On the Edge of Dispensability, the Chloroplast ndh Genes
Abstract
1. Introduction
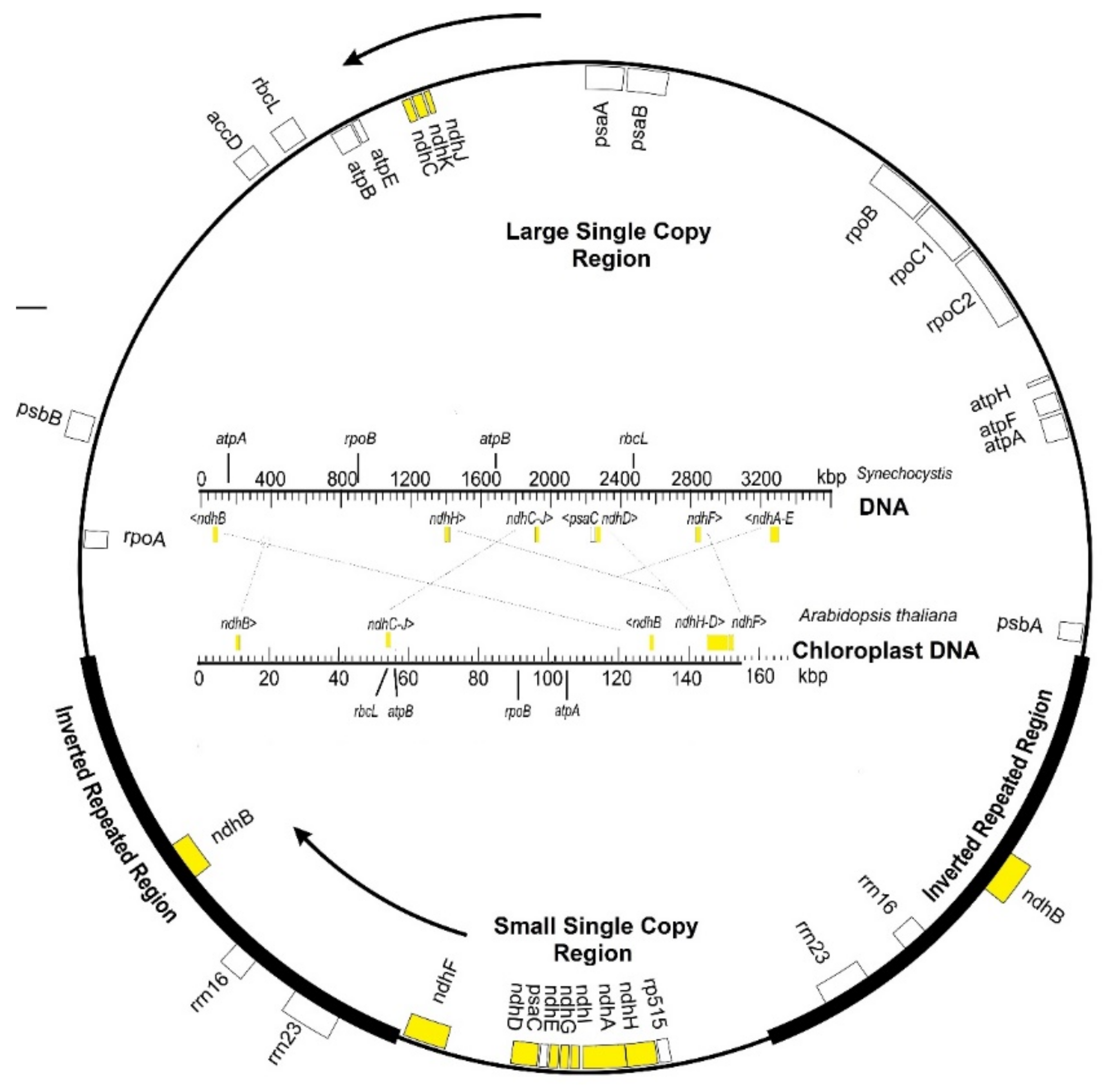
2. Chloroplast ndh Genes
3. Functional Role of the Thylakoid Ndh Complex
4. Dispensing with the Role of the ndh Genes
5. Plant Death or Species Extinction, a ndh Dilemma?
6. Concluding Remarks and Future Perspectives
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ohyama, K.; Fukuzawa, H.; Kochi, T.; Shira, H.; Sano, T.; Umesono, K.; Shiki, Y.; Takeuchi, M.; Chang, Z.; Aota, S.; et al. Chloroplast gene organization deduced from complete sequence of liverwort Marchantia polymorpha chloroplast DNA. Nature 1986, 322, 572–574. [Google Scholar] [CrossRef]
- Shinozaki, K.; Ohme, M.; Tanaka, M.; Wakasugi, T.; Hayashida, N.; Matsubayashi, T.; Zaita, N.; Chunwongse, J.; Obokata, J.; Yamaguchi-Shinozaki, K.; et al. The complete nucleotide sequence of the tobacco chloroplast genome. Its gene organization and expression. EMBO J. 1986, 5, 2043–2049. [Google Scholar] [CrossRef]
- Martin, W.; Herrmann, R.G. Gene transfer from organelles to nucleus: How much, what happens, and why? Plant Physiol. 1998, 118, 9–17. [Google Scholar] [CrossRef]
- Martin, W.; Rujan, T.; Richly, E.; Hansen, A.; Cornelsen, S.; Lins, T.; Leister, D.; Stoebe, B.; Hasegawa, M.; Penny, D. Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc. Natl. Acad. Sci. USA 2002, 99, 12246–12251. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Morden, C.W.; Palmer, J.D. Function and evolution of a minimal plastid genome from a non-photosynthetic parasitic plant. Proc. Natl. Acad. Sci. USA 1992, 89, 10648–10652. [Google Scholar] [CrossRef]
- Haberhausen, G.; Zetsche, K. Functional loss of all ndh genes in an otherwise relatively unaltered plastid genome of the holoparasitic flowering plant Cuscuta reflexa. Plant Mol. Biol. 1994, 24, 217–222. [Google Scholar] [CrossRef]
- Funk, H.T.; Berg, S.; Krupinska, K.; Maier, U.G.; Krause, K. Complete DNA sequences of the plastid genomes of two parasitic flowering plant species, Cuscuta reflexa and Cuscuta gronovii. BMC Plant Biol. 2007, 7, 45. [Google Scholar] [CrossRef]
- Sazanov, L.; Burrows, P.A.; Nixon, P.J. The plastid ndh genes code for an NADH-specific dehydrogenase: Purification and characterization of a mitochondrial-like complex I from pea thylakoid membranes. Proc. Natl. Acad. Sci. USA 1998, 95, 1319–1324. [Google Scholar] [CrossRef]
- Casano, L.M.; Zapata, J.M.; Martín, M.; Sabater, B. Chlororespiration and poising of cyclic electron transport: Plastoquinone as electron transporter between thylakoid NADH dehydrogenase and peroxidase. J. Biol. Chem. 2000, 275, 942–948. [Google Scholar] [CrossRef]
- Martín, M.; Sabater, B. Plastid ndh genes in plant evolution. Plant Physiol. Biochem. 2010, 48, 636–645. [Google Scholar] [CrossRef]
- Martín, M.; Casano, L.M.; Sabater, B. Identification of the product of ndhA gene as a thylakoid protein synthesized in response to photooxidative treatment. Plant Cell Physiol. 1996, 37, 293–298. [Google Scholar] [CrossRef]
- Martín, M.; Casano, L.M.; Zapata, J.M.; Guéra, A.; del Campo, E.M.; Schmitz-Linneweber, C.; Maier, R.M.; Sabater, B. Role of thylakoid Ndh complex and peroxidase in the protection against photo-oxidative stress: Fluorescence and enzyme activities in wild-type and ndhF-deficient tobacco. Physiol. Plant. 2004, 122, 443–452. [Google Scholar] [CrossRef]
- Friedrich, T.; Scheide, D. The respiratory complex I of bacteria, archaea and eukarya and its module common with membrane-bound multisubunit hydrogenases. FEBS Lett. 2000, 479, 1–5. [Google Scholar] [CrossRef]
- Freyer, R.; López, C.; Maier, R.M.; Martín, M.; Sabater, B.; Kossel, H. Editing of chloroplast ndhB encoded transcript shows divergence between closely related members of the grass family (poaceae). Plant Mol. Biol. 1995, 29, 679–684. [Google Scholar] [CrossRef]
