Functionalization of Human Serum Albumin by Tyrosine Click
Abstract
:1. Introduction
2. Results
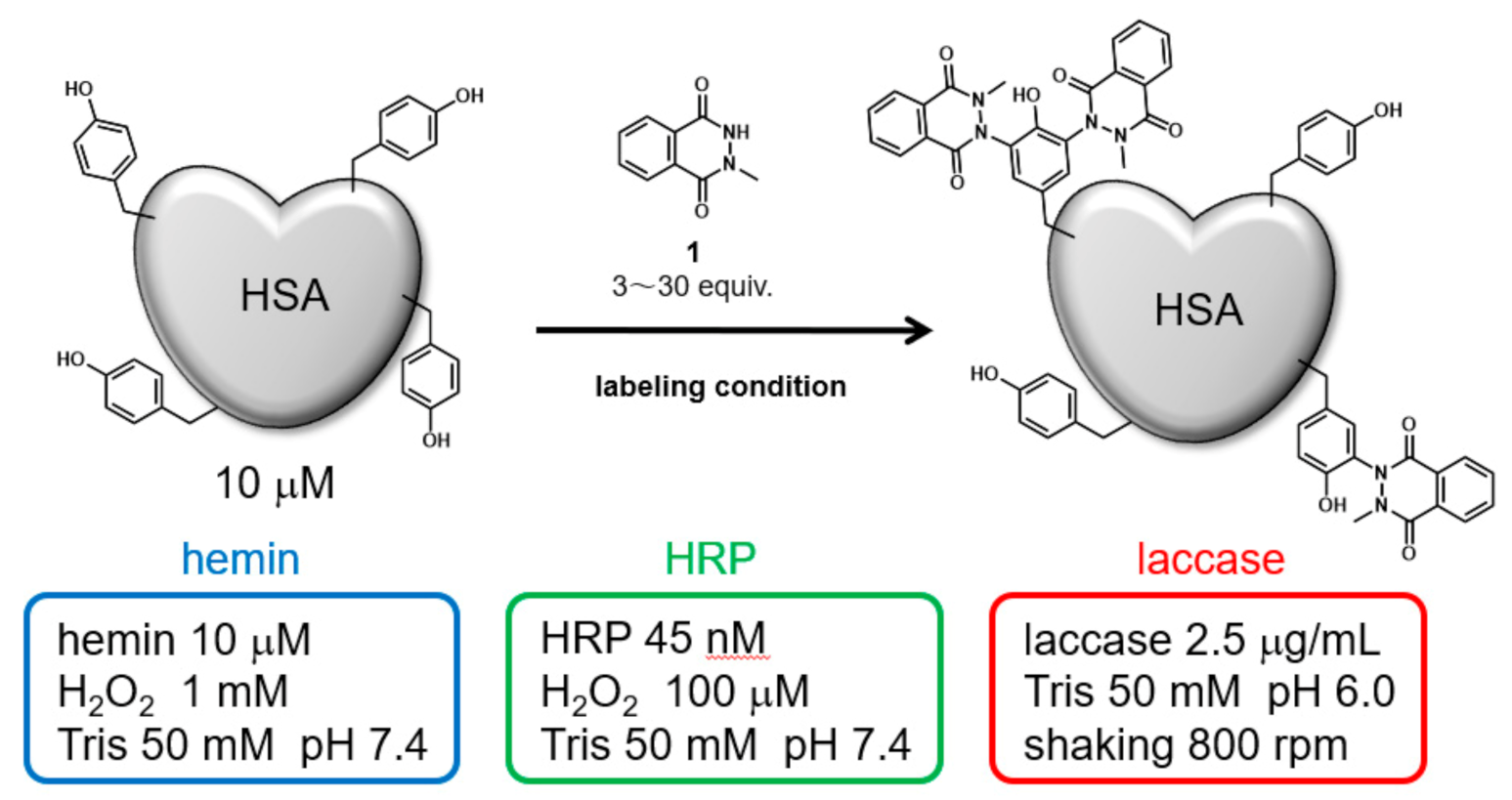
2.1. Selection of Conditions for Tyrosine Residue Modification of HSA
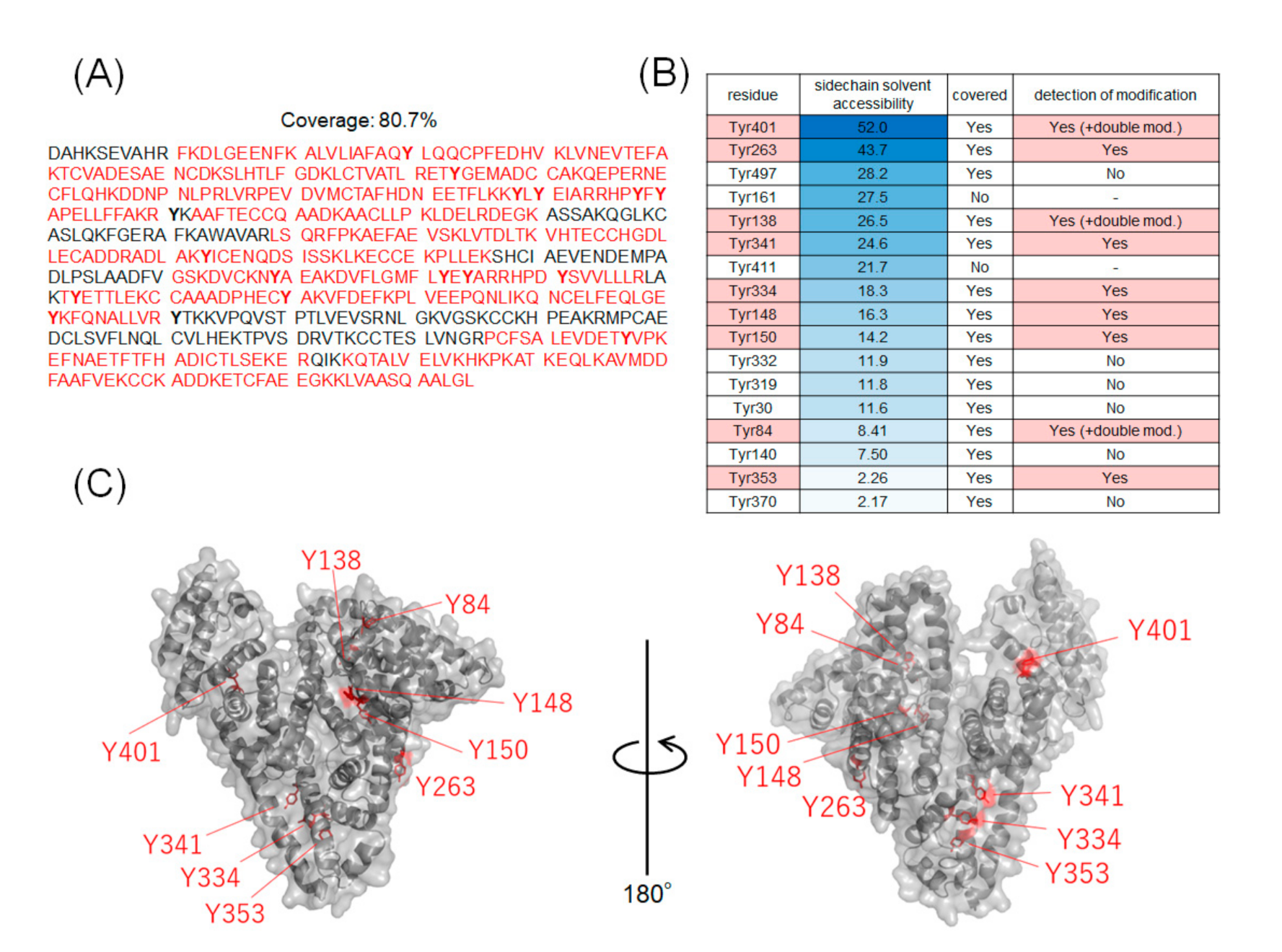
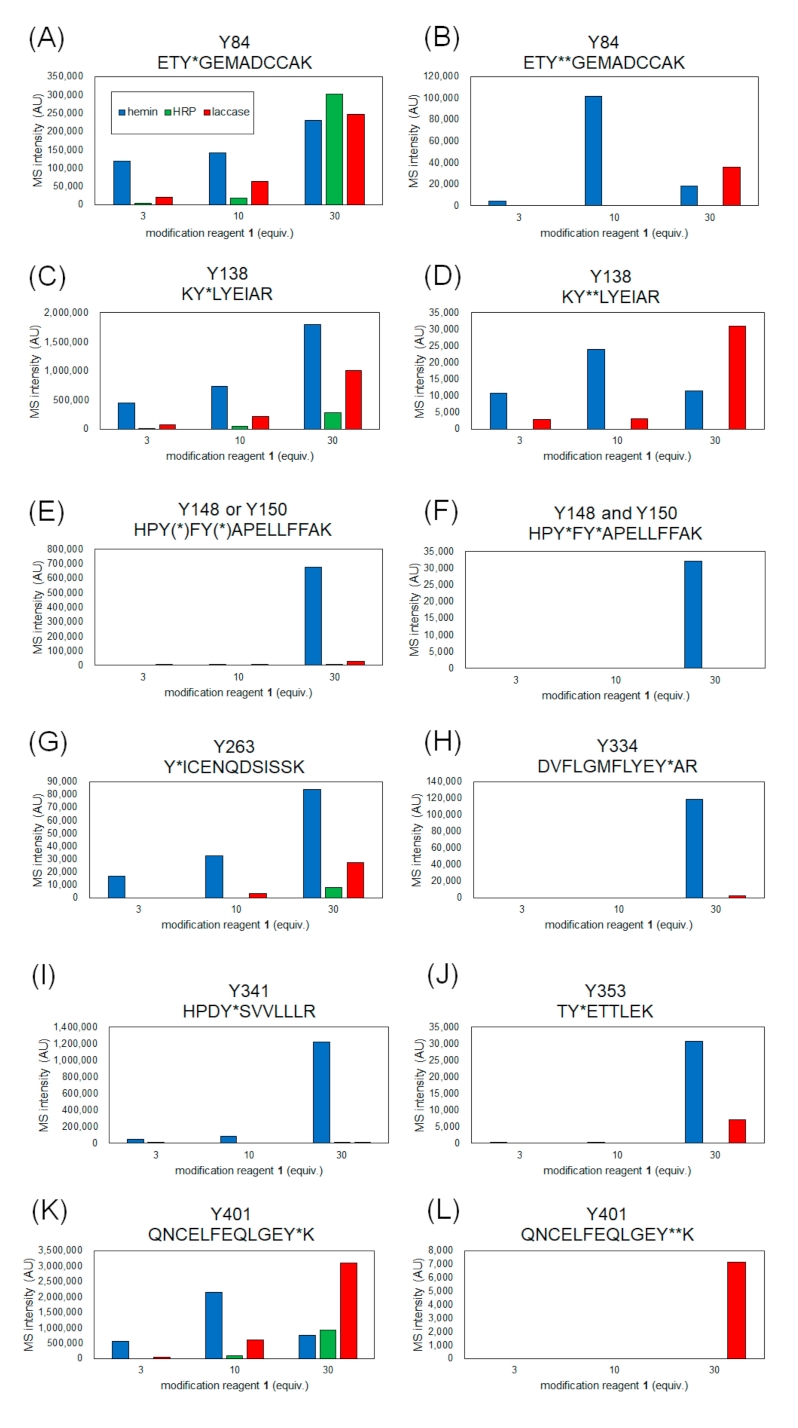
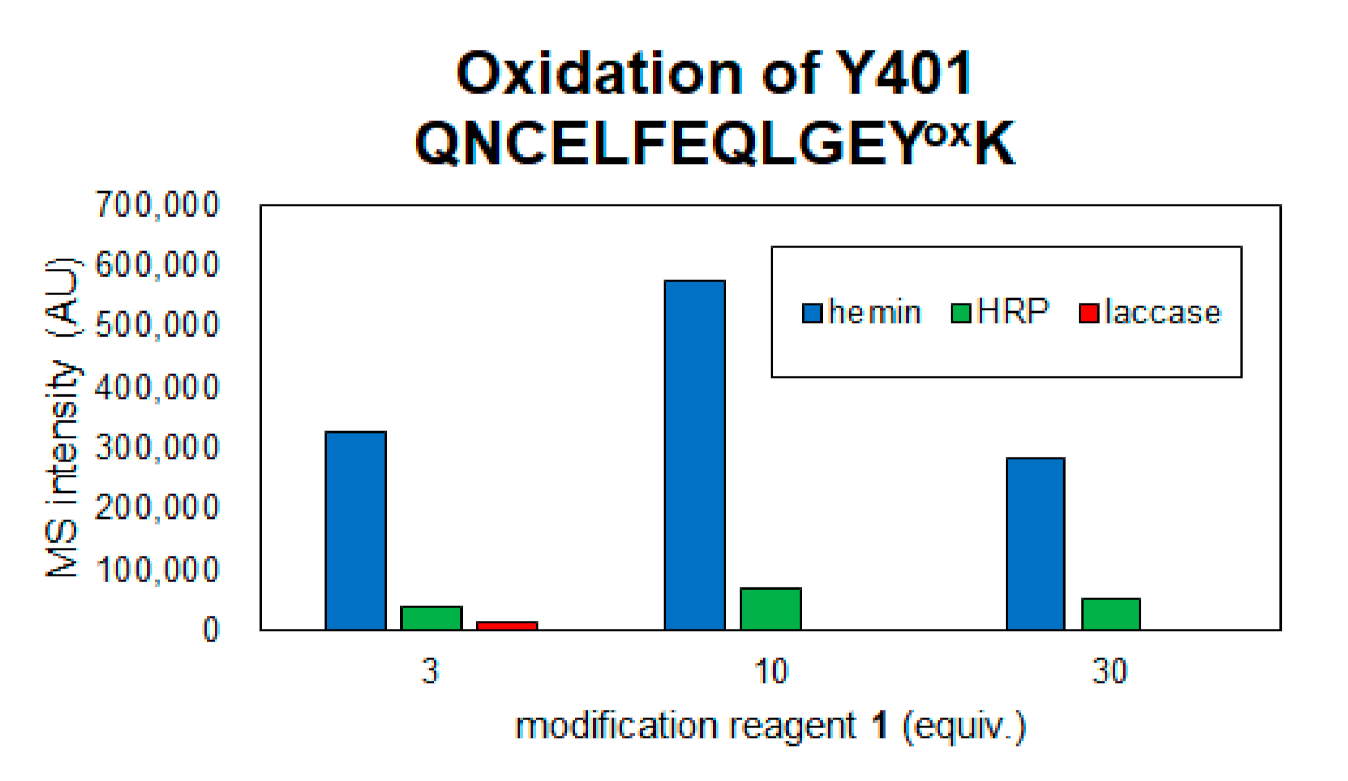
2.2. Identification of Modification Sites by Mass Spectrometry and Comparison of Modification Efficiency under Various Reaction Conditions
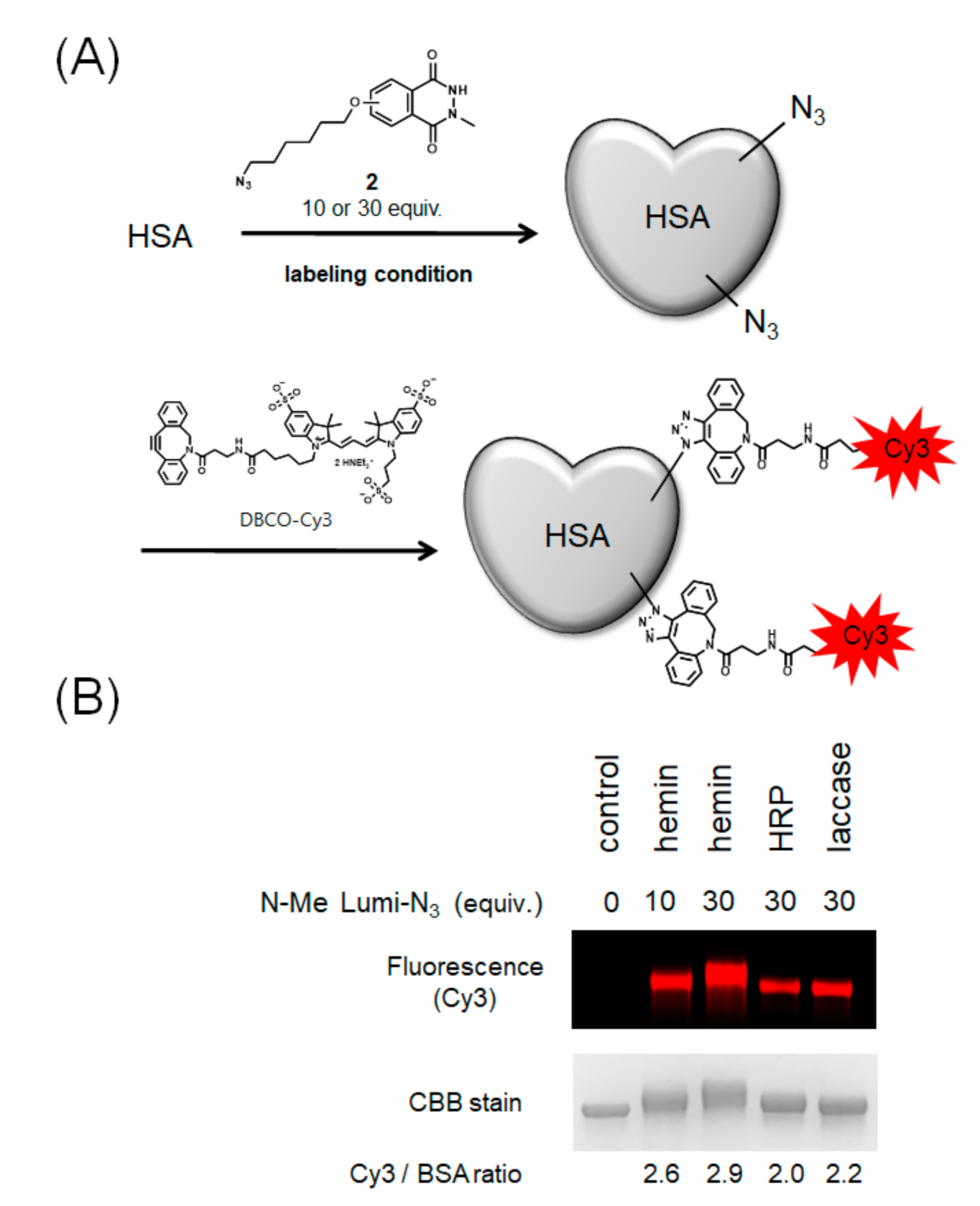
2.3. Modification with Azide-Conjugated Modification Reagent and Evaluation of Modification Efficiencies
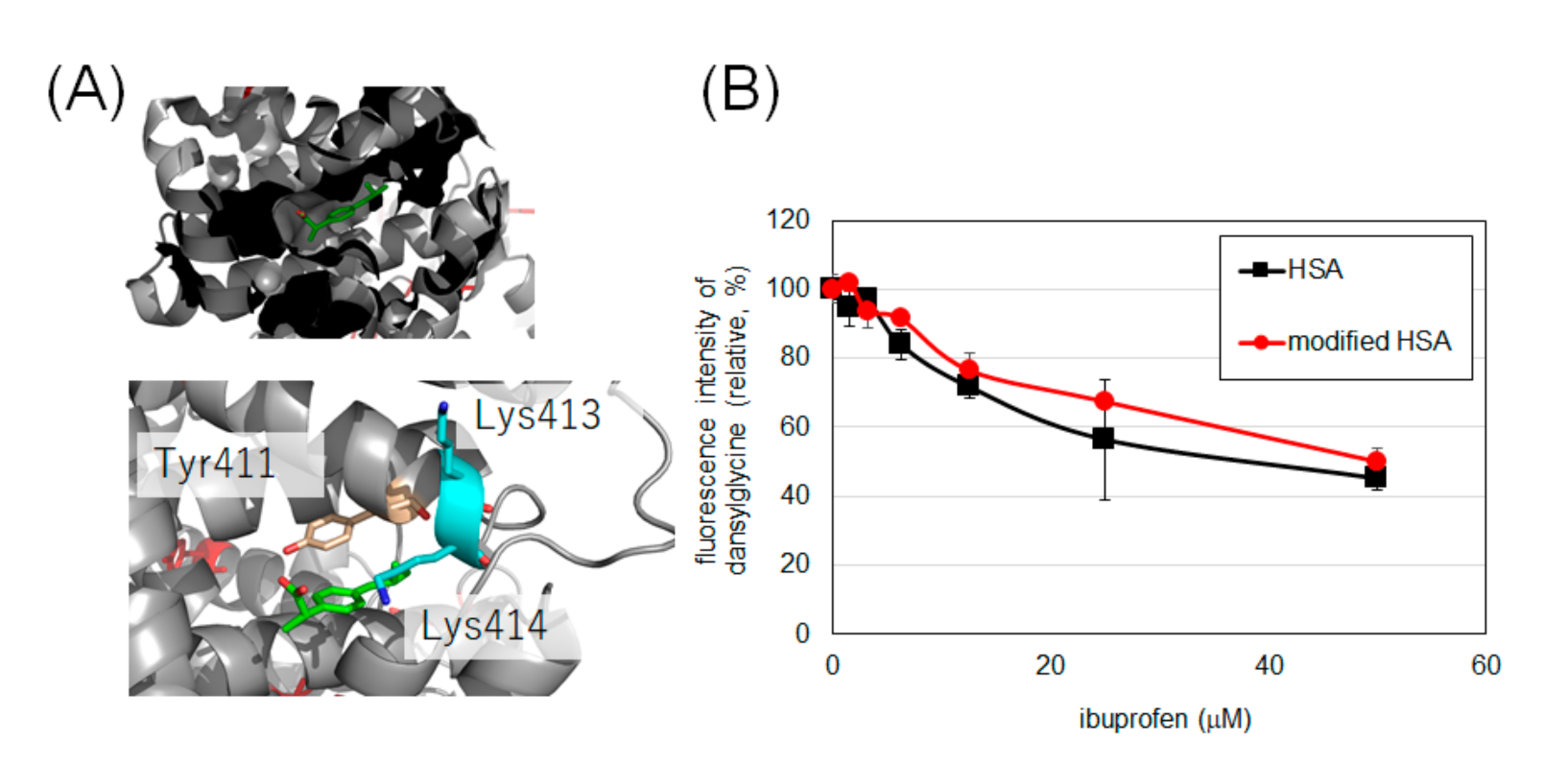
2.4. Evaluation of Drug Binding Ability of Tyrosine-Modified HSA
3. Discussion
4. Material and Methods
4.1. HSA Modification Using Hemin
4.2. HSA Modification Using HRP
4.3. HSA Modification Using Laccase
4.4. In Gel Digestion
4.5. NanoLC-MS/MS Analysis
4.6. Evaluation of Labeling Efficiency
4.7. Evaluation of Ligand Binding Ability of Labeled HSA
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fanali, G.; Di Masi, A.; Trezza, V.; Marino, M.; Fasano, M.; Ascenzi, P. Human serum albumin: From bench to bedside. Mol. Aspects Med. 2012, 33, 209–290. [Google Scholar] [CrossRef]
