Chelerythrine Chloride: A Potential Rumen Microbial Urease Inhibitor Screened by Targeting UreG
Abstract
:1. Introduction
2. Results
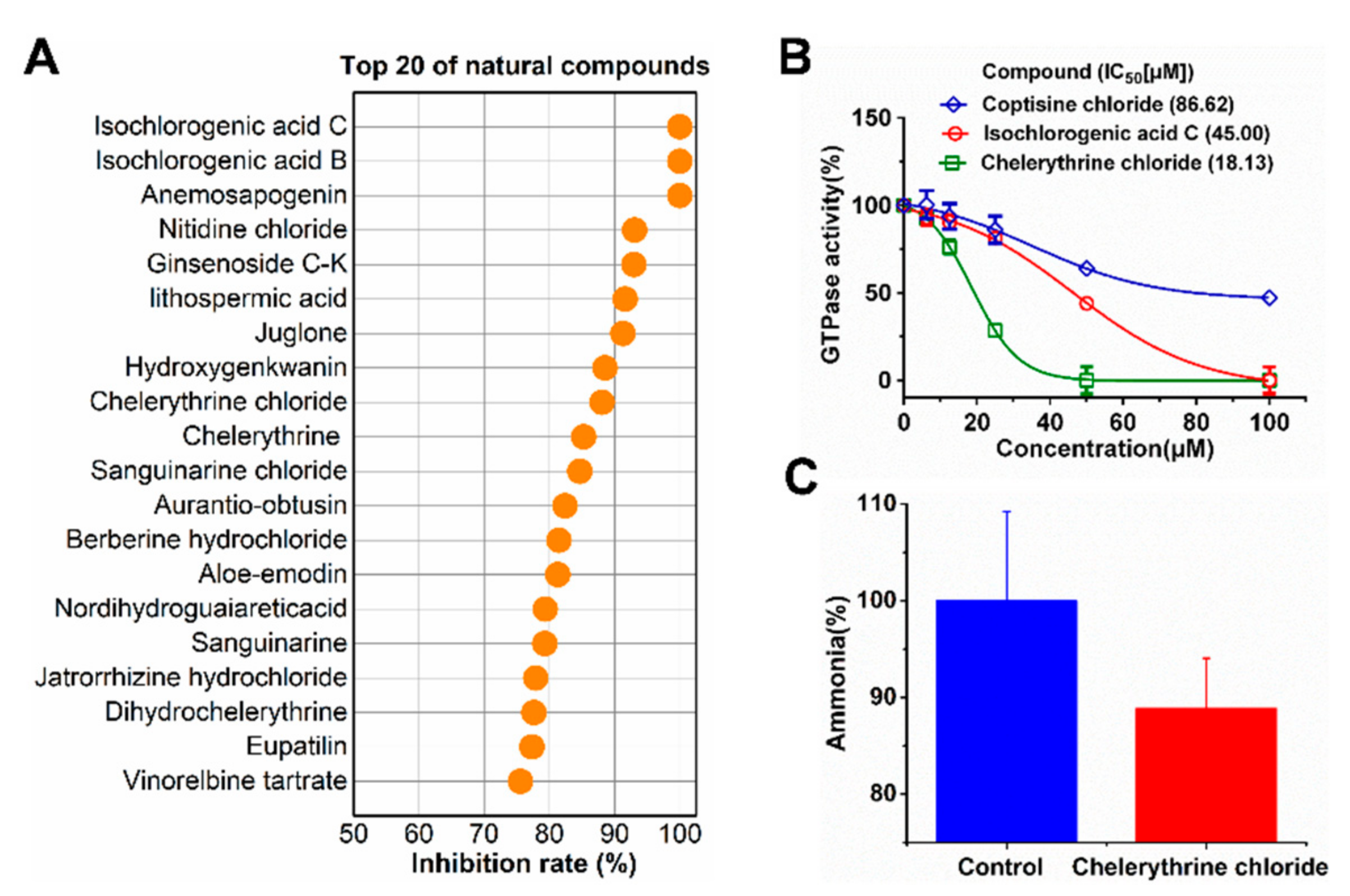
2.1. Screening of Urease Inhibitor
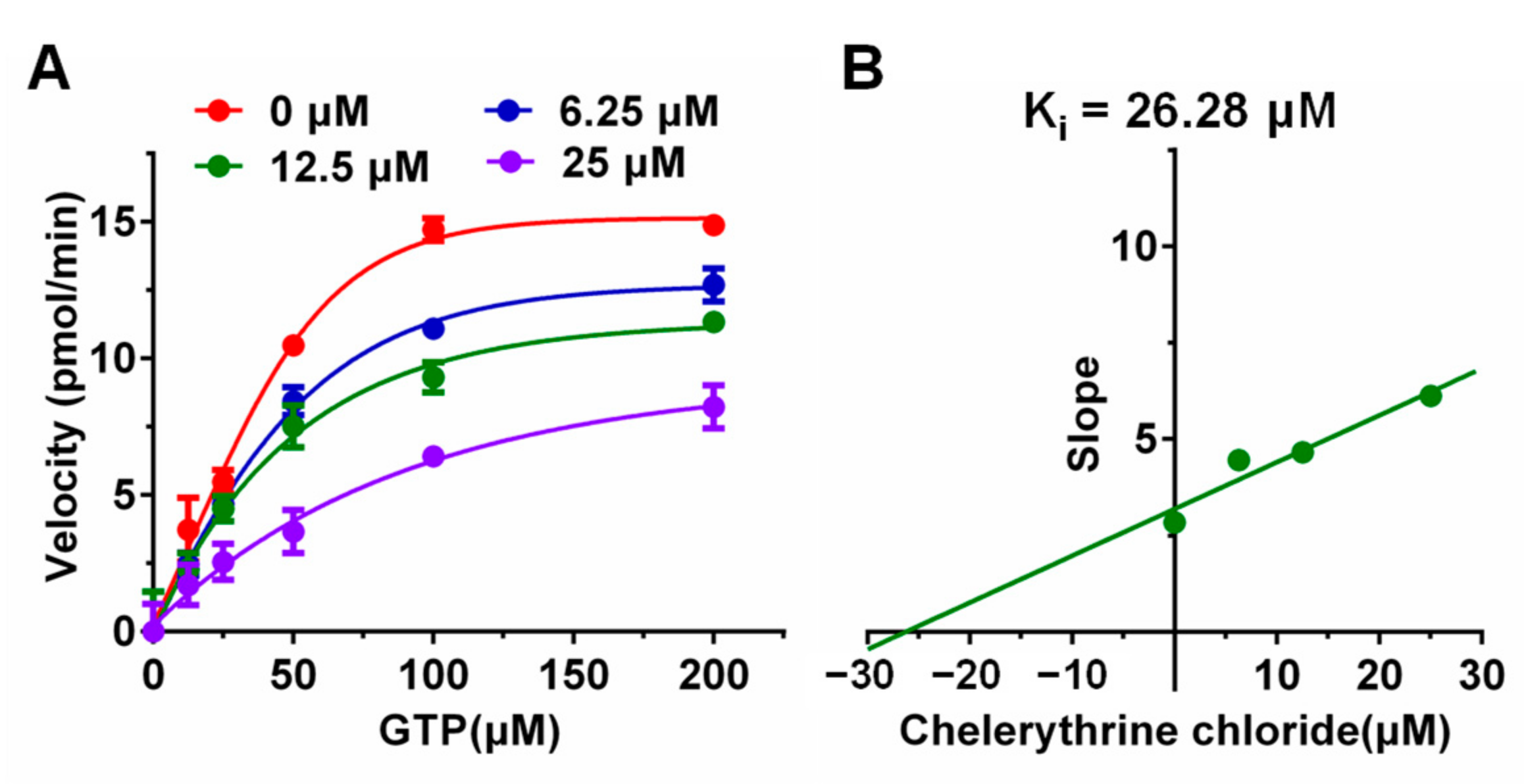
2.2. Kinetic Study of UreG Inhibition by Chelerythrine Chloride
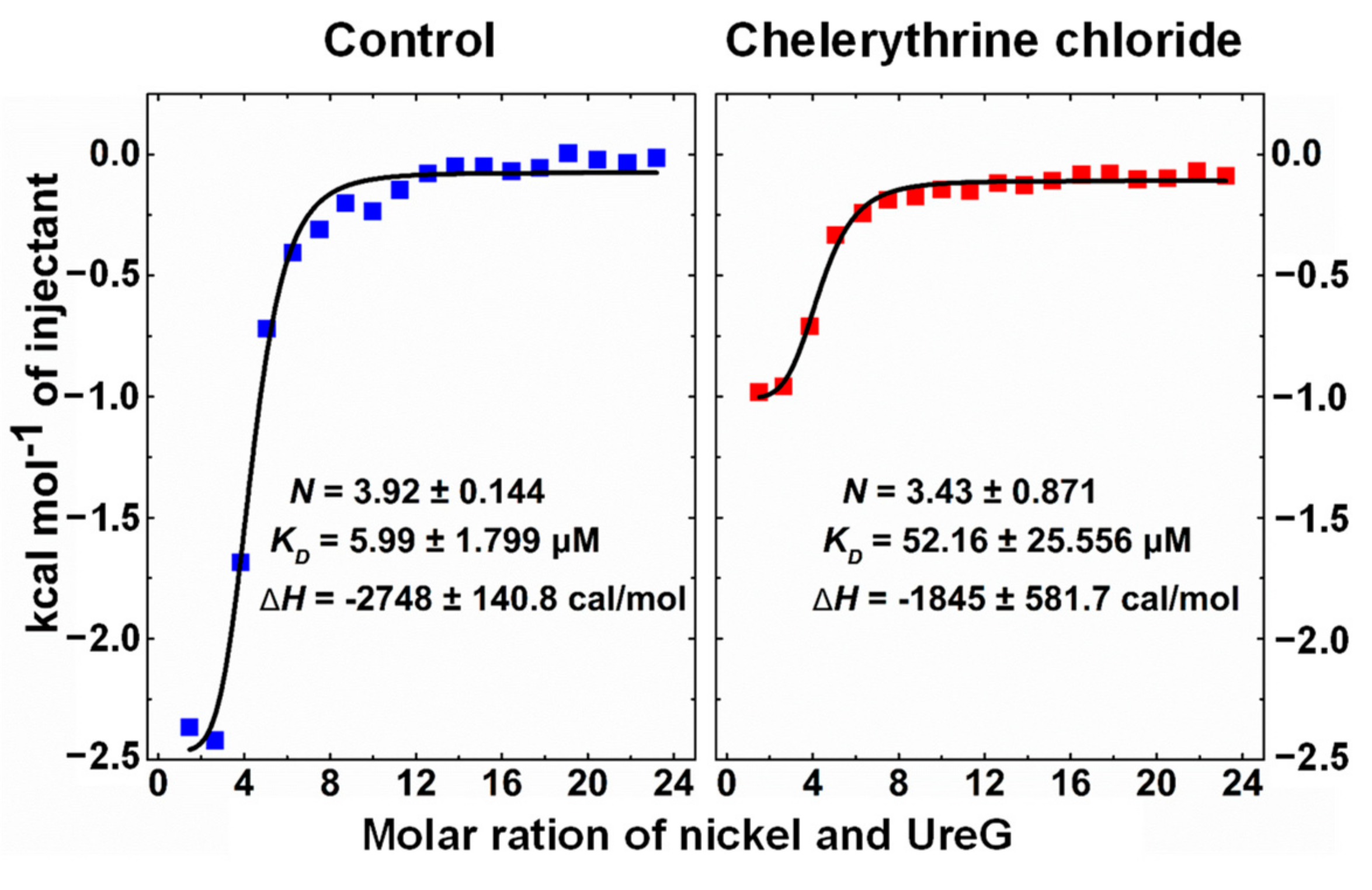
2.3. Chelerythrine Chloride Inhibited Nickel Binding to UreG
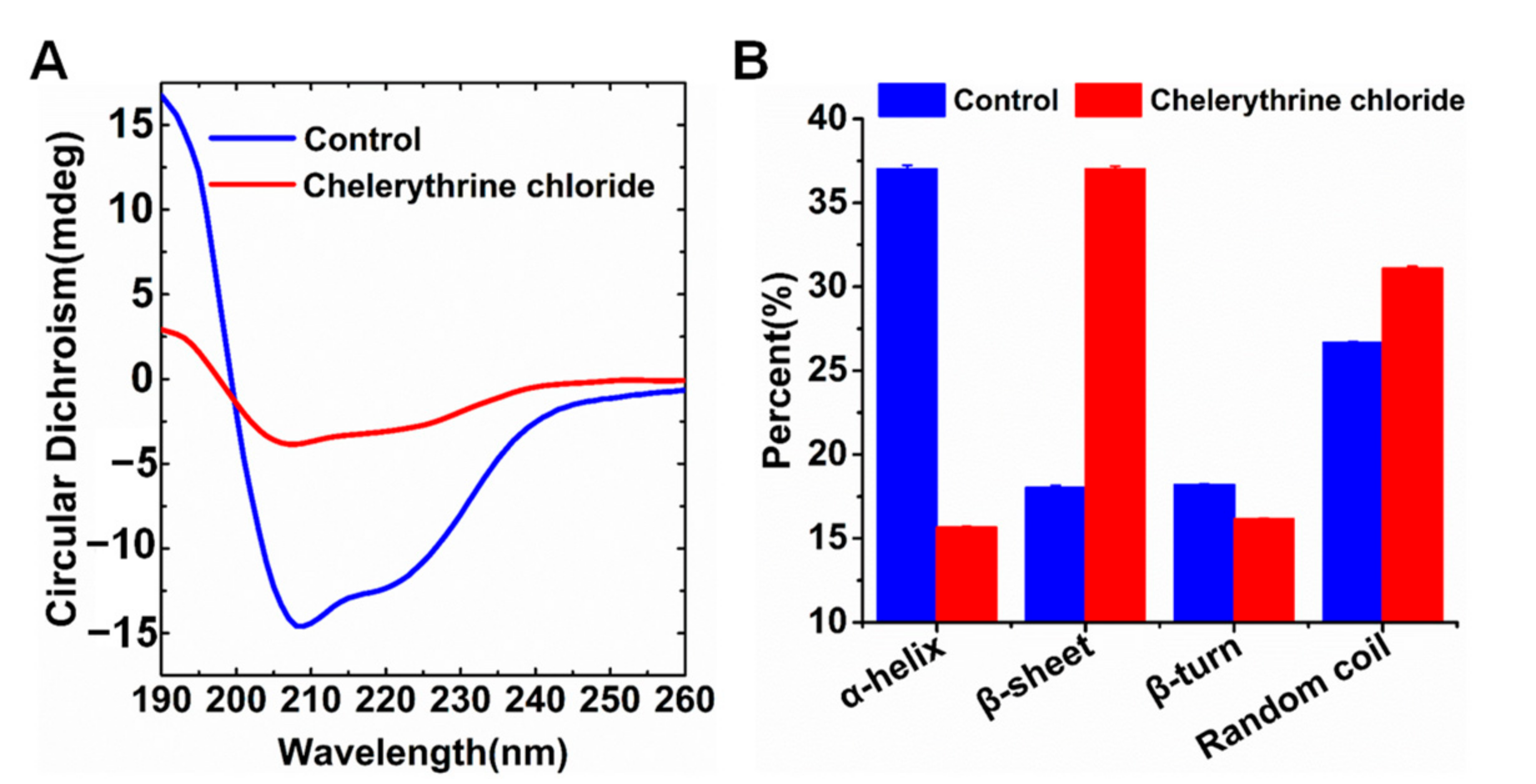
2.4. Chelerythrine Chloride-Induced Secondary Structure Change of UreG
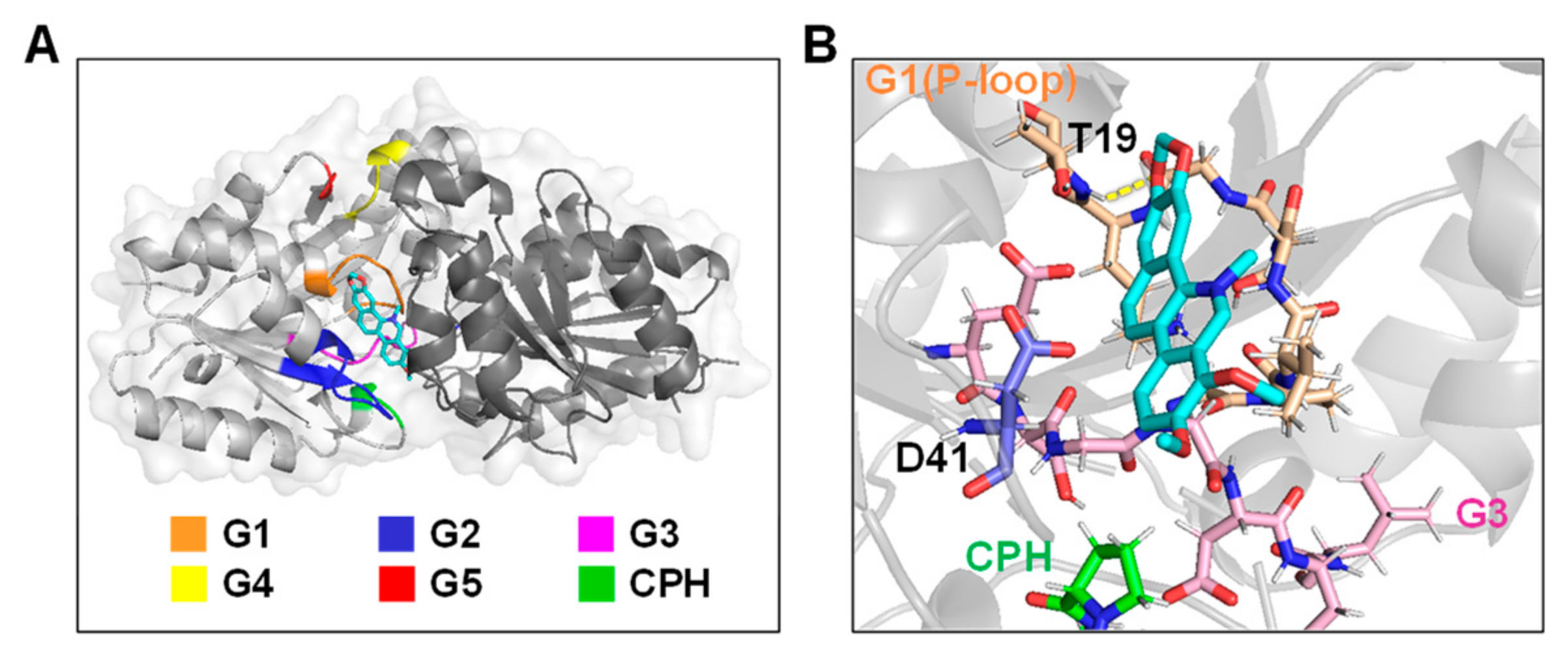
2.5. Molecular Docking of Chelerythrine Chloride towards UreG
3. Discussion
4. Materials and Methods
4.1. Preparation of Urease Accessory Protein UreG
4.2. GTPase Activity of UreG
4.3. Preparation of Ruminal Microbial Crude Protein and Measurement of Ammonia Released
4.4. Kinetic Study
4.5. Molecular Docking
4.6. ITC Measurements
4.7. CD Spectroscopy
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Matthews, C.; Crispie, F.; Lewis, E.; Reid, M.; O’Toole, P.W.; Cotter, P.D. The rumen microbiome: A crucial consideration when optimising milk and meat production and nitrogen utilisation efficiency. Gut Microbes 2019, 10, 115–132. [Google Scholar] [CrossRef] [PubMed]
- Kertz, A.F. Review: Urea feeding to dairy cattle: A historical perspective and review. Prof. Anim. Sci. 2010, 26, 257–272. [Google Scholar] [CrossRef]
- Lin, W.; Mathys, V.; Ang, E.L.; Koh, V.H.; Martínez Gómez, J.M.; Ang, M.L.; Zainul Rahim, S.Z.; Tan, M.P.; Pethe, K.; Alonso, S. Urease activity represents an alternative pathway for Mycobacterium tuberculosis nitrogen metabolism. Infect. Immun. 2012, 80, 2771–2779. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reynolds, C.K.; Kristensen, N.B. Nitrogen recycling through the gut and the nitrogen economy of ruminants: An asynchronous symbiosis. J. Anim. Sci. 2008, 86, E293–E305. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Powell, J.M.; Wattiaux, M.A.; Broderick, G.A. Short communication: Evaluation of milk urea nitrogen as a management tool to reduce ammonia emissions from dairy farms. J. Dairy Sci. 2011, 94, 4690–4694. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K. Urea/Ammonia Metabolism in the Rumen and Toxicity in Ruminants. In Rumen Microbiology: From Evolution to Revolution; Puniya, A.K., Singh, R., Eds.; Springer: New Delhi, India, 2015; pp. 329–341. [Google Scholar]
- Uwizeye, A.; de Boer, I.J.M.; Opio, C.I.; Schulte, R.P.O.; Falcucci, A.; Tempio, G.; Teillard, F.; Casu, F.; Rulli, M.; Galloway, J.N.; et al. Nitrogen emissions along global livestock supply chains. Nat. Food 2020, 1, 436–437. [Google Scholar] [CrossRef]
- Kohn, A.; Dinneen, M.; Russek-Cohen, E. Using blood urea nitrogen to predict nitrogen excretion and efficiency of nitrogen utilization in cattle, sheep, goats, horses, pigs, and rats. J. Anim. Sci. Technol. 2005, 83, 879–889. [Google Scholar] [CrossRef] [Green Version]
- Huhtanen, P.; Hristov, N. A meta-analysis of the effects of dietary protein concentration and degradability on milk protein yield and milk N efficiency in dairy cows. J. Dairy Sci. 2009, 92, 3222–3232. [Google Scholar] [CrossRef]
- Flachowsky, G.; Lebzien, P. Possibilities for reduction of nitrogen (N) excretion from ruminants and the need for further research—A review. Landbauforsch. Volkenrode 2006, 56, 19–30. [Google Scholar]
- Estiu, G.; Merz, K.M. Enzymatic catalysis of urea decomposition: Elimination or hydrolysis? J. Am. Chem. Soc. 2004, 126, 11832–11842. [Google Scholar] [CrossRef]
- Krajewska, B.; Zaborska, W.; Leszko, M. Inhibition of chitosan-immobilized urease by slow-binding inhibitors: Ni2+, F− and acetohydroxamic acid. J. Mol. Catal. B Enzym. 2001, 14, 101–109. [Google Scholar] [CrossRef]
- Domínguez, M.J.; Sanmartín, C.; Font, M.; Palop, J.A.; San Francisco, S.; Urrutia, O.; Houdusse, F.; García-Mina, J. Design, synthesis, and biological evaluation of phosphoramide derivatives as urease inhibitors. J. Agric. Food Chem. 2008, 56, 3721–3731. [Google Scholar] [CrossRef] [PubMed]
