Best Practices and Progress in Precision-Cut Liver Slice Cultures
Abstract
1. Introduction
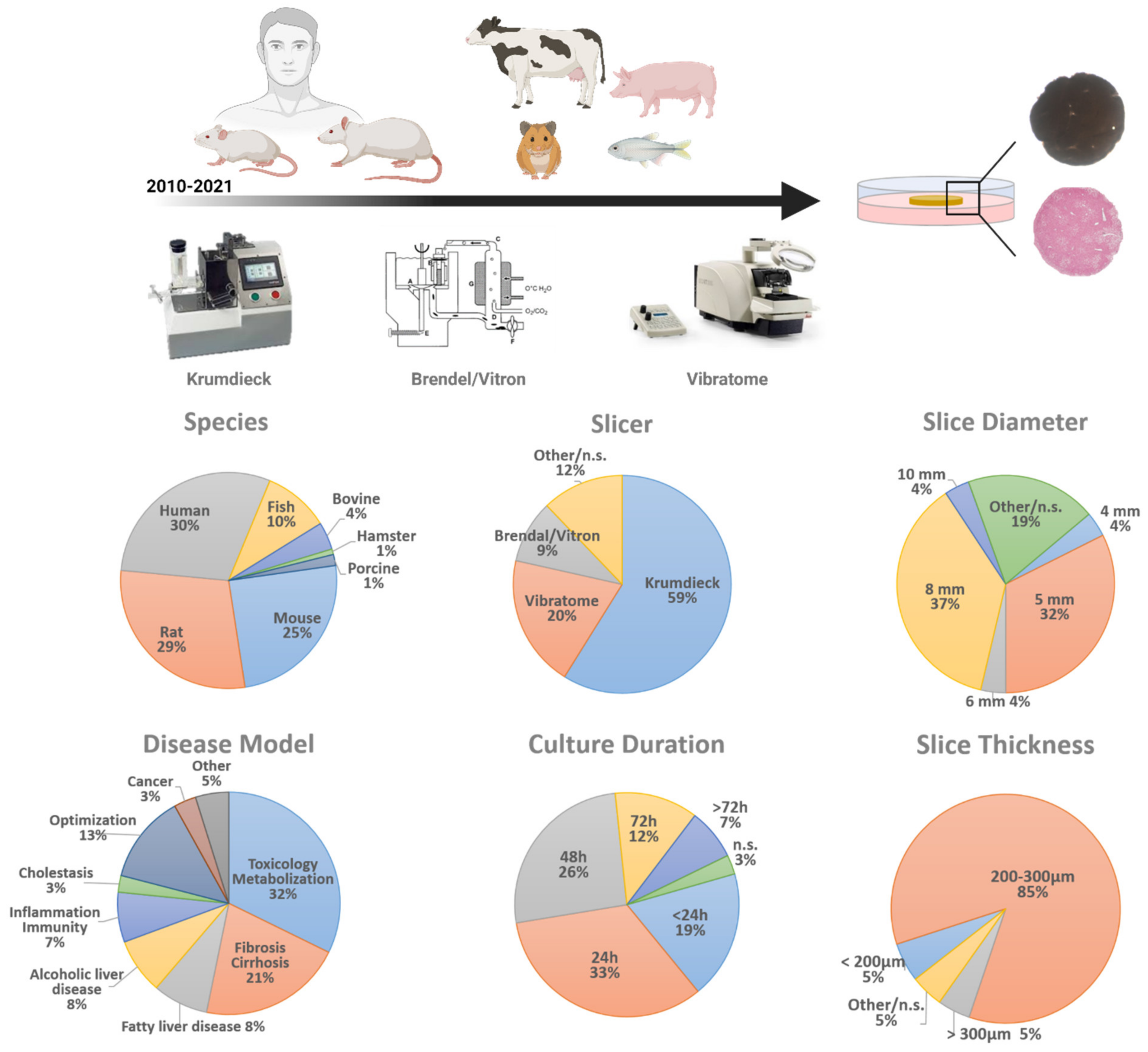
2. Precision-Cut Liver Slice Parameters
2.1. Tissue Slicer
2.2. Liver Slice Thickness
2.3. Liver Slice Diameter
2.4. Liver Slice Culture Conditions
2.5. Species
2.6. Disease Model
3. Precision-Cut Liver Slice Read-Outs
3.1. Culture Assessment
3.1.1. Viability and Cytotoxicity
| Read-Out | Advantages | Disadvantages/Difficulties | Analysis Time | Cost per Data Point |
|---|---|---|---|---|
| Viability and Cytotoxicity | ||||
| LDH leakage | Slice specific follow-up over time High throughput | Reader required | ±1 h | € |
| ATP assay | High throughput | End point analysis only Total tissue lysis required Reader required | ±1 h | € |
| Morphology | Evaluation of liver structure | Interpretation not always straightforward Embedding technically challenging | ±3 h | € |
| Slice analysis | ||||
| qRT-PCR m(i)RNA analysis | Evaluation of multiple m(i)RNA’s | Selective m(i)RNA measurements | ±3 h | €€ |
| Gene expression profiling | Generation of unbiased and complete mRNA profile | Need for bioinformatic skills Need for (nucleomics) infrastructure | Several days | €€€€ |
| Western Blot | Medium sensitivity High specificity | Low throughput High tissue input required | 1 day | €€€ |
| Hydroxyproline assay | Quantitative | High tissue input required Accurate measurement of input | 1 day | €€ |
| Immunohistochemical stainings | Can provide more detailed insight into zonation and cell type specific protein expression | Embedding technically challenging Relative low throughput Limited amount of stainings per slice | >1 day | €€ |
| Triglyceride quantification | Fatty/metabolic liver disease specific | Total homogenization of tissue required | ±2 h | €€ |
| GSH content | Functional | Affected by fat storage [46] | ±1 h | €€ |
| Medium analysis | ||||
| ALT, AST measurements | Straightforward Direct indicator of liver damage | Concentration of medium might be required/sufficient level of damage required to exceed threshold Low throughput | Minutes | € |
| ELISA | Quantitative Multiple proteins can be tested for 1 slice | Low throughput | Hours | €€€ |
| Protein Profiling | Generation of unbiased and complete protein profile | Need for bioinformatic skills Need for (proteomics) infrastructure | Several days | €€€€ |
3.1.2. Morphology
3.2. Slice Analysis
3.2.1. RNA Analysis
3.2.2. Protein Analysis
3.2.3. Other Tissue Analyses
3.3. Medium Analysis
3.4. Other Applications
| Liver Disease | Recommended Read-Outs |
|---|---|
| Toxicology | Hepatocyte viability by LDH leakage or ATP assay [86] Hepatocyte viability by functionality through Albumin ELISA [73] |
| Fibrosis | LDH leakage or ATP assay (hepatocyte viability) [71] mRNA levels of fibrosis markers by qRT-PCR [24] Protein levels of fibrosis markers (WB, staining or ELISA) [24] Sirius Red staining for cross-linked collagen [24] |
| Fatty liver disease | LDH leakage or ATP assay [58] Lipid Staining (e.g., ORO) [58] Triglyceride analysis/quantification [58] qRT-PCR for fibrosis mRNA markers [87] |
| Alcoholic liver disease | LDH leakage or ATP assay [42] Medium analysis for metabolites [42,88] qRT-PCR for fibrosis mRNA markers [42,85] Electron microscopy [65] * |
| Inflammation/Immunity | ELISA on medium for inflammatory markers [48,72] qRT-PCR for inflammatory mRNA markers [89] |
| Cholestasis | LDH leakage or ATP assay [90,91] qRT-PCR for cholestatic mRNA markers [91] Bile acid determination [91] |
| Cancer | LDH leakage or ATP assay [21] qRT-PCR for cancer-related mRNA markers [21] H&E staining (tumor morphology) [54] Staining for Ki67 (proliferation) [54] |
4. Future Perspectives
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Krumdieck, C.L.; dos Santos, J.E.; Ho, K.J. A new instrument for the rapid preparation of tissue slices. Anal. Biochem. 1980, 104, 118–123. [Google Scholar] [CrossRef]
- Smith, P.F.; Gandolfi, A.J.; Krumdieck, C.L.; Putnam, C.W.; Zukoski, C.F., 3rd; Davis, W.M.; Brendel, K. Dynamic organ culture of precision liver slices for in vitro toxicology. Life Sci. 1985, 36, 1367–1375. [Google Scholar] [CrossRef]
- Beigel, J.; Fella, K.; Kramer, P.J.; Kroeger, M.; Hewitt, P. Genomics and proteomics analysis of cultured primary rat hepatocytes. Toxicol. In Vitro 2008, 22, 171–181. [Google Scholar] [CrossRef]
