Developments of Riboswitches and Toehold Switches for Molecular Detection—Biosensing and Molecular Diagnostics
Abstract
1. Introduction
2. Operating Principles of Riboswitches and Toehold Switches
2.1. Riboswitches
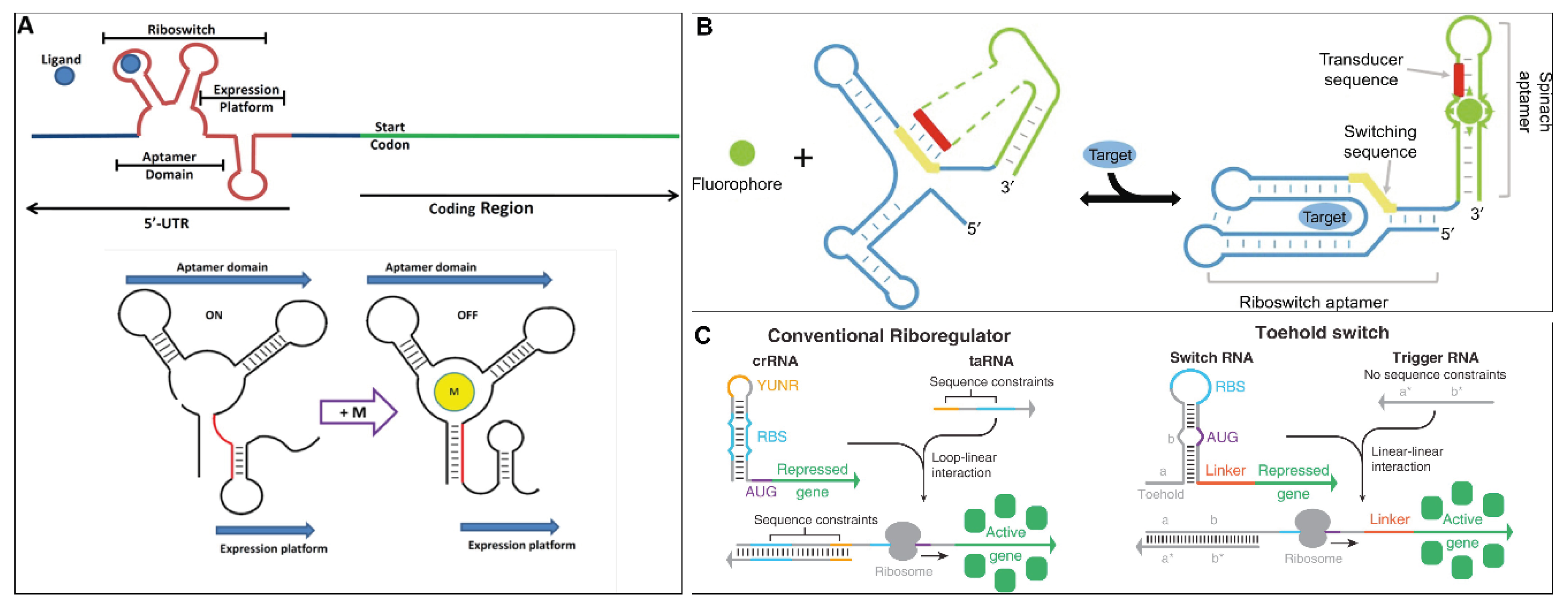
2.2. Toehold Switches
3. Design Approaches for the Use of Riboswitches and Toehold Switches in Molecular Detection
3.1. Design Principles of Riboswitches
3.1.1. Aptamer Screening Approaches
3.1.2. Design Principles
3.2. Design Principles of Toehold Switches
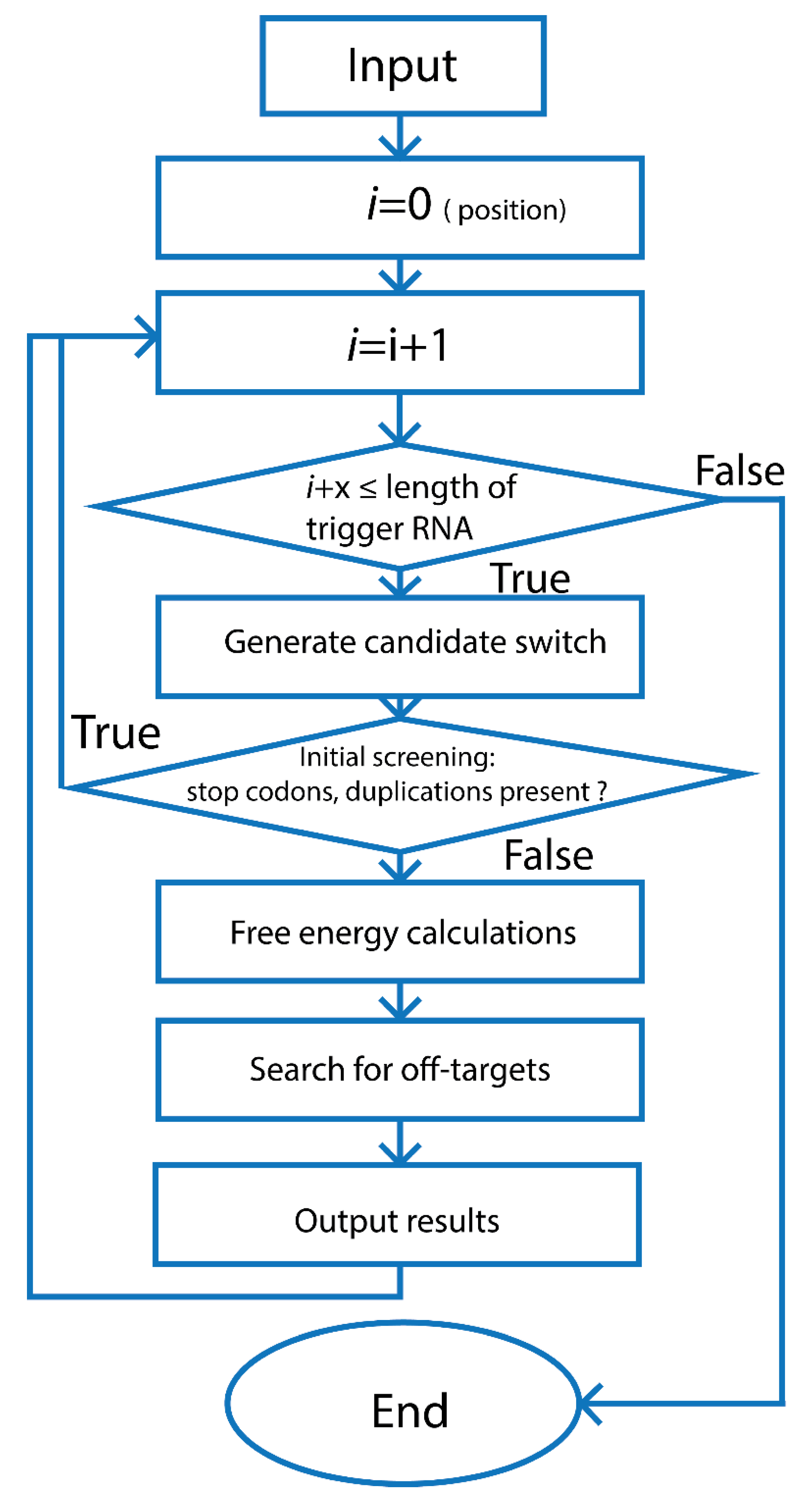
3.2.1. Design Principles
3.2.2. Modifying Toehold Switch Parameters for the Fine-Tuning of Gene Expression
4. Applications of Riboswitches and Toehold Switches in Molecular Detection
4.1. Applications of Riboswitches
4.2. Applications of Toehold Switches
5. Conclusions and Perspective
Author Contributions
Funding
Conflicts of Interest
References
- Harbaugh, S.; Goodson, M.; Chushak, Y.; Chávez, J.; Kelley-Loughnane, N. Riboswitches as sensor entities. In Handbook of Cell Biosensors; Thouand, G., Ed.; Springer: Cham, Switzerland, 2019; pp. 1–30. [Google Scholar] [CrossRef]
- Rodríguez-Serrano, A.F.; Hsing, I.M. 110th Anniversary: Engineered ribonucleic acid control elements as biosensors for in vitro diagnostics. Ind. Eng. Chem. Res. 2019, 58, 17174–17181. [Google Scholar] [CrossRef]
- Hallberg, Z.F.; Su, Y.; Kitto, R.Z.; Hammond, M.C. Engineering and in vivo applications of riboswitches. Annu. Rev. Biochem. 2017, 86, 515–539. [Google Scholar] [CrossRef] [PubMed]
- Villa, J.K.; Su, Y.; Contreras, L.M.; Hammond, M.C. Synthetic biology of small RNAs and riboswitches. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef]
- Contreras, L.M. Methods and advances in RNA characterization and design. Methods 2018, 143, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Moon, T.S. Design rules of synthetic non-coding RNAs in bacteria. Methods 2018, 143, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Jensen, M.K.; Keasling, J.D. Development of biosensors and their application in metabolic engineering. Curr. Opin. Chem. Biol. 2015, 28, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Berens, C.; Suess, B. Riboswitch engineering—Making the all-important second and third steps. Curr. Opin. Biotechnol. 2015, 31, 10–15. [Google Scholar] [CrossRef]
