Identification of EIL and ERF Genes Related to Fruit Ripening in Peach
Abstract
1. Introduction
2. Results
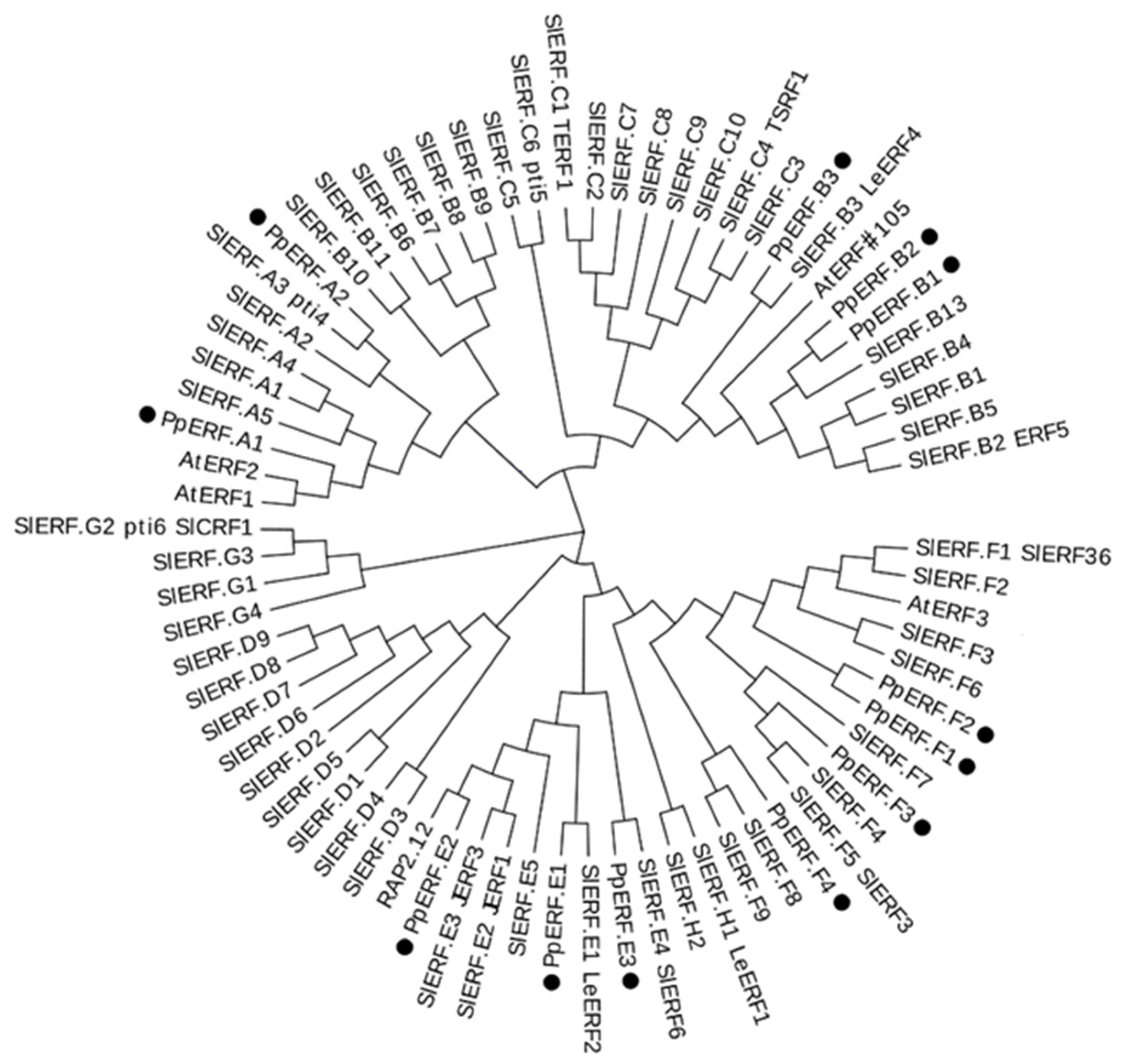
2.1. Identification of EIL and ERF Genes That Are Expressed in Peach Fruit
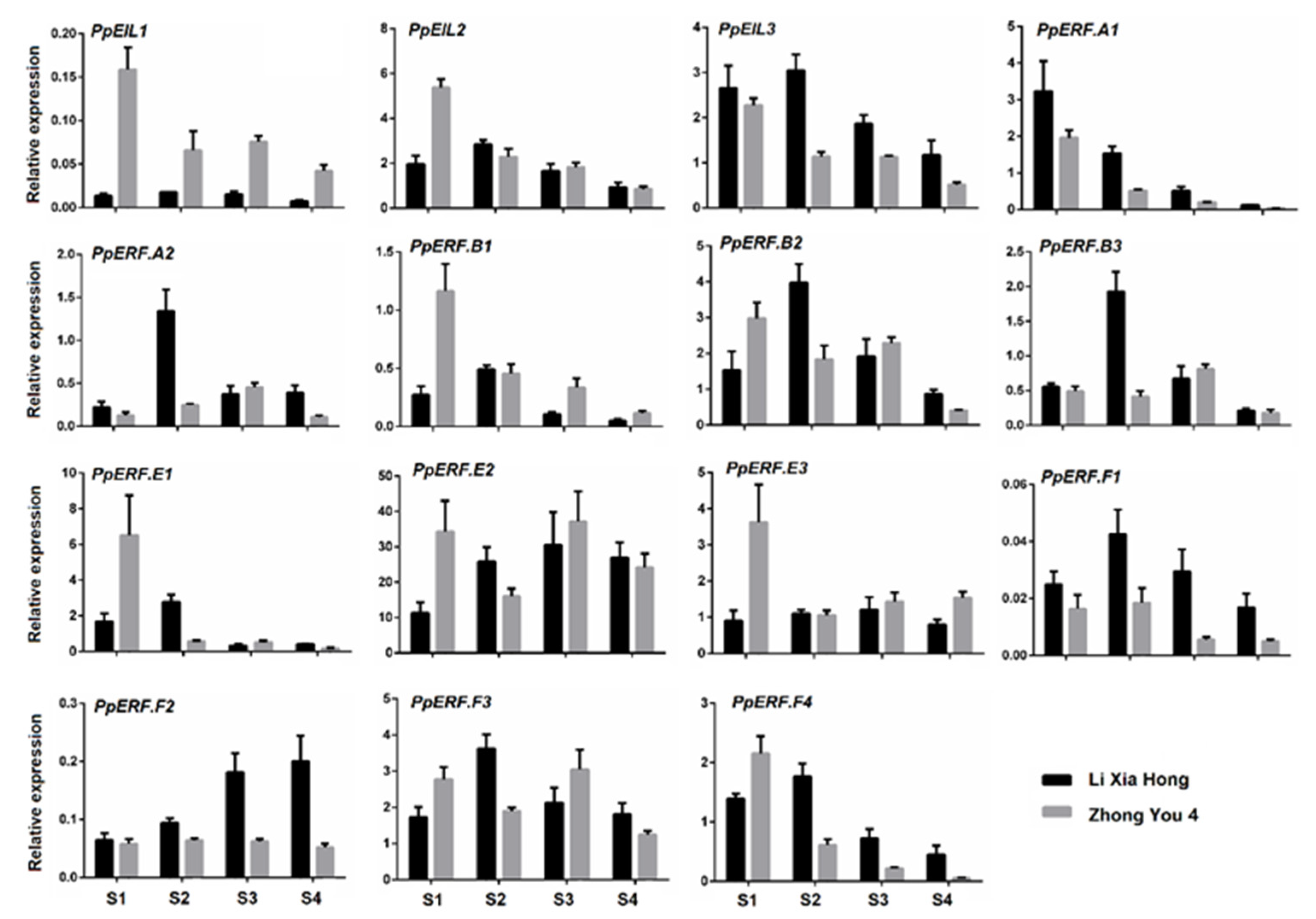
2.2. Expression Profile of EIL and ERF Genes in Peach Fruit
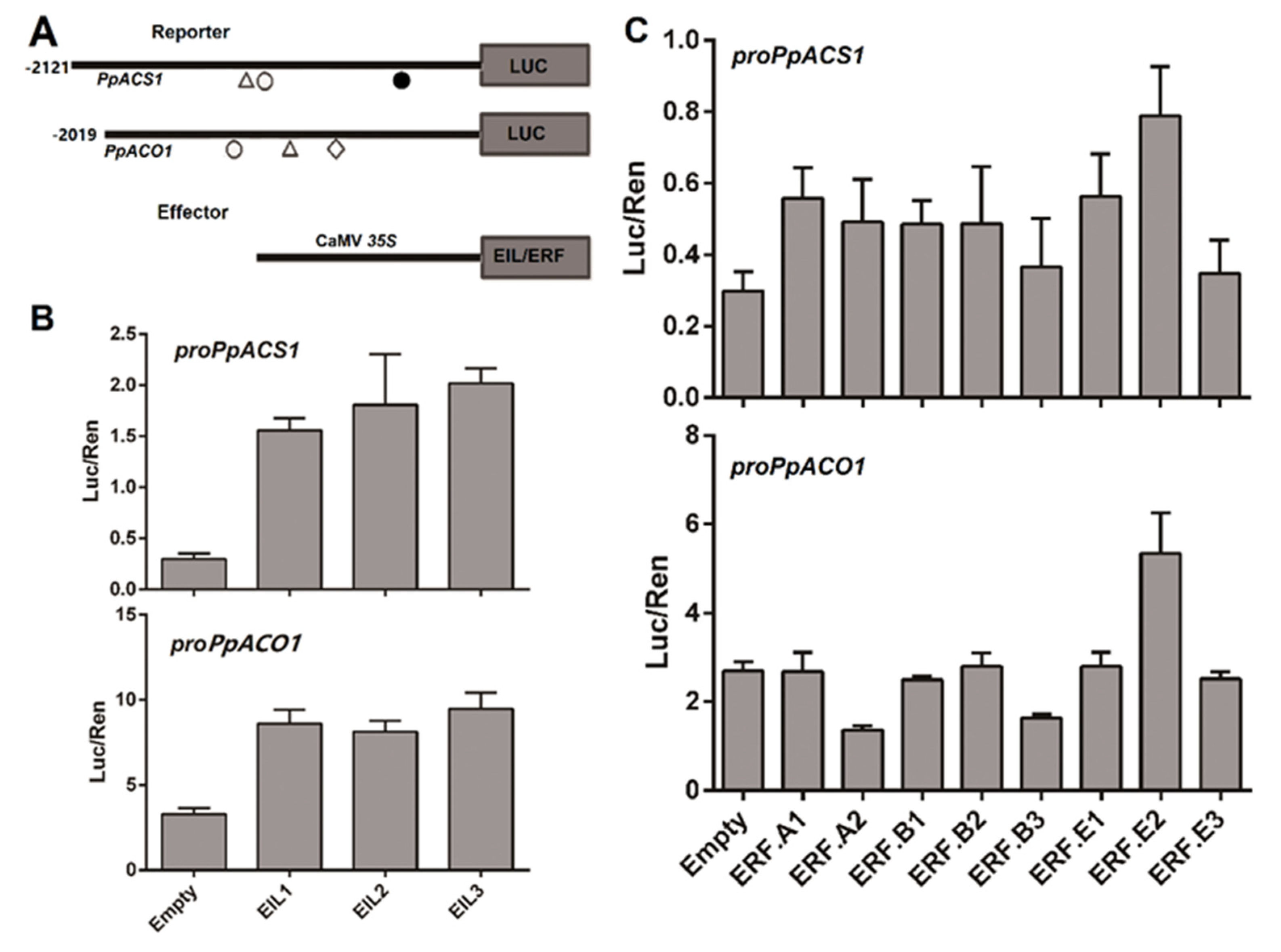
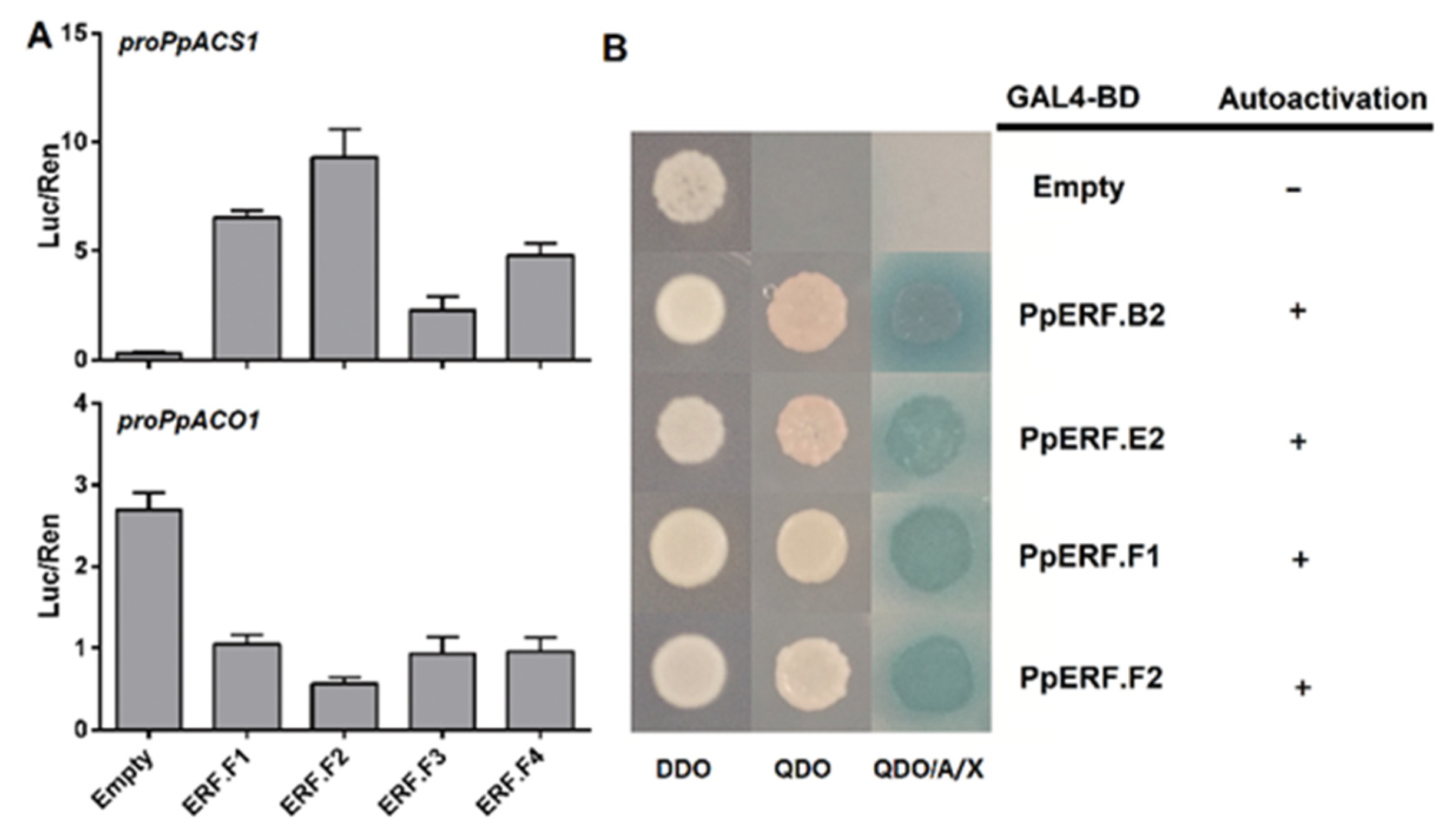
2.3. Roles of PpEILs and PpERFs in Transcriptional Regulation of Ethylene Biosynthesis Genes
2.4. Analysis of Transcriptional Activation Ability of PpERF Proteins in Yeast
2.5. Identification of Cis-Elements in the PpACS1 Promoter That Interact with EIL and ERF TFs
3. Discussion
4. Methods
4.1. Plant Material
4.2. Heatmap Analysis of the ERF Gene Family