- Azri, S.; Mata, H.P.; Reid, L.L.; Gandolfi, A.J.; Brendel, K. Further examination of the selective toxicity of CCl4 in rat liver slices. Toxicol. Appl. Pharmacol. 1992, 112, 81–86. [Google Scholar] [CrossRef]
- Obatomi, D.K.; Brant, S.; Anthonypillai, V.; Bach, P.H. Toxicity of atractyloside in precision-cut rat and porcine renal and hepatic tissue slices. Toxicol. Appl. Pharmacol. 1998, 148, 35–45. [Google Scholar] [CrossRef] [PubMed]
- De Kanter, R.; De Jager, M.H.; Draaisma, A.L.; Jurva, J.U.; Olinga, P.; Meijer, D.K.; Groothuis, G.M. Drug-metabolizing activity of human and rat liver, lung, kidney and intestine slices. Xenobiotica 2002, 32, 349–362. [Google Scholar] [CrossRef] [PubMed]
- Van de Bovenkamp, M.; Groothuis, G.M.; Meijer, D.K.; Olinga, P. Precision-cut fibrotic rat liver slices as a new model to test the effects of anti-fibrotic drugs in vitro. J. Hepatol. 2006, 45, 696–703. [Google Scholar] [CrossRef]
- De Graaf, I.A.; Olinga, P.; de Jager, M.H.; Merema, M.T.; de Kanter, R.; van de Kerkhof, E.G.; Groothuis, G.M. Preparation and incubation of precision-cut liver and intestinal slices for application in drug metabolism and toxicity studies. Nat. Protoc. 2010, 5, 1540–1551. [Google Scholar] [CrossRef]
- Vickers, A.E.; Saulnier, M.; Cruz, E.; Merema, M.T.; Rose, K.; Bentley, P.; Olinga, P. Organ slice viability extended for pathway characterization: An in vitro model to investigate fibrosis. Toxicol. Sci. 2004, 82, 534–544. [Google Scholar] [CrossRef]
- Othman, A.; Ehnert, S.; Dropmann, A.; Ruoß, M.; Nüssler, A.K.; Hammad, S. Precision-cut liver slices as an alternative method for long-term hepatotoxicity studies. Arch. Toxicol. 2020, 94, 2889–2891. [Google Scholar] [CrossRef]
- Palma, E.; Doornebal, E.J.; Chokshi, S. Precision-cut liver slices: A versatile tool to advance liver research. Hepatol. Int. 2019, 13, 51–57. [Google Scholar] [CrossRef]
- Godoy, P.; Hewitt, N.J.; Albrecht, U.; Andersen, M.E.; Ansari, N.; Bhattacharya, S.; Bode, J.G.; Bolleyn, J.; Borner, C.; Böttger, J.; et al. Recent advances in 2D and 3D in vitro systems using primary hepatocytes, alternative hepatocyte sources and non-parenchymal liver cells and their use in investigating mechanisms of hepatotoxicity, cell signaling and ADME. Arch. Toxicol. 2013, 87, 1315–1530. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, M.; Lampe, J.; Lange, S.; Smirnow, I.; Königsrainer, A.; Hann-von-Weyhern, C.; Fend, F.; Gregor, M.; Bitzer, M.; Lauer, U.M. Improved reproducibility in preparing precision-cut liver tissue slices. Cytotechnology 2009, 61, 145–152. [Google Scholar] [CrossRef]
- Price, R.J.; Ball, S.E.; Renwick, A.B.; Barton, P.T.; Beamand, J.A.; Lake, B.G. Use of precision-cut rat liver slices for studies of xenobiotic metabolism and toxicity: Comparison of the Krumdieck and Brendel tissue slicers. Xenobiotica 1998, 28, 361–371. [Google Scholar] [CrossRef]
- Fisher, R.L.; Judith, U.; Paul, N.; Klaus, B. Histological and biochemical evaluation of precision-cut liver slices. Toxicol. Methods 2001, 11, 59–79. [Google Scholar] [CrossRef]
- De Graaf, I.A.; de Kanter, R.; de Jager, M.H.; Camacho, R.; Langenkamp, E.; van de Kerkhof, E.G.; Groothuis, G.M. Empirical validation of a rat in vitro organ slice model as a tool for in vivo clearance prediction. Drug Metab. Dispos. 2006, 34, 591–599. [Google Scholar] [CrossRef]
- Fisher, R.L.; Hasal, S.J.; Sanuik, J.T.; Gandolfi, A.J.; Brendel, K. Determination of Optimal Incubation Media and Suitable Slice Diameters in Precision-Cut Liver Slices: Optimization of Tissue Slice Culture, Part 2. Toxicol. Methods 1995, 5, 115–130. [Google Scholar] [CrossRef]
- Schumacher, K.; Khong, Y.M.; Chang, S.; Ni, J.; Sun, W.; Yu, H. Perfusion culture improves the maintenance of cultured liver tissue slices. Tissue Eng. 2007, 13, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Szalowska, E.; Stoopen, G.; Rijk, J.C.; Wang, S.; Hendriksen, P.J.; Groot, M.J.; Ossenkoppele, J.; Peijnenburg, A.A. Effect of oxygen concentration and selected protocol factors on viability and gene expression of mouse liver slices. Toxicol. In Vitro 2013, 27, 1513–1524. [Google Scholar] [CrossRef]
- Kidambi, S.; Yarmush, R.S.; Novik, E.; Chao, P.; Yarmush, M.L.; Nahmias, Y. Oxygen-mediated enhancement of primary hepatocyte metabolism, functional polarization, gene expression, and drug clearance. Proc. Natl. Acad. Sci. USA 2009, 106, 15714–15719. [Google Scholar] [CrossRef] [PubMed]