- Jang, S.; Jang, S.; Yang, J.; Seo, S.W.; Jung, G.Y. RNA-based dynamic genetic controllers: Development strategies and applications. Curr. Opin. Biotechnol. 2018, 53, 1–11. [Google Scholar] [CrossRef]
- Machtel, P.; Bakowska-Zywicka, K.; Zywicki, M. Emerging applications of riboswitches—From antibacterial targets to molecular tools. J. Appl. Genet. 2016, 57, 531–541. [Google Scholar] [CrossRef]
- Mehdizadeh Aghdam, E.; Hejazi, M.S.; Barzegar, A. Riboswitches: From living biosensors to novel targets of antibiotics. Gene 2016, 592, 244–259. [Google Scholar] [CrossRef]
- Kawasaki, S.; Fujita, Y.; Nagaike, T.; Tomita, K.; Saito, H. Synthetic mRNA devices that detect endogenous proteins and distinguish mammalian cells. Nucleic Acids. Res. 2017, 45, e117. [Google Scholar] [CrossRef] [PubMed]
- Yokobayashi, Y. Aptamer-based and aptazyme-based riboswitches in mammalian cells. Curr. Opin. Chem. Biol. 2019, 52, 72–78. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, F.J.; Dwyer, D.J.; Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 2006, 24, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, F.J.; Dwyer, D.J.; Ding, C.; Pervouchine, D.D.; Cantor, C.R.; Collins, J.J. Engineered riboregulators enable post-transcriptional control of gene expression. Nat. Biotechnol. 2004, 22, 841. [Google Scholar] [CrossRef] [PubMed]
- Green, A.A.; Silver, P.A.; Collins, J.J.; Yin, P. Toehold switches: De-novo-designed regulators of gene expression. Cell 2014, 159, 925–939. [Google Scholar] [CrossRef]
- Pardee, K.; Green, A.A.; Ferrante, T.; Cameron, D.E.; DaleyKeyser, A.; Yin, P.; Collins, J.J. Paper-based synthetic gene networks. Cell 2014, 159, 940–954. [Google Scholar] [CrossRef]
- Pardee, K.; Green, A.A.; Takahashi, M.K.; Braff, D.; Lambert, G.; Lee, J.W.; Ferrante, T.; Ma, D.; Donghia, N.; Fan, M.; et al. Rapid, low-cost detection of zika virus using programmable biomolecular components. Cell 2016, 165, 1255–1266. [Google Scholar] [CrossRef]
