrative Review. Nutrients 2021, 13, 486. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular Vesicles: Exosomes, Microvesicles, and Friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [Green Version]
- Robbins, P.D.; Morelli, A.E. Regulation of Immune Responses by Extracellular Vesicles. Nat. Rev. Immunol. 2014, 14, 195–208. [Google Scholar] [CrossRef] [Green Version]
- Mentkowski, K.I.; Snitzer, J.D.; Rusnak, S.; Lang, J.K. Therapeutic Potential of Engineered Extracellular Vesicles. AAPS J. 2018, 20, 50. [Google Scholar] [CrossRef] [Green Version]
- Ma, Z.; Wang, Y.; Li, H. Applications of Extracellular Vesicles in Tissue Regeneration. Biomicrofluidics 2020, 14, 011501. [Google Scholar] [CrossRef]
- Vella, L.J.; Hill, A.F.; Cheng, L. Focus on Extracellular Vesicles: Exosomes and Their Role in Protein Trafficking and Biomarker Potential in Alzheimer’s and Parkinson’s Disease. Int. J. Mol. Sci. 2016, 17, 173. [Google Scholar] [CrossRef]
- Lucchetti, D.; Ricciardi Tenore, C.; Colella, F.; Sgambato, A. Extracellular Vesicles and Cancer: A Focus on Metabolism, Cytokines, and Immunity. Cancers 2020, 12, 171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, X.; Yuan, T.; Tschannen, M.; Sun, Z.; Jacob, H.; Du, M.; Liang, M.; Dittmar, R.L.; Liu, Y.; Liang, M.; et al. Characterization of Human Plasma-Derived Exosomal RNAs by Deep Sequencing. BMC Genom. 2013, 14, 319. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Greening, D.W.; Gopal, S.K.; Xu, R.; Simpson, R.J.; Chen, W. Exosomes and Their Roles in Immune Regulation and Cancer. Semin. Cell Dev. Biol. 2015, 40, 72–81. [Google Scholar] [CrossRef]
- Dickhout, A.; Koenen, R.R. Extracellular Vesicles as Biomarkers in Cardiovascular Disease; Chances and Risks. Front. Cardiovasc. Med. 2018, 5, 113. [Google Scholar] [CrossRef]
- Burrello, J.; Monticone, S.; Gai, C.; Gomez, Y.; Kholia, S.; Camussi, G. Stem Cell-Derived Extracellular Vesicles and Immune-Modulation. Front. Cell Dev. Biol. 2016, 4, 83. [Google Scholar] [CrossRef] [Green Version]
- Aryani, A.; Denecke, B. Exosomes as a Nanodelivery System: A Key to the Future of Neuromedicine? Mol. Neurobiol. 2016, 53, 818–834. [Google Scholar] [CrossRef] [Green Version]
- Capomaccio, S.; Cappelli, K.; Bazzucchi, C.; Coletti, M.; Gialletti, R.; Moriconi, F.; Passamonti, F.; Pepe, M.; Petrini, S.; Mecocci, S.; et al. Equine Adipose-Derived Mesenchymal Stromal Cells Release Extracellular Vesicles Enclosing Different Subsets of Small RNAs. Stem Cells Int. 2019, 2019, 1–12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding Light on the Cell Biology of Extracellular Vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Kalluri, R.; LeBleu, V.S. The Biology, Function, and Biomedical Applications of Exosomes. Science 2020, 367, eaau6977. [Google Scholar] [CrossRef]
- Melnik, B.C.; Stremmel, W.; Weiskirchen, R.; John, S.M.; Schmitz, G. Exosome-Derived MicroRNAs of Human Milk and Their Effects on Infant Health and Development. Biomolecules 2021, 11, 851. [Google Scholar] [CrossRef]
- Wang, Q.; Zhuang, X.; Mu, J.; Deng, Z.-B.; Jiang, H.; Zhang, L.; Xiang, X.; Wang, B.; Yan, J.; Miller, D.; et al. Delivery of Therapeutic Agents by Nanoparticles Made of Grapefruit-Derived Lipids. Nat. Commun. 2013, 4, 1867. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Munagala, R.; Aqil, F.; Jeyabalan, J.; Gupta, R.C. Bovine Milk-Derived Exosomes for Drug Delivery. Cancer Lett. 2016, 371, 48–61. [Google Scholar] [CrossRef] [Green Version]