- Maier, R.M.; Neckermann, K.; Igloi, G.L.; Kossel, H. Complete sequence of the maize chloroplast genome: Gene content, hotspots of divergence and fine tuning of genetic information by transcript editing. J. Mol. Biol. 1995, 251, 614–628. [Google Scholar] [CrossRef]
- Hiratsuka, J.; Shimada, H.; Whittier, R.; Ishibashi, T.; Sakamoto, M.; Mori, M.; Kondo, C.; Honji, Y.; Sun, C.-R.; Meng, B.-Y.; et al. The complete sequence of the rice (Oryza sativa) chloroplast genome: Intermolecular recombination between distinct tRNA genes accounts for a major plastid DNA inversion during the evolution of the cereals. Mol. Gen. Genet. 1989, 217, 185–194. [Google Scholar] [CrossRef]
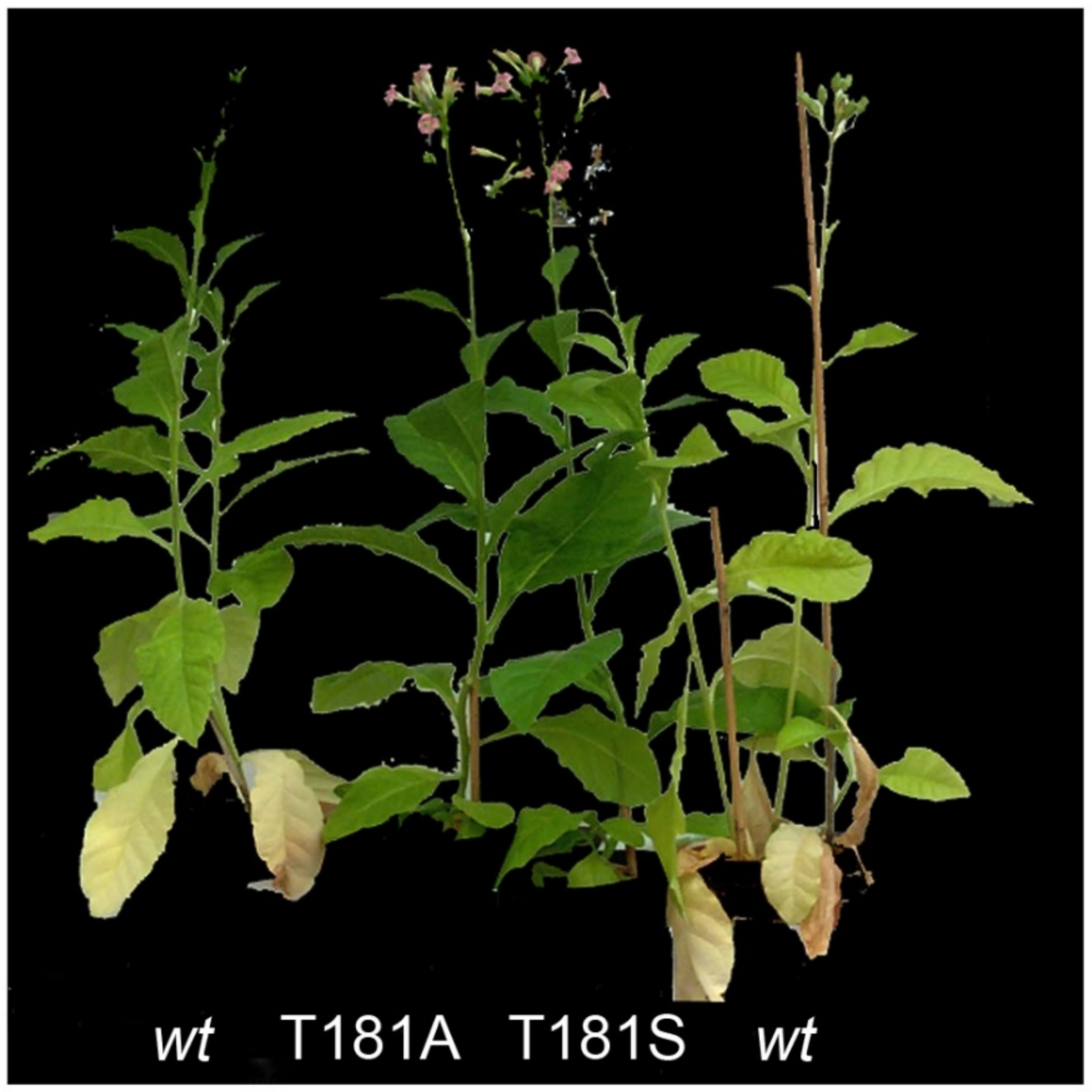
- Martín, M.; Funk, H.T.; Serrot, P.H.; Poltnigg, P.; Sabater, B. Functional characterization of the thylakoid Ndh complex phosphorylation by site-directed mutations in the ndhF gene. Biochim. Biophys. Acta BBA Bioenerg. 2009, 1787, 920–928. [Google Scholar] [CrossRef]
- Martínez, P.; López, C.; Roldán, M.; Sabater, B.; Martín, M. Plastid DNA of five ecotypes of Arabidopsis thaliana: Sequence of ndhG gene and maternal inheritance. Plant Sci. 1997, 123, 113–122. [Google Scholar] [CrossRef]
- Berger, S.; Ellersiek, U.; Westhoff, P.; Steinmuller, K. Studies on the expression of NDH-H, a subunit of the NAD(P)H-plastoquinone-oxidoreductase of higher plant chloroplasts. Planta 1993, 190, 25–31. [Google Scholar] [CrossRef]
- Burrows, P.A.; Sazanov, L.A.; Svab, Z.; Maliga, P.; Nixon, P.J. Identification of a functional respiratory complex in chloroplasts through analysis of tobacco mutants containing disrupted plastid ndh genes. EMBO J. 1998, 17, 868–871. [Google Scholar] [CrossRef]
- Barrett, C.F.; Freudenstein, J.V.; Li, J.; Mayfield-Jones, D.R.; Perez, L.; Santos, C. Investigating the path of plastid genome degradation in an early-transitional clade of heterotrophic orchids, and implications for heterotrophic angiosperms. Mol. Biol. Evol. 2014, 31, 3095–3112. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Hou, B.-W.; Niu, Z.-T.; Liu, W.; Xue, Q.-Y.; Ding, X.-Y. Comparative chloroplast genomes of photosynthetic orchids: Insights into evolution of the Orchidaceae and development of molecular markers for phylogenetic applications. PLoS ONE 2014, 9, e99016. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.S.; Chen, J.J.; Huang, Y.T.; Chan, M.T.; Daniell, H.; Chang, W.J.; Hsu, C.T.; Liao, D.C.; Wu, F.H.; Lin, S.Y.; et al. The location and translocation of ndh genes of chloroplast origin in the Orchidaceae family. Sci. Rep. 2015, 5, 9040. [Google Scholar] [CrossRef] [PubMed]
- Wakasugi, T.; Tsudzuki, J.; Ito, S.; Nakashima, K.; Tsudzuki, T.; Sugiura, M. Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc. Natl. Acad. Sci. USA 1994, 91, 9794–9798. [Google Scholar] [CrossRef]
- Braukmann, T.W.A.; Kuzmina, M.; Stefanovíc, S. Loss of all ndh genes in Gnetales and conifers: Extent and evolutionary significance for seed plant phylogeny. Curr. Genet. 2009, 55, 323–337. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.S.; Wang, Y.N.; Liu, S.M.; Chaw, S.M. Chloroplast genome (cpDNA) of Cycas taitungensis and 56 Cp Protein-coding genes of Gnetum parvifolium: Insights into CpDNA evolution and phylogeny of extant seed plants. Mol. Biol. Evol. 2007, 24, 1366–1379. [Google Scholar] [CrossRef] [PubMed]
- Hirao, T.; Watanabe, A.; Kurita, M.; Kondo, T.; Takata, K. Complete nucleotide sequence of the Cryptomeria japonica D. Don. chloroplast genome and comparative chloroplast genomics: Diversified genomic structure of coniferous species. BMC Plant Biol. 2008, 8, 70. [Google Scholar] [CrossRef] [PubMed]