- Dincer, C.; Bruch, R.; Kling, A.; Dittrich, P.S.; Urban, G.A. Multiplexed point-of-care testing—xPOCT. Trends Biotechnol. 2017, 35, 728–742. [Google Scholar] [CrossRef]
- Nguyen, H.V.; Nguyen, V.D.; Nguyen, H.Q.; Chau, T.H.T.; Lee, E.Y.; Seo, T.S. Nucleic acid diagnostics on the total integrated lab-on-a-disc for point-of-care testing. Biosens. Bioelectron. 2019, 141, 111466. [Google Scholar] [CrossRef]
- Nguyen, H.V.; Nguyen, V.D.; Lee, E.Y.; Seo, T.S. Point-of-care genetic analysis for multiplex pathogenic bacteria on a fully integrated centrifugal microdevice with a large-volume sample. Biosens. Bioelectron. 2019, 136, 132–139. [Google Scholar] [CrossRef]
- Slomovic, S.; Pardee, K.; Collins, J.J. Synthetic biology devices for in vitro and in vivo diagnostics. Proc. Natl. Acad. Sci. USA 2015, 112, 14429–14435. [Google Scholar] [CrossRef] [PubMed]
- Pardee, K. Perspective: Solidifying the impact of cell-free synthetic biology through lyophilization. Biochem. Eng. J. 2018, 138, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Soltani, M.; Davis, B.R.; Ford, H.; Nelson, J.A.D.; Bundy, B.C. Reengineering cell-free protein synthesis as a biosensor: Biosensing with transcription, translation, and protein-folding. Biochem. Eng. J. 2018, 138, 165–171. [Google Scholar] [CrossRef]
- Pardee, K.; Slomovic, S.; Nguyen, P.Q.; Lee, J.W.; Donghia, N.; Burrill, D.; Ferrante, T.; McSorley, F.R.; Furuta, Y.; Vernet, A.; et al. Portable, on-demand biomolecular manufacturing. Cell 2016, 167, 248–259. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Shen, L.; Wu, K.; Diehnelt, C.W.; Green, A.A. Low-cost detection of norovirus using paper-based cell-free systems and synbody-based viral enrichment. Synth. Biol. 2018, 3, ysy018. [Google Scholar] [CrossRef]
- Urdea, M.; Penny, L.A.; Olmsted, S.S.; Giovanni, M.Y.; Kaspar, P.; Shepherd, A.; Wilson, P.; Dahl, C.A.; Buchsbaum, S.; Moeller, G.; et al. Requirements for high impact diagnostics in the developing world. Nature 2006, 444, 73. [Google Scholar] [CrossRef]
- Grand View Research. Available online: https://www.grandviewresearch.com/industry-analysis/biosensors-market (accessed on 27 April 2020).