- Mazzei, L.; Cianci, M.; Musiani, F.; Ciurli, S. Inactivation of urease by 1,4-benzoquinone: Chemistry at the protein surface. Dalton Trans. 2016, 45, 5455–5459. [Google Scholar] [CrossRef] [Green Version]
- Mazzei, L.; Cianci, M.; Musiani, F.; Lente, G.; Palombo, M.; Ciurli, S. Inactivation of urease by catechol: Kinetics and structure. J. Inorg. Biochem. 2017, 166, 182–189. [Google Scholar] [CrossRef] [Green Version]
- Shaw, W.H.R.; Raval, D.N. The inhibition of urease by metal ions at pH 8. 9. J. Am. Chem. Soc. 1961, 83, 3184–3187. [Google Scholar] [CrossRef]
- Kosikowska, P.; Berlicki, L. Urease inhibitors as potential drugs for gastric and urinary tract infections: A patent review. Expert Opin. Ther. Pat. 2011, 21, 945–957. [Google Scholar] [CrossRef] [PubMed]
- Williams, J.J.; Rodman, J.S.; Peterson, C.M. A randomized double-blind study of acetohydroxamic acid in struvite nephrolithiasis. N. Engl. J. Med. 1984, 311, 760–764. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K.; Aschenbach, J.R. Ureases in the gastrointestinal tracts of ruminant and monogastric animals and their implication in urea-N/ammonia metabolism: A review. J. Adv. Res. 2018, 13, 39–50. [Google Scholar] [CrossRef]
- Shang, X.; Pan, H.; Li, M.; Miao, X.; Ding, H. Lonicera japonica Thunb.: Ethnopharmacology, phytochemistry and pharmacology of an important traditional Chinese medicine. J. Ethnopharmacol. 2011, 138, 1–21. [Google Scholar] [CrossRef]
- Zhang, M.; Wang, J.; Zhu, L.; Li, T.; Jiang, W.; Zhou, J.; Peng, W.; Wu, C. Zanthoxylum bungeanum maxim. (rutaceae): A systematic review of its traditional uses, botany, phytochemistry, pharmacology, pharmacokinetics, and toxicology. Int. J. Mol. Sci. 2017, 18, 2172. [Google Scholar] [CrossRef]
- Wu, J.; Luo, Y.; Deng, D.; Su, S.; Li, S.; Xiang, L.; Hu, Y.; Wang, P.; Meng, X. Coptisine from coptis chinensis exerts diverse beneficial properties: A concise review. J. Cell. Mol. Med. 2019, 23, 946–7960. [Google Scholar] [CrossRef] [Green Version]