- Koch, A.; Saran, S.; Tran, D.D.; Klebba-Färber, S.; Thiesler, H.; Sewald, K.; Schindler, S.; Braun, A.; Klopfleisch, R.; Tamura, T. Murine precision-cut liver slices (PCLS): A new tool for studying tumor microenvironments and cell signaling ex vivo. Cell Commun. Signal. 2014, 12, 73. [Google Scholar] [CrossRef]
- Starokozhko, V.; Abza, G.B.; Maessen, H.C.; Merema, M.T.; Kuper, F.; Groothuis, G.M. Viability, function and morphological integrity of precision-cut liver slices during prolonged incubation: Effects of culture medium. Toxicol. In Vitro 2015, 30, 288–299. [Google Scholar] [CrossRef]
- Starokozhko, V.; Vatakuti, S.; Schievink, B.; Merema, M.T.; Asplund, A.; Synnergren, J.; Aspegren, A.; Groothuis, G.M.M. Maintenance of drug metabolism and transport functions in human precision-cut liver slices during prolonged incubation for 5 days. Arch. Toxicol. 2017, 91, 2079–2092. [Google Scholar] [CrossRef] [PubMed]
- Paish, H.L.; Reed, L.H.; Brown, H.; Bryan, M.C.; Govaere, O.; Leslie, J.; Barksby, B.S.; Garcia Macia, M.; Watson, A.; Xu, X.; et al. A Bioreactor Technology for Modeling Fibrosis in Human and Rodent Precision-Cut Liver Slices. Hepatology 2019, 70, 1377–1391. [Google Scholar] [CrossRef]
- Wu, X.; Roberto, J.B.; Knupp, A.; Kenerson, H.L.; Truong, C.D.; Yuen, S.Y.; Brempelis, K.J.; Tuefferd, M.; Chen, A.; Horton, H.; et al. Precision-cut human liver slice cultures as an immunological platform. J. Immunol. Methods 2018, 455, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Kartasheva-Ebertz, D.; Gaston, J.; Lair-Mehiri, L.; Massault, P.P.; Scatton, O.; Vaillant, J.C.; Morozov, V.A.; Pol, S.; Lagaye, S. Adult human liver slice cultures: Modelling of liver fibrosis and evaluation of new anti-fibrotic drugs. World J. Hepatol. 2021, 13, 187–217. [Google Scholar] [CrossRef]
- Kang, C.; Qiao, Y.; Li, G.; Baechle, K.; Camelliti, P.; Rentschler, S.; Efimov, I.R. Human Organotypic Cultured Cardiac Slices: New Platform For High Throughput Preclinical Human Trials. Sci. Rep. 2016, 6, 28798. [Google Scholar] [CrossRef]
- Watson, S.A.; Scigliano, M.; Bardi, I.; Ascione, R.; Terracciano, C.M.; Perbellini, F. Preparation of viable adult ventricular myocardial slices from large and small mammals. Nat. Protoc. 2017, 12, 2623–2639. [Google Scholar] [CrossRef] [PubMed]
- Stribos, E.G.D.; Luangmonkong, T.; Leliveld, A.M.; De Jong, I.J.; Van Son, W.J.; Hillebrands, J.-L.; Seelen, M.A.; Van Goor, H.; Olinga, P.; Mutsaers, H.A.M. Precision-cut human kidney slices as a model to elucidate the process of renal fibrosis. Transl. Res. 2016, 170, 8–16. [Google Scholar] [CrossRef]
- Poosti, F.; Pham, B.T.; Oosterhuis, D.; Poelstra, K.; Van Goor, H.; Olinga, P.; Hillebrands, J.L. Precision-cut kidney slices (PCKS) to study development of renal fibrosis and efficacy of drug targeting ex vivo. DMM Dis. Models Mech. 2015, 8, 1227–1236. [Google Scholar] [CrossRef]
- Uhl, F.E.; Vierkotten, S.; Wagner, D.E.; Burgstaller, G.; Costa, R.; Koch, I.; Lindner, M.; Meiners, S.; Eickelberg, O.; Königshoff, M. Preclinical validation and imaging of Wnt-induced repair in human 3D lung tissue cultures. Eur. Respir. J. 2015, 46, 1150–1166. [Google Scholar] [CrossRef] [PubMed]
- Klouda, T.; Kim, H.; Kim, J.; Visner, G.; Yuan, K. Precision Cut Lung Slices as an Efficient Tool for Ex vivo Pulmonary Vessel Structure and Contractility Studies. J. Vis. Exp. 2021. [Google Scholar] [CrossRef] [PubMed]
- Verwer, R.W.H.; Hermens, W.T.J.M.C.; Dijkhuizen, P.A.; Brake, O.T.; Baker, R.E.; Salehi, A.; Sluiter, A.A.; Kok, M.J.M.; Müller, A.L.J.; Verhaagen, J.; et al. Cells in human postmortem brain tissue slices remain alive for several weeks in culture. FASEB J. 2002, 16, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.L.; Gandolfi, A.J.; Brendel, K. Human liver quality is a dominant factor in the outcome of in vitro studies. Cell Biol. Toxicol. 2001, 17, 179–189. [Google Scholar] [CrossRef] [PubMed]
- Jetten, M.J.; Claessen, S.M.; Dejong, C.H.; Lahoz, A.; Castell, J.V.; van Delft, J.H.; Kleinjans, J.C. Interindividual variation in response to xenobiotic exposure established in precision-cut human liver slices. Toxicology 2014, 323, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Meyer, C.; Xu, C.; Weng, H.; Hellerbrand, C.; Dijke, P.; Dooley, S. Animal Models of Chronic Liver Diseases. Am. J. Physiol. Gastrointest. Liver Physiol. 2012, 304, G449–G468. [Google Scholar] [CrossRef]
- Vatakuti, S.; Schoonen, W.G.; Elferink, M.L.; Groothuis, G.M.; Olinga, P. Acute toxicity of CCl4 but not of paracetamol induces a transcriptomic signature of fibrosis in precision-cut liver slices. Toxicol. In Vitro 2015, 29, 1012–1020. [Google Scholar] [CrossRef]
