The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus
Abstract
:1. Introduction
2. Results
2.1. HapX Is Rapidly Degraded during Adaptation from Iron-Depleted to Iron-Replete Conditions in a Proteasome Dependent Manner
2.2. HapX Turnover Is Very Fast during Iron Limitation
2.3. HapX Affinity Purification Studies Identified Protein Interactors and Post-Translational Modifications during the Adaptation from Iron-Deficiency to Iron-Replete Conditions
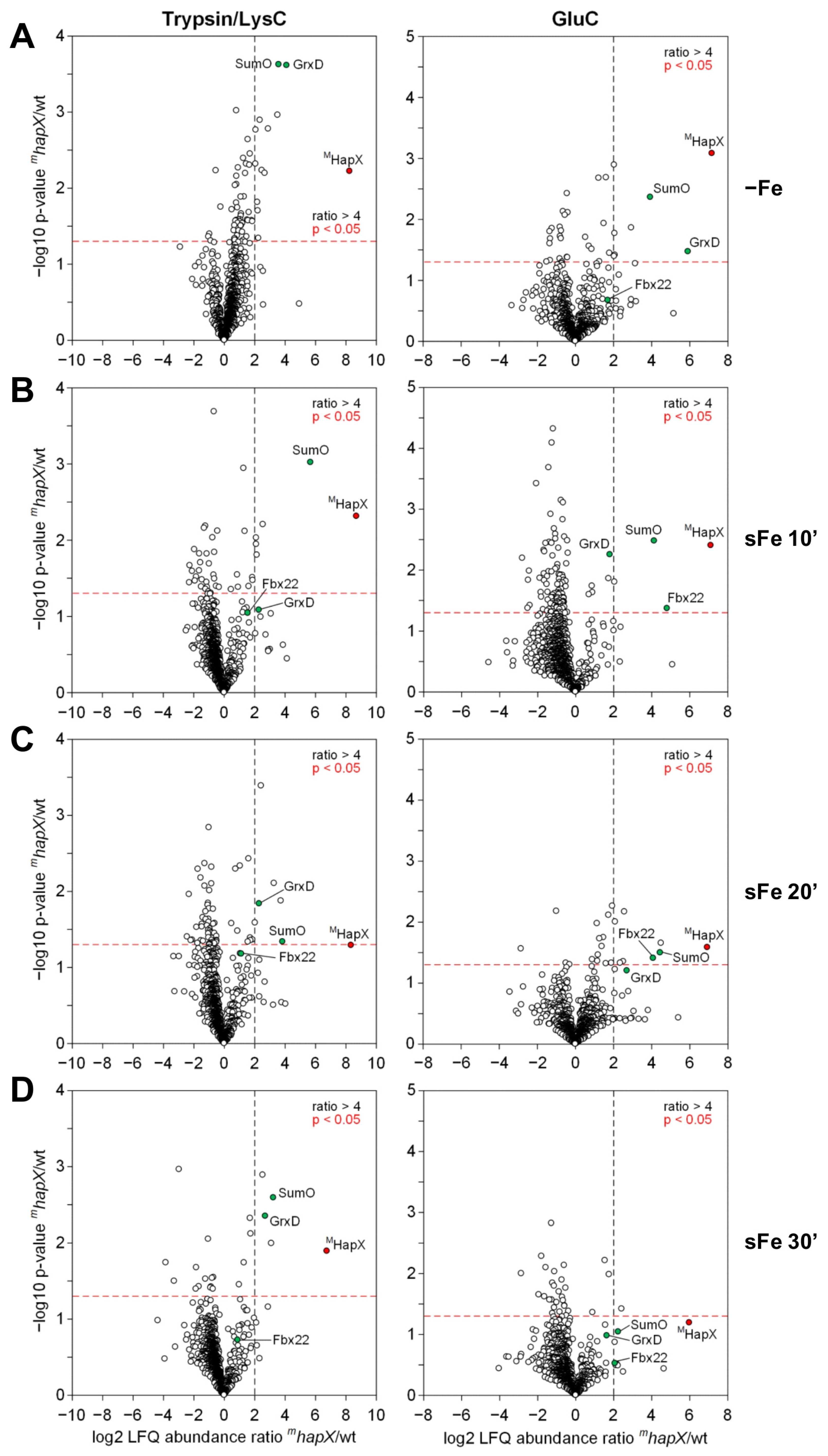
2.3.1. HapX Specifically Interacts with Fbx22 and SumO during sFe Conditions
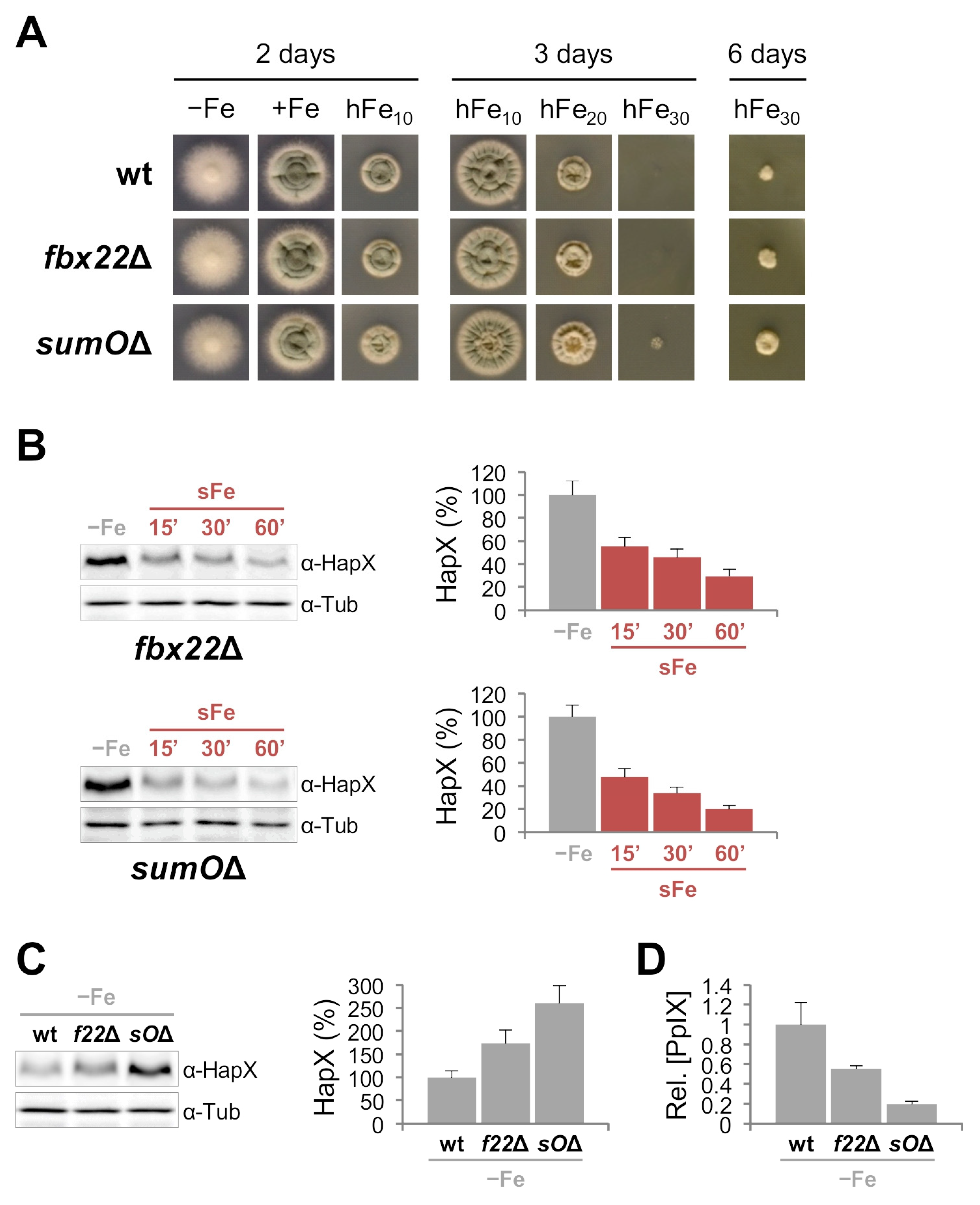
Inactivation of Fbx22 and SumO Affects Iron Resistance, HapX Levels and Degradation and Heme Biosynthesis
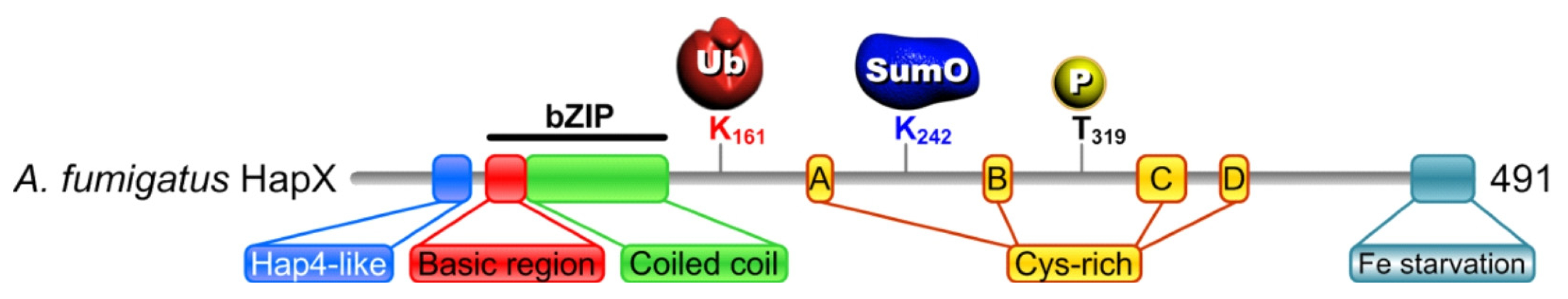
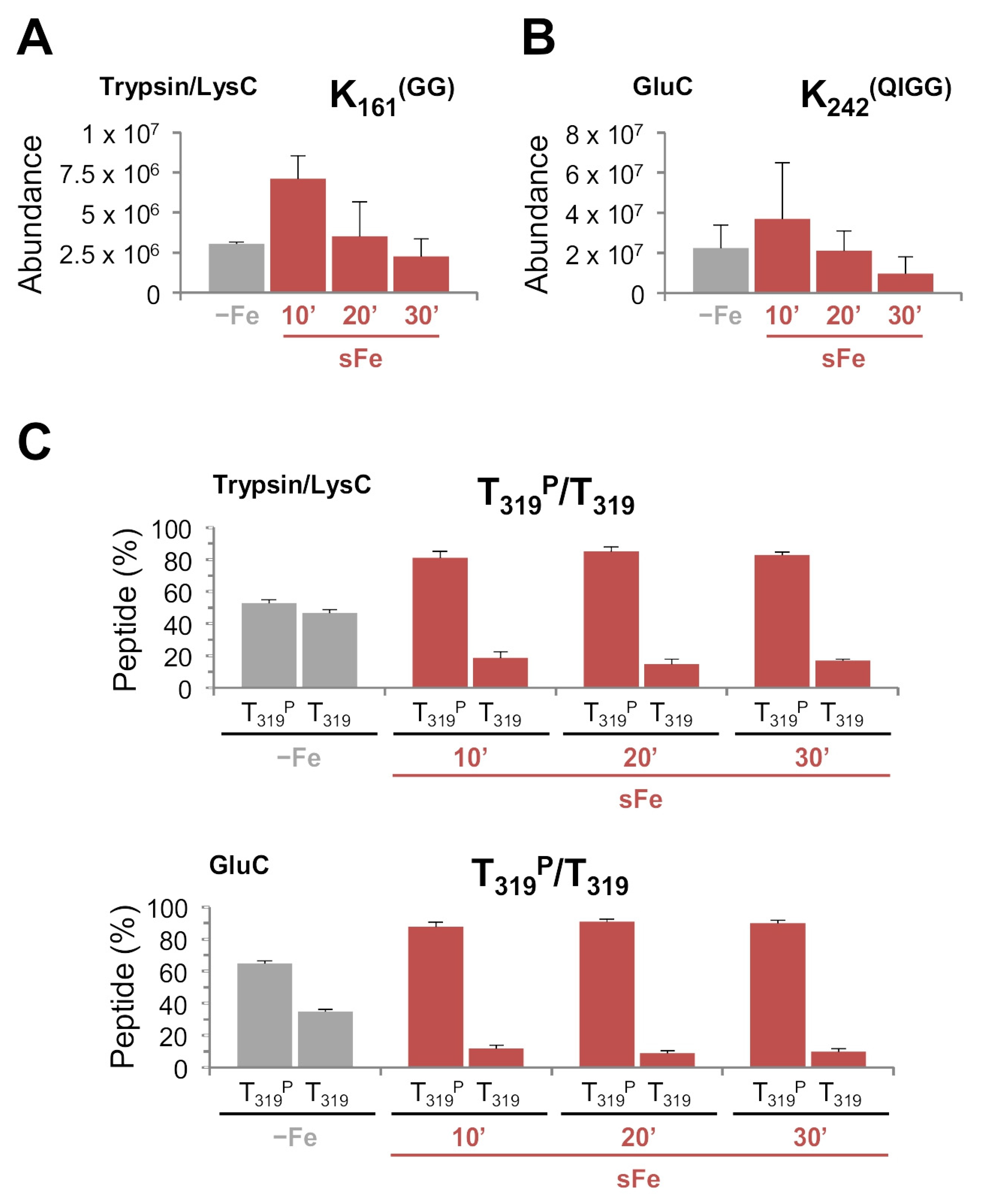
2.3.2. HapX Is Ubiquitinated, Sumoylated and Phosphorylated during sFe Conditions
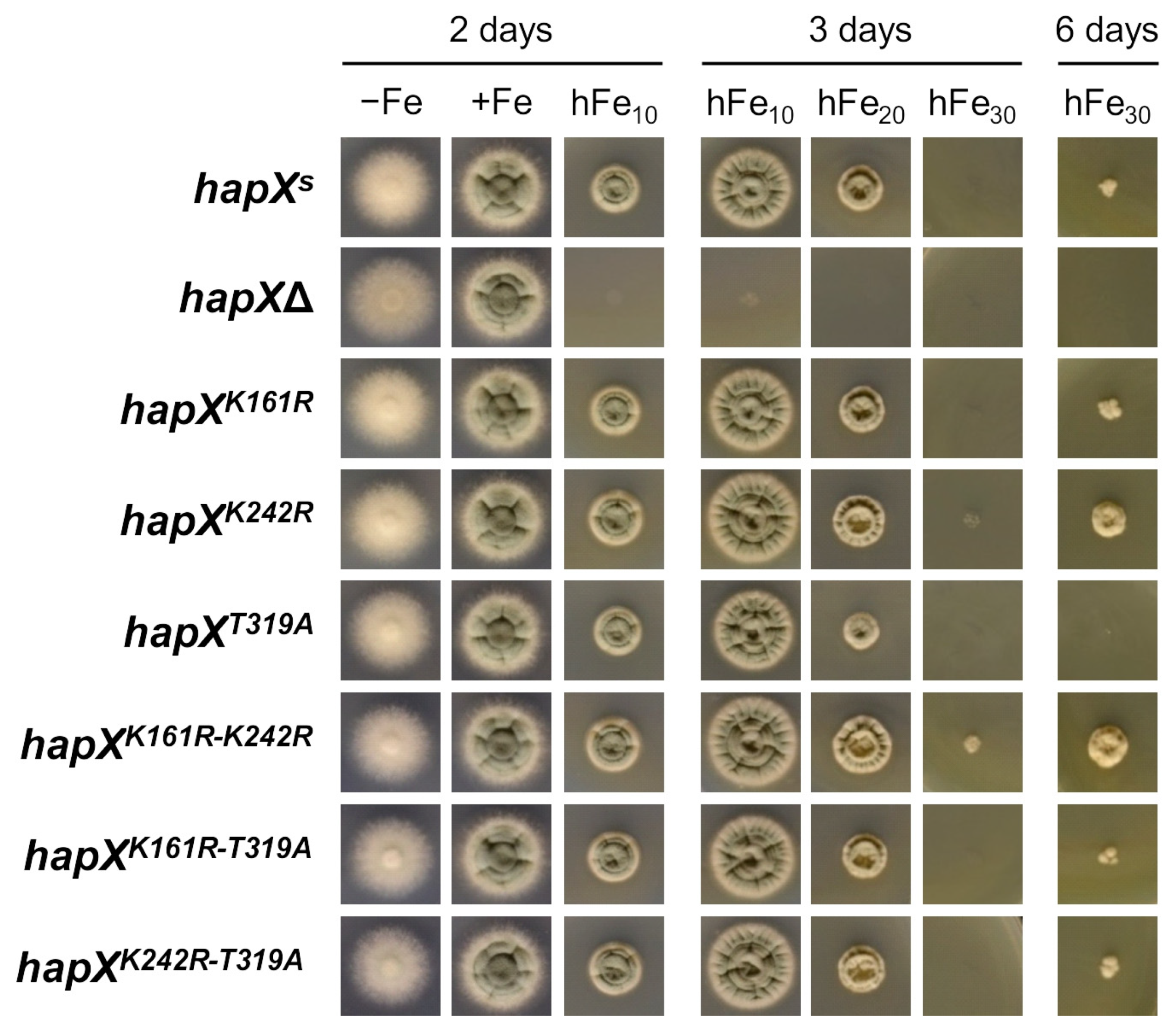
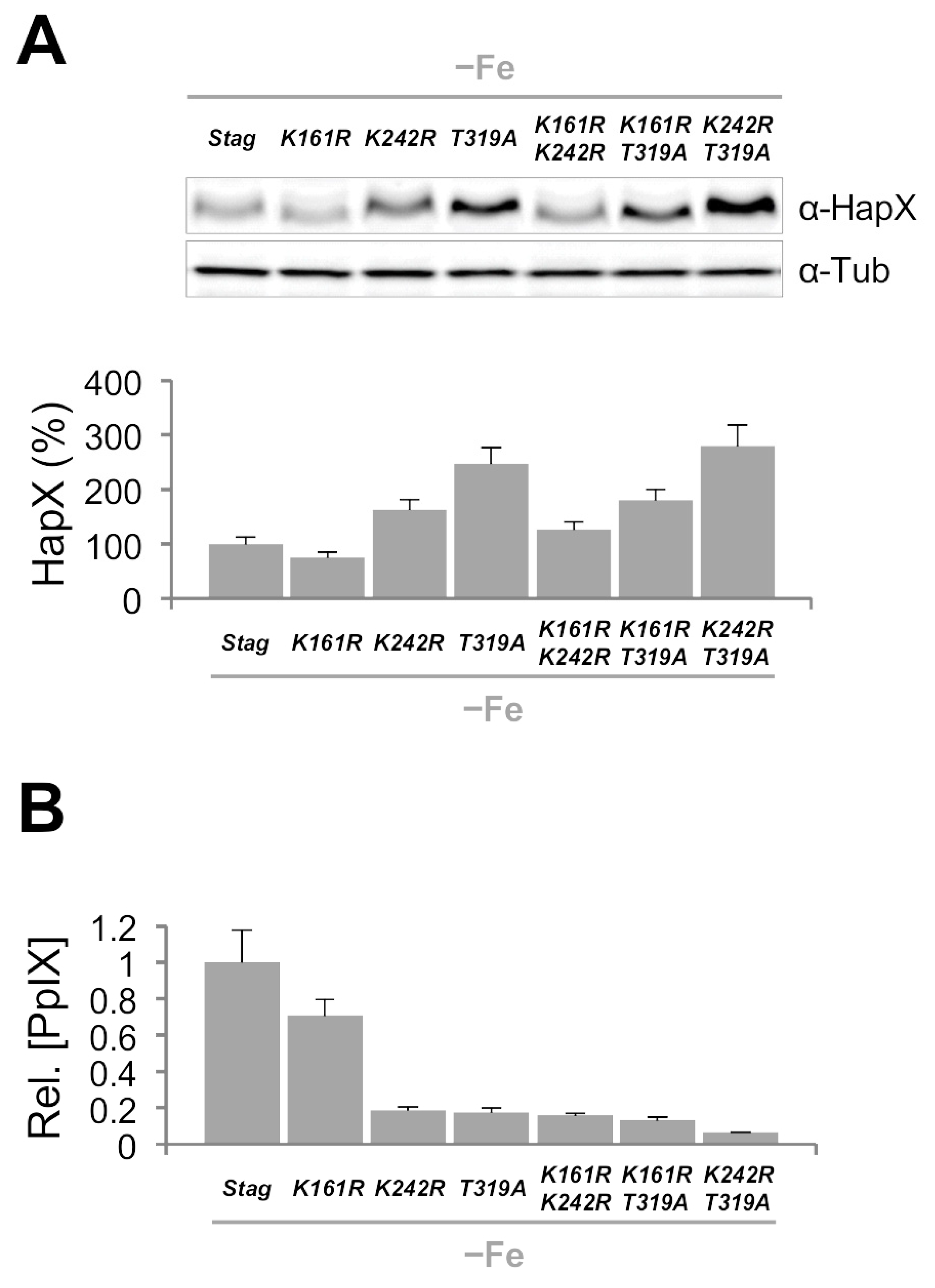
Point Mutations in HapX K161, K242 and T319 Affect Iron Resistance, HapX Levels and Degradation and Heme Biosynthesis
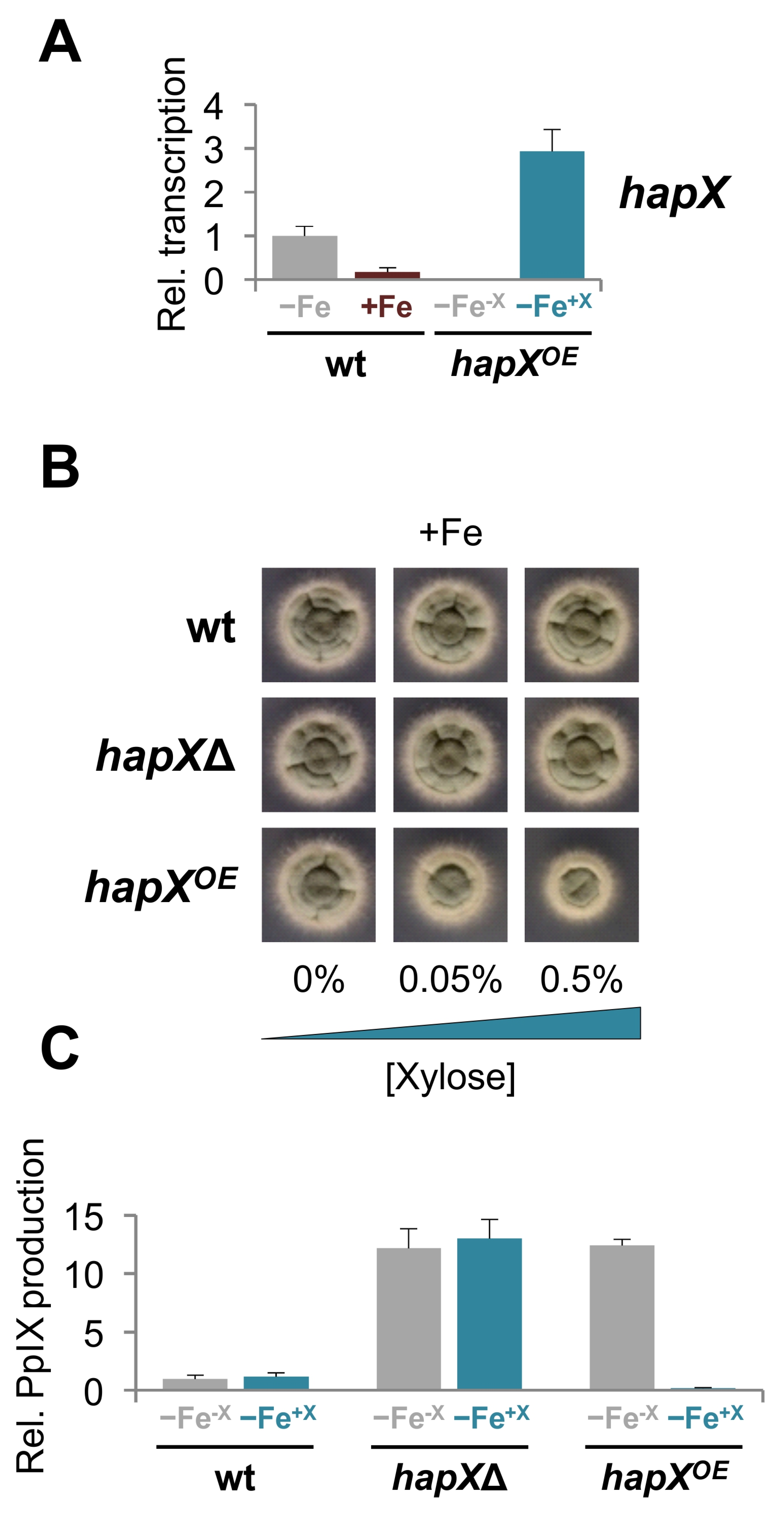
2.4. Overexpression of hapX Negatively Affects Growth and Production of Iron-Related Metabolites
3. Discussion
3.1. HapX Is Regulated at the Protein Level
3.2. HapX Interacts with Fbx22 and SumO and Is Prost-Translationally Modified Specifically under sFe Conditions
4. Materials and Methods
4.1. Fungal Isolates and Culture Conditions
4.2. Generation of Mutant Strains
4.3. Nucleic Acid Manipulations, Quantitative Real-Time RT-PCR and PpIX Analysis
4.4. Western Blotting
4.5. Myc-Trap Immunoprecipitation of MHapX Fusion Protein
4.6. In-Solution Digest of Myc-Trap Eluates
4.7. nLC-MS/MS Analysis
4.8. Protein Database Search
4.9. Western Blot Detection of Proteins after Myc-Trap
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| RIA | Reductive iron assimilation |
| CBC | CCAAT-binding complex |
| CRR | Cysteine rich region |
| −Fe | Iron-deplete |
| +Fe | Iron-replete |
| sFe | Shift from iron-depleted to iron-replete |
| FsC | Fusarinine C |
| TAFC | Triacetylfusarinine C |
| chx | Cycloheximide |
| LFQ | Label-free quantification abundances |
| SCF | Skp, Cullin, F-box |
| hFe | high iron |
| PpIX | Protoporphyrin IX |
| NDSM | Negatively charged amino acid-dependent SUMOylation motif |