- Serrot, P.H.; Sabater, B.; Martín, M. Activity, polypeptide and gene identification of thylakoid Ndh complex in trees: Potential physiological relevance of fluorescence assays. Physiol. Plant. 2012, 146, 110–120. [Google Scholar] [CrossRef]
- Blazier, J.C.; Guisinger, M.M.; Jansen, R.K. Recent loss of plastid-encoded ndh genes within Erodium (Geraniaceae). Plant Mol. Biol. 2011, 76, 263–272. [Google Scholar] [CrossRef]
- Braukmann, T.W.A.; Stefanovíc, S. Plastid genome evolution in mycoheterotrophic Ericaceae. Plant Mol. Biol. 2012, 79, 5–20. [Google Scholar] [CrossRef]
- Peredo, E.L.; King, U.M.; Les, D.H. The plastid genome of Najas flexilis: Adaptation to submersed environments is accompanied by the complete loss of the NDH Complex in an Aquatic Angiosperm. PLoS ONE 2013, 8, e68591. [Google Scholar] [CrossRef]
- Ruhlman, T.A.; Chang, W.J.; Chen, J.J.; Huang, Y.T.; Chan, M.T.; Zhang, J.; Liao, D.C.; Blazier, J.C.; Jin, X.; Shih, M.C.; et al. NDH expression marks major transitions in plant evolution and reveals coordinate intracellular gene loss. BMC Plant Biol. 2015, 15, 100. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.-L.; Wang, R.-N.; Zhang, N.-Y.; Fan, W.-B.; Fang, M.-F.; Li, Z.-H. Molecular evolution of chloroplast genomes of orchid species: Insights into phylogenetic relationship and adaptive evolution. Int. J. Mol. Sci. 2018, 19, 716. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.-S.; Fu, P.-C.; Zhou, X.-J.; Cheng, Y.-W.; Zhang, F.-Q.; Chen, S.-L.; Gao, Q.-B. The complete plastome sequences of seven species in Gentiana sect. Kudoa (Gentianaceae): Insights into plastid gene loss and molecular evolution. Front. Plant Sci. 2018, 9, 493. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Wang, J.; Jia, Y.; Li, W.; Fusheng Xu, F.; Wang, X. Comparative chloroplast genome analyses of species in Gentiana section Cruciata (Gentianaceae) and the development of authentication markers. Int. J. Mol. Sci. 2018, 19, 1962. [Google Scholar] [CrossRef] [PubMed]
- Mower, J.P.; Guo, W.; Partha, R.; Fan, W.; Levsen, N.; Wolff, K.; Jacqueline, M.; Nugent, J.M.; Pabón-Mora, N.; González, F. Plastomes from tribe Plantagineae (Plantaginaceae) reveal infrageneric structural synapormorphies and localized hypermutation for Plantago and functional loss of ndh genes from Littorella. Mol. Phylogenet. Evol. 2021, 162, 107217. [Google Scholar] [CrossRef] [PubMed]
- Ross, T.G.; Barrett, C.F.; Gomez, M.S.; Lam, V.K.Y.; Henriquez, C.L.; Les, D.H.; Davis, J.I.; Cuenca, A.; Petersen, G.; Seberg, O.; et al. Plastid phylogenomics and molecular evolution of Alismatales. Cladistics 2016, 32, 160–178. [Google Scholar] [CrossRef]
- Maurya, S.; Darshetkar, A.M.; Datar, M.N.; Tamhankar, S.; Li, P.; Choudhary, R.K. Plastome data provide insights into intra and interspecific diversity and ndh gene loss in Capparis (Capparaceae). Phytotaxa 2020, 432, 206–220. [Google Scholar] [CrossRef]
- Fajardo, D.; Senalik, D.; Ames, M.; Zhu, H.; Steffan, S.A.; Harbut, R.; Polashock, J.; Vorsa, N.; Gillespie, E.; Kron, K.; et al. Complete Plastid Genome Sequence of Vaccinium macrocarpon: Structure, Gene Content, and Rearrangements Revealed by Next Generation Sequencing. Tree Gen. Genom. 2013, 9, 489–498. [Google Scholar] [CrossRef]
- Szczecinska, M.; Gomolinska, A.; Szkudlarz, P.; Sawick, J. Plastid and nuclear genomic resources of a relict and endangered plant species: Chamaedaphne calyculata (L.) Moench (Ericaceae). Turk. J. Bot. 2014, 38, 1229–1238. [Google Scholar] [CrossRef]
- Sanderson, M.J.; Copetti, D.; Búrquez, A.; Bustamante, E.; Charboneau, J.L.M.; Luis, E.; Eguiarte, L.E.; Kumar, S.; Lee, H.O.; Lee, J.; et al. Exceptional reduction of the plastid genome of saguaro cactus (Carnegiea gigantea): Loss of the ndh gene suite and inverted repeat. Am. J. Bot. 2015, 102, 1115–1127. [Google Scholar] [CrossRef] [PubMed]