- Lin, D.; Chen, T.; Xie, M.; Li, M.; Zeng, B.; Sun, R.; Zhu, Y.; Ye, D.; Wu, J.; Sun, J.; et al. Oral Administration of Bovine and Porcine Milk Exosome Alter MiRNAs Profiles in Piglet Serum. Sci. Rep. 2020, 10, 6983. [Google Scholar] [CrossRef] [PubMed]
- Izumi, H.; Tsuda, M.; Sato, Y.; Kosaka, N.; Ochiya, T.; Iwamoto, H.; Namba, K.; Takeda, Y. Bovine Milk Exosomes Contain MicroRNA and MRNA and Are Taken up by Human Macrophages. J. Dairy Sci. 2015, 98, 2920–2933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- El-Andaloussi, S.; Lee, Y.; Lakhal-Littleton, S.; Li, J.; Seow, Y.; Gardiner, C.; Alvarez-Erviti, L.; Sargent, I.L.; Wood, M.J.A. Exosome-Mediated Delivery of SiRNA in Vitro and in Vivo. Nat. Protoc. 2012, 7, 2112–2126. [Google Scholar] [CrossRef] [PubMed]
- Benmoussa, A.; Laugier, J.; Beauparlant, C.J.; Lambert, M.; Droit, A.; Provost, P. Complexity of the MicroRNA Transcriptome of Cow Milk and Milk-Derived Extracellular Vesicles Isolated via Differential Ultracentrifugation. J. Dairy Sci. 2020, 103, 16–29. [Google Scholar] [CrossRef]
- Admyre, C.; Johansson, S.M.; Qazi, K.R.; Filen, J.-J.; Lahesmaa, R.; Norman, M.; Neve, E.P.A.; Scheynius, A.; Gabrielsson, S. Exosomes with Immune Modulatory Features Are Present in Human Breast Milk. J. Immunol. 2007, 179, 1969–1978. [Google Scholar] [CrossRef]
- Hata, T.; Murakami, K.; Nakatani, H.; Yamamoto, Y.; Matsuda, T.; Aoki, N. Isolation of Bovine Milk-Derived Microvesicles Carrying MRNAs and MicroRNAs. Biochem. Biophys. Res. Commun. 2010, 396, 528–533. [Google Scholar] [CrossRef]
- Chen, T.; Xie, M.-Y.; Sun, J.-J.; Ye, R.-S.; Cheng, X.; Sun, R.-P.; Wei, L.-M.; Li, M.; Lin, D.-L.; Jiang, Q.-Y.; et al. Porcine Milk-Derived Exosomes Promote Proliferation of Intestinal Epithelial Cells. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Badawy, A.A.; El-Magd, M.A.; AlSadrah, S.A. Therapeutic Effect of Camel Milk and Its Exosomes on MCF7 Cells In Vitro and In Vivo. Integr. Cancer Ther. 2018, 17, 1235–1246. [Google Scholar] [CrossRef] [Green Version]
- Baddela, V.S.; Nayan, V.; Rani, P.; Onteru, S.K.; Singh, D. Physicochemical Biomolecular Insights into Buffalo Milk-Derived Nanovesicles. Appl. Biochem. Biotechnol. 2016, 178, 544–557. [Google Scholar] [CrossRef]
- Quan, S.; Nan, X.; Wang, K.; Jiang, L.; Yao, J.; Xiong, B. Characterization of Sheep Milk Extracellular Vesicle-MiRNA by Sequencing and Comparison with Cow Milk. Animals 2020, 10, 331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Golan-Gerstl, R.; Elbaum Shiff, Y.; Moshayoff, V.; Schecter, D.; Leshkowitz, D.; Reif, S. Characterization and Biological Function of Milk-Derived MiRNAs. Mol. Nutr. Food Res. 2017, 61, 1700009. [Google Scholar] [CrossRef]
- Zhao, S.; Wang, Y.; Liang, Y.; Zhao, M.; Long, H.; Ding, S.; Yin, H.; Lu, Q. MicroRNA-126 Regulates DNA Methylation in CD4+ T Cells and Contributes to Systemic Lupus Erythematosus by Targeting DNA Methyltransferase 1. Arthritis Rheum. 2011, 63, 1376–1386. [Google Scholar] [CrossRef]
- Quintanilha, B.; Reis, B.; Duarte, G.; Cozzolino, S.; Rogero, M. Nutrimiromics: Role of MicroRNAs and Nutrition in Modulating Inflammation and Chronic Diseases. Nutrients 2017, 9, 1168. [Google Scholar] [CrossRef] [PubMed]
- Trinchese, G.; Cavaliere, G.; De Filippo, C.; Aceto, S.; Prisco, M.; Chun, J.T.; Penna, E.; Negri, R.; Muredda, L.; Demurtas, A.; et al. Human Milk and Donkey Milk, Compared to Cow Milk, Reduce Inflammatory Mediators and Modulate Glucose and Lipid Metabolism, Acting on Mitochondrial Function and Oleylethanolamide Levels in Rat Skeletal Muscle. Front. Physiol. 2018, 9, 32. [Google Scholar] [CrossRef] [Green Version]