| AMM | Aspergillus minimal medium |
| TCA | Trichloroacetic acid |
References
- De Hoog, G.S.; Guarro, J. Atlas of Clinical Fungi; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 1995. [Google Scholar]
- Tekaia, F.; Latge, J.P. Aspergillus fumigatus: Saprophyte or pathogen? Curr. Opin. Microbiol. 2005, 8, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Denning, D.W.; Gow, N.A.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden killers: Human fungal infections. Sci. Transl. Med. 2012, 4, 165rv13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bongomin, F.; Gago, S.; Oladele, R.O.; Denning, D.W. Global and Multi-National Prevalence of Fungal Diseases-Estimate Precision. J. Fungi 2017, 3, 57. [Google Scholar] [CrossRef] [PubMed]
- Brakhage, A.A. Systemic fungal infections caused by Aspergillus species: Epidemiology, infection process and virulence determinants. Curr. Drug Targets 2005, 6, 875–886. [Google Scholar] [CrossRef] [PubMed]
- Fisher, M.C.; Henk, D.A.; Briggs, C.J.; Brownstein, J.S.; Madoff, L.C.; McCraw, S.L.; Gurr, S.J. Emerging fungal threats to animal, plant and ecosystem health. Nature 2012, 484, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Calo, S.; Shertz-Wall, C.; Lee, S.C.; Bastidas, R.J.; Nicolas, F.E.; Granek, J.A.; Mieczkowski, P.; Torres-Martinez, S.; Ruiz-Vazquez, R.M.; Cardenas, M.E.; et al. Antifungal drug resistance evoked via RNAi-dependent epimutations. Nature 2014, 513, 555–558. [Google Scholar] [CrossRef] [Green Version]
- Robbins, N.; Caplan, T.; Cowen, L.E. Molecular Evolution of Antifungal Drug Resistance. Annu. Rev. Microbiol. 2017, 71, 753–775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halliwell, B.; Gutteridge, J.M.C. Oxygen-toxicity, oxygen radicals, transition-metals and disease. Biochem. J. 1984, 219, 1–14. [Google Scholar] [CrossRef]
- Lin, H.; Li, L.; Jia, X.; Ward, D.M.; Kaplan, J. Genetic and biochemical analysis of high iron toxicity in yeast: Iron toxicity is due to the accumulation of cytosolic iron and occurs under both aerobic and anaerobic conditions. J. Biol. Chem. 2011, 286, 3851–3862. [Google Scholar] [CrossRef] [Green Version]
- Schrettl, M.; Beckmann, N.; Varga, J.; Heinekamp, T.; Jacobsen, I.D.; Joechl, C.; Moussa, T.A.; Wang, S.; Gsaller, F.; Blatzer, M.; et al. HapX-Mediated Adaption to Iron Starvation Is Crucial for Virulence of Aspergillus fumigatus. PLoS Pathog. 2010, 6, e1001124. [Google Scholar] [CrossRef] [Green Version]