- Pérard-Viret, J.; Quteishat, L.; Alsalim, R.; Royer, J.; Dumas, F. Cephalotaxus alkaloids. Alkaloids Chem. Biol. 2017, 78, 205–352. [Google Scholar]
- Modolo, L.V.; de Souza, A.X.; Horta, L.P.; Araujo, D.P.; de Fátima, Â. An overview on the potential of natural products as ureases inhibitors: A review. J. Adv. Res. 2015, 6, 35–44. [Google Scholar] [CrossRef] [Green Version]
- Gnetegha Ayemele, A.; Ma, L.; Park, T.; Xu, J.; Yu, Z.; Bu, D. Giant milkweed (Calotropis gigantea): A new plant resource to inhibit protozoa and decrease ammoniagenesis of rumen microbiota in vitro without impairing fermentation. Sci. Total Environ. 2020, 743, 140665. [Google Scholar] [CrossRef]
- Hassan, S.T.; Žemlička, M. Plant-Derived Urease Inhibitors as Alternative Chemotherapeutic Agents. Arch. Pharm. (Weinheim) 2016, 349, 507–522. [Google Scholar] [CrossRef] [PubMed]
- Jabri, E.; Carr, M.B.; Hausinger, R.P.; Karplus, P.A. The crystal structure of urease from Klebsiella aerogenes. Science 1995, 268, 998–1004. [Google Scholar] [CrossRef]
- Ha, N.C.; Oh, S.T.; Sung, J.Y.; Cha, K.A.; Lee, M.H.; Oh, B.H. Supramolecular assembly and acid resistance of Helicobacter pylori urease. Nat. Struct. Biol. 2001, 8, 505–509. [Google Scholar] [CrossRef] [PubMed]
- Leipe, D.D.; Wolf, Y.I.; Koonin, E.V.; Aravind, L. Classification and evolution of P-loop GTPases and related ATPases. J. Mol. Biol. 2002, 317, 41–72. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; Li, H.; Lai, T.P.; Sun, H. UreE-UreG complex facilitates nickel transfer and preactivates GTPase of UreG in Helicobacter pylori. J. Biol. Chem. 2015, 290, 12474–12485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuen, M.H.; Fong, Y.H.; Nim, Y.S.; Lau, P.H.; Wong, K.B. Structural insights into how GTP-dependent conformational changes in a metallochaperone UreG facilitate urease maturation. Proc. Natl. Acad. Sci. USA 2017, 114, E10890–E10898. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Farrugia, M.A.; Macomber, L.; Hausinger, R.P. Biosynthesis of the urease metallocenter. J. Biol. Chem. 2013, 288, 13178–13185. