Mass Spectrometry-Based Proteomic Characterization of Cutaneous Melanoma Ectosomes Reveals the Presence of Cancer-Related Molecules
Abstract
1. Introduction
2. Results
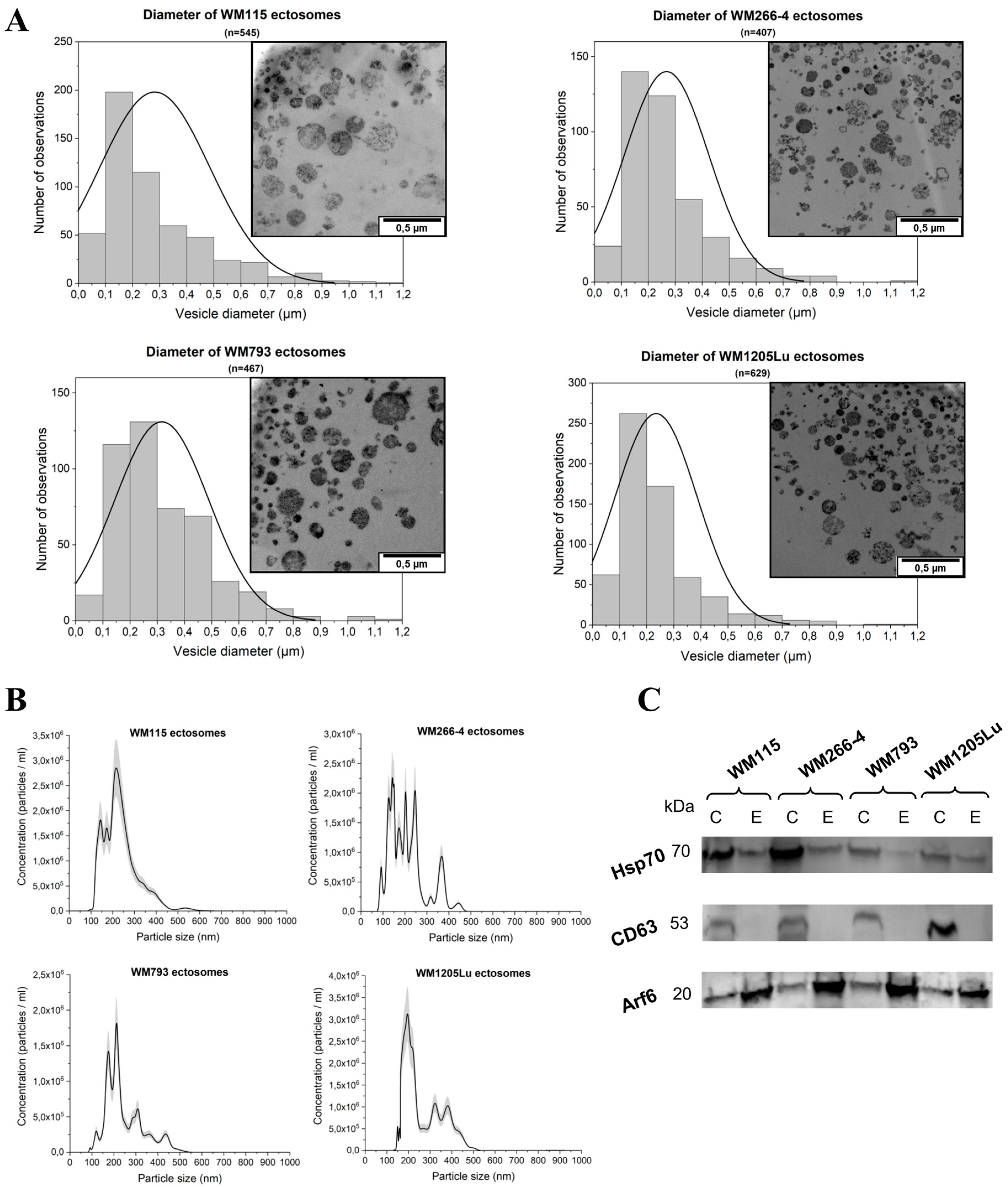
2.1. Size and Marker Characterization of CM Ectosomes
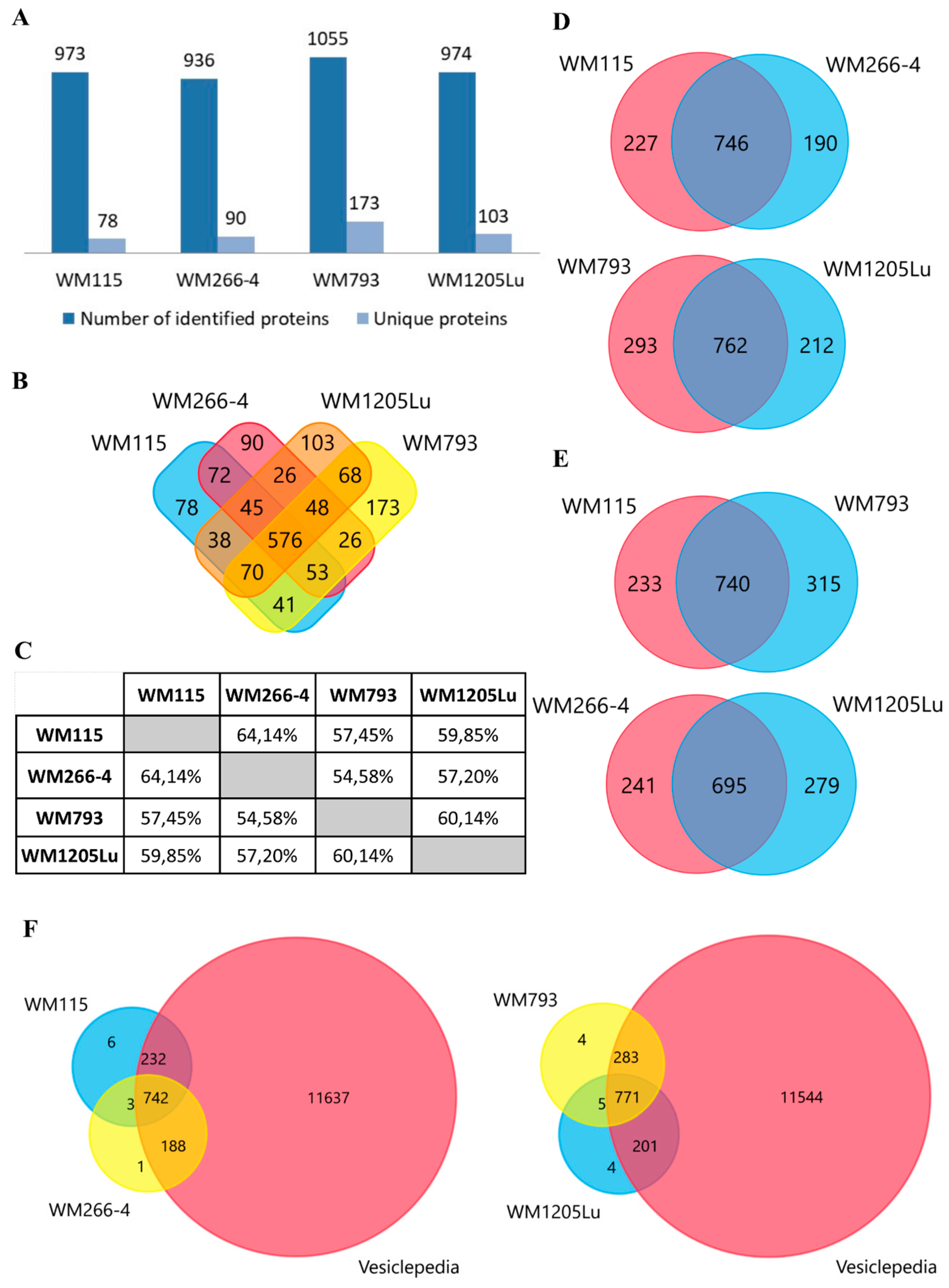
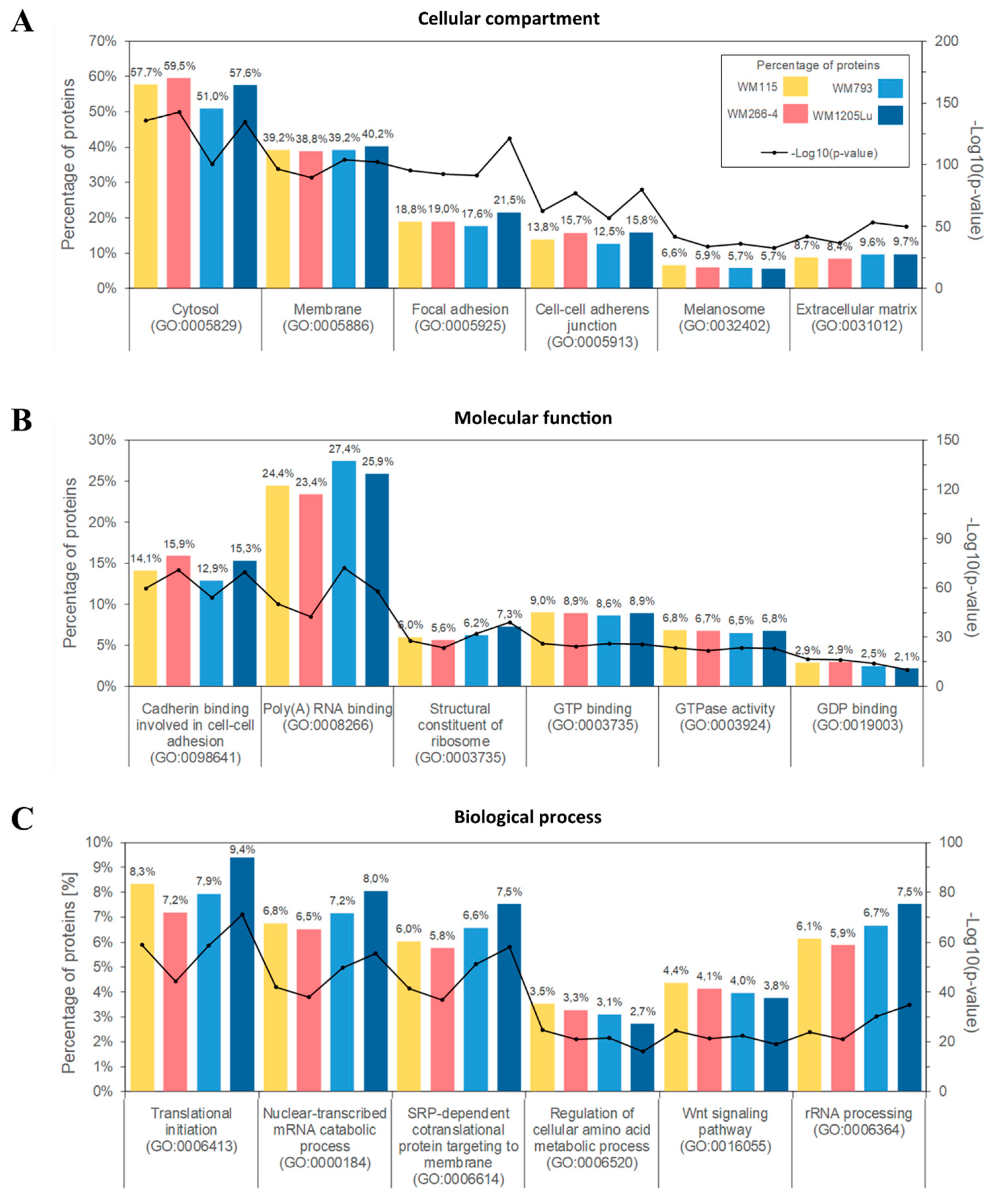
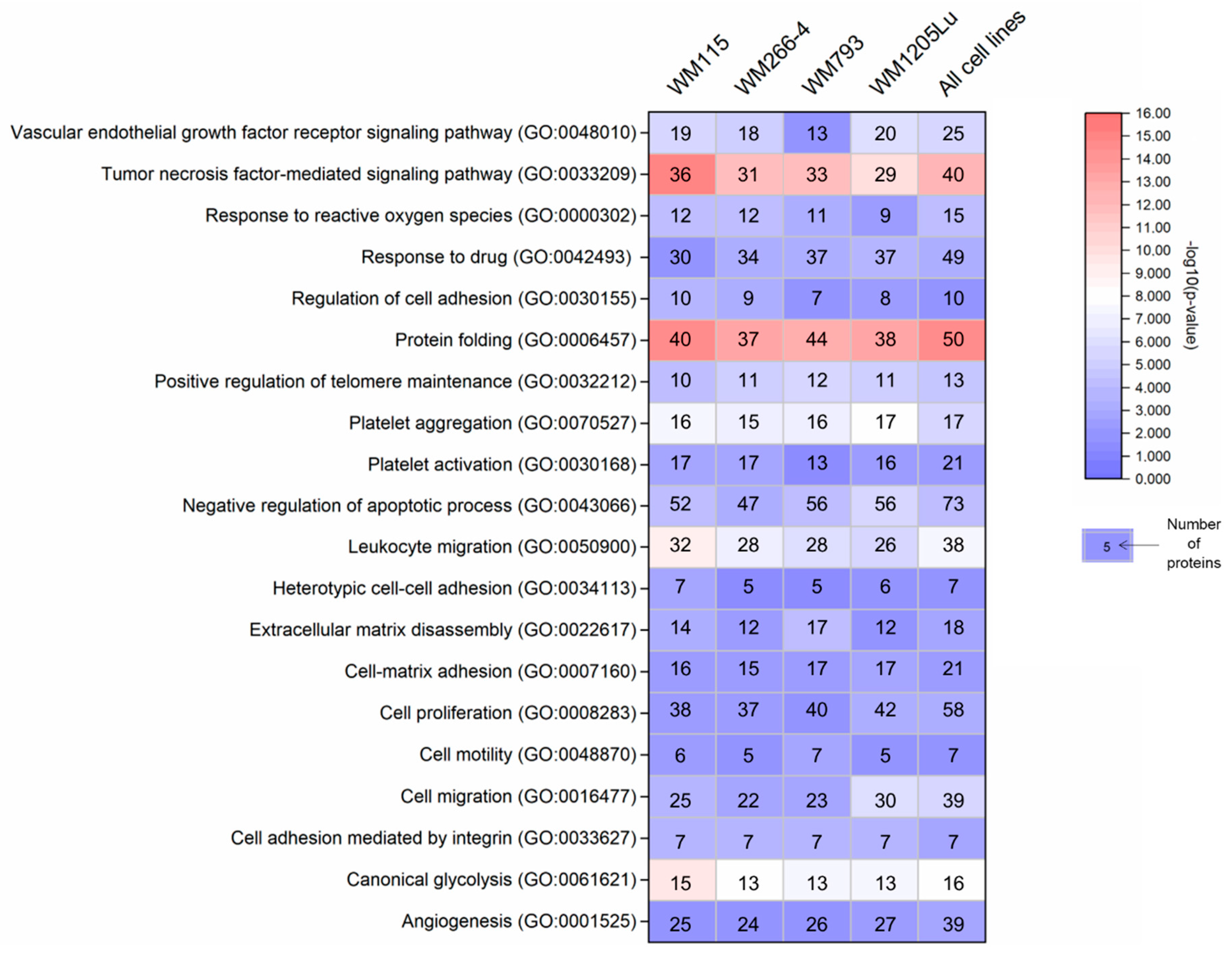
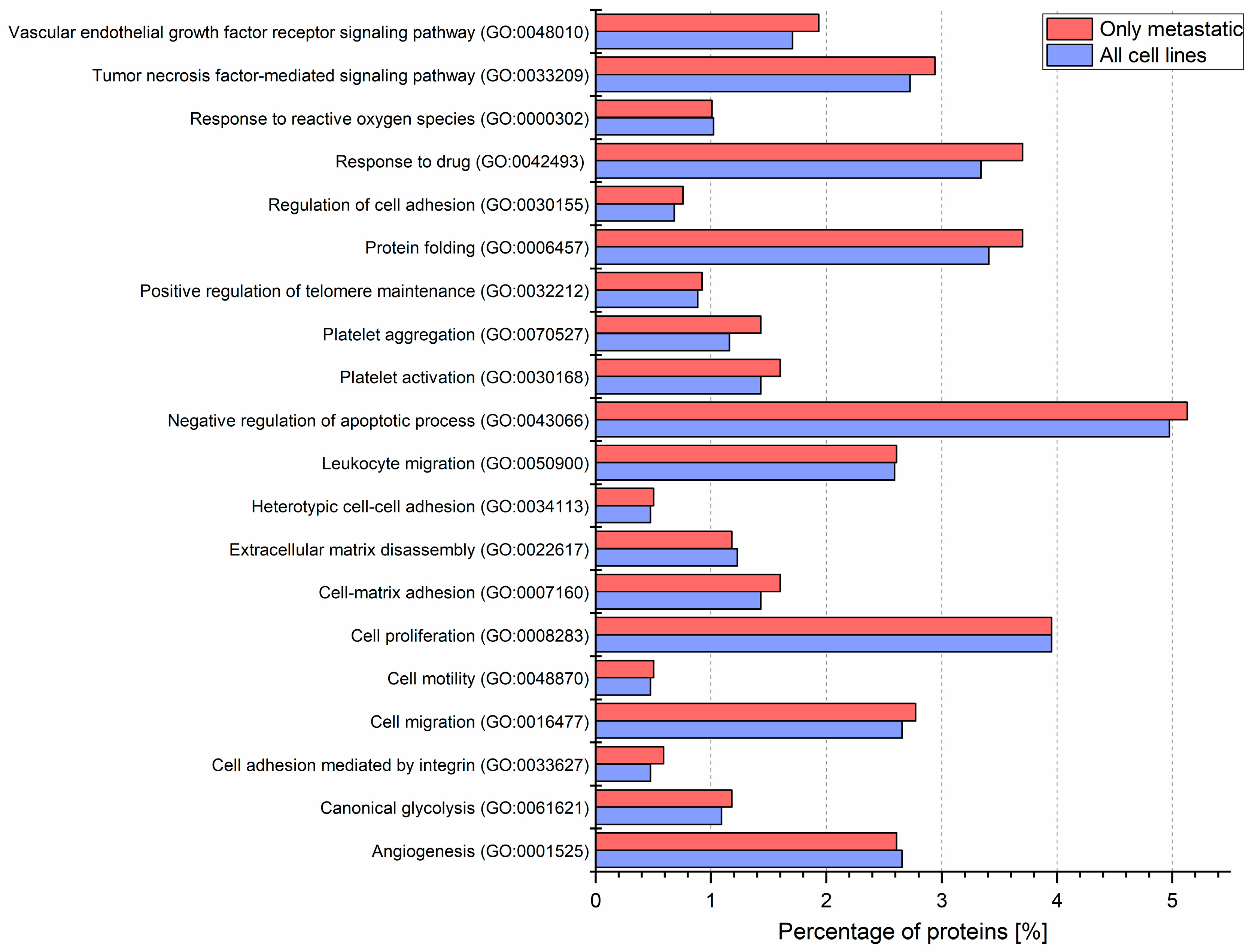
2.2. Identified Proteins of CM Ectosomes and Their Functional Classification
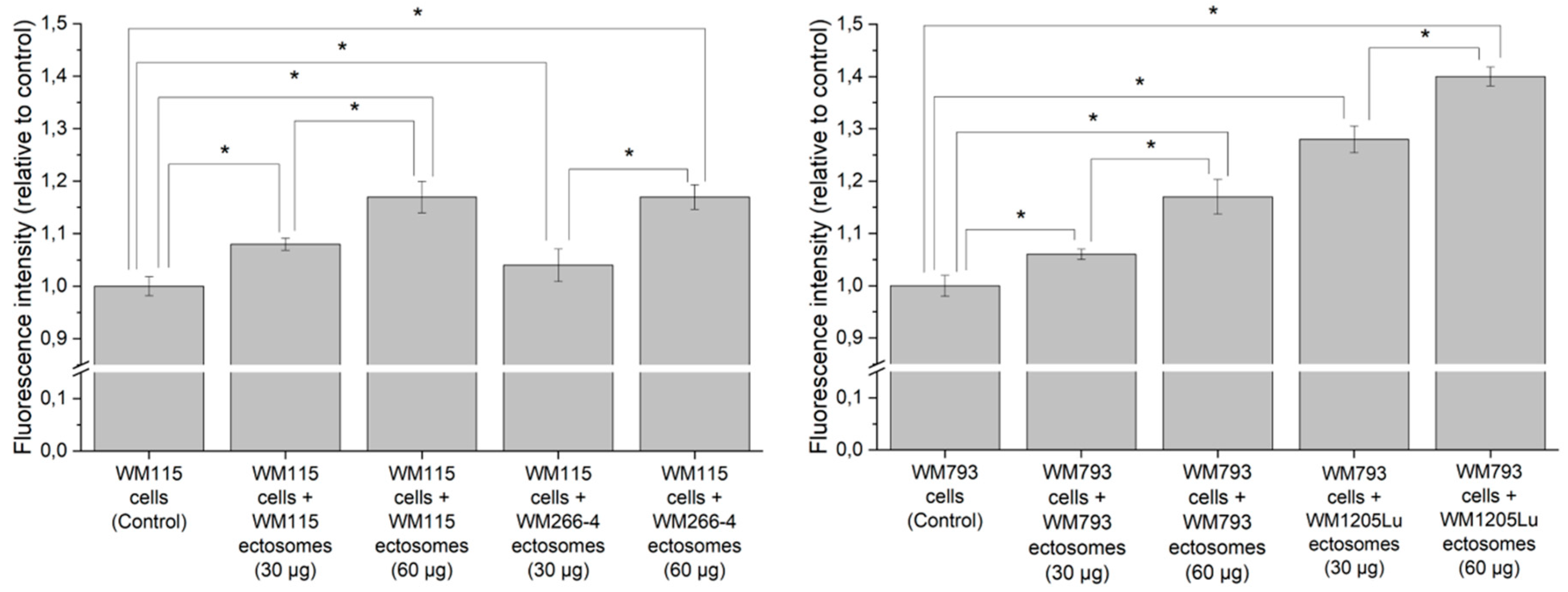
2.3. Cancer-Promoting Effect of Ectosomes Derived from Metastatic Cells on Less Invasive Cells
3. Discussion
3.1. Proteins Involved in Cancer Progression Detected in CM-derived Ectosomes by LC–MS/MS
3.2. Melanoma-Derived Ectosomes as a Source of Potential Disease Biomarkers
4. Materials and Methods
4.1. Materials
4.2. Cell Lines and Cell Culture Conditions
4.3. Isolation of Ectosomes and Assessment of the Purity of the Ectosome Samples
4.4. LC–MS/MS Proteomics
4.4.1. Ectosome Lysis
4.4.2. Sample Preparation for Mass Spectrometric Analysis
4.4.3. Liquid Chromatography and Tandem Mass Spectrometry (LC–MS/MS)
4.4.4. Analysis of Proteomic Data
4.5. Bioinformatic Analysis
4.6. Wound Healing Assay
4.7. Alamar Blue Cell Viability Assay
4.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| CM | Cutaneous Melanoma |
| EV | Extracellular Vesicles |
| LC–MS/MS | Liquid Chromatography and Mass Spectrometry |
| NTA | Nanoparticle Tracking Analysis |
| TEM | Transmission Electron Microscopy |
References
- Karimkhani, C.; Green, A.C.; Nijsten, T.; Weinstock, M.A.; Dellavalle, R.P.; Naghavi, M.; Fitzmaurice, C. The global burden of melanoma: Results from the Global Burden of Disease Study 2015. Br. J. Dermatol. 2017, 177, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell. Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Zaborowski, M.P.; Balaj, L.; Breakefield, X.O.; Lai, C.P. Extracellular vesicles: Composition, biological relevance, and methods of study. Bioscience 2015, 65, 783–797. [Google Scholar] [CrossRef] [PubMed]