- Lopez-Berges, M.S.; Capilla, J.; Turra, D.; Schafferer, L.; Matthijs, S.; Joechl, C.; Cornelis, P.; Guarro, J.; Haas, H.; Di Pietro, A. HapX-mediated iron homeostasis is essential for rhizosphere competence and virulence of the soilborne pathogen Fusarium oxysporum. Plant Cell 2012, 24, 3805–3822. [Google Scholar] [CrossRef] [Green Version]
- Hsu, P.-C.; Yang, C.-Y.; Lan, C.-Y. Candida albicans Hap43 is a repressor induced under low-iron conditions and is essential for iron-responsive transcriptional regulation and virulence. Eukaryot. Cell 2011, 10, 207–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jung, W.H.; Saikia, S.; Hu, G.; Wang, J.; Fung, C.K.-Y.; D’Souza, C.; White, R.; Kronstad, J.W. HapX Positively and Negatively Regulates the Transcriptional Response to Iron Deprivation in Cryptococcus neoformans. PLoS Pathog. 2010, 6, e1001209. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Deng, C.; Tian, L.; Xiong, D.; Tian, C.; Klosterman, S.J. The Transcription Factor VdHapX Controls Iron Homeostasis and Is Crucial for Virulence in the Vascular Pathogen Verticillium dahliae. Msphere 2018, 3, e00400-18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrettl, M.; Kim, H.S.; Eisendle, M.; Kragl, C.; Nierman, W.C.; Heinekamp, T.; Werner, E.R.; Jacobsen, I.; Illmer, P.; Yi, H.; et al. SreA-mediated iron regulation in Aspergillus fumigatus. Mol. Microbiol. 2008, 70, 27–43. [Google Scholar] [CrossRef] [Green Version]
- Gsaller, F.; Hortschansky, P.; Beattie, S.R.; Klammer, V.; Tuppatsch, K.; Lechner, B.E.; Rietzschel, N.; Werner, E.R.; Vogan, A.A.; Chung, D.; et al. The Janus transcription factor HapX controls fungal adaptation to both iron starvation and iron excess. EMBO J. 2014, 33, 2261–2276. [Google Scholar] [CrossRef]
- Misslinger, M.; Scheven, M.T.; Hortschansky, P.; Lopez-Berges, M.S.; Heiss, K.; Beckmann, N.; Heigl, T.; Hermann, M.; Kruger, T.; Kniemeyer, O.; et al. The monothiol glutaredoxin GrxD is essential for sensing iron starvation in Aspergillus fumigatus. PLoS Genet. 2019, 15, e1008379. [Google Scholar] [CrossRef] [Green Version]
- Misslinger, M.; Lechner, B.E.; Bacher, K.; Haas, H. Iron-sensing is governed by mitochondrial, not by cytosolic iron-sulfur cluster biogenesis in Aspergillus fumigatus. Met. Integr. Biometal Sci. 2018, 10, 1687–1700. [Google Scholar] [CrossRef] [Green Version]