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Zhao, S.; He, Y.; Zheng, N.; Yan, X.; Wang, J. Substitution of residues in UreG to investigate UreE interactions and nickel binding in a predominant urease gene cluster from the ruminal metagenome. Int. J. Biol. Macromol. 2020, 161, 1591–1601. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Huang, P.; Wong, K.; Xu, Y.; Tan, L.; Chen, H.; Lu, Q.; Luo, C.; Tam, C.; Zhu, L.; et al. Coptisine-induced inhibition of Helicobacter pylori: Elucidation of specific mechanisms by probing urease active site and its maturation process. J. Enzyme Inhib. Med. Chem. 2018, 33, 1362–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dvorakova, M.; Landa, P. Anti-inflammatory activity of natural stilbenoids: A review. Pharmacol. Res. 2017, 124, 126–145. [Google Scholar] [CrossRef]
- Lichota, A.; Gwozdzinski, K. Anticancer Activity of Natural Compounds from Plant and Marine Environment. Int. J. Mol. Sci. 2018, 19, 3533. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mani, J.S.; Johnson, J.B.; Steel, J.C.; Broszczak, D.A.; Neilsen, P.M.; Walsh, K.B.; Naiker, M. Natural product-derived phytochemicals as potential agents against coronaviruses: A review. Virus Res. 2020, 284, 197989. [Google Scholar] [CrossRef]
- Patra, A.K.; Saxena, J. The effect and mode of action of saponins on the microbial populations and fermentation in the rumen and ruminant production. Nutr. Res. Rev. 2009, 22, 204–219. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K.; Saxena, J. A new perspective on the use of plant secondary metabolites to inhibit methanogenesis in the rumen. Phytochemistry 2010, 71, 1198–1222. [Google Scholar] [CrossRef] [PubMed]
- Cobellis, G.; Trabalza-Marinucci, M.; Yu, Z. Critical evaluation of essential oils as rumen modifiers in ruminant nutrition: A review. Sci. Total Environ. 2016, 545, 556–568. [Google Scholar] [CrossRef]