- Sabater, B.; Martín, M.; Schmitz-Linneweber, C.; Maier, R.M. Is clustering of plastid RNA editing sites a consequence of transitory loss of gene function?—Implications for past environmental and evolutionary events in plants. Perspect. Plant Ecol. Evol. Syst. 2002, 5, 81–90. [Google Scholar] [CrossRef]
- Marín, D.; Martín, M.; Serrot, P.; Sabater, B. Thermodynamic balance of photosynthesis and transpiration at increasing CO2 concentrations and rapid light fluctuations. BioSystems 2014, 116, 21–26. [Google Scholar] [CrossRef]
- Martín, M.; Marín, D.; Serrot, P.H.; Sabater, B. Evolutionary reversion of editing sites of ndh genes suggests their origin in the Permian-Triassic before the increase of atmospheric CO2. Front. Ecol. Evol. 2015, 3, 81. [Google Scholar] [CrossRef]
- Zhe, M.; Zhang, L.; Liu, F.; Huang, Y.; Fan, W.; Yang, J.; Zhu, A. Plastid RNA editing reduction accompanied with genetic variations in Cymbidium, a genus with diverse lifestyle modes. Plant Divers. 2021, in press. [Google Scholar] [CrossRef]
- Ni, Z.; Ye, Y.; Bai, T.; Xu, M.; Xu, L.A. Complete chloroplast genome of Pinus massoniana (Pinaceae): Gene rearrangements, loss of ndh genes, and short inverted repeats contraction, expansion. Molecules 2017, 22, 1528. [Google Scholar] [CrossRef] [PubMed]
- Zeb, U.; Dong, W.-L.; Zhang, T.-T.; Wang, R.-N.; Shahzad, K.; Ma, X.-F.; Li, Z.-H. Comparative plastid genomics of Pinus species: Insights into sequence variations and phylogenetic relationships. J. Syst. Evol. 2020, 58, 118–132. [Google Scholar] [CrossRef]
- Sokołowska, J.; Fuchs, H.; Celínski, K. New insight into taxonomy of European mountain pines, Pinus mugo complex, based on complete chloroplast genomes sequencing. Plants 2021, 10, 1331. [Google Scholar] [CrossRef]
- Casano, L.M.; Lascano, H.R.; Martín, M.; Sabater, B. Topology of the plastid Ndh complex and its NDH-F subunit in thylakoid membranes. Biochem. J. 2004, 382, 145–155. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lennon, A.M.; Prommeenate, P.; Nixon, P.J. Location, expression, and orientation of the putative chlororespiratory enzymes, Ndh, and IMMUTANS, in higher plant plastids. Planta 2003, 218, 254–260. [Google Scholar] [CrossRef] [PubMed]
- Cuello, J.; Quiles, M.J.; Albacete, M.E.; Sabater, B. Properties of a large complex of NADH dehydrogenase from barley leaves. Plant Cell Physiol. 1995, 36, 265–271. [Google Scholar] [CrossRef]
- Corneille, S.; Courmac, L.; Guedeney, G.; Havaux, M.; Peltier, G. Reduction of the plastoquinone pool by exogenous NADH and NADPH in higher plant chloroplasts. Characterization of a NAD(P)H-plastoquinone oxidoreductase activity. Biochim. Biophys. Acta BBA Bioenerg. 1998, 1363, 59–69. [Google Scholar] [CrossRef]
- Díaz, M.; de Haro, V.; Muñoz, R.; Quiles, M.J. Chlororespiration is involved in the adaptation of Brassica plants to heat and high light intensity. Plant Cell Environ. 2007, 30, 1578–1585. [Google Scholar] [CrossRef]
- Joët, T.; Cournac, L.; Peltier, G.; Havaux, M. Cyclic electron flow around Photosystem I in C3 plants. In vivo control by the redox state of chloroplast and involvement of the NADH-dehydrogenase complex. Plant Physiol. 2002, 128, 760–769. [Google Scholar] [CrossRef]
- Rumeau, D.; Peltier, G.; Cournac, L. Chlororespiration and cyclic electron flow around PSI during photosynthesis and plant stress response. Plant Cell Environ. 2007, 30, 1041–1051. [Google Scholar] [CrossRef] [PubMed]