- Morel, C.; McClure, L.; Edwards, S.; Goodfellow, V.; Sandberg, D.; Thomas, J.; Mossialos, E. Ensuring Innovation in Diagnostics for Bacterial Infection: Implications for Policy; European Observatory on Health Systems and Policies: Copenhagen, Denmark, 2016; ISBN 978-92-890-50364. [Google Scholar]
- Parkhey, P.; Mohan, S.V. Chapter 6.1—Biosensing applications of microbial fuel cell: Approach toward miniaturization. In Microbial Electrochemical Technology; Mohan, S.V., Varjani, S., Pandey, A., Eds.; Elsevier: Amsterdam, The Netherlands, 2019; pp. 977–997. [Google Scholar] [CrossRef]
- Geraldi, A.; Giri-Rachman, E.A. Synthetic biology-based portable in vitro diagnostic platforms. Alex. J. Med. 2019, 54, 423–428. [Google Scholar] [CrossRef]
- Su, L.; Jia, W.; Hou, C.; Lei, Y. Microbial biosensors: A review. Biosens. Bioelectron. 2011, 26, 1788–1799. [Google Scholar] [CrossRef]
- Findeiß, S.; Etzel, M.; Will, S.; Mörl, M.; Stadler, P.F. Design of artificial riboswitches as biosensors. Sensors 2017, 17, 1990. [Google Scholar] [CrossRef]
- Serganov, A.; Nudler, E. A decade of riboswitches. Cell 2013, 152, 17–24. [Google Scholar] [CrossRef]
- Gusarov, I.; Nudler, E. The mechanism of intrinsic transcription termination. Mol. Cell. 1999, 3, 495–504. [Google Scholar] [CrossRef]
- Martin, F.H.; Tinoco, I., Jr. DNA-RNA hybrid duplexes containing oligo (dA: rU) sequences are exceptionally unstable and may facilitate termination of transcription. Nucleic Acids Res. 1980, 8, 2295–2300. [Google Scholar] [CrossRef] [PubMed]
- Skordalakes, E.; Berger, J.M. Structure of the Rho transcription terminator: Mechanism of mRNA recognition and helicase loading. Cell 2003, 114, 135–146. [Google Scholar] [CrossRef]
- Gu, H.; Duan, N.; Xia, Y.; Hun, X.; Wang, H.; Wang, Z. Magnetic separation-based multiple SELEX for effectively selecting aptamers against saxitoxin, domoic acid, and tetrodotoxin. J. Agric. Food Chem. 2018, 66, 9801–9809. [Google Scholar] [CrossRef] [PubMed]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef] [PubMed]
- Filonov, G.S.; Moon, J.D.; Svensen, N.; Jaffrey, S.R. Broccoli: Rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 2014, 136, 16299–16308. [Google Scholar] [CrossRef]
- Song, W.; Filonov, G.S.; Kim, H.; Hirsch, M.; Li, X.; Moon, J.D.; Jaffrey, S.R. Imaging RNA polymerase III transcription using a photostable RNA–fluorophore complex. Nat. Chem. Biol. 2017, 13, 1187. [Google Scholar] [CrossRef]
- Trachman, R.J., 3rd; Truong, L.; Ferre-D’Amare, A.R. Structural principles of fluorescent RNA aptamers. Trends Pharmacol. Sci. 2017, 38, 928–939. [Google Scholar] [CrossRef]
- Jaffrey, S.R. Chapter Nine—RNA-based fluorescent biosensors for detecting metabolites in vitro and in living cells. In Advances in Pharmacology; Pasternak, G.W., Coyle, J.T., Eds.; Academic Press: Cambridge, MA, USA, 2018; Volume 82, pp. 187–203. [Google Scholar] [CrossRef]
- You, M.; Litke, J.L.; Jaffrey, S.R. Imaging metabolite dynamics in living cells using a spinach-based riboswitch. Proc. Natl. Acad. Sci. USA 2015, 112, E2756–E2765. [Google Scholar] [CrossRef]
- Carpenter, A.C.; Paulsen, I.T.; Williams, T.C. Blueprints for biosensors: Design, limitations, and applications. Genes 2018, 9, 375. [Google Scholar] [CrossRef]
- Liang, J.C.; Bloom, R.J.; Smolke, C.D. Engineering biological systems with synthetic RNA molecules. Mol. Cell. 2011, 43, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Ausländer, S.; Fussenegger, M. Toehold gene switches make big footprints. Nature 2014, 516, 333. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Evans, T.; Zhang, F. Applications and advances of metabolite biosensors for metabolic engineering. Metab. Eng. 2015, 31, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Galizi, R.; Jaramillo, A. Engineering CRISPR guide RNA riboswitches for in vivo applications. Curr. Opin. Biotechnol. 2019, 55, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818. [Google Scholar] [CrossRef] [PubMed]
- Szeto, K.; Latulippe, D.R.; Ozer, A.; Pagano, J.M.; White, B.S.; Shalloway, D.; Lis, J.T.; Craighead, H.G. Rapid-SELEX for RNA aptamers. PLoS ONE 2013, 8, e82667. [Google Scholar] [CrossRef] [PubMed]
- Szeto, K.; Craighead, H.G. Devices and approaches for generating specific high-affinity nucleic acid aptamers. Appl. Phys. Rev. 2014, 1, 031103. [Google Scholar] [CrossRef]