- Benmoussa, A.; Lee, C.H.C.; Laffont, B.; Savard, P.; Laugier, J.; Boilard, E.; Gilbert, C.; Fliss, I.; Provost, P. Commercial Dairy Cow Milk MicroRNAs Resist Digestion under Simulated Gastrointestinal Tract Conditions. J. Nutr. 2016, 146, 2206–2215. [Google Scholar] [CrossRef] [PubMed]
- Kusuma, R.J.; Manca, S.; Friemel, T.; Sukreet, S.; Nguyen, C.; Zempleni, J. Human Vascular Endothelial Cells Transport Foreign Exosomes from Cow’s Milk by Endocytosis. Am. J. Physiol. Cell Physiol. 2016, 310, C800–C807. [Google Scholar] [CrossRef] [Green Version]
- Alsaweed, M.; Hartmann, P.; Geddes, D.; Kakulas, F. MicroRNAs in Breastmilk and the Lactating Breast: Potential Immunoprotectors and Developmental Regulators for the Infant and the Mother. Int. J. Environ. Res. Public Health 2015, 12, 13981–14020. [Google Scholar] [CrossRef] [Green Version]
- Liao, Y.; Du, X.; Li, J.; Lönnerdal, B. Human Milk Exosomes and Their MicroRNAs Survive Digestion in Vitro and Are Taken up by Human Intestinal Cells. Mol. Nutr. Food Res. 2017, 61, 1700082. [Google Scholar] [CrossRef]
- Hou, J.; An, X.; Song, Y.; Cao, B.; Yang, H.; Zhang, Z.; Shen, W.; Li, Y. Detection and Comparison of MicroRNAs in the Caprine Mammary Gland Tissues of Colostrum and Common Milk Stages. BMC Genet. 2017, 18, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, R.; Dudemaine, P.-L.; Zhao, X.; Lei, C.; Ibeagha-Awemu, E.M. Comparative Analysis of the MiRNome of Bovine Milk Fat, Whey and Cells. PloS ONE 2016, 11, e0154129. [Google Scholar] [CrossRef] [Green Version]
- Sedykh, S.E.; Purvinish, L.V.; Monogarov, A.S.; Burkova, E.E.; Grigor’eva, A.E.; Bulgakov, D.V.; Dmitrenok, P.S.; Vlassov, V.V.; Ryabchikova, E.I.; Nevinsky, G.A. Purified Horse Milk Exosomes Contain an Unpredictable Small Number of Major Proteins. Biochim. Open 2017, 4, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C.; Schmitz, G. MicroRNAs: Milk’s Epigenetic Regulators. Best Pract. Res. Clin. Endocrinol. Metab. 2017, 31, 427–442. [Google Scholar] [CrossRef]
- Wang, J.; Bian, Y.; Wang, Z.; Li, D.; Wang, C.; Li, Q.; Gao, X. MicroRNA-152 Regulates DNA Methyltransferase 1 and Is Involved in the Development and Lactation of Mammary Glands in Dairy Cows. PLoS ONE 2014, 9, e101358. [Google Scholar] [CrossRef] [PubMed]
- Contreras, J.; Rao, D.S. MicroRNAs in Inflammation and Immune Responses. Leukemia 2012, 26, 404–413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hulsmans, M.; De Keyzer, D.; Holvoet, P. MicroRNAs Regulating Oxidative Stress and Inflammation in Relation to Obesity and Atherosclerosis. FASEB J. 2011, 25, 2515–2527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ocansey, D.K.W.; Zhang, L.; Wang, Y.; Yan, Y.; Qian, H.; Zhang, X.; Xu, W.; Mao, F. Exosome-mediated Effects and Applications in Inflammatory Bowel Disease. Biol. Rev. 2020, 95, 1287–1307. [Google Scholar] [CrossRef]
- Ghafouri-Fard, S.; Eghtedarian, R.; Taheri, M. The Crucial Role of Non-Coding RNAs in the Pathophysiology of Inflammatory Bowel Disease. Biomed. Pharmacother. 2020, 129, 110507. [Google Scholar] [CrossRef]
- Lin, L.; Zhou, G.; Chen, P.; Wang, Y.; Han, J.; Chen, M.; He, Y.; Zhang, S. Which Long Noncoding RNAs and Circular RNAs Contribute to Inflammatory Bowel Disease? Cell Death Dis. 2020, 11, 456. [Google Scholar] [CrossRef]
- Harrison’s Principles of Internal Medicine, 18e|AccessMedicine|McGraw-Hill Medical. Available online: http://accessmedicine.mhmedical.com/book.aspx?bookid=331 (accessed on 26 March 2018).