- Martín, M.; Marín, D.; Serrot, P.H.; Sabater, B. The rise of the photosynthetic rate when light intensity increases is delayed in ndh gene-defective tobacco at high but not at low CO2 concentrations. Front. Plant Sci. 2015, 6, 34. [Google Scholar] [CrossRef] [PubMed]
- Heber, U.; Walker, D. Concerning a dual function of coupled cyclic electron transport in leaves. Plant Physiol. 1992, 100, 1621–1626. [Google Scholar] [CrossRef] [PubMed]
- Casano, L.M.; Martín, M.; Sabater, B. Sensitivity of superoxide dismutase transcript levels and activities to oxidative stress is lower in mature-senescent than in young barley leaves. Plant Physiol. 1994, 106, 1033–1039. [Google Scholar] [CrossRef]
- Casano, L.M.; Martín, M.; Zapata, J.M.; Sabater, B. Leaf age- and paraquat concentration-dependent effects on the levels of enzymes protecting against photooxidative stress. Plant Sci. 1999, 149, 13–22. [Google Scholar] [CrossRef]
- Abarca, D.; Martín, M.; Sabater, B. Differential leaf stress responses in young and senescent plants. Physiol. Plant. 2001, 113, 409–415. [Google Scholar] [CrossRef]
- Abarca, D.; Roldán, M.; Martín, M.; Sabater, B. Arabidopsis thaliana ecotype Cvi shows an increased tolerance to photo-oxidative stress and contains a new chloroplastic copper/zinc superoxide dismutase isoenzyme. J. Exp. Bot. 2001, 52, 1417–1425. [Google Scholar] [CrossRef]
- Zapata, J.M.; Guera, A.; Esteban-Carrasco, A.; Martín, M.; Sabater, B. Chloroplasts regulate leaf senescence: Delayed senescence in transgenic ndhF-defective tobacco. Cell Death Differ. 2005, 12, 1277–1284. [Google Scholar] [CrossRef] [PubMed]
- Sabater, B.; Martín, M. Chloroplast control of leaf senescence. In Plastid Development in Leaves during Growth and Senescence. Advances in Photosynthesis and Respiration; Biswal, B., Krupinska, K., Biswal, U., Eds.; Springer Science+Business Media: Dordrecht, The Netherlands, 2013; Volume 36, pp. 529–550. [Google Scholar] [CrossRef]
- Sabater, B.; Martín, M. Hypothesis: Increase of the ratio singlet oxygen plus superoxide radical to hydrogen peroxide changes stress defense response to programmed leaf death. Front. Plant Sci. 2013, 4, 479. [Google Scholar] [CrossRef] [PubMed]
- Sabater, B.; Martín, M. Control of leaf cell death by reactive oxygen species. In Programmed Cell Death in Plants and Animals; Rice, J., Ed.; Nova Science Publishers, Inc.: New York, NY, USA, 2016; Chapter 3; pp. 75–93. ISBN 978-1-63484-505-2. [Google Scholar]
- Yamamoto, H.; Peng, L.; Fukao, Y.; Shikanai, T. A Src homology 3 domain-like fold protein forms a ferredoxin binding site for the chloroplast NADH dehydrogenase-like complex of Arabidopsis. Plant Cell 2011, 23, 1480–1493. [Google Scholar] [CrossRef] [PubMed]
- Kurisu, G.; Zhang, H.; Smith, J.L.; Cramer, W.A. Structure of the cytochrome b6f complex of oxygenic photosynthesis: Tuning the cavity. Science 2003, 302, 1009–1014. [Google Scholar] [CrossRef]
- Silva, S.R.; Diaz, Y.C.; Penha, H.A.; Pinheiro, D.G.; Fernandes, C.C.; Miranda, V.F.; Michael, T.P.; Varani, A.M. The Chloroplast Genome of Utricularia reniformis Sheds Light on the Evolution of the ndh Gene Complex of Terrestrial Carnivorous Plants from the Lentibulariaceae Family. PLoS ONE 2016, 11, e0165176. [Google Scholar] [CrossRef]
- Gruzdev, E.V.; Kadnikov, V.V.; Beletsky, A.V.; Kochieva, E.Z.; Mardanov, A.V.; Skryabin, K.G.; Ravin, N.V. Plastid genomes of carnivorous plants Drosera rotundifolia and Nepenthes × ventrata reveal evolutionary patterns resembling those observed in parasitic plants. Int. J. Mol. Sci. 2019, 20, 4107. [Google Scholar] [CrossRef] [PubMed]
- Horváth, E.M.; Peter, S.O.; Joët, T.; Rumeau, D.; Cournac, L.; Horváth, G.V.; Kavanagh, T.A.; Schäfer, C.; Peltier, G.; Medgyesy, P. Targeted inactivation of the plastid ndhB gene in tobacco results in an enhanced sensitivity of photosynthesis to moderate stomatal closure. Plant Physiol. 2000, 123, 1337–1349. [Google Scholar] [CrossRef]