- Luo, X.; McKeague, M.; Pitre, S.; Dumontier, M.; Green, J.; Golshani, A.; Derosa, M.C.; Dehne, F. Computational approaches toward the design of pools for the in vitro selection of complex aptamers. RNA 2010, 16, 2252–2262. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Urak, K.T.; Shore, S.; Rockey, W.M.; Chen, S.-J.; McCaffrey, A.P.; Giangrande, P.H. In vitro RNA SELEX for the generation of chemically-optimized therapeutic RNA drugs. Methods 2016, 103, 167–174. [Google Scholar] [CrossRef]
- Lauridsen, L.H.; Doessing, H.B.; Long, K.S.; Nielsen, A.T. A capture-SELEX strategy for multiplexed selection of RNA aptamers against small molecules. In Synthetic Metabolic Pathways; Springer: Cham, Switzerland, 2018; pp. 291–306. [Google Scholar] [CrossRef]
- Dong, L.; Tan, Q.; Ye, W.; Liu, D.; Chen, H.; Hu, H.; Wen, D.; Liu, Y.; Cao, Y.; Kang, J.; et al. Screening and Identifying A Novel ssDNA aptamer against alpha-fetoprotein using CE-SELEX. Sci. Rep. 2015, 5, 15552. [Google Scholar] [CrossRef] [PubMed]
- Lamberti, I.; Scarano, S.; Esposito, C.L.; Antoccia, A.; Antonini, G.; Tanzarella, C.; De Franciscis, V.; Minunni, M. In vitro selection of RNA aptamers against CA125 tumor marker in ovarian cancer and its study by optical biosensing. Methods 2016, 97, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Turner, D.H.; Mathews, D.H. NNDB: The nearest neighbor parameter database for predicting stability of nucleic acid secondary structure. Nucleic Acids Res. 2009, 38, D280–D282. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M.; Jaeger, J.A.; Turner, D.H. A comparison of optimal and suboptimal RNA secondary structures predicted by free energy minimization with structures determined by phylogenetic comparison. Nucleic Acids Res. 1991, 19, 2707–2714. [Google Scholar] [CrossRef]
- Xia, T.; SantaLucia, J., Jr.; Burkard, M.E.; Kierzek, R.; Schroeder, S.J.; Jiao, X.; Cox, C.; Turner, D.H. Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with watson—Crick base pairs. Biochemistry 1998, 37, 14719–14735. [Google Scholar] [CrossRef]
- Wu, J.C.; Gardner, D.P.; Ozer, S.; Gutell, R.R.; Ren, P. Correlation of RNA secondary structure statistics with thermodynamic stability and applications to folding. J. Mol. Biol. 2009, 391, 769–783. [Google Scholar] [CrossRef]
- Wachsmuth, M.; Domin, G.; Lorenz, R.; Serfling, R.; Findeiß, S.; Stadler, P.F.; Mörl, M. Design criteria for synthetic riboswitches acting on transcription. RNA Biology 2015, 12, 221–231. [Google Scholar] [CrossRef]
- Peters, J.M.; Vangeloff, A.D.; Landick, R. Bacterial transcription terminators: The RNA 3′-end chronicles. J. Mol. Biol. 2011, 412, 793–813. [Google Scholar] [CrossRef]
- Mathews, D.H.; Turner, D.H.; Zuker, M. RNA secondary structure prediction. Curr. Protoc. Nucleic Acid Chem. 2007. [Google Scholar] [CrossRef]
- Proctor, J.R.; Meyer, I.M. COFOLD: An RNA secondary structure prediction method that takes co-transcriptional folding into account. Nucleic Acids Res. 2013, 41, e102. [Google Scholar] [CrossRef]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Markham, N.R.; Zuker, M. UNAFold. In Bioinformatics; Springer: Cham, Switzerlands, 2008; pp. 3–31. [Google Scholar] [CrossRef]
- Hofacker, I.L. Vienna RNA secondary structure server. Nucleic Acids Res. 2003, 31, 3429–3431. [Google Scholar] [CrossRef] [PubMed]
- Reuter, J.S.; Mathews, D.H. RNAstructure: Software for RNA secondary structure prediction and analysis. BMC Bioinform. 2010, 11, 129. [Google Scholar] [CrossRef] [PubMed]
- Zadeh, J.N.; Steenberg, C.D.; Bois, J.S.; Wolfe, B.R.; Pierce, M.B.; Khan, A.R.; Dirks, R.M.; Pierce, N.A. NUPACK: Analysis and design of nucleic acid systems. J. Comput. Chem. 2011, 32, 170–173. [Google Scholar] [CrossRef]
- Busch, A.; Richter, A.S.; Backofen, R. IntaRNA: Efficient prediction of bacterial sRNA targets incorporating target site accessibility and seed regions. Bioinformatics 2008, 24, 2849–2856. [Google Scholar] [CrossRef]
- Wu, M.J.; Andreasson, J.O.; Kladwang, W.; Greenleaf, W.; Das, R. Automated design of diverse stand-alone riboswitches. ACS Synth. Biol. 2019, 8, 1838–1846. [Google Scholar] [CrossRef]
- Meagher, R.J.; Negrete, O.A.; Van Rompay, K.K. Engineering paper-based sensors for zika virus. Trends Mol. Med. 2016, 22, 529–530. [Google Scholar] [CrossRef]