- Qendro, V.; Lundgren, D.H.; Rezaul, K.; Mahony, F.; Ferrell, N.; Bi, A.; Latifi, A.; Chowdhury, D.; Gygi, S.; Haas, W.; et al. Large-scale proteomic characterization of melanoma expressed proteins reveals nestin and vimentin as biomarkers that can potentially distinguish melanoma subtypes. J. Proteome. Res. 2014, 13, 5031–5040. [Google Scholar] [CrossRef]
- Lazar, I.; Clement, E.; Ducoux-Petit, M.; Denat, L.; Soldan, V.; Dauvillier, S.; Balor, S.; Burlet-Schiltz, O.; Larue, L.; Muller, C.; et al. Proteome characterization of melanoma exosomes reveals a specific signature for metastatic cell lines. Pigment Cell Melanoma Res. 2015, 28, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Liao, C.F.; Lin, S.H.; Chen, H.C.; Tai, C.J.; Chang, C.C.; Li, L.T.; Yeh, C.M.; Yeh, K.T.; Chen, Y.C.; Hsu, T.H.; et al. CSE1L, a novel microvesicle membrane protein, mediates Ras-triggered microvesicle generation and metastasis of tumor cells. Mol. Med. 2012, 18, 1269–1280. [Google Scholar] [CrossRef] [PubMed]
- Hatanaka, M.; Higashi, Y.; Fukushige, T.; Baba, N.; Kawai, K.; Hashiguchi, T.; Su, J.; Zeng, W.; Chen, X.; Kanekura, T. Cleaved CD147 shed from the surface of malignant melanoma cells activates MMP2 produced by fibroblasts. Anticancer Res. 2014, 34, 7091–7096. [Google Scholar]
- Clancy, J.W.; Sedgwick, A.; Rosse, C.; Muralidharan-Chari, V.; Raposo, G.; Method, M.; Chavrier, P.; D’Souza-Schorey, C. Regulated delivery of molecular cargo to invasive tumour-derived microvesicles. Nature Commun. 2015, 6, 6919. [Google Scholar] [CrossRef]
- Lima, L.G.; Oliveira, A.S.; Campos, L.C.; Bonamino, M.; Chammas, R.; Werneck, C.; Vicente, C.P.; Barcinski, M.A.; Petersen, L.C.; Monteiro, R.Q. Malignant transformation in melanocytes is associated with increased production of procoagulant microvesicles. Thromb. Haemost. 2011, 106, 712–723. [Google Scholar] [CrossRef]