- Ortega, F.J.; Cardona-Alvarado, M.I.; Mercader, J.M.; Moreno-Navarrete, J.M.; Moreno, M.; Sabater, M.; Fuentes-Batllevell, N.; Ramírez-Chávez, E.; Ricart, W.; Molina-Torres, J.; et al. Circulating Profiling Reveals the Effect of a Polyunsaturated Fatty Acid-Enriched Diet on Common MicroRNAs. J. Nutr. Biochem. 2015, 26, 1095–1101. [Google Scholar] [CrossRef]
- Tian, L.; Song, Z.; Shao, W.; Du, W.W.; Zhao, L.R.; Zeng, K.; Yang, B.B.; Jin, T. Curcumin Represses Mouse 3T3-L1 Cell Adipogenic Differentiation via Inhibiting MiR-17-5p and Stimulating the Wnt Signalling Pathway Effector Tcf7l2. Cell Death Dis. 2017, 8, e2559. [Google Scholar] [CrossRef] [Green Version]
- Tomé-Carneiro, J.; Larrosa, M.; Yáñez-Gascón, M.J.; Dávalos, A.; Gil-Zamorano, J.; Gonzálvez, M.; García-Almagro, F.J.; Ruiz Ros, J.A.; Tomás-Barberán, F.A.; Espín, J.C.; et al. One-Year Supplementation with a Grape Extract Containing Resveratrol Modulates Inflammatory-Related MicroRNAs and Cytokines Expression in Peripheral Blood Mononuclear Cells of Type 2 Diabetes and Hypertensive Patients with Coronary Artery Disease. Pharmacol. Res. 2013, 72, 69–82. [Google Scholar] [CrossRef]
- Braud, M.; Magee, D.A.; Park, S.D.E.; Sonstegard, T.S.; Waters, S.M.; MacHugh, D.E.; Spillane, C. Genome-Wide MicroRNA Binding Site Variation between Extinct Wild Aurochs and Modern Cattle Identifies Candidate MicroRNA-Regulated Domestication Genes. Front. Genet. 2017, 8, 3. [Google Scholar] [CrossRef] [Green Version]
- Mecocci, S.; Gevi, F.; Pietrucci, D.; Cavinato, L.; Luly, F.R.; Pascucci, L.; Petrini, S.; Ascenzioni, F.; Zolla, L.; Chillemi, G.; et al. Anti-Inflammatory Potential of Cow, Donkey and Goat Milk Extracellular Vesicles as Revealed by Metabolomic Profile. Nutrients 2020, 12, 2908. [Google Scholar] [CrossRef] [PubMed]
- Teng, F.; Fussenegger, M. Shedding Light on Extracellular Vesicle Biogenesis and Bioengineering. Adv. Sci. 2021, 8, 2003505. [Google Scholar] [CrossRef]
- Zempleni, J.; Aguilar-Lozano, A.; Sadri, M.; Sukreet, S.; Manca, S.; Wu, D.; Zhou, F.; Mutai, E. Biological Activities of Extracellular Vesicles and Their Cargos from Bovine and Human Milk in Humans and Implications for Infants. J. Nutr. 2017, 147, 3–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, T.; Xi, Q.-Y.; Sun, J.-J.; Ye, R.-S.; Cheng, X.; Sun, R.-P.; Wang, S.-B.; Shu, G.; Wang, L.-N.; Zhu, X.-T.; et al. Revelation of MRNAs and Proteins in Porcine Milk Exosomes by Transcriptomic and Proteomic Analysis. BMC Vet. Res. 2017, 13, 101. [Google Scholar] [CrossRef]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Time-Dependent Expression Profiles of MicroRNAs and MRNAs in Rat Milk Whey. PLoS ONE 2014, 9, e88843. [Google Scholar] [CrossRef]
- Herrera, M.B.; Fonsato, V.; Gatti, S.; Deregibus, M.C.; Sordi, A.; Cantarella, D.; Calogero, R.; Bussolati, B.; Tetta, C.; Camussi, G. Human Liver Stem Cell-Derived Microvesicles Accelerate Hepatic Regeneration in Hepatectomized Rats. J. Cell Mol. Med. 2010, 14, 1605–1618. [Google Scholar] [CrossRef] [Green Version]
- Valadi, H.; Ekström, K.; Bossios, A.; Sjöstrand, M.; Lee, J.J.; Lötvall, J.O. Exosome-Mediated Transfer of MRNAs and MicroRNAs Is a Novel Mechanism of Genetic Exchange between Cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wei, Z.; Batagov, A.O.; Schinelli, S.; Wang, J.; Wang, Y.; El Fatimy, R.; Rabinovsky, R.; Balaj, L.; Chen, C.C.; Hochberg, F.; et al. Coding and Noncoding Landscape of Extracellular RNA Released by Human Glioma Stem Cells. Nat. Commun. 2017, 8, 1145. [Google Scholar] [CrossRef] [Green Version]