- Pham, H.L.; Wong, A.; Chua, N.; Teo, W.S.; Yew, W.S.; Chang, M.W. Engineering a riboswitch-Based genetic platform for the self-directed evolution of acid-tolerant phenotypes. Nat. Commun. 2017, 8, 411. [Google Scholar] [CrossRef]
- Wang, S.; Emery, N.J.; Liu, A.P. A novel synthetic toehold switch for microRNA detection in mammalian cells. ACS Synth. Biol. 2019, 8, 1079–1088. [Google Scholar] [CrossRef]
- To, A.C.-Y.; Chu, D.H.-T.; Wang, A.R.; Li, F.C.-Y.; Chiu, A.W.-O.; Gao, D.Y.; Choi, C.H.J.; Kong, S.-K.; Chan, T.-F.; Chan, K.-M.; et al. A comprehensive web tool for toehold switch design. Bioinformatics 2018, 34, 2862–2864. [Google Scholar] [CrossRef]
- Serra, M.J.; Turner, D.H. Predicting thermodynamic properties of RNA. In Methods in enzymology; Elsevier: Amsterdam, The Netherlands, 1995; Volume 259, pp. 242–261. [Google Scholar] [CrossRef]
- Mathews, D.H.; Sabina, J.; Zuker, M.; Turner, D.H. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J. Mol. Biol. 1999, 288, 911–940. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, M.K.; Tan, X.; Dy, A.J.; Braff, D.; Akana, R.T.; Furuta, Y.; Donghia, N.; Ananthakrishnan, A.; Collins, J.J. A low-cost paper-based synthetic biology platform for analyzing gut microbiota and host biomarkers. Nat. Commun. 2018, 9, 3347. [Google Scholar] [CrossRef] [PubMed]
- Fowler, C.C.; Brown, E.D.; Li, Y. Using a riboswitch sensor to examine coenzyme B12 metabolism and transport in E. coli. Chem. Biol. 2010, 17, 756–765. [Google Scholar] [CrossRef] [PubMed]
- Fowler, C.C.; Sugiman-Marangos, S.; Junop, M.S.; Brown, E.D.; Li, Y. Exploring intermolecular interactions of a substrate binding protein using a riboswitch-based sensor. Chem. Biol. 2013, 20, 1502–1512. [Google Scholar] [CrossRef]
- Verhounig, A.; Karcher, D.; Bock, R. Inducible gene expression from the plastid genome by a synthetic riboswitch. Proc. Natl. Acad. Sci. USA 2010, 107, 6204–6209. [Google Scholar] [CrossRef]
- Michener, J.K.; Smolke, C.D. High-throughput enzyme evolution in saccharomyces cerevisiae using a synthetic RNA switch. Metab. Eng. 2012, 14, 306–316. [Google Scholar] [CrossRef]
- Yang, J.; Seo, S.W.; Jang, S.; Shin, S.-I.; Lim, C.H.; Roh, T.-Y.; Jung, G.Y. Synthetic RNA devices to expedite the evolution of metabolite-producing microbes. Nat. Commun. 2013, 4, 1413. [Google Scholar] [CrossRef]
- Nomura, Y.; Kumar, D.; Yokobayashi, Y. Synthetic mammalian riboswitches based on guanine aptazyme. Chem. Commun. 2012, 48, 7215–7217. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Chen, C.; Whiteley, A.T.; Portnoy, D.A.; Hammond, M.C. RNA-Based fluorescent biosensors for live cell imaging of second messenger cyclic Di-AMP. J. Am. Chem. Soc. 2015, 137, 6432–6435. [Google Scholar] [CrossRef]
- Su, Y.; Hickey, S.F.; Keyser, S.G.; Hammond, M.C. In Vitro and In vivo enzyme activity screening via RNA-based fluorescent biosensors for S-Adenosyl-L-Homocysteine (SAH). J. Am. Chem. Soc. 2016, 138, 7040–7047. [Google Scholar] [CrossRef]
- Hollands, K.; Proshkin, S.; Sklyarova, S.; Epshtein, V.; Mironov, A.; Nudler, E.; Groisman, E.A. Riboswitch control of Rho-dependent transcription termination. Proc. Natl. Acad. Sci. USA 2012, 109, 5376–5381. [Google Scholar] [CrossRef] [PubMed]
- Haller, A.; Rieder, U.; Aigner, M.; Blanchard, S.C.; Micura, R. Conformational capture of the SAM-II riboswitch. Nat. Chem. Biol. 2011, 7, 393. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.G.; Sudarsan, N.; Breaker, R.R. Mechanism for gene control by a natural allosteric group I ribozyme. RNA 2011, 17, 1967–1972. [Google Scholar] [CrossRef] [PubMed]
- Cheah, M.T.; Wachter, A.; Sudarsan, N.; Breaker, R.R. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 2007, 447, 497. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Wilson, S.C.; Hickey, S.F.; Gonzalez, T.L.; Su, Y.; Hallberg, Z.F.; Brewer, T.F.; Iavarone, A.T.; Carlson, H.K.; Hsieh, Y.-F.; et al. GEMM-I riboswitches from geobacter sense the bacterial second messenger cyclic AMP-GMP. Proc. Natl. Acad. Sci. USA 2015, 112, 5383–5388. [Google Scholar] [CrossRef]
- Weinberg, Z.; Nelson, J.W.; Lunse, C.E.; Sherlock, M.E.; Breaker, R.R. Bioinformatic analysis of riboswitch structures uncovers variant classes with altered ligand specificity. Proc. Natl. Acad. Sci. USA 2017, 114, E2077–E2085. [Google Scholar] [CrossRef]