- Hortschansky, P.; Eisendle, M.; Al-Abdallah, Q.; Schmidt, A.D.; Bergmann, S.; Thon, M.; Kniemeyer, O.; Abt, B.; Seeber, B.; Werner, E.R.; et al. Interaction of HapX with the CCAAT-binding complex—A novel mechanism of gene regulation by iron. Embo J. 2007, 26, 3157–3168. [Google Scholar] [CrossRef] [Green Version]
- Hortschansky, P.; Haas, H.; Huber, E.M.; Groll, M.; Brakhage, A.A. The CCAAT-binding complex (CBC) in Aspergillus species. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 560–570. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, T.; Scheven, M.T.; Misslinger, M.; Zhao, C.; Hoefgen, S.; Gsaller, F.; Lau, J.; Jochl, C.; Donaldson, I.; Valiante, V.; et al. The fungal CCAAT-binding complex and HapX display highly variable but evolutionary conserved synergetic promoter-specific DNA recognition. Nucleic Acids Res. 2020, 48, 3567–3590. [Google Scholar] [CrossRef] [PubMed]
- Schrettl, M.; Haas, H. Iron homeostasis-Achilles’ heel of Aspergillus fumigatus? Curr. Opin. Microbiol. 2011, 14, 400–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schrettl, M.; Bignell, E.; Kragl, C.; Sabiha, Y.; Loss, O.; Eisendle, M.; Wallner, A.; Arst, H.N., Jr.; Haynes, K.; Haas, H. Distinct roles for intra- and extracellular siderophores during Aspergillus fumigatus infection. PLoS Pathog. 2007, 3, e128. [Google Scholar] [CrossRef] [PubMed]
- Yasmin, S.; Alcazar-Fuoli, L.; Gruendlinger, M.; Puempel, T.; Cairns, T.; Blatzer, M.; Lopez, J.F.; Grimalt, J.O.; Bignell, E.; Haas, H. Mevalonate governs interdependency of ergosterol and siderophore biosyntheses in the fungal pathogen Aspergillus fumigatus. Proc. Natl. Acad. Sci. USA 2012, 109, E497–E504. [Google Scholar] [CrossRef] [Green Version]
- Lee, D.H.; Goldberg, A.L. Selective inhibitors of the proteasome-dependent and vacuolar pathways of protein degradation in Saccharomyces cerevisiae. J. Biol. Chem. 1996, 271, 27280–27284. [Google Scholar] [CrossRef] [Green Version]
- Schneider-Poetsch, T.; Ju, J.; Eyler, D.E.; Dang, Y.; Bhat, S.; Merrick, W.C.; Green, R.; Shen, B.; Liu, J.O. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nat. Chem. Biol. 2010, 6, 209–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Deshpande, I.; Keusch, J.J.; Challa, K.; Iesmantavicius, V.; Gasser, S.M.; Gut, H. The Sir4 H-BRCT domain interacts with phospho-proteins to sequester and repress yeast heterochromatin. EMBO J. 2019, 38, e101744. [Google Scholar] [CrossRef]