- Matsumura, Y.; Maeda, H. A New Concept for Macromolecular Therapeutics in Cancer Chemotherapy: Mechanism of Tumoritropic Accumulation of Proteins and the Antitumor Agent Smancs. Cancer Res. 1986, 46, 6387–6392. [Google Scholar]
- Stehle, G.; Sinn, H.; Wunder, A.; Schrenk, H.H.; Stewart, J.C.M.; Hartung, G.; Maier-Borst, W.; Heene, D.L. Plasma protein (albumin) catabolism by the tumor itself—Implications for tumor metabolism and the genesis of cachexia. Crit. Rev. Oncol. Hematol. 1997, 26, 77–100. [Google Scholar] [CrossRef]
- Sinn, H.; Schrenk, H.H.; Friedrich, E.A.; Schilling, U.; Maier-Borst, W. Design of compounds having an enhanced tumour uptake, using serum albumin as a carrier. Part I. Int. J. Radiat. Appl. Instrum. 1990, 17, 819–826. [Google Scholar] [CrossRef]
- Schilling, U.; Friedrich, E.A.; Sinn, H.; Schrenk, H.H.; Clorius, J.H.; Maier-Borst, W. Design of compounds having enhanced tumour uptake, using serum albumin as a carrier-part II. In vivo studies. Int. J. Radiat. Appl. Instrum. 1992, 19, 685–695. [Google Scholar] [CrossRef]