- Sun, Y.; Geng, Q.; Du, Y.; Yang, X.; Zhai, H. Induction of cyclic electron flow around photosystem I during heat stress in grape leaves. Plant Sci. 2017, 256, 65–71. [Google Scholar] [CrossRef]
- Strand, D.D.; Fisher, N.; Kramer, D.M. The higher plant plastid NAD(P)H dehydrogenase-like complex (NDH) is a high efficiency proton pump that increases ATP production by cyclic electron flow. J. Biol. Chem. 2017, 292, 11850–11860. [Google Scholar] [CrossRef]
- Paredes, M.; Quiles, M.J. Stimulation of chlororespiration by drought under heat and high illumination in Rosa meillandina. J. Plant Physiol. 2013, 170, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Segura, M.V.; Quiles, M.J. Involvement of chlororespiration in chilling stress in the tropical species Spathiphyllum wallisü. Plant Cell Environ. 2015, 38, 525–533. [Google Scholar] [CrossRef]
- Li, Q.; Yao, Z.J.; Mi, H. Alleviation of photoinhibition by co-ordination of chlororespiration and cyclic electron flow mediated by NDH under heat stressed condition in tobacco. Front. Plant Sci. 2016, 7, 285. [Google Scholar] [CrossRef]
- Alafari, H.A.; Abd-Elgawadab, M.E. Differential expression gene/protein contribute to heat stress-responsive in Tetraena propinqua in Saudi Arabia. Saudi J. Biol. Sci. 2021, 28, 5017–5027. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Zhang, Q.S.; Zhao, W.; Liu, Z.; Ma, M.Y.; Zhong, M.Y.; Wang, M.X.; Xu, B. Chlororespiration serves as photoprotection for the photo-inactivated oxygen-evolving complex in Zostera marina, a marine angiosperm. Plant Cell Physiol. 2020, 61, 1517–1529. [Google Scholar] [CrossRef]
- Zhao, J.; Yu, W.; Zhang, L.; Liu, J. Chlororespiration protects the photosynthetic apparatus against photoinhibition by alleviating inhibition of photodamaged-PSII repair in Haematococcus pluvialis at the green motile stage. Algal Res. 2021, 54, 102140. [Google Scholar] [CrossRef]
- Sun, Y.; Deng, T.; Zhang, A.; Moore, M.J.; Landis, J.B.; Lin, N.; Zhang, H.; Zhang, X.; Huang, J.; Zhang, X.; et al. Genome sequencing of the endangered Kingdonia uniflora (Circaeasteraceae, Ranunculales) reveals potential mechanisms of evolutionary specialization. iScience 2020, 23, 101124. [Google Scholar] [CrossRef]
- Folk, R.A.; Sewnath, N.; Xiang, C.-L.; Sinn, B.T.; Guralnick, R.P. Degradation of key photosynthetic genes in the critically endangered semi-aquatic flowering plant Saniculiphyllum guangxiense (Saxifragaceae). BMC Plant Biol. 2020, 20, 324. [Google Scholar] [CrossRef]
- Su, Y.; Huang, L.; Wang, Z.; Wang, T. Comparative chloroplast genomics between the invasive weed Mikania micrantha and its indigenous congener Mikania cordata: Structure variation, identification of highly divergent regions, divergence time estimation, and phylogenetic analysis. Mol. Phylogenet. Evol. 2018, 126, 181–195. [Google Scholar] [CrossRef]
- Scobeyeva, V.A.; Artyushin, I.V.; Krinitsina, A.A.; Nikitin, P.A.; Antipin, M.I.; Kuptsov, S.V.; Belenikin, M.S.; Omelchenko, D.O.; Logacheva, M.D.; Konorov, E.A.; et al. Gene loss, pseudogenization in plastomes of genus Allium (Amaryllidaceae), and putative selection for adaptation to environmental conditions. Front. Genet. 2021, 12, 674783. [Google Scholar] [CrossRef] [PubMed]
- Ivanova, Z.; Sablok, G.; Daskalova, E.; Zahmanova, G.; Apostolova, E.; Yahubyan, G.; Baev, V. Chloroplast genome analysis of resurrection tertiary relict Haberlea rhodopensis highlights genes important for desiccation stress response. Front. Plant Sci. 2017, 8, 204. [Google Scholar] [CrossRef] [PubMed]