- Batagov, A.O.; Kurochkin, I.V. Exosomes Secreted by Human Cells Transport Largely MRNA Fragments That Are Enriched in the 3′-Untranslated Regions. Biol. Direct. 2013, 8, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Liegro, C.M.; Schiera, G.; Di Liegro, I. Extracellular Vesicle-Associated RNA as a Carrier of Epigenetic Information. Genes 2017, 8, 240. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Plotnikov, A.; Zehorai, E.; Procaccia, S.; Seger, R. The MAPK Cascades: Signaling Components, Nuclear Roles and Mechanisms of Nuclear Translocation. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2011, 1813, 1619–1633. [Google Scholar] [CrossRef] [Green Version]
- Qi, H.; Wang, Y.; Fa, S.; Yuan, C.; Yang, L. Extracellular Vesicles as Natural Delivery Carriers Regulate Oxidative Stress Under Pathological Conditions. Front. Bioeng. Biotechnol. 2021, 9, 752019. [Google Scholar] [CrossRef]
- Sanwlani, R.; Fonseka, P.; Chitti, S.V.; Mathivanan, S. Milk-Derived Extracellular Vesicles in Inter-Organism, Cross-Species Communication and Drug Delivery. Proteomes 2020, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Driedonks, T.A.P.; Nolte-’t Hoen, E.N.M. Circulating Y-RNAs in Extracellular Vesicles and Ribonucleoprotein Complexes; Implications for the Immune System. Front. Immunol. 2019, 9, 3164. [Google Scholar] [CrossRef]
- Benmoussa, A.; Provost, P. Milk MicroRNAs in Health and Disease. Compr. Rev. Food Sci. Food Saf. 2019, 18, 703–722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tingö, L.; Ahlberg, E.; Johansson, L.; Pedersen, S.A.; Chawla, K.; Sætrom, P.; Cione, E.; Simpson, M.R. Non-Coding RNAs in Human Breast Milk: A Systematic Review. Front. Immunol. 2021, 12, 725323. [Google Scholar] [CrossRef]
- Van Herwijnen, M.J.C.; Driedonks, T.A.P.; Snoek, B.L.; Kroon, A.M.T.; Kleinjan, M.; Jorritsma, R.; Pieterse, C.M.J.; Nolte-‘t Hoen, E.N.M.; Wauben, M.H.M. Abundantly Present MiRNAs in Milk-Derived Extracellular Vesicles Are Conserved Between Mammals. Front. Nutr. 2018, 5, 81. [Google Scholar] [CrossRef]
- Sheedy, F.J. Turning 21: Induction of MiR-21 as a Key Switch in the Inflammatory Response. Front. Immunol. 2015, 6, 19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Manca, S.; Upadhyaya, B.; Mutai, E.; Desaulniers, A.T.; Cederberg, R.A.; White, B.R.; Zempleni, J. Milk Exosomes Are Bioavailable and Distinct MicroRNA Cargos Have Unique Tissue Distribution Patterns. Sci. Rep. 2018, 8, 11321. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Su, X.; Xu, S.; Wang, H.; Han, D.; Li, J.; Huang, M.; Cao, X. MicroRNA in Vivo Precipitation Identifies MiR-151-3p as a Computational Unpredictable MiRNA to Target Stat3 and Inhibits Innate IL-6 Production. Cell Mol. Immunol. 2018, 15, 99–110. [Google Scholar] [CrossRef]
- Kakimoto, Y.; Matsushima, Y.; Tanaka, M.; Hayashi, H.; Wang, T.; Yokoyama, K.; Ochiai, E.; Osawa, M. MicroRNA Profiling of Gastric Content from Breast-Fed and Formula-Fed Infants to Estimate Last Feeding: A Pilot Study. Int. J. Legal Med. 2020, 134, 903–909. [Google Scholar] [CrossRef]
- Carrillo-Lozano, E.; Sebastián-Valles, F.; Knott-Torcal, C. Circulating MicroRNAs in Breast Milk and Their Potential Impact on the Infant. Nutrients 2020, 12, 3066. [Google Scholar] [CrossRef]
- Zempleni, J. Milk Exosomes: Beyond Dietary MicroRNAs. Genes Nutr. 2017, 12, 1–4. [Google Scholar] [CrossRef]
- Friedrich, M.; Pracht, K.; Mashreghi, M.-F.; Jäck, H.-M.; Radbruch, A.; Seliger, B. The Role of the MiR-148/-152 Family in Physiology and Disease. Eur. J. Immunol. 2017, 47, 2026–2038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szczepankiewicz, D.; Langwiński, W.; Kołodziejski, P.; Pruszyńska-Oszmałek, E.; Sassek, M.; Nowakowska, J.; Chmurzyńska, A.; Nowak, K.W.; Szczepankiewicz, A. Allergic Inflammation Alters MicroRNA Expression Profile in Adipose Tissue in the Rat. Genes 2020, 11, 1034. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.-H.; Shrestha, S.; Yang, C.-D.; Chang, N.-W.; Lin, Y.-L.; Liao, K.-W.; Huang, W.-C.; Sun, T.-H.; Tu, S.-J.; Lee, W.-H.; et al. MiRTarBase Update 2018: A Resource for Experimentally Validated MicroRNA-Target Interactions. Nucleic Acids Res. 2018, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]