- van de Bovenkamp, M.; Groothuis, G.M.; Draaisma, A.L.; Merema, M.T.; Bezuijen, J.I.; van Gils, M.J.; Meijer, D.K.; Friedman, S.L.; Olinga, P. Precision-cut liver slices as a new model to study toxicity-induced hepatic stellate cell activation in a physiologic milieu. Toxicol. Sci. 2005, 85, 632–638. [Google Scholar] [CrossRef]
- Van de Bovenkamp, M.; Groothuis, G.M.; Meijer, D.K.; Olinga, P. Liver slices as a model to study fibrogenesis and test the effects of anti-fibrotic drugs on fibrogenic cells in human liver. Toxicol. In Vitro 2008, 22, 771–778. [Google Scholar] [CrossRef]
- Westra, I.M.; Oosterhuis, D.; Groothuis, G.M.; Olinga, P. Precision-cut liver slices as a model for the early onset of liver fibrosis to test antifibrotic drugs. Toxicol. Appl. Pharmacol. 2014, 274, 328–338. [Google Scholar] [CrossRef]
- Gore, E.; Bigaeva, E.; Oldenburger, A.; Jansen, Y.J.M.; Schuppan, D.; Boersema, M.; Rippmann, J.F.; Broermann, A.; Olinga, P. Investigating fibrosis and inflammation in an ex vivo NASH murine model. Am. J. Physiol. Gastrointest. Liver Physiol. 2020, 318, G336–G351. [Google Scholar] [CrossRef]
- Duryee, M.J.; Willis, M.S.; Schaffert, C.S.; Reidelberger, R.D.; Dusad, A.; Anderson, D.R.; Klassen, L.W.; Thiele, G.M. Precision-cut liver slices from diet-induced obese rats exposed to ethanol are susceptible to oxidative stress and increased fatty acid synthesis. Am. J. Physiol. Gastrointest. Liver Physiol. 2014, 306, G208–G217. [Google Scholar] [CrossRef]
- Crouch, S.P.; Kozlowski, R.; Slater, K.J.; Fletcher, J. The use of ATP bioluminescence as a measure of cell proliferation and cytotoxicity. J. Immunol. Methods 1993, 160, 81–88. [Google Scholar] [CrossRef]
- Chan, F.K.; Moriwaki, K.; De Rosa, M.J. Detection of necrosis by release of lactate dehydrogenase activity. Methods Mol. Biol. 2013, 979, 65–70. [Google Scholar] [CrossRef] [PubMed]
- Riss, T.L.; Moravec, R.A. Use of multiple assay endpoints to investigate the effects of incubation time, dose of toxin, and plating density in cell-based cytotoxicity assays. Assay Drug Dev. Technol. 2004, 2, 51–62. [Google Scholar] [CrossRef]
- Huang, J.; Jia, Y.; Li, Q.; Son, K.; Hamilton, C.; Burris, W.R.; Bridges, P.J.; Stromberg, A.J.; Matthews, J.C. Glutathione content and expression of proteins involved with glutathione metabolism differs in longissimus dorsi, subcutaneous adipose, and liver tissues of finished vs. growing beef steers. J. Anim. Sci. 2018, 96, 5152–5165. [Google Scholar] [CrossRef] [PubMed]
- Bigaeva, E.; Gore, E.; Mutsaers, H.A.M.; Oosterhuis, D.; Kim, Y.O.; Schuppan, D.; Bank, R.A.; Boersema, M.; Olinga, P. Exploring organ-specific features of fibrogenesis using murine precision-cut tissue slices. Biochim. Biophys. Acta Mol. Basis. Dis. 2020, 1866, 165582. [Google Scholar] [CrossRef]
- Beaumont, M.; Neyrinck, A.M.; Olivares, M.; Rodriguez, J.; de Rocca Serra, A.; Roumain, M.; Bindels, L.B.; Cani, P.D.; Evenepoel, P.; Muccioli, G.G.; et al. The gut microbiota metabolite indole alleviates liver inflammation in mice. FASEB J. 2018, 32, fj201800544. [Google Scholar] [CrossRef] [PubMed]
- Ijssennagger, N.; Janssen, A.W.F.; Milona, A.; Ramos Pittol, J.M.; Hollman, D.A.A.; Mokry, M.; Betzel, B.; Berends, F.J.; Janssen, I.M.; van Mil, S.W.C.; et al. Gene expression profiling in human precision cut liver slices in response to the FXR agonist obeticholic acid. J. Hepatol. 2016, 64, 1158–1166. [Google Scholar] [CrossRef] [PubMed]
- Harvey, T.N.; Sandve, S.R.; Jin, Y.; Vik, J.O.; Torgersen, J.S. Liver slice culture as a model for lipid metabolism in fish. PeerJ 2019, 7, e7732. [Google Scholar] [CrossRef] [PubMed]
- Aluru, N.; Hallanger, I.; McMonagle, H.; Harju, M. Hepatic gene expression profiling of OPFR exposed Atlantic cod (Gadus morhua) liver revealed altered cholesterol biosynthesis and lipid metabolism. Environ. Toxicol. Chem. 2021, 40, 1639–1648. [Google Scholar] [CrossRef] [PubMed]
- Yadetie, F.; Zhang, X.; Hanna, E.M.; Aranguren-Abadía, L.; Eide, M.; Blaser, N.; Brun, M.; Jonassen, I.; Goksøyr, A.; Karlsen, O.A. RNA-Seq analysis of transcriptome responses in Atlantic cod (Gadus morhua) precision-cut liver slices exposed to benzo[a]pyrene and 17α-ethynylestradiol. Aquat. Toxicol. 2018, 201, 174–186. [Google Scholar] [CrossRef] [PubMed]