- Skowyra, D.; Craig, K.L.; Tyers, M.; Elledge, S.J.; Harper, J.W. F-box proteins are receptors that recruit phosphorylated substrates to the SCF ubiquitin-ligase complex. Cell 1997, 91, 209–219. [Google Scholar] [CrossRef] [Green Version]
- Jonkers, W.; Rep, M. Lessons from fungal F-box proteins. Eukaryot Cell 2009, 8, 677–695. [Google Scholar] [CrossRef] [Green Version]
- de Assis, L.J.; Ulas, M.; Ries, L.N.A.; El Ramli, N.A.M.; Sarikaya-Bayram, O.; Braus, G.H.; Bayram, O.; Goldman, G.H. Regulation of Aspergillus nidulans CreA-Mediated Catabolite Repression by the F-Box Proteins Fbx23 and Fbx47. MBio 2018, 9, e00840-18. [Google Scholar] [CrossRef] [Green Version]
- Welchman, R.L.; Gordon, C.; Mayer, R.J. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat. Rev. Mol. Cell Biol. 2005, 6, 599–609. [Google Scholar] [CrossRef] [PubMed]
- Harting, R.; Bayram, O.; Laubinger, K.; Valerius, O.; Braus, G.H. Interplay of the fungal sumoylation network for control of multicellular development. Mol. Microbiol. 2013, 90, 1125–1145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wong, K.H.; Todd, R.B.; Oakley, B.R.; Oakley, C.E.; Hynes, M.J.; Davis, M.A. Sumoylation in Aspergillus nidulans: sumO inactivation, overexpression and live-cell imaging. Fungal Genet. Biol. 2008, 45, 728–737. [Google Scholar] [CrossRef] [Green Version]
- Sahu, M.S.; Patra, S.; Kumar, K.; Kaur, R. SUMOylation in Human Pathogenic Fungi: Role in Physiology and Virulence. J. Fungi 2020, 6, 32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, S.H.; Galanis, A.; Witty, J.; Sharrocks, A.D. An extended consensus motif enhances the specificity of substrate modification by SUMO. EMBO J. 2006, 25, 5083–5093. [Google Scholar] [CrossRef] [Green Version]
- Zadra, I.; Abt, B.; Parson, W.; Haas, H. xylP promoter-based expression system and its use for antisense downregulation of the Penicillium chrysogenum nitrogen regulator NRE. Appl. Environ. Microbiol. 2000, 66, 4810–4816. [Google Scholar] [CrossRef] [Green Version]
- Tanaka, A.; Kato, M.; Nagase, T.; Kobayashi, T.; Tsukagoshi, N. Isolation of genes encoding novel transcription factors which interact with the Hap complex from Aspergillus species. Biochim. Biophys. Acta Gene Struct. Expr. 2002, 1576, 176–182. [Google Scholar] [CrossRef]