- Xie, D.-F.; Yu, Y.; Deng, Y.-Q.; Li, J.; Liu, H.-Y.; Dong Zhou, S.-D.; He, X.-J. Comparative analysis of the chloroplast genomes of the Chinese endemic genus Urophysa and their contribution to chloroplast phylogeny and adaptive evolution. Int. J. Mol. Sci. 2018, 19, 1847. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.-L.; Fan, W.-B.; Wang, N.; Dong, P.-B.; Zhang, T.-T.; Yue, M.; Li, Z.-H. Evolutionary analysis of plastid genomes of seven Lonicera L. species: Implications for sequence divergence and phylogenetic relationships. Int. J. Mol. Sci. 2018, 19, 4039. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Zhong, S.; Zhang, M.; Liang, Y.; Gong, G.; Chang, X.; Tan, F.; Yang, H.; Qiu, X.; Luo, L.; et al. Potential role of photosynthesis in the regulation of reactive oxygen species and defence responses to Blumeria graminis f. sp. tritici in wheat. Int. J. Mol. Sci. 2020, 21, 5767. [Google Scholar] [CrossRef]
- Liu, Y.; Ren, D.; Pike, S.; Pallardy, S.; Gassmann, W.; Zhang, S. Chloroplast-generated reactive oxygen species are involved in hypersensitive response-like cell death mediated by a mitogen-activated protein kinase cascade. Plant J. 2007, 51, 941–954. [Google Scholar] [CrossRef]
- Reape, T.J.; Molony, E.M.; McCabe, P.F. Programmed cell death in plants: Distinguishing between different modes. J. Exp. Bot. 2008, 59, 435–444. [Google Scholar] [CrossRef]
- Doyle, S.M.; Diamond, M.; McCabe, P.F. Chloroplast and reactive oxygen species involvement in apoptotic-like programmed cell death in Arabidopsis suspension cultures. J. Exp. Bot. 2010, 61, 473–482. [Google Scholar] [CrossRef]
- Huang, M.S.; Braun, D.M. Genetic analyses of cell death in maize (Zea mays, poaceae) leaves reveal a distinct pathway operating in the camouflage1 mutant. Am. J. Bot. 2010, 97, 357–364. [Google Scholar] [CrossRef]
- Nilo, R.; Saffie, C.; Lilley, K.; Baeza-Yates, R.; Cambiazo, V.; Campos-Vargas, R.; González, M.; Meisel, L.A.; Retamales, J.; Silva, H.; et al. Proteomic analysis of peach fruit mesocarp softening and chilling injury using difference gel electrophoresis (DIGE). BMC Genom. 2010, 11, 43. [Google Scholar] [CrossRef]
- Mayta, M.L.; Hajirezaei, M.-R.; Carrillo, N.; Lodeyro, A.F. Leaf senescence: The chloroplast connection comes of age. Plants 2019, 8, 495. [Google Scholar] [CrossRef]
- McCoy, S.R.; Kuehl, J.V.; Boore, J.L.; Raubeson, L.A. The complete plastid genome sequence of Welwitschia mirabilis: An unusually compact plastome with accelerated divergence rates. BMC Evol. Biol. 2008, 8, 130. [Google Scholar] [CrossRef] [PubMed]
- Gechev, T.S.; Van Breusegem, F.; Stone, J.M.; Denev, J.; Laloi, C. Reactive oxygen species as signals that modulate plant stress responses and programmed cell death. BioEssays 2006, 28, 1091–1101. [Google Scholar] [CrossRef] [PubMed]
- Mubarakshina, M.M.; Ivanov, B.N.; Naydov, I.A.; Hillier, W.; Badger, M.R.; Krieger-Liszkay, A. Production and diffusion of chloroplastic H2O2 and its implication to signalling. J. Exp. Bot. 2010, 61, 3577–3587. [Google Scholar] [CrossRef] [PubMed]
- Wiciarz, M.; Gubernator, B.; Kruk, J.; Niewiadomska, E. Enhanced chloroplastic generation of H2O2 in stress resistant Thellungiella salsuginea in comparison to Arabidopsis thaliana. Physiol. Plant. 2015, 153, 467–476. [Google Scholar] [CrossRef] [PubMed]
- Bowler, C.; Van Montagu, M.; Inzé, D. Superoxide dismutase and stress tolerance. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1992, 43, 83–116. [Google Scholar] [CrossRef]
- Kurepa, J.; Hérouart, D.; Van Montagu, M.; Inzé, D. Differential expression of CuZn- and Fe-superoxide dismutase genes of tobacco during development, oxidative stress and hormonal treatments. Plant Cell Physiol. 1997, 38, 463–470. [Google Scholar] [CrossRef]