- Bigaeva, E.; Gore, E.; Simon, E.; Zwick, M.; Oldenburger, A.; de Jong, K.P.; Hofker, H.S.; Schlepütz, M.; Nicklin, P.; Boersema, M.; et al. Transcriptomic characterization of culture-associated changes in murine and human precision-cut tissue slices. Arch. Toxicol. 2019, 93, 3549–3583. [Google Scholar] [CrossRef] [PubMed]
- Kenerson, H.L.; Sullivan, K.M.; Seo, Y.D.; Stadeli, K.M.; Ussakli, C.; Yan, X.; Lausted, C.; Pillarisetty, V.G.; Park, J.O.; Riehle, K.J.; et al. Tumor slice culture as a biologic surrogate of human cancer. Ann. Transl. Med. 2020, 8, 114. [Google Scholar] [CrossRef]
- Westra, I.M.; Oosterhuis, D.; Groothuis, G.M.; Olinga, P. The effect of antifibrotic drugs in rat precision-cut fibrotic liver slices. PLoS ONE 2014, 9, e95462. [Google Scholar] [CrossRef]
- Bartucci, R.; van der Meer, A.Z.; Boersma, Y.L.; Olinga, P.; Salvati, A. Nanoparticle-induced inflammation and fibrosis in ex vivo murine precision-cut liver slices and effects of nanoparticle exposure conditions. Arch. Toxicol. 2021. [Google Scholar] [CrossRef]
- Daga, M.; de Graaf, I.A.M.; Argenziano, M.; Barranco, A.S.M.; Loeck, M.; Al-Adwi, Y.; Cucci, M.A.; Caldera, F.; Trotta, F.; Barrera, G.; et al. Glutathione-responsive cyclodextrin-nanosponges as drug delivery systems for doxorubicin: Evaluation of toxicity and transport mechanisms in the liver. Toxicol. In Vitro 2020, 65, 104800. [Google Scholar] [CrossRef]
- Prins, G.H.; Luangmonkong, T.; Oosterhuis, D.; Mutsaers, H.A.M.; Dekker, F.J.; Olinga, P. A Pathophysiological Model of Non-Alcoholic Fatty Liver Disease Using Precision-Cut Liver Slices. Nutrients 2019, 11, 507. [Google Scholar] [CrossRef]
- Aoudjehane, L.; Gautheron, J.; Le Goff, W.; Goumard, C.; Gilaizeau, J.; Nget, C.S.; Savier, E.; Atif, M.; Lesnik, P.; Morichon, R.; et al. Novel defatting strategies reduce lipid accumulation in primary human culture models of liver steatosis. Dis. Model Mech. 2020, 13. [Google Scholar] [CrossRef]
- Shepherd, E.L.; Karim, S.; Newsome, P.N.; Lalor, P.F. Inhibition of vascular adhesion protein-1 modifies hepatic steatosis in vitro and in vivo. World J. Hepatol. 2020, 12, 931–948. [Google Scholar] [CrossRef]
- Bosisio, F.M.; Antoranz, A.; van Herck, Y.; Bolognesi, M.M.; Marcelis, L.; Chinello, C.; Wouters, J.; Magni, F.; Alexopoulos, L.; Stas, M.; et al. Functional heterogeneity of lymphocytic patterns in primary melanoma dissected through single-cell multiplexing. Elife 2020, 9, e53008. [Google Scholar] [CrossRef] [PubMed]
- Bolognesi, M.M.; Manzoni, M.; Scalia, C.R.; Zannella, S.; Bosisio, F.M.; Faretta, M.; Cattoretti, G. Multiplex Staining by Sequential Immunostaining and Antibody Removal on Routine Tissue Sections. J. Histochem. Cytochem. 2017, 65, 431–444. [Google Scholar] [CrossRef] [PubMed]
- Zielinski, J.M.; Luke, J.J.; Guglietta, S.; Krieg, C. High Throughput Multi-Omics Approaches for Clinical Trial Evaluation and Drug Discovery. Front. Immunol. 2021, 12, 590742. [Google Scholar] [CrossRef]
- Dragoni, S.; Franco, G.; Regoli, M.; Bracciali, M.; Morandi, V.; Sgaragli, G.; Bertelli, E.; Valoti, M. Gold nanoparticles uptake and cytotoxicity assessed on rat liver precision-cut slices. Toxicol. Sci. 2012, 128, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Palma, E.; Riva, A.; Moreno, C.; Odena, G.; Mudan, S.; Manyakin, N.; Miquel, R.; Degré, D.; Trepo, E.; Sancho-Bru, P.; et al. Perturbations in Mitochondrial Dynamics Are Closely Involved in the Progression of Alcoholic Liver Disease. Alcohol. Clin. Exp. Res. 2020, 44, 856–865. [Google Scholar] [CrossRef]
- Sadasivan, S.K.; Siddaraju, N.; Khan, K.M.; Vasamsetti, B.; Kumar, N.R.; Haridas, V.; Reddy, M.B.; Baggavalli, S.; Oommen, A.M.; Pralhada Rao, R. Developing an in vitro screening assay platform for evaluation of antifibrotic drugs using precision-cut liver slices. Fibrogenesis Tissue Repair. 2015, 8, 1. [Google Scholar] [CrossRef]
- Ookhtens, M.; Kaplowitz, N. Role of the liver in interorgan homeostasis of glutathione and cyst(e)ine. Semin. Liver Dis. 1998, 18, 313–329. [Google Scholar] [CrossRef]
- Hissin, P.J.; Hilf, R. A fluorometric method for determination of oxidized and reduced glutathione in tissues. Anal. Biochem. 1976, 74, 214–226. [Google Scholar] [CrossRef]
- Lőrincz, T.; Szarka, A. The determination of hepatic glutathione at tissue and subcellular level. J. Pharmacol. Toxicol. Methods 2017, 88, 32–39. [Google Scholar] [CrossRef]
- Pratt, D.S.; Kaplan, M.M. Evaluation of abnormal liver-enzyme results in asymptomatic patients. N. Engl. J. Med. 2000, 342, 1266–1271. [Google Scholar] [CrossRef]