- Misslinger, M.; Hortschansky, P.; Brakhage, A.A.; Haas, H. Fungal iron homeostasis with a focus on Aspergillus fumigatus. Biochim. Biophys. Acta Mol. Cell Res. 2021, 1868, 118885. [Google Scholar] [CrossRef]
- Bairwa, G.; Hee Jung, W.; Kronstad, J.W. Iron acquisition in fungal pathogens of humans. Met. Integr. Biometal Sci. 2017, 9, 215–227. [Google Scholar] [CrossRef] [Green Version]
- Gerwien, F.; Skrahina, V.; Kasper, L.; Hube, B.; Brunke, S. Metals in fungal virulence. FEMS Microbiol. Rev. 2018, 42, fux050. [Google Scholar] [CrossRef] [Green Version]
- Labbe, S.; Khan, M.G.; Jacques, J.F. Iron uptake and regulation in Schizosaccharomyces pombe. Curr. Opin. Microbiol. 2013, 16, 669–676. [Google Scholar] [CrossRef] [PubMed]
- Cooper, G.M. The Cell: A Molecular Approach, 8th ed.; Oxford University Press: New York, NY, USA, 2019. [Google Scholar]
- Collins, G.A.; Gomez, T.A.; Deshaies, R.J.; Tansey, W.P. Combined chemical and genetic approach to inhibit proteolysis by the proteasome. Yeast 2010, 27, 965–974. [Google Scholar] [CrossRef] [Green Version]
- Impens, F.; Radoshevich, L.; Cossart, P.; Ribet, D. Mapping of SUMO sites and analysis of SUMOylation changes induced by external stimuli. Proc. Natl. Acad. Sci. USA 2014, 111, 12432–12437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nie, X.; Yu, S.; Qiu, M.; Wang, X.; Wang, Y.; Bai, Y.; Zhang, F.; Wang, S. Aspergillus flavus SUMO Contributes to Fungal Virulence and Toxin Attributes. J. Agric. Food Chem. 2016, 64, 6772–6782. [Google Scholar] [CrossRef]
- Horta, M.A.C.; Thieme, N.; Gao, Y.; Burnum-Johnson, K.E.; Nicora, C.D.; Gritsenko, M.A.; Lipton, M.S.; Mohanraj, K.; de Assis, L.J.; Lin, L.; et al. Broad Substrate-Specific Phosphorylation Events Are Associated with the Initial Stage of Plant Cell Wall Recognition in Neurospora crassa. Front. Microbiol. 2019, 10, 2317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hartwell, L.H. Genetic control of the cell division cycle in yeast. II. Genes controlling DNA replication and its initiation. J. Mol. Biol. 1971, 59, 183–194. [Google Scholar] [CrossRef]