- Zhao, X.P.; Wang, M.; Song, Y.; Song, K.; Yan, T.L.; Wang, L.; Liu, K.; Shang, Z.J. Membrane microvesicles as mediators for melanoma-fibroblasts communication: Roles of the VCAM-1/VLA-4 axis and the ERK1/2 signal pathway. Cancer Lett. 2015, 360, 125–133. [Google Scholar] [CrossRef]
- Martínez-Lorenzo, M.J.; Anel, A.; Alava, M.A.; Piñeiro, A.; Naval, J.; Lasierra, P.; Larrad, L. The human melanoma cell line MelJuSo secretes bioactive FasL and APO2L/TRAIL on the surface of microvesicles. Possible contribution to tumor counterattack. Exp. Cell. Res. 2004, 295, 315–329. [Google Scholar] [CrossRef]
- Johnson, D.B.; Estrada, M.V.; Salgado, R.; Sanchez, V.; Doxie, D.B.; Opalenik, S.R.; Vilgelm, A.E.; Feld, E.; Johnson, A.S.; Greenplate, A.R. Melanoma-specific MHC-II expression represents a tumour-autonomous phenotype and predicts response to anti-PD-1/PD-L1 therapy. Nature Commun. 2016, 7, 10582. [Google Scholar] [CrossRef] [PubMed]
- Muhsin-Sharafaldine, M.R.; Saunderson, S.C.; Dunn, A.C.; Faed, J.M.; Kleffmann, T.; McLellan, A.D. Procoagulant and immunogenic properties of melanoma exosomes, microvesicles and apoptotic vesicles. Oncotarget. 2016, 7, 56279–56294. [Google Scholar] [CrossRef] [PubMed]
- Hsu, M.Y.; Shih, D.T.; Meier, F.E.; Van Belle, P.; Hsu, J.Y.; Elder, D.E.; Buck, C.A.; Herlyn, M. Adenoviral gene transfer of beta3 integrin subunit induces conversion from radial to vertical growth phase in primary human melanoma. Am. J. Pathol. 1998, 153, 1435–1442. [Google Scholar] [CrossRef]
- Klein, C.E.; Dressel, D.; Steinmayer, T.; Mauch, C.; Eckes, B.; Krieg, T.; Bankert, R.B.; Weber, L. Integrin alpha 2 beta 1 is upregulated in fibroblasts and highly aggressive melanoma cells in three-dimensional collagen lattices and mediates the reorganization of collagen I fibrils. J. Cell Biol. 1991, 115, 1427–1436. [Google Scholar] [CrossRef]
- Melchiori, A.; Mortarini, R.; Carlone, S.; Marchisio, P.C.; Anichini, A.; Noonan, D.M.; Albini, A. The alpha 3 beta 1 integrin is involved in melanoma cell migration and invasion. Exp. Cell Res. 1995, 219, 233–242. [Google Scholar] [CrossRef]