- Gradishar, W.J.; Krasnojon, D.; Cheporov, S.; Makhson, A.N.; Manikhas, G.M.; Clawson, A.; Bhar, P. Significantly longer progression-free survival with nab-paclitaxel compared with docetaxel as first-line therapy for metastatic breast cancer. J. Clin. Oncol. 2009, 27, 3611–3619. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, M.J.; Soon-Shiong, P.; Desai, N. Protein nanoparticles as drug carriers in clinical medicine. Adv. Drug Deliv. Rev. 2008, 60, 876–885. [Google Scholar] [CrossRef]
- Elsadek, B.; Kratz, F. Impact of albumin on drug delivery—New applications on the horizon. J. Control. Release 2012, 157, 4–28. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Chen, X. Simple bioconjugate chemistry serves great clinical advances: Albumin as a versatile platform for diagnosis and precision therapy. Chem. Soc. Rev. 2016, 45, 1432–1456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Toyo’oka, T.; Sano, A.; Kuriki, T.; Suzuki, N. Covalent binding of cefotaxime to human serum albumin. J Pharm. Dyn. 1983, 6, 139–140. [Google Scholar] [CrossRef]
- Kikuchi, S.; Kanoh, D.; Sato, S.; Sakurai, Y.; Suzuki, M.; Nakamura, H. Maleimide-functionalized closo-dodecaborate albumin conjugates (MID-AC): Unique ligation at cysteine and lysine residues enables efficient boron delivery to tumor for neutron capture therapy. J. Control. Release 2016, 237, 160–167. [Google Scholar] [CrossRef] [PubMed]
- Ishii, S.; Sato, S.; Asami, H.; Hasegawa, T.; Kohno, J.Y.; Nakamura, H. Design of S-S bond containing maleimide-conjugated: Closo -dodecaborate (SSMID): Identification of unique modification sites on albumin and investigation of intracellular uptake. Org. Biomol. Chem. 2019, 17, 5496–5499. [Google Scholar] [CrossRef]
- Kawai, K.; Nishimura, K.; Okada, S.; Sato, S.; Suzuki, M.; Takata, T.; Nakamura, H. Cyclic RGD-Functionalized closo-Dodecaborate Albumin Conjugates as Integrin Targeting Boron Carriers for Neutron Capture Therapy. Mol. Pharm. 2020, 17, 3740–3747. [Google Scholar] [CrossRef] [PubMed]
- Fujiki, K.; Tanaka, K. Biomolecular labeling based on lysine-clickable 6π-azaelectrocyclization toward innovative cancer theranostics. Bioorg. Med. Chem. 2021, 42, 116238. [Google Scholar] [CrossRef] [PubMed]