- Zhang, C. Essential functions of iron-requiring proteins in DNA replication, repair and cell cycle control. Protein Cell 2014, 5, 750–760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sadowski, M.; Suryadinata, R.; Tan, A.R.; Roesley, S.N.; Sarcevic, B. Protein monoubiquitination and polyubiquitination generate structural diversity to control distinct biological processes. IUBMB Life 2012, 64, 136–142. [Google Scholar] [CrossRef]
- Giaever, G.; Chu, A.M.; Ni, L.; Connelly, C.; Riles, L.; Veronneau, S.; Dow, S.; Lucau-Danila, A.; Anderson, K.; Andre, B.; et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002, 418, 387–391. [Google Scholar] [CrossRef]
- Rosonina, E.; Akhter, A.; Dou, Y.; Babu, J.; Sri Theivakadadcham, V.S. Regulation of transcription factors by sumoylation. Transcription 2017, 8, 220–231. [Google Scholar] [CrossRef] [Green Version]
- Boulanger, M.; Chakraborty, M.; Tempe, D.; Piechaczyk, M.; Bossis, G. SUMO and Transcriptional Regulation: The Lessons of Large-Scale Proteomic, Modifomic and Genomic Studies. Molecules 2021, 26, 828. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, K.; Nishide, J.; Okazaki, K.; Kato, H.; Niwa, O.; Nakagawa, T.; Matsuda, H.; Kawamukai, M.; Murakami, Y. Characterization of a fission yeast SUMO-1 homologue, pmt3p, required for multiple nuclear events, including the control of telomere length and chromosome segregation. Mol. Cell Biol. 1999, 19, 8660–8672. [Google Scholar] [CrossRef] [Green Version]
- Hunter, T. The age of crosstalk: Phosphorylation, ubiquitination, and beyond. Mol. Cell 2007, 28, 730–738. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, T.; Dumig, M.; Jaber, B.M.; Szewczyk, E.; Olbermann, P.; Morschhauser, J.; Krappmann, S. Validation of a self-excising marker in the human pathogen Aspergillus fumigatus by employing the beta-rec/six site-specific recombination system. Appl. Environ. Microbiol. 2010, 76, 6313–6317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pontecorvo, G.; Roper, J.A.; Hemmons, L.M.; Macdonald, K.D.; Bufton, A.W. The genetics of Aspergillus nidulans. Adv. Genet. 1953, 5, 141–238. [Google Scholar]
- Szewczyk, E.; Nayak, T.; Oakley, C.E.; Edgerton, H.; Xiong, Y.; Taheri-Talesh, N.; Osmani, S.A.; Oakley, B.R. Fusion PCR and gene targeting in Aspergillus nidulans. Nat. Protoc. 2006, 1, 3111–3120. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.G.; Young, L.; Chuang, R.Y.; Venter, J.C.; Hutchison, C.A., 3rd; Smith, H.O. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 2009, 6, 343–345. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbour Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Lopez-Berges, M.S.; Rispail, N.; Prados-Rosales, R.C.; Di Pietro, A. A Nitrogen Response Pathway Regulates Virulence Functions in Fusarium oxysporum via the Protein Kinase TOR and the bZIP Protein MeaB. Plant Cell 2010, 22, 2459–2475. [Google Scholar] [CrossRef] [Green Version]
- Hervas-Aguilar, A.; Penalva, M.A. Endocytic machinery protein SlaB is dispensable for polarity establishment but necessary for polarity maintenance in hyphal tip cells of Aspergillus nidulans. Eukaryot Cell 2010, 9, 1504–1518. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.L.; Osmani, A.H.; Ukil, L.; Son, S.; Markossian, S.; Shen, K.F.; Govindaraghavan, M.; Varadaraj, A.; Hashmi, S.B.; De Souza, C.P.; et al. Single-step affinity purification for fungal proteomics. Eukaryot Cell 2010, 9, 831–833. [Google Scholar] [CrossRef] [Green Version]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
López-Berges, M.S.; Scheven, M.T.; Hortschansky, P.; Misslinger, M.; Baldin, C.; Gsaller, F.; Werner, E.R.; Krüger, T.; Kniemeyer, O.; Weber, J.; et al. The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus. Int. J. Mol. Sci. 2021, 22, 7739. https://doi.org/10.3390/ijms22147739
López-Berges MS, Scheven MT, Hortschansky P, Misslinger M, Baldin C, Gsaller F, Werner ER, Krüger T, Kniemeyer O, Weber J, et al. The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus. International Journal of Molecular Sciences. 2021; 22(14):7739. https://doi.org/10.3390/ijms22147739
Chicago/Turabian StyleLópez-Berges, Manuel Sánchez, Mareike Thea Scheven, Peter Hortschansky, Matthias Misslinger, Clara Baldin, Fabio Gsaller, Ernst R. Werner, Thomas Krüger, Olaf Kniemeyer, Jakob Weber, and et al. 2021. "The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus" International Journal of Molecular Sciences 22, no. 14: 7739. https://doi.org/10.3390/ijms22147739
APA StyleLópez-Berges, M. S., Scheven, M. T., Hortschansky, P., Misslinger, M., Baldin, C., Gsaller, F., Werner, E. R., Krüger, T., Kniemeyer, O., Weber, J., Brakhage, A. A., & Haas, H. (2021). The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus. International Journal of Molecular Sciences, 22(14), 7739. https://doi.org/10.3390/ijms22147739