- Maroney, M.J.; Ciurli, S. Nonredox nickel enzymes. Chem. Rev. 2014, 114, 4206–4228. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Luo, C.; Shan, C.; You, Q.; Lu, J.; Elf, S.; Zhou, Y.; Wen, Y.; Vinkenborg, J.L.; Fan, J.; et al. Inhibition of human copper trafficking by a small molecule significantly attenuates cancer cell proliferation. Nat. Chem. 2015, 7, 968–979. [Google Scholar] [CrossRef] [Green Version]
- Lu, Q.; Li, C.; Wu, G. Insight into the inhibitory effects of Zanthoxylum nitidum against Helicobacter pylori urease and jack bean urease: Kinetics and mechanism. J. Ethnopharmacol. 2020, 249, 112419. [Google Scholar] [CrossRef]
- Mahady, G.B.; Pendland, S.L.; Stoia, A.; Chadwick, L.R. In vitro susceptibility of Helicobacter pylori to isoquinoline alkaloids from Sanguinaria canadensis and Hydrastis canadensis. Phytother. Res. 2003, 17, 217–221. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Koohi-Moghadam, M.; Wang, R.; Chang, Y.Y.; Woo, P.C.Y.; Wang, J.; Li, H.; Sun, H. Metallochaperone UreG serves as a new target for design of urease inhibitor: A novel strategy for development of antimicrobials. PLoS Biol. 2018, 16, e2003887. [Google Scholar] [CrossRef]
- Kosina, P.; Gregorova, J.; Gruz, J.; Vacek, J.; Kolar, M.; Vogel, M.; Roos, W.; Naumann, K.; Simanek, V.; Ulrichova, J. Phytochemical and antimicrobial characterization of Macleaya cordata herb. Fitoterapia 2010, 81, 1006–1012. [Google Scholar] [CrossRef]
- Hu, N.X.; Chen, M.; Liu, Y.S.; Shi, Q.; Yang, B.; Zhang, H.C.; Cheng, P.; Tang, Q.; Liu, Z.Y.; Zeng, J.G. Pharmacokinetics of sanguinarine, chelerythrine, and their metabolites in broiler chickens following oral and intravenous administration. J. Vet. Pharmacol. Ther. 2019, 42, 197–206. [Google Scholar] [CrossRef]
- Zhang, X.; Zhao, S.; He, Y.; Zheng, N.; Yan, X.; Wang, J. Pipeline for Targeted Meta-Proteomic Analyses to Assess the Diversity of Cattle Rumen Microbial Urease. Front. Microbiol. 2020, 11, 573414. [Google Scholar] [CrossRef]
- Weatherburn, M. Phenol-hypochlorite reaction for determination of ammonia. Anal. Chem. 1967, 39, 971–973. [Google Scholar] [CrossRef]