- Asano, S.; Patterson, J.T.; Gaj, T.; Iii, C.F.B. Site-Selective Labeling of a Lysine Residue in Human Serum Albumin. Angew. Chem. Int. Ed. 2014, 53, 11783–11786. [Google Scholar] [CrossRef]
- Yuaan, C.; Shen, C.; Mun, S.; Azizah, A.; Halim, A.; Tayyab, S. Lysine modification of human serum albumin and its effect on protein conformation and nalidixic acid binding. J. Indian Chem. Soc. 2021, 98, 100031. [Google Scholar]
- Rahman, M.H.; Maruyama, T.; Okada, T.; Yamasaki, K.; Otagiri, M. Study of interaction of carprofen and its enantiomers with human serum albumin-I. Biochem. Pharmacol. 1993, 46, 1721–1731. [Google Scholar] [CrossRef]
- Bertucci, C.; Barsotti, M.C.; Ra, A.; Salvadori, P. Binding properties of human albumin modi ¢ ed by covalent binding of penicillin. Biochim. Biophys. Acta Protein Struct. Mol. Enzymol. 2001, 1544, 386–392. [Google Scholar] [CrossRef]
- Sato, S.; Nakamura, H. Labeling of Peroxide-Induced Oxidative Stress Hotspots by Hemin-Catalyzed Tyrosine Click. Chem. Pharm. Bull. 2020, 68, 885–890. [Google Scholar] [CrossRef]
- Sato, S.; Nakamura, K.; Nakamura, H. Tyrosine-Specific Chemical Modification with in Situ Hemin-Activated Luminol Derivatives. ACS Chem. Biol. 2015, 10, 2633–2640. [Google Scholar] [CrossRef]