- Hsu, M.Y.; Meier, F.E.; Nesbit, M.; Hsu, J.Y.; Van Belle, P.; Elder, D.E.; Herlyn, M. E-cadherin expression in melanoma cells restores keratinocyte-mediated growth control and down-regulates expression of invasion-related adhesion receptors. Am. J. Pathol. 2000, 156, 1515–1525. [Google Scholar] [CrossRef]
- Yan, S.; Holderness, B.M.; Li, Z.; Seidel, G.D.; Gui, J.; Fisher, J.L.; Ernstoff, M.S. Epithelial-mesenchymal expression phenotype of primary melanoma and matched metastases and relationship with overall survival. Anticancer Res. 2016, 36, 6449–6456. [Google Scholar] [CrossRef]
- Franzen, C.A.; Blackwell, R.H.; Todorovic, V.; Greco, K.A.; Foreman, K.E.; Flanigan, R.C.; Kuo, P.C.; Gupta, G.N. Urothelial cells undergo epithelial-to-mesenchymal transition after exposure to muscle invasive bladder cancer exosomes. Oncogenesis 2015, 4, e163. [Google Scholar] [CrossRef]
- Galindo-Hernandez, O.; Gonzales-Vazquez, C.; Cortes-Reynosa, P.; Reyes-Uribe, E.; Chavez-Ocaña, S.; Reyes-Hernandez, O.; Sierra-Martinez, M.; Salazar, E.P. Extracellular vesicles from women with breast cancer promote an epithelial-mesenchymal transition-like process in mammary epithelial cells MCF10A. Tumour Biol. 2015, 36, 9649–9659. [Google Scholar] [CrossRef]
- Al-Nedawi, K.; Meehan, B.; Kerbel, R.S.; Allison, A.C.; Rak, J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proc. Natl. Acad. Sci. USA 2009, 106, 3794–3799. [Google Scholar] [CrossRef] [PubMed]
- Millimaggi, D.; Mari, M.; D’Ascenzo, S.; Carosa, E.; Jannini, E.A.; Zucker, S.; Carta, G.; Pavan, A.; Dolo, V. Tumor vesicle-associated CD147 modulates the angiogenic capability of endothelial cells. Neoplasia 2007, 9, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhu, X.J.; Zeng, C.; Wu, P.H.; Wang, H.X.; Chen, Z.C.; Li, Q.B. Microvesicles secreted from human multiple myeloma cells promote angiogenesis. Acta Pharmacol. Sin. 2014, 35, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Mezouar, S.; Darbousset, R.; Dignat-George, F.; Panicot-Dubois, L.; Dubois, C. Inhibition of platelet activation prevents the P-selectin and integrin-dependent accumulation of cancer cell microparticles and reduces tumor growth and metastasis in vivo. Int. J. Cancer 2015, 136, 462–475. [Google Scholar] [CrossRef] [PubMed]