- Pion, M.; Jaramillo-Ruiz, D.; Martínez, A.; Muñoz-Fernández, M.A.; Correa-Rocha, R. HIV Infection of Human Regulatory T Cells Downregulates Foxp3 Expression by Increasing DNMT3b Levels and DNA Methylation in the FOXP3 Gene. AIDS 2013, 27, 2019–2029. [Google Scholar] [CrossRef]
- Arntz, O.J.; Pieters, B.C.H.; Oliveira, M.C.; Broeren, M.G.A.; Bennink, M.B.; de Vries, M.; van Lent, P.L.E.M.; Koenders, M.I.; van den Berg, W.B.; van der Kraan, P.M.; et al. Oral Administration of Bovine Milk Derived Extracellular Vesicles Attenuates Arthritis in Two Mouse Models. Mol. Nutr. Food Res. 2015, 59, 1701–1712. [Google Scholar] [CrossRef]
- Melnik, B.C. Milk Exosomal MiRNAs: Potential Drivers of AMPK-to-MTORC1 Switching in β-Cell de-Differentiation of Type 2 Diabetes Mellitus. Nutr. Metab. 2019, 16, 85. [Google Scholar] [CrossRef] [PubMed]
- Stephen, B.J.; Pareek, N.; Saeed, M.; Kausar, M.A.; Rahman, S.; Datta, M. Xeno-MiRNA in Maternal-Infant Immune Crosstalk: An Aid to Disease Alleviation. Front. Immunol. 2020, 11, 404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Z.; Yu, S.; Xu, M.; Li, P. Effects of Microwave on Extracellular Vesicles and MicroRNA in Milk. J. Dairy Sci. 2018, 101, 2932–2940. [Google Scholar] [CrossRef]
- Sun, Q.; Chen, X.; Yu, J.; Zen, K.; Zhang, C.-Y.; Li, L. Immune Modulatory Function of Abundant Immune-Related MicroRNAs in Microvesicles from Bovine Colostrum. Protein Cell 2013, 4, 197–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; Xiao, Y.; Ge, W.; Zhou, K.; Wen, J.; Yan, W.; Wang, Y.; Wang, B.; Qu, C.; Wu, J.; et al. MiR-200b Inhibits TGF-Β1-Induced Epithelial-Mesenchymal Transition and Promotes Growth of Intestinal Epithelial Cells. Cell Death Dis. 2013, 4, e541. [Google Scholar] [CrossRef] [Green Version]
- Malhotra, N.; Kang, J. SMAD Regulatory Networks Construct a Balanced Immune System. Immunology 2013, 139, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Nejad, C.; Stunden, H.J.; Gantier, M.P. A Guide to MiRNAs in Inflammation and Innate Immune Responses. FEBS J. 2018, 285, 3695–3716. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.S.; Choi, Y.E.; Hwang, S.J.; Han, Y.-H.; Park, M.-J.; Bae, I.H. IL-4, a Direct Target of MiR-340/429, Is Involved in Radiation-Induced Aggressive Tumor Behavior in Human Carcinoma Cells. Oncotarget 2016, 7, 86836–86856. [Google Scholar] [CrossRef] [Green Version]
- Bian, J.; Liu, R.; Fan, T.; Liao, L.; Wang, S.; Geng, W.; Wang, T.; Shi, W.; Ruan, Q. MiR-340 Alleviates Psoriasis in Mice through Direct Targeting of IL-17A. J. Immunol. 2018, 201, 1412–1420. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, H.; Neurath, M.F.; Atreya, R. Role of the IL23/IL17 Pathway in Crohn’s Disease. Front. Immunol. 2021, 12, 622934. [Google Scholar] [CrossRef] [PubMed]
- Xie, W.; Li, M.; Xu, N.; Lv, Q.; Huang, N.; He, J.; Zhang, Y. MiR-181a Regulates Inflammation Responses in Monocytes and Macrophages. PLoS ONE 2013, 8, e58639. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Su, Y.; Yuan, J.; Zhang, F.; Lei, Q.; Zhang, T.; Li, K.; Guo, J.; Hong, Y.; Bu, G.; Lv, X.; et al. MicroRNA-181a-5p and MicroRNA-181a-3p Cooperatively Restrict Vascular Inflammation and Atherosclerosis. Cell Death Dis. 2019, 10, 365. [Google Scholar] [CrossRef] [Green Version]
- Jaiswal, A.; Reddy, S.S.; Maurya, M.; Maurya, P.; Barthwal, M.K. MicroRNA-99a Mimics Inhibit M1 Macrophage Phenotype and Adipose Tissue Inflammation by Targeting TNFα. Cell Mol. Immunol. 2019, 16, 495–507. [Google Scholar] [CrossRef]
- Zhu, M.; Li, Y.; Sun, K. MicroRNA-182-5p Inhibits Inflammation in LPS-Treated RAW264.7 Cells by Mediating the TLR4/NF-ΚB Signaling Pathway. Int. J. Clin. Exp. Pathol. 2018, 11, 5725–5734. [Google Scholar]