- Barcena-Varela, M.; Paish, H.; Alvarez, L.; Uriarte, I.; Latasa, M.U.; Santamaria, E.; Recalde, M.; Garate, M.; Claveria, A.; Colyn, L.; et al. Epigenetic mechanisms and metabolic reprogramming in fibrogenesis: Dual targeting of G9a and DNMT1 for the inhibition of liver fibrosis. Gut 2021, 70, 388–400. [Google Scholar] [CrossRef]
- Morán-Salvador, E.; Titos, E.; Rius, B.; González-Périz, A.; García-Alonso, V.; López-Vicario, C.; Miquel, R.; Barak, Y.; Arroyo, V.; Clària, J. Cell-specific PPARγ deficiency establishes anti-inflammatory and anti-fibrogenic properties for this nuclear receptor in non-parenchymal liver cells. J. Hepatol. 2013, 59, 1045–1053. [Google Scholar] [CrossRef]
- Granitzny, A.; Knebel, J.; Schaudien, D.; Braun, A.; Steinberg, P.; Dasenbrock, C.; Hansen, T. Maintenance of high quality rat precision cut liver slices during culture to study hepatotoxic responses: Acetaminophen as a model compound. Toxicol. In Vitro 2017, 42, 200–213. [Google Scholar] [CrossRef] [PubMed]
- Karim, S.; Liaskou, E.; Hadley, S.; Youster, J.; Faint, J.; Adams, D.H.; Lalor, P.F. An in vitro model of human acute ethanol exposure that incorporates CXCR3- and CXCR4-dependent recruitment of immune cells. Toxicol. Sci. 2013, 132, 131–141. [Google Scholar] [CrossRef][Green Version]
- Duryee, M.J.; Wiese, B.M.; Bowman, J.R.; Vanlandingham, J.D.; Klassen, L.W.; Thiele, G.E.; Hunter, C.D.; Anderson, D.R.; Mikuls, T.R.; Thiele, G.M. Liver tissue metabolically transformed by alcohol induces immune recognition of liver self-proteins but not in vivo inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 2018, 314, G418–G430. [Google Scholar] [CrossRef]
- Suriguga, S.; Luangmonkong, T.; Mutsaers, H.A.M.; Groothuis, G.M.M.; Olinga, P. Host microbiota dictates the proinflammatory impact of LPS in the murine liver. Toxicol. In Vitro 2020, 67, 104920. [Google Scholar] [CrossRef] [PubMed]
- Luangmonkong, T.; Suriguga, S.; Bigaeva, E.; Boersema, M.; Oosterhuis, D.; de Jong, K.P.; Schuppan, D.; Mutsaers, H.A.M.; Olinga, P. Evaluating the antifibrotic potency of galunisertib in a human ex vivo model of liver fibrosis. Br. J. Pharmacol. 2017, 174, 3107–3117. [Google Scholar] [CrossRef] [PubMed]
- Hadi, M.; Dragovic, S.; van Swelm, R.; Herpers, B.; van de Water, B.; Russel, F.G.; Commandeur, J.N.; Groothuis, G.M. AMAP, the alleged non-toxic isomer of acetaminophen, is toxic in rat and human liver. Arch. Toxicol. 2013, 87, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Van Swelm, R.P.; Hadi, M.; Laarakkers, C.M.; Masereeuw, R.; Groothuis, G.M.; Russel, F.G. Proteomic profiling in incubation medium of mouse, rat and human precision-cut liver slices for biomarker detection regarding acute drug-induced liver injury. J. Appl. Toxicol. 2014, 34, 993–1001. [Google Scholar] [CrossRef] [PubMed]
- Lambrecht, J.; Jan Poortmans, P.; Verhulst, S.; Reynaert, H.; Mannaerts, I.; van Grunsven, L.A. Circulating ECV-Associated miRNAs as Potential Clinical Biomarkers in Early Stage HBV and HCV Induced Liver Fibrosis. Front. Pharmacol. 2017, 8, 56. [Google Scholar] [CrossRef]
- Lambrecht, J.; Mannaerts, I.; van Grunsven, L.A. The role of miRNAs in stress-responsive hepatic stellate cells during liver fibrosis. Front. Physiol. 2015, 6, 209. [Google Scholar] [CrossRef]
- Lambrecht, J.; Verhulst, S.; Reynaert, H.; van Grunsven, L.A. The miRFIB-Score: A Serological miRNA-Based Scoring Algorithm for the Diagnosis of Significant Liver Fibrosis. Cells 2019, 8, 1003. [Google Scholar] [CrossRef]
- Zárybnický, T.; Matoušková, P.; Ambrož, M.; Šubrt, Z.; Skálová, L.; Boušová, I. The Selection and Validation of Reference Genes for mRNA and microRNA Expression Studies in Human Liver Slices Using RT-qPCR. Genes 2019, 10, 763. [Google Scholar] [CrossRef]
- Okun, J.G.; Conway, S.; Schmidt, K.V.; Schumacher, J.; Wang, X.; de Guia, R.; Zota, A.; Klement, J.; Seibert, O.; Peters, A.; et al. Molecular regulation of urea cycle function by the liver glucocorticoid receptor. Mol. Metab. 2015, 4, 732–740. [Google Scholar] [CrossRef] [PubMed]
- Stärkel, P.; Schnabl, B.; Leclercq, S.; Komuta, M.; Bataller, R.; Argemi, J.; Palma, E.; Chokshi, S.; Hellerbrand, C.; Maccioni, L.; et al. Deficient IL-6/Stat3 Signaling, High TLR7, and Type I Interferons in Early Human Alcoholic Liver Disease: A Triad for Liver Damage and Fibrosis. Hepatol. Commun. 2019, 3, 867–882. [Google Scholar] [CrossRef]