- Schroder, W.A.; Major, L.D.; Le, T.T.; Gardner, J.; Sweet, M.J.; Janciauskiene, S.; Suhrbier, A. Tumor cell-expressed SerpinB2 is present on microparticles and inhibits metastasis. Cancer Med. 2014, 3, 500–513. [Google Scholar] [CrossRef] [PubMed]
- Bosserhoff, A.K.; Ellmann, L.; Quast, A.S.; Eberle, J.; Boyle, G.M.; Kuphal, S. Loss of T-cadherin (CDH-13) regulates AKT signaling and desensitizes cells to apoptosis in melanoma. Mol. Carcinog. 2014, 53, 635–647. [Google Scholar] [CrossRef]
- Baj-Krzyworzeka, M.; Weglarczyk, K.; Mytar, B.; Szatanek, R.; Baran, J.; Zembala, M. Tumour-derived microvesicles contain interleukin-8 and modulate production of chemokines by human monocytes. Anticancer Res. 2011, 31, 1329–1335. [Google Scholar]
- Valenti, R.; Huber, V.; Filipazzi, P.; Pilla, L.; Sovena, G.; Villa, A.; Corbelli, A.; Fais, S.; Parmiani, G.; Rivoltini, L. Human tumor-released microvesicles promote the differentiation of myeloid cells with transforming growth factor-beta-mediated suppressive activity on T lymphocytes. Cancer Res. 2006, 66, 9290–9298. [Google Scholar] [CrossRef]
- Jorfi, S.; Ansa-Addo, E.A.; Kholia, S.; Stratton, S.; Valley, S.; Lange, S.; Inal, J. Inhibition of microvesiculation sensitizes prostate cancer cells to chemotherapy and reduces docetaxel dose required to limit tumor growth in vivo. Sci. Rep. 2015, 5, 1–13. [Google Scholar] [CrossRef]
- Walsh, N.; Kennedy, S.; Larkin, A.M.; Tryfonopoulos, D.; Eustace, A.J.; Mahgoub, T.; Conway, C.; Oglesby, I.; Collins, D.; Ballot, J.; et al. Membrane transport proteins in human melanoma: Associations with tumour aggressiveness and metastasis. Br. J. Cancer 2010, 102, 1157–1162. [Google Scholar] [CrossRef]