- Kawasaki, H.; Amano, H. Anti-Inflammatory Role of MicroRNA-429 in Human Gingival Epithelial Cells-Inhibition of IL-8 Production through Direct Binding to IKKβ MRNA. Mol. Med. Rep. 2021, 24, 581. [Google Scholar] [CrossRef] [PubMed]
- Mo, J.-S.; Alam, K.J.; Kim, H.-S.; Lee, Y.-M.; Yun, K.-J.; Chae, S.-C. MicroRNA 429 Regulates Mucin Gene Expression and Secretion in Murine Model of Colitis. J. Crohn’s Colitis 2016, 10, 837–849. [Google Scholar] [CrossRef] [Green Version]
- López de las Hazas, M.-C.; del Pozo-Acebo, L.; Hansen, M.S.; Gil-Zamorano, J.; Mantilla-Escalante, D.C.; Gómez-Coronado, D.; Marín, F.; Garcia-Ruiz, A.; Rasmussen, J.T.; Dávalos, A. Dietary Bovine Milk MiRNAs Transported in Extracellular Vesicles Are Partially Stable during GI Digestion, Are Bioavailable and Reach Target Tissues but Need a Minimum Dose to Impact on Gene Expression. Eur. J. Nutr. 2021. online ahead of print. [Google Scholar] [CrossRef]
- Del Pozo-Acebo, L.; López de las Hazas, M.-C.; Tomé-Carneiro, J.; Gil-Cabrerizo, P.; San-Cristobal, R.; Busto, R.; García-Ruiz, A.; Dávalos, A. Bovine Milk-Derived Exosomes as a Drug Delivery Vehicle for MiRNA-Based Therapy. IJMS Int. J. Mol. Sci. 2021, 22, 1105. [Google Scholar] [CrossRef]
- Vaswani, K.; Mitchell, M.D.; Holland, O.J.; Qin Koh, Y.; Hill, R.J.; Harb, T.; Davies, P.S.W.; Peiris, H. A Method for the Isolation of Exosomes from Human and Bovine Milk. J. Nutr. Metab. 2019, 2019, 5764740. [Google Scholar] [CrossRef]
- Botta, L.; Filippi, S.; Bizzarri, B.M.; Meschini, R.; Caputo, M.; Proietti-De-Santis, L.; Iside, C.; Nebbioso, A.; Gualandi, G.; Saladino, R. Oxidative Nucleophilic Substitution Selectively Produces Cambinol Derivatives with Antiproliferative Activity on Bladder Cancer Cell Lines. Bioorg. Med. Chem. Lett. 2019, 29, 78–82. [Google Scholar] [CrossRef]
- Cavinato, L.; Genise, E.; Luly, F.R.; Di Domenico, E.G.; Del Porto, P.; Ascenzioni, F. Escaping the Phagocytic Oxidative Burst: The Role of SODB in the Survival of Pseudomonas Aeruginosa Within Macrophages. Front. Microbiol. 2020, 11, 326. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Howe, K.L.; Achuthan, P.; Allen, J.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Azov, A.G.; Bennett, R.; Bhai, J.; et al. Ensembl 2021. Nucleic Acids Res. 2021, 49, D884–D891. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [Green Version]
- Langmead, B.; Salzberg, S.L. Fast Gapped-Read Alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. MiRBase: From MicroRNA Sequences to Function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef] [PubMed]
- Leinonen, R.; Sugawara, H.; Shumway, M. The Sequence Read Archive. Nucleic Acids Res. 2011, 39, D19–D21. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Wang, S.; Li, W. RSeQC: Quality Control of RNA-Seq Experiments. Bioinformatics 2012, 28, 2184–2185. [Google Scholar] [CrossRef] [Green Version]
- Darbellay, F.; Necsulea, A. Comparative Transcriptomics Analyses across Species, Organs, and Developmental Stages Reveal Functionally Constrained LncRNAs. Mol. Biol. Evol. 2020, 37, 240–259. [Google Scholar] [CrossRef] [Green Version]
- Brawand, D.; Soumillon, M.; Necsulea, A.; Julien, P.; Csárdi, G.; Harrigan, P.; Weier, M.; Liechti, A.; Aximu-Petri, A.; Kircher, M.; et al. The Evolution of Gene Expression Levels in Mammalian Organs. Nature 2011, 478, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15. [Google Scholar] [CrossRef] [Green Version]
- Blighe, K.; Rana, S.; Turkes, E.; Ostendorf, B.; Grioni, A.; Lewis, M. EnhancedVolcano: Publication-Ready Volcano Plots with Enhanced Colouring and Labeling Bioconductor Version Release (3.13); 2021; Available online: https://www.bioconductor.org/packages/release/bioc/vignettes/EnhancedVolcano/inst/doc/EnhancedVolcano.htmL (accessed on 1 October 2021).