- Prochazkova, D.; Sairam, R.K.; Srivastava, G.C.; Singh, D.V. Oxidative stress and antioxidant activity as the basis of senescence in maize. Plant Sci. 2001, 161, 765–771. [Google Scholar] [CrossRef]
- Ohe, M.; Rapolu, M.; Mieda, T.; Miyagawa, Y.; Yabuta, Y.; Yoshimura, K.; Shigeoka, S. Decline of leaf photooxidative stress tolerance with age in tobacco. Plant Sci. 2005, 168, 1487–1493. [Google Scholar] [CrossRef]
- Orr, W.C.; Sohal, R.S. Extension of lifespan by overexpression of superoxide dismutase and catalase in Drosophila melanogaster. Science 1994, 263, 1128–1130. [Google Scholar] [CrossRef]
- Lascano, H.R.; Casano, L.M.; Martín, M.; Sabater, B. The activity of the chloroplastic Ndh complex is regulated by phosphorylation of the NDH-F subunit. Plant Physiol. 2003, 132, 256–262. [Google Scholar] [CrossRef]
- Nashilevitz, S.; Melamed-Bessudo, C.; Izkovich, Y.; Rogachev, I.; Osorio, S.; Itkin, M.; Adato, A.; Pankratov, J.; Hirschberg, J.; Femie, A.R.; et al. An orange ripening mutant links plastid NAD(P)H dehydrogenase complex activity to central and specialized metabolism during tomato fruit maturation. Plant. Cell 2010, 22, 1977–1997. [Google Scholar] [CrossRef] [PubMed]
- Krieger-Liszkay, A.; Trosch, M.; Krupinska, K. Generation of reactive oxygen species in thylakoids from senescing flag leaves of the barley varieties Lomerit and Carina. Planta 2015, 241, 1497–1508. [Google Scholar] [CrossRef] [PubMed]
- Kane, D.J.; Sarafian, T.A.; Anton, R.; Hahn, H.; Gralla, E.B.; Valentine, J.S.; Örd, T.; Bredesen, D.E. Bcl-2 inhibition of neural death: Decreased generation of reactive oxygen species. Science 1993, 262, 1274–1277. [Google Scholar] [CrossRef] [PubMed]
- Green, D.R.; Reed, J.C. Mitochondria and apoptosis. Science 1998, 281, 1309–1312. [Google Scholar] [CrossRef]
- Yu, Z.-H.; Perdue, T.D.; Heimer, Y.M.; Jones, A.M. Mitochondrial involvement in tracheary element programmed cell death. Cell Death Differ. 2002, 9, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Lam, E.; Kato, N.; Lawton, M. Programmed cell death, mitochondria and the plant hypersensitive response. Nature 2001, 411, 848–853. [Google Scholar] [CrossRef]
- Ruiz-Pesini, E.; Mishmar, D.; Brandon, M.; Procaccio, V.; Wallace, D.C. Effects of purifying and adaptive selection on regional variation in human mtDNA. Science 2004, 303, 223–226. [Google Scholar] [CrossRef] [PubMed]


| Ndh Gene | Encoded Polypeptide | Homologous Polypeptides in Respiratory Complex I (References) |
|---|---|---|
| ndhA | NDH-A | ND1/NuoH/FpoH, EchB, NQ08 [11,13] |
| ndhB | NDH-B | ND2/NuoN/FpoO, NQO14 [13,14,15] |
| ndhC | NDH-C | ND3/NuoA/FpoA, NQ07 [14,15,16] |
| ndhD | NDH-D | ND4/NuoM/FpoM, NQ013 [13] |
| ndhE | NDH-E | ND4L/NuoK/FpoK, NQ011 [13,15,16] |
| ndhF | NDH-F | ND5/NuoL/FpoL, NQ012 [13,17] |
| ndhG | NDH-G | ND6/NuoJ/FpoJ, NQ010 [13,18] |
| ndhH | NDH-H | 49(IP)/NuoD/FpoD, EchE, NQ05 [13,19,20] |
| ndhI | NDH-I | TYKY/NuoI/FpoI, EchF, NQ09 [13,20] |
| ndhJ | NDH-J | 30(IP)/NuoC/FpoC, EchD, NQ04 [13,20] |
| ndhK | NDH-K | PSTT/NuoB/FpoB, EchC, NQ06 [13,20] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sabater, B. On the Edge of Dispensability, the Chloroplast ndh Genes. Int. J. Mol. Sci. 2021, 22, 12505. https://doi.org/10.3390/ijms222212505
Sabater B. On the Edge of Dispensability, the Chloroplast ndh Genes. International Journal of Molecular Sciences. 2021; 22(22):12505. https://doi.org/10.3390/ijms222212505
Chicago/Turabian StyleSabater, Bartolomé. 2021. "On the Edge of Dispensability, the Chloroplast ndh Genes" International Journal of Molecular Sciences 22, no. 22: 12505. https://doi.org/10.3390/ijms222212505
APA StyleSabater, B. (2021). On the Edge of Dispensability, the Chloroplast ndh Genes. International Journal of Molecular Sciences, 22(22), 12505. https://doi.org/10.3390/ijms222212505