- Kruijff, S.; Hoekstra, H.J. The current status of S-100B as a biomarker in melanoma. Eur. J. Surg. Oncol. 2012, 38, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Barak, V.; Leibovici, V.; Peretz, T.; Kalichman, I.; Lotem, M.; Merims, S. Assessing response to new treatments and prognosis in melanoma patients, by the biomarker S-100β. Anticancer Res. 2015, 35, 6755–6760. [Google Scholar] [PubMed]
- Jurisic, V.; Radenkovic, S.; Konjevic, G. The Actual Role of LDH as Tumor Marker, Biochemical and Clinical Aspects. Adv. Exp. Med. Biol. 2015, 867, 115–124. [Google Scholar] [PubMed]
- Findeisen, P.; Zapatka, M.; Peccerella, T.; Matzk, H.; Neumaier, M.; Schadendorf, D.; Ugurel, S. Serum amyloid A as a prognostic marker in melanoma identified by proteomic profiling. J. Clin. Oncol. 2009, 27, 2199–2208. [Google Scholar] [CrossRef]
- Suzuki, A.; Iizuka, A.; Komiyama, M.; Takikawa, M.; Kume, A.; Tai, S.; Ohshita, C.; Kurusu, A.; Nakamura, Y.; Yamamoto, A.; et al. Identification of melanoma antigens using a Serological Proteome Approach (SERPA). Cancer Genomics Proteomics 2010, 7, 17–23. [Google Scholar]
- Caputo, E.; Maiorana, L.; Vasta, V.; Pezzino, F.M.; Sunkara, S.; Wynne, K.; Elia, G.; Marincola, F.M.; McCubrey, J.A.; Libra, M.; et al. Characterization of human melanoma cell lines and melanocytes by proteome analysis. Cell Cycle 2011, 10, 2924–2936. [Google Scholar] [CrossRef]
- Liu, H.; Ong, S.E.; Badu-Nkansah, K.; Schindler, J.; White, F.M.; Hynes, R.O. CUB-domaincontaining protein 1 (CDCP1) activates Src to promote melanoma metastasis. Proc. Natl. Acad. Sci. USA 2011, 108, 1379–1384. [Google Scholar] [CrossRef]
- Al-Ghoul, M.; Brück, T.B.; Lauer-Fields, J.L.; Asirvatham, V.S.; Zapata, C.; Kerr, R.G.; Fields, G.B. Comparative proteomic analysis of matched primary and metastatic melanoma cell lines. J. Proteome Res. 2008, 7, 4107–4118. [Google Scholar] [CrossRef]
- Rondepierre, F.; Bouchon, B.; Papon, J.; Bonnet-Duquennoy, M.; Kintossou, R.; Moins, N.; Maublant, J.; Madelmont, J.C.; D’Incan, M.; Degoul, F. Proteomic studies of B16 lines: Involvement of annexin A1 in melanoma dissemination. Biochim. Biophys. Acta 2009, 1794, 61–69. [Google Scholar] [CrossRef]