- Bindea, G.; mLecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.-H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape Plug-in to Decipher Functionally Grouped Gene Ontology and Pathway Annotation Networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [Green Version]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- The IMEx Consortium Curators; del-Toro, N.; Duesbury, M.; Koch, M.; Perfetto, L.; Shrivastava, A.; Ochoa, D.; Wagih, O.; Piñero, J.; Kotlyar, M.; et al. Capturing Variation Impact on Molecular Interactions in the IMEx Consortium Mutations Data Set. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Morris, J.H.; Apeltsin, L.; Newman, A.M.; Baumbach, J.; Wittkop, T.; Su, G.; Bader, G.D.; Ferrin, T.E. ClusterMaker: A Multi-Algorithm Clustering Plugin for Cytoscape. BMC Bioinform. 2011, 12, 436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
| Enclosing mEVs | No. of miRNAs | miRNAs | ||
|---|---|---|---|---|
| Cow–Donkey–Goat | 10 | miR-151a miR-191 miR-30b let-7i | miR-27b miR-30d miR-186 | miR-200b miR-148b miR-21 |
| Cow–Goat | 15 | miR-141 miR-23b miR-93 miR-25 miR-361 | let-7b miR-660 let-7g miR-30a miR-200a | miR-125a miR-20a miR-29a miR-140 miR-200c |
| Donkey–Goat | 6 | miR-148a miR-143 | miR-26b miR-30e | miR-223 miR-155 |
| Cow–Donkey | 1 | miR-375 | ||
| Cow | 10 | miR-340 mir181a miR-362 miR-99a | miR-532 miR-26b miR-182 | miR-429 miR-425 miR-34a |
| Donkey | 10 | miR-374a miR-152 miR-101 miR-98 | miR-590 miR-320a let-7d | miR-23a miR-145 let-7c |
| Goat | 8 | miR-26a let-7f miR-27a | miR-22 miR-224 miR-16 | miR-125b miR-190a |
| Species | 3′-UTR Targets | 5′-UTR Targets | CDS Targets | Total Targets | Filtered Targets |
|---|---|---|---|---|---|
| Cow | 1351 | 221 | 1196 | 2296 | 3 |
| Donkey | 814 | 109 | 603 | 1313 | 15 |
| Goat | 1537 | 256 | 1283 | 2568 | 14 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mecocci, S.; Pietrucci, D.; Milanesi, M.; Pascucci, L.; Filippi, S.; Rosato, V.; Chillemi, G.; Capomaccio, S.; Cappelli, K. Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential. Int. J. Mol. Sci. 2021, 22, 12759. https://doi.org/10.3390/ijms222312759
Mecocci S, Pietrucci D, Milanesi M, Pascucci L, Filippi S, Rosato V, Chillemi G, Capomaccio S, Cappelli K. Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential. International Journal of Molecular Sciences. 2021; 22(23):12759. https://doi.org/10.3390/ijms222312759
Chicago/Turabian StyleMecocci, Samanta, Daniele Pietrucci, Marco Milanesi, Luisa Pascucci, Silvia Filippi, Vittorio Rosato, Giovanni Chillemi, Stefano Capomaccio, and Katia Cappelli. 2021. "Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential" International Journal of Molecular Sciences 22, no. 23: 12759. https://doi.org/10.3390/ijms222312759
APA StyleMecocci, S., Pietrucci, D., Milanesi, M., Pascucci, L., Filippi, S., Rosato, V., Chillemi, G., Capomaccio, S., & Cappelli, K. (2021). Transcriptomic Characterization of Cow, Donkey and Goat Milk Extracellular Vesicles Reveals Their Anti-Inflammatory and Immunomodulatory Potential. International Journal of Molecular Sciences, 22(23), 12759. https://doi.org/10.3390/ijms222312759