- Szalowska, E.; Stoopen, G.; Groot, M.J.; Hendriksen, P.J.; Peijnenburg, A.A. Treatment of mouse liver slices with cholestatic hepatotoxicants results in down-regulation of Fxr and its target genes. BMC Med. Genom. 2013, 6, 39. [Google Scholar] [CrossRef] [PubMed]
- De Chiara, F.; Thomsen, K.L.; Habtesion, A.; Jones, H.; Davies, N.; Gracia-Sancho, J.; Manicardi, N.; Hall, A.; Andreola, F.; Paish, H.L.; et al. Ammonia Scavenging Prevents Progression of Fibrosis in Experimental Nonalcoholic Fatty Liver Disease. Hepatology 2020, 71, 874–892. [Google Scholar] [CrossRef]
- Villalobos-García, D.; Hernández-Muñoz, R. Catalase increases ethanol oxidation through the purine catabolism in rat liver. Biochem. Pharmacol. 2017, 137, 107–112. [Google Scholar] [CrossRef] [PubMed]
- Iswandana, R.; Pham, B.T.; van Haaften, W.T.; Luangmonkong, T.; Oosterhuis, D.; Mutsaers, H.A.; Olinga, P. Organ- and species-specific biological activity of rosmarinic acid. Toxicol. In Vitro 2016, 32, 261–268. [Google Scholar] [CrossRef]
- Vatakuti, S.; Pennings, J.L.; Gore, E.; Olinga, P.; Groothuis, G.M. Classification of Cholestatic and Necrotic Hepatotoxicants Using Transcriptomics on Human Precision-Cut Liver Slices. Chem. Res. Toxicol. 2016, 29, 342–351. [Google Scholar] [CrossRef]
- Starokozhko, V.; Greupink, R.; van de Broek, P.; Soliman, N.; Ghimire, S.; de Graaf, I.A.M.; Groothuis, G.M.M. Rat precision-cut liver slices predict drug-induced cholestatic injury. Arch. Toxicol. 2017, 91, 3403–3413. [Google Scholar] [CrossRef]
- Van Midwoud, P.M.; Groothuis, G.M.; Merema, M.T.; Verpoorte, E. Microfluidic biochip for the perifusion of precision-cut rat liver slices for metabolism and toxicology studies. Biotechnol. Bioeng. 2010, 105, 184–194. [Google Scholar] [CrossRef]
- Van Midwoud, P.M.; Merema, M.T.; Verpoorte, E.; Groothuis, G.M. A microfluidic approach for in vitro assessment of interorgan interactions in drug metabolism using intestinal and liver slices. Lab. Chip 2010, 10, 2778–2786. [Google Scholar] [CrossRef]
- Van Midwoud, P.M.; Merema, M.T.; Verweij, N.; Groothuis, G.M.; Verpoorte, E. Hydrogel embedding of precision-cut liver slices in a microfluidic device improves drug metabolic activity. Biotechnol. Bioeng. 2011, 108, 1404–1412. [Google Scholar] [CrossRef]
- Wang, K.; Lin, B.; Brems, J.J.; Gamelli, R.L. Hepatic apoptosis can modulate liver fibrosis through TIMP1 pathway. Apoptosis 2013, 18, 566–577. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, Y.; Tian, Y.; Chen, S.; Liu, H. Knockdown of TRIM15 inhibits the activation of hepatic stellate cells. J. Mol. Histol. 2021. [Google Scholar] [CrossRef]
- Ruigrok, M.J.R.; El Amasi, K.E.M.; Leeming, D.J.; Sand, J.M.B.; Frijlink, H.W.; Hinrichs, W.L.J.; Olinga, P. Silencing Heat Shock Protein 47 (HSP47) in Fibrogenic Precision-Cut Lung Slices: A Surprising Lack of Effects on Fibrogenesis? Front. Med. 2021, 8, 607962. [Google Scholar] [CrossRef] [PubMed]
- Ruigrok, M.J.R.; Maggan, N.; Willaert, D.; Frijlink, H.W.; Melgert, B.N.; Olinga, P.; Hinrichs, W.L.J. siRNA-Mediated RNA Interference in Precision-Cut Tissue Slices Prepared from Mouse Lung and Kidney. AAPS J. 2017, 19, 1855–1863. [Google Scholar] [CrossRef]
- Tateno, C.; Yoshizane, Y.; Saito, N.; Kataoka, M.; Utoh, R.; Yamasaki, C.; Tachibana, A.; Soeno, Y.; Asahina, K.; Hino, H.; et al. Near completely humanized liver in mice shows human-type metabolic responses to drugs. Am. J. Pathol. 2004, 165, 901–912. [Google Scholar] [CrossRef]
- Nagarajan, P.; Mahesh Kumar, M.J.; Venkatesan, R.; Majundar, S.S.; Juyal, R.C. Genetically modified mouse models for the study of nonalcoholic fatty liver disease. World J. Gastroenterol. 2012, 18, 1141–1153. [Google Scholar] [CrossRef]
- Pascale, R.M.; Simile, M.M.; Peitta, G.; Seddaiu, M.A.; Feo, F.; Calvisi, D.F. Experimental Models to Define the Genetic Predisposition to Liver Cancer. Cancers 2019, 11, 1450. [Google Scholar] [CrossRef]
- Li, A.P.; Bode, C.; Sakai, Y. A novel in vitro system, the integrated discrete multiple organ cell culture (IdMOC) system, for the evaluation of human drug toxicity: Comparative cytotoxicity of tamoxifen towards normal human cells from five major organs and MCF-7 adenocarcinoma breast cancer cells. Chem. Biol. Interact. 2004, 150, 129–136. [Google Scholar] [CrossRef] [PubMed]
- Jeon, J.W.; Lee, S.H.; Kim, D.; Sung, J.H. In vitro hepatic steatosis model based on gut-liver-on-a-chip. Biotechnol. Prog. 2021, e3121. [Google Scholar] [CrossRef]