- Bernard, K.; Litman, E.; Fitzpatrick, J.L.; Shellman, Y.G.; Argast, G.; Polvinen, K.; Everett, A.D.; Fukasawa, K.; Norris, D.A.; Ahn, N.G.; et al. Functional proteomic analysis of melanoma progression. Cancer Res. 2003, 63, 6716–6725. [Google Scholar]
- Alegre, E.; Zubiri, L.; Perez-Gracia, J.L.; González-Cao, M.; Soria, L.; Martín-Algarra, S.; González, A. Circulating melanoma exosomes as diagnostic and prognosis biomarkers. Clin. Chim. Acta 2016, 454, 28–32. [Google Scholar] [CrossRef] [PubMed]
- Crescitelli, R.; Lässer, C.; Jang, S.C.; Cvjetkovic, A.; Malmhäll, C.; Karimi, N.; Höög, J.L.; Johansson, I.; Fuchs, J.; Thorsell, A.; et al. Subpopulations of extracellular vesicles from human metastatic melanoma tissue identified by quantitative proteomics after optimized isolation. J Extracell Vesicles. 2020, 9, 1722433. [Google Scholar] [PubMed]
- Jang, S.C.; Crescitelli, R.; Cvjetkovic, A.; Belgrano, V.; Olofsson Bagge, R.; Sundfeldt, K.; Ochiya, T.; Kalluri, R.; Lötvall, J. Mitochondrial protein enriched extracellular vesicles discovered in human melanoma tissues can be detected in patient plasma. J Extracell Vesicles. 2019, 8, 1635420. [Google Scholar] [CrossRef] [PubMed]
- Herlyn, M.; Balaban, G.; Bennicelli, J.; Guerry, D., IV; Halaban, R.; Herlyn, D.; Elder, D.E.; Maul, G.G.; Steplewski, Z.; Nowell, P.C. Primary melanoma cells of the vertical growth phase: Similarities to metastatic cells. J. Natl. Cancer Inst. 1985, 74, 283–289. [Google Scholar]
- Cornil, I.; Theodorescu, D.; Man, S.; Herlyn, M.; Jambrosic, J.; Kerbel, R.S. Fibroblast cell interactions with human melanoma cells affect tumor cell growth as a function of tumor progression. Proc. Natl. Acad. Sci. USA 1991, 88, 6028–6032. [Google Scholar] [CrossRef]
- Juhasz, I.; Albelda, S.M.; Elder, D.E.; Murphy, G.F.; Adachi, K.; Herlyn, D.; Valyi-Nagy, I.T.; Herlyn, M. Growth and invasion of human melanomas in human skin grafted to immunodeficient mice. Am. J. Pathol. 1993, 143, 528–537. [Google Scholar]