- García S anchez, F.; NavasDíaz, A.; Ramos Peinado, M.C.; Belledone, C. Free and sol-gel immobilized alkaline phosphatase-based biosensor for the determination of pesticides and inorganic compounds. Anal. Chim. Acta 2003, 484, 45–51. [Google Scholar] [CrossRef]
- Mascarello, A.; Chiaradia, L.D.; Vernal, J.; Villarino, A.; Guido, R.V.C.; Perizzolo, P.; Poirier, V.; Wong, D.; Martins, P.G.A.; Nunes, R.J.; et al. Inhibition of Mycobacterium tuberculosis tyrosine phosphatase PtpA by synthetic chalcones: Kinetics, molecular modeling, toxicity and effect on growth. Bioorgan. Med. Chem. 2010, 18, 3783–3789. [Google Scholar] [CrossRef] [PubMed]
- Turner, T.L.; Nguyen, V.H.; McLauchlan, C.C.; Dymon, Z.; Dorsey, B.M.; Hooker, J.D.; Jones, M.A. Inhibitory effects of decavanadate on several enzymes and leishmania tarentolae in vitro. J. Inorg. Biochem. 2012, 108, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Jaiswal, P.; Singh, V.K.; Singh, D.K. Enzyme inhibition by molluscicidal component of Areca catechu and Carica papaya in the nervous tissue of vector snail Lymnaea acuminate. Pest. Biochem. Physiol. 2008, 92, 164–168. [Google Scholar] [CrossRef]
- Kari, J.; Schiano-di-Cola, C.; Hansen, S.F.; Badino, S.F.; Sørensen, T.H.; Cavaleiro, A.M.; Bzrch, K.; Westh, P. A steady-state approach for inhibition of heterogeneous enzyme reactions. Biochem. J. 2020, 477, 1971–1982. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.R.; Yang, K.; Li, W.T.; Pang, Z.H.; Zhang, Q.; Wang, J.J.; Dang, W.X.; Jia, R.Y.; Fu, Z.W.; Li, Y.X.; et al. Evaluation of the inhibition of chlorophenols towards human cytochrome P450 3A4 and differences among various species. Sci. Total Environ. 2020, 724, 138187. [Google Scholar] [CrossRef] [PubMed]
- Anand, U.; Jash, C.; Mukherjee, S. Protein unfolding and subsequent refolding: A spectroscopic investigation. Phys. Chem. Chem. Phys. 2011, 13, 20418–20426. [Google Scholar] [CrossRef] [PubMed]
- Anand, U.; Jash, C.; Mukherjee, S. Spectroscopic probing of the microenvironment in a protein-surfactant assembly. J. Phys. Chem. B 2010, 114, 15839–15845. [Google Scholar] [CrossRef] [PubMed]