- Dou, M.; Clair, G.; Tsai, C.F.; Xu, K.; Chrisler, W.B.; Sontag, R.L.; Zhao, R.; Moore, R.J.; Liu, T.; Pasa-Tolic, L.; et al. High-Throughput Single Cell Proteomics Enabled by Multiplex Isobaric Labeling in a Nanodroplet Sample Preparation Platform. Anal. Chem. 2019, 91, 13119–13127. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Scheibinger, M.; Ellwanger, D.C.; Krey, J.F.; Choi, D.; Kelly, R.T.; Heller, S.; Barr-Gillespie, P.G. Single-cell proteomics reveals changes in expression during hair-cell development. Elife 2019, 8, e50777. [Google Scholar] [CrossRef]
- Piltti, K.M.; Cummings, B.J.; Carta, K.; Manughian-Peter, A.; Worne, C.L.; Singh, K.; Ong, D.; Maksymyuk, Y.; Khine, M.; Anderson, A.J. Live-cell time-lapse imaging and single-cell tracking of in vitro cultured neural stem cells—Tools for analyzing dynamics of cell cycle, migration, and lineage selection. Methods 2018, 133, 81–90. [Google Scholar] [CrossRef]
- Krenkel, O.; Hundertmark, J.; Ritz, T.P.; Weiskirchen, R.; Tacke, F. Single Cell RNA Sequencing Identifies Subsets of Hepatic Stellate Cells and Myofibroblasts in Liver Fibrosis. Cells 2019, 8, 503. [Google Scholar] [CrossRef]
- MacParland, S.A.; Liu, J.C.; Ma, X.Z.; Innes, B.T.; Bartczak, A.M.; Gage, B.K.; Manuel, J.; Khuu, N.; Echeverri, J.; Linares, I.; et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 2018, 9, 4383. [Google Scholar] [CrossRef]
- Lombardo, J.A.; Aliaghaei, M.; Nguyen, Q.H.; Kessenbrock, K.; Haun, J.B. Microfluidic platform accelerates tissue processing into single cells for molecular analysis and primary culture models. Nat. Commun. 2021, 12, 2858. [Google Scholar] [CrossRef]
- Morán-Salvador, E.; López-Parra, M.; García-Alonso, V.; Titos, E.; Martínez-Clemente, M.; González-Périz, A.; López-Vicario, C.; Barak, Y.; Arroyo, V.; Clària, J. Role for PPARγ in obesity-induced hepatic steatosis as determined by hepatocyte- and macrophage-specific conditional knockouts. FASEB J. 2011, 25, 2538–2550. [Google Scholar] [CrossRef]
- Charlier, N.; Neyrinck, A.M.; Beghein, N.; Delzenne, N.M.; Gallez, B. Assessment of liver phagocytic activity using EPR spectrometry and imaging. Magn. Reson. Imaging 2009, 27, 565–569. [Google Scholar] [CrossRef] [PubMed]
- Neyrinck, A.M.; Taper, H.S.; Gevers, V.; Declerck, B.; Delzenne, N.M. Inhibition of Kupffer cell activity induces hepatic triglyceride synthesis in fasted rats, independent of lipopolysaccharide challenge. J. Hepatol. 2002, 36, 466–473. [Google Scholar] [CrossRef]
- Neyrinck, A.M.; Gomez, C.; Delzenne, N.M. Precision-cut liver slices in culture as a tool to assess the physiological involvement of Kupffer cells in hepatic metabolism. Comp. Hepatol. 2004, 3 (Suppl. 1), S45. [Google Scholar] [CrossRef][Green Version]
- Vilaseca, M.; García-Calderó, H.; Lafoz, E.; Ruart, M.; López-Sanjurjo, C.I.; Murphy, M.P.; Deulofeu, R.; Bosch, J.; Hernández-Gea, V.; Gracia-Sancho, J.; et al. Mitochondria-targeted antioxidant mitoquinone deactivates human and rat hepatic stellate cells and reduces portal hypertension in cirrhotic rats. Liver Int. 2017, 37, 1002–1012. [Google Scholar] [CrossRef] [PubMed]
- Page, A.; Paoli, P.P.; Hill, S.J.; Howarth, R.; Wu, R.; Kweon, S.M.; French, J.; White, S.; Tsukamoto, H.; Mann, D.A.; et al. Alcohol directly stimulates epigenetic modifications in hepatic stellate cells. J. Hepatol. 2015, 62, 388–397. [Google Scholar] [CrossRef]
- Van Dijk, F.; Hazelhoff, C.M.; Post, E.; Prins, G.G.H.; Rombouts, K.; Poelstra, K.; Olinga, P.; Beljaars, L. Design of a Gene Panel to Expose the Versatile Role of Hepatic Stellate Cells in Human Liver Fibrosis. Pharmaceutics 2020, 12, 278. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dewyse, L.; Reynaert, H.; van Grunsven, L.A. Best Practices and Progress in Precision-Cut Liver Slice Cultures. Int. J. Mol. Sci. 2021, 22, 7137. https://doi.org/10.3390/ijms22137137
Dewyse L, Reynaert H, van Grunsven LA. Best Practices and Progress in Precision-Cut Liver Slice Cultures. International Journal of Molecular Sciences. 2021; 22(13):7137. https://doi.org/10.3390/ijms22137137
Chicago/Turabian StyleDewyse, Liza, Hendrik Reynaert, and Leo A. van Grunsven. 2021. "Best Practices and Progress in Precision-Cut Liver Slice Cultures" International Journal of Molecular Sciences 22, no. 13: 7137. https://doi.org/10.3390/ijms22137137
APA StyleDewyse, L., Reynaert, H., & van Grunsven, L. A. (2021). Best Practices and Progress in Precision-Cut Liver Slice Cultures. International Journal of Molecular Sciences, 22(13), 7137. https://doi.org/10.3390/ijms22137137