- Sato, S.; Nakamura, K.; Nakamura, H. Horseradish-Peroxidase-Catalyzed Tyrosine Click Reaction. ChemBioChem 2017, 18, 475–478. [Google Scholar] [CrossRef]
- Sato, S.; Nakamura, H. Ligand-directed selective protein modification based on local single-electron-transfer catalysis. Angew. Chemie Int. Ed. 2013, 52, 8681–8684. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Ishii, S.; Nakamura, H. Development of Albumin-closo-Dodecaborate Conjugates as Boron Carriers for Neutron-Capture Therapy by Ru(bpy)3-Photocatalyzed Modification of Tyrosine. Eur. J. Inorg. Chem. 2017, 2017, 4406–4410. [Google Scholar] [CrossRef] [Green Version]
- Sato, S.; Nakane, K.; Nakamura, H. A laccase-catalysed tyrosine click reaction. Org. Biomol. Chem. 2020, 18, 3664–3668. [Google Scholar] [CrossRef] [PubMed]
- Sudlow, G.; Birkett, D.J.; Wade, D.N. The Characterization of Two Specific Drug Binding Sites on Human Serum. Mol. Pharmacol. 1975, 11, 824–832. [Google Scholar]
- Sudlow, G.; Birkett, D.J.; Wade, D.N. Further Characterization of Specific Drug Binding Site on Human Serum Albumin. Mol. Pharmacol. 1976, 12, 1052–1061. [Google Scholar]