- Anand, U.; Ray, S.; Ghosh, S.; Banerjee, R.; Mukherjee, S. Structural aspects of a protein-surfactant assembly: Native and reduced States of human serum albumin. Protein J. 2015, 34, 147–157. [Google Scholar] [CrossRef]
- Mondal, R.; Ghosh, N.; Paul, B.K.; Mukherjee, S. Triblock-Copolymer-Assisted Mixed-Micelle Formation Results in the Refolding of Unfolded Protein. Langmuir 2018, 34, 896–903. [Google Scholar] [CrossRef]
| Compound | CAS Number | Source | Structure | Binding Energy (kcal/mol) | Compound (Ki) |
|---|---|---|---|---|---|
| Coptisine chloride | 6020-18-4 | Coptidis rhizoma | 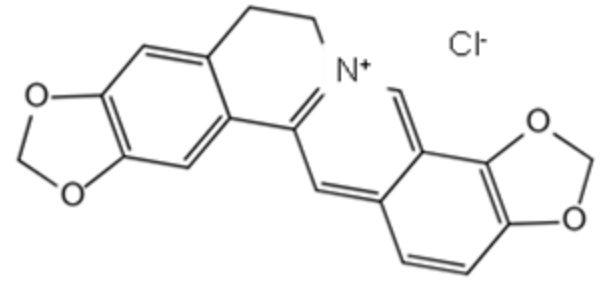 | −6.47 | 17.97 μM |
| Isochlorogenic acid C | 32451-88-0 | Honeysuckle | 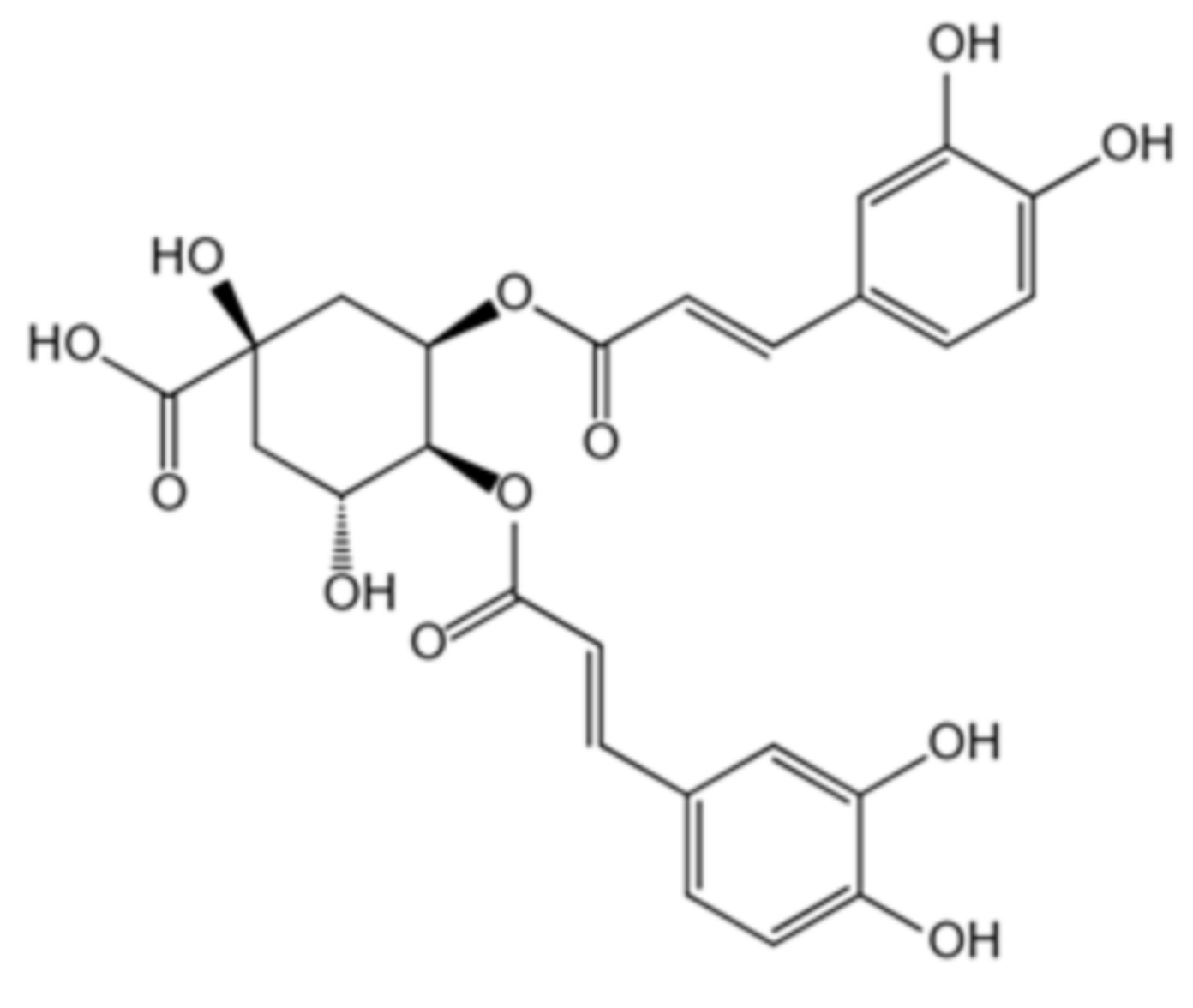 | −2.82 | 8.56 mM |
| Chelerythrine chloride | 3895-92-9 | Celandine |  | −6.72 | 11.88 μM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.; He, Y.; Xiong, Z.; Li, M.; Li, M.; Zheng, N.; Zhao, S.; Wang, J. Chelerythrine Chloride: A Potential Rumen Microbial Urease Inhibitor Screened by Targeting UreG. Int. J. Mol. Sci. 2021, 22, 8212. https://doi.org/10.3390/ijms22158212
Zhang X, He Y, Xiong Z, Li M, Li M, Zheng N, Zhao S, Wang J. Chelerythrine Chloride: A Potential Rumen Microbial Urease Inhibitor Screened by Targeting UreG. International Journal of Molecular Sciences. 2021; 22(15):8212. https://doi.org/10.3390/ijms22158212
Chicago/Turabian StyleZhang, Xiaoyin, Yue He, Zhanbo Xiong, Min Li, Ming Li, Nan Zheng, Shengguo Zhao, and Jiaqi Wang. 2021. "Chelerythrine Chloride: A Potential Rumen Microbial Urease Inhibitor Screened by Targeting UreG" International Journal of Molecular Sciences 22, no. 15: 8212. https://doi.org/10.3390/ijms22158212
APA StyleZhang, X., He, Y., Xiong, Z., Li, M., Li, M., Zheng, N., Zhao, S., & Wang, J. (2021). Chelerythrine Chloride: A Potential Rumen Microbial Urease Inhibitor Screened by Targeting UreG. International Journal of Molecular Sciences, 22(15), 8212. https://doi.org/10.3390/ijms22158212