- McCown, P.J.; Corbino, K.A.; Stav, S.; Sherlock, M.E.; Breaker, R.R. Riboswitch diversity and distribution. RNA 2017, 23, 995–1011. [Google Scholar] [CrossRef]
- Atilho, R.M.; Perkins, K.R.; Breaker, R.R. Rare variants of the FMN riboswitch class in clostridium difficile and other bacteria exhibit altered ligand specificity. RNA 2019, 25, 23–34. [Google Scholar] [CrossRef]
- Page, K.; Shaffer, J.; Lin, S.; Zhang, M.; Liu, J.M. Engineering riboswitches in vivo using dual genetic selection and fluorescence-activated cell sorting. ACS Synth. Biol. 2018, 7, 2000–2006. [Google Scholar] [CrossRef]
- Groher, F.; Bofill-Bosch, C.; Schneider, C.; Braun, J.; Jager, S.; Geissler, K.; Hamacher, K.; Suess, B. Riboswitching with ciprofloxacin-development and characterization of a novel RNA regulator. Nucleic Acids Res. 2018, 46, 2121–2132. [Google Scholar] [CrossRef]
- Ausländer, S.; Fussenegger, M. Synthetic RNA-based switches for mammalian gene expression control. Curr. Opin. Biotechnol. 2017, 48, 54–60. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, S.; Caliskan, N.; Rodnina, M.V.; Murata, A.; Nakatani, K. Small synthetic molecule-stabilized RNA pseudoknot as an activator for—1 ribosomal frameshifting. Nucleic acids Res. 2018, 46, 8079–8089. [Google Scholar] [CrossRef] [PubMed]
- Auslander, S.; Ketzer, P.; Hartig, J.S. A ligand-dependent hammerhead ribozyme switch for controlling mammalian gene expression. Mol. Biosyst. 2010, 6, 807–814. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, A. Rational design of artificial riboswitches based on ligand-dependent modulation of Internal ribosome entry in wheat germ extract and their applications as label-free biosensors. RNA 2011, 17, 478–488. [Google Scholar] [CrossRef]
- Thavarajah, W.; Silverman, A.D.; Verosloff, M.; Kelley-Loughnane, N.; Jewett, M.C.; Lucks, J.B. Point-of-use detection of environmental fluoride via a cell-free riboswitch-based biosensor. ACS Synth. Biol. 2019. [Google Scholar] [CrossRef]
- Wang, X.C.; Wilson, S.C.; Hammond, M.C. Next-generation RNA-based fluorescent biosensors enable anaerobic detection of cyclic Di-GMP. Nucleic Acids Res. 2016, 44, e139. [Google Scholar] [CrossRef]
- Paige, J.S.; Nguyen-Duc, T.; Song, W.; Jaffrey, S.R. Fluorescence imaging of cellular metabolites with RNA. Science 2012, 335, 1194. [Google Scholar] [CrossRef]
- Kellenberger, C.A.; Wilson, S.C.; Sales-Lee, J.; Hammond, M.C. RNA-based fluorescent biosensors for live cell imaging of second messengers cyclic Di-GMP and cyclic AMP-GMP. J. Am. Chem. Soc. 2013, 135, 4906–4909. [Google Scholar] [CrossRef]
- Bouhedda, F.; Autour, A.; Ryckelynck, M. Light-up RNA aptamers and their cognate fluorogens: From their development to their applications. Int. J. Mol. Sci. 2017, 19, 44. [Google Scholar] [CrossRef]
- Mahony, J.B.; Petrich, A.; Louie, L.; Song, X.; Chong, S.; Smieja, M.; Chernesky, M.; Loeb, M.; Richardson, S. Performance and cost evaluation of one commercial and six in-house conventional and real-time reverse transcription-PCR assays for detection of severe acute respiratory syndrome coronavirus. J. Clin. Microbiol. 2004, 42, 1471–1476. [Google Scholar] [CrossRef]
- Faye, O.; Faye, O.; Dupressoir, A.; Weidmann, M.; Ndiaye, M.; Sall, A.A. One-step RT-PCR for detection of Zika virus. J. Clin. Virol. 2008, 43, 96–101. [Google Scholar] [CrossRef] [PubMed]
- Ro, Y.T.; Ticer, A.; Carrion, R., Jr.; Patterson, J.L. Rapid detection and quantification of Ebola Zaire Virus by one-step real-time quantitative reverse transcription-polymerase chain reaction. Microbiol. Immunol. 2017, 61, 130–137. [Google Scholar] [CrossRef] [PubMed]
- Shen, M.; Zhou, Y.; Ye, J.; Abdullah Al-maskri, A.A.; Kang, Y.; Zeng, S.; Cai, S. Recent advances and perspectives of nucleic acid detection for coronavirus. J. Pharm. Anal. 2020. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Zhou, Y.; Carlson, P.D.; Teichmann, M.; Chaudhary, S.; Simmel, F.C.; Silver, P.A.; Collins, J.J.; Lucks, J.B.; Yin, P.; et al. De novo-designed translation-repressing riboregulators for multi-input cellular logic. Nat. Chem. Biol. 2019, 15, 1173–1182. [Google Scholar] [CrossRef]
- Hong, F.; Ma, D.; Wu, K.; Mina, L.A.; Luiten, R.C.; Liu, Y.; Yan, H.; Green, A.A. Precise and programmable detection of mutations using ultraspecific riboregulators. Cell 2020. [Google Scholar] [CrossRef] [PubMed]