- Surman, M.; Hoja-Łukowicz, D.; Szwed, S.; Drożdż, A.; Stępień, E.; Przybyło, M. Human melanoma-derived ectosomes are enriched with specific glycan epitopes. Life Sci. 2018, 207, 395–411. [Google Scholar] [CrossRef]
- Surman, M.; Hoja-Łukowicz, D.; Szwed, S.; Kędracka-Krok, S.; Jankowska, U.; Kurtyka, M.; Drożdż, A.; Lityńska, A.; Stępień, E.; Przybyło, M. An insight into the proteome of uveal melanoma-derived ectosomes reveals the presence of potentially useful biomarkers. Int. J. Mol. Sci. 2019, 20, 3789. [Google Scholar] [CrossRef]
- Hughes, C.S.; Foehr, S.; Garfield, D.A.; Furlong, E.E.; Steinmetz, L.M.; Krijgsveld, J. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 2014, 10, 757. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Surman, M.; Kędracka-Krok, S.; Hoja-Łukowicz, D.; Jankowska, U.; Drożdż, A.; Stępień, E.Ł.; Przybyło, M. Mass Spectrometry-Based Proteomic Characterization of Cutaneous Melanoma Ectosomes Reveals the Presence of Cancer-Related Molecules. Int. J. Mol. Sci. 2020, 21, 2934. https://doi.org/10.3390/ijms21082934
Surman M, Kędracka-Krok S, Hoja-Łukowicz D, Jankowska U, Drożdż A, Stępień EŁ, Przybyło M. Mass Spectrometry-Based Proteomic Characterization of Cutaneous Melanoma Ectosomes Reveals the Presence of Cancer-Related Molecules. International Journal of Molecular Sciences. 2020; 21(8):2934. https://doi.org/10.3390/ijms21082934
Chicago/Turabian StyleSurman, Magdalena, Sylwia Kędracka-Krok, Dorota Hoja-Łukowicz, Urszula Jankowska, Anna Drożdż, Ewa Ł. Stępień, and Małgorzata Przybyło. 2020. "Mass Spectrometry-Based Proteomic Characterization of Cutaneous Melanoma Ectosomes Reveals the Presence of Cancer-Related Molecules" International Journal of Molecular Sciences 21, no. 8: 2934. https://doi.org/10.3390/ijms21082934
APA StyleSurman, M., Kędracka-Krok, S., Hoja-Łukowicz, D., Jankowska, U., Drożdż, A., Stępień, E. Ł., & Przybyło, M. (2020). Mass Spectrometry-Based Proteomic Characterization of Cutaneous Melanoma Ectosomes Reveals the Presence of Cancer-Related Molecules. International Journal of Molecular Sciences, 21(8), 2934. https://doi.org/10.3390/ijms21082934