- Stewart, A.J.; Blindauer, C.A.; Berezenko, S.; Sleep, D.; Tooth, D.; Sadler, P.J. Role of Tyr84 in controlling the reactivity of Cys34 of human albumin. FEBS J. 2005, 272, 353–362. [Google Scholar] [CrossRef] [PubMed]
- Ashrafi-Kooshk, M.R.; Ebrahimi, F.; Ranjbar, S.; Ghobadi, S.; Moradi, N.; Khodarahmi, R. Comparative studies on drug binding to the purified and pharmaceutical-grade human serum albumins: Bridging between basic research and clinical applications of albumin. Biologicals 2015, 43, 333–343. [Google Scholar] [CrossRef] [PubMed]
- Tayyab, S.; Haq, S.K.; Sabeeha; Aziz, M.A.; Khan, M.M.; Muzammil, S. Effect of lysine modification on the conformation and indomethacin binding properties of human serum albumin. Int. J. Biol. Macromol. 1999, 8130, 173–180. [Google Scholar]
- Kohita, H.; Matsushita, Y.; Moriguchi, I. Binding of Carprofen to human and bovine serum albumin. Chem. Pharm. Bull. 1994, 42, 937–940. [Google Scholar] [CrossRef] [Green Version]
- Yamasaki, K.; Nishi, K.; Anraku, M.; Taguchi, K.; Maruyama, T. Metal-catalyzed oxidation of human serum albumin does not alter the interactive binding to the two principal drug binding sites. Biochem. Biophys. Rep. 2018, 14, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Obara, S.; Nakane, K.; Fujimura, C.; Tomoshige, S.; Ishikawa, M.; Sato, S. Functionalization of Human Serum Albumin by Tyrosine Click. Int. J. Mol. Sci. 2021, 22, 8676. https://doi.org/10.3390/ijms22168676
Obara S, Nakane K, Fujimura C, Tomoshige S, Ishikawa M, Sato S. Functionalization of Human Serum Albumin by Tyrosine Click. International Journal of Molecular Sciences. 2021; 22(16):8676. https://doi.org/10.3390/ijms22168676
Chicago/Turabian StyleObara, Satsuki, Keita Nakane, Chizu Fujimura, Shusuke Tomoshige, Minoru Ishikawa, and Shinichi Sato. 2021. "Functionalization of Human Serum Albumin by Tyrosine Click" International Journal of Molecular Sciences 22, no. 16: 8676. https://doi.org/10.3390/ijms22168676
APA StyleObara, S., Nakane, K., Fujimura, C., Tomoshige, S., Ishikawa, M., & Sato, S. (2021). Functionalization of Human Serum Albumin by Tyrosine Click. International Journal of Molecular Sciences, 22(16), 8676. https://doi.org/10.3390/ijms22168676