- Garst, A.D.; Edwards, A.L.; Batey, R.T. Riboswitches: Structures and mechanisms. Cold Spring Harb. Perspect. Biol. 2011, 3, a003533. [Google Scholar] [CrossRef]
- Awwad, A.M.; McKeague, M. Riboswitches and synthetic aptamers: A head-to-head comparison. Aptamers 2018, 2, 1–10. [Google Scholar]
- Mousavi, P.S.; Smith, S.J.; Chen, J.B.; Karlikow, M.; Tinafar, A.; Robinson, C.; Liu, W.; Ma, D.; Green, A.A.; Kelley, S.O.; et al. A multiplexed, electrochemical interface for gene-circuit-based sensors. Nat. Chem. 2020, 12, 48–55. [Google Scholar] [CrossRef]
- Angenent-Mari, N.M.; Garruss, A.S.; Soenksen, L.R.; Church, G.; Collins, J.J. Deep learning for RNA synthetic biology. Biorxiv 2019, 872077. [Google Scholar] [CrossRef]
- Valeri, J.A.; Collins, K.M.; Lepe, B.A.; Lu, T.K.; Camacho, D.M. Sequence-to-function deep learning frameworks for synthetic biology. Biorxiv 2019, 870055. [Google Scholar] [CrossRef]

| Class | Sensor | Type | Interaction | Application | Reference |
|---|---|---|---|---|---|
| Riboswitches | Adenosylcobalamin (coenzyme B12) sensor | Natural | Ligand-RNA | Examination of the metabolism and transportation of coenzyme B12 in Escherichia coli | [81] |
| Adenosylcobalamin (coenzyme B12) sensor | Natural | Ligand-RNA | Investigation of the coenzyme B12 transporter in E. coli | [82] | |
| Thiamin pyrophosphate (TPP) and theophylline sensors | Synthetic | Ligand-RNA | Translational regulator of gene expression in plastids | [83] | |
| Theophylline sensor | Synthetic | Ligand-RNA | High-throughput in vivo screening system of Saccharomyces cerevisiae for enzyme engineering | [84] | |
| l-Lysine sensor | synthetic | Ligand-RNA | High-throughput screening platform for the evolution of metabolite-producing E. coli | [85] | |
| pH-based sensor | Synthetic | Ligand-RNA | Precise control of E. coli gene expression under different pH conditions | [75] | |
| Guanine-based sensor | Synthetic | Ligand-RNA | Control gene expression in mammalian cells | [86] | |
| RNA-based fluorescent biosensors | TPP, guanine, adenine and SAM sensors | Synthetic | Ligand-RNA | Live imaging of metabolite dynamic changes in E. coli living cells | [44] |
| Cyclic di-GMP and cyclic AMP-GMP sensor | Synthetic | Ligand-RNA | Live imaging of cyclic dinucleotides in E. coli living cells | [87] | |
| S-adenosyl-l-homocysteine (SAH) sensors | Synthetic | Ligand-RNA | Direct detection of SAH both in vivo and in vitro | [88] | |
| Toehold switches | Ebola RNA sensor | Synthetic | RNA-RNA | Diagnosis of the Ebola virus in clinical samples | [17] |
| Zika RNA sensor | Synthetic | RNA-RNA | Diagnosis of the Zika virus in clinical samples | [25] | |
| Gut microbiota RNA sensor | Synthetic | RNA-RNA | Analysis of the gut microbiota | [80] | |
| Norovirus RNA sensor | Synthetic | RNA-RNA | Diagnosis of the norovirus in stool samples | [26] | |
| microRNA (miRNA) sensor | Synthetic | miRNA-RNA | Detection of microRNAs in the mammalian cells | [76] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoang Trung Chau, T.; Hoang Anh Mai, D.; Ngoc Pham, D.; Thi Quynh Le, H.; Yeol Lee, E. Developments of Riboswitches and Toehold Switches for Molecular Detection—Biosensing and Molecular Diagnostics. Int. J. Mol. Sci. 2020, 21, 3192. https://doi.org/10.3390/ijms21093192
Hoang Trung Chau T, Hoang Anh Mai D, Ngoc Pham D, Thi Quynh Le H, Yeol Lee E. Developments of Riboswitches and Toehold Switches for Molecular Detection—Biosensing and Molecular Diagnostics. International Journal of Molecular Sciences. 2020; 21(9):3192. https://doi.org/10.3390/ijms21093192
Chicago/Turabian StyleHoang Trung Chau, Tin, Dung Hoang Anh Mai, Diep Ngoc Pham, Hoa Thi Quynh Le, and Eun Yeol Lee. 2020. "Developments of Riboswitches and Toehold Switches for Molecular Detection—Biosensing and Molecular Diagnostics" International Journal of Molecular Sciences 21, no. 9: 3192. https://doi.org/10.3390/ijms21093192
APA StyleHoang Trung Chau, T., Hoang Anh Mai, D., Ngoc Pham, D., Thi Quynh Le, H., & Yeol Lee, E. (2020). Developments of Riboswitches and Toehold Switches for Molecular Detection—Biosensing and Molecular Diagnostics. International Journal of Molecular Sciences, 21(9), 3192. https://doi.org/10.3390/ijms21093192