4.3. RNA Extraction and Quantitative Real-Time PCR (qRT-PCR)
4.4. Dual Luciferase Reporter Assay
4.5. Measurement of Transcriptional Activation Activity of ERFs in Yeast
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Giovannoni, J. Molecular biology of fruit maturation and ripening. Annu. Rev. Plant Biol. 2001, 52, 725–749. [Google Scholar] [CrossRef] [PubMed]
- Giovannoni, J.J. Fruit ripening mutants yield insights into ripening control. Curr. Opin. Plant Biol. 2007, 10, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Seymour, G.B.; Østergaard, L.; Chapman, N.; Knapp, S.; Martin, C. Fruit Development and Ripening. Annu. Rev. Plant Biol. 2013, 64, 219–241. [Google Scholar] [CrossRef] [PubMed]
- McMurchie, E.J.; McGlasson, W.B.; Eaks, I.L. Treatment of fruit with propylene gives information about the biogenesis of ethylene. Nature 1972, 237, 235–236. [Google Scholar] [CrossRef]
- Wang, K.L.-C.; Li, H.; Ecker, J.R. Ethylene Biosynthesis and Signaling Networks. Plant Cell 2002, 14, s131–s151. [Google Scholar] [CrossRef]
- Barry, C. The Regulation of 1-Aminocyclopropane-1-Carboxylic Acid Synthase Gene Expression during the Transition from System-1 to System-2 Ethylene Synthesis in Tomato. Plant Physiol. 2000, 123, 979–986. [Google Scholar] [CrossRef]
- Yokotani, N.; Nakano, R.; Imanishi, S.; Nagata, M.; Inaba, A.; Kubo, Y. Ripening-associated ethylene biosynthesis in tomato fruit is autocatalytically and developmentally regulated. J. Exp. Bot. 2009, 60, 3433–3442. [Google Scholar] [CrossRef]
- Barry, C.; Blume, B.; Hamilton, A.J.; Bouzayen, M.; Cooper, W.; Grierson, D. Differential expression of the 1-aminocyclopropane-1-carboxylate oxidase gene family of tomato. Plant J. 1996, 9, 525–535. [Google Scholar] [CrossRef]
- Nakatsuka, A.; Murachi, S.; Okunishi, H.; Shiomi, S.; Nakano, R.; Kubo, Y.; Inaba, A. Differential expression and internal feedback regulation of 1-aminocyclopropane-1-carboxylate synthase, 1-aminocyclopropane-1-carboxylate oxidase, and ethylene receptor genes in tomato fruit during development and ripening. Plant Physiol. 1998, 118, 1295–1305. [Google Scholar] [CrossRef]
- Chao, Q.; Rothenberg, M.; Solano, R.; Roman, G.; Terzaghi, W.; Ecker, J.R. Activation of the ethylene gas response pathway in Arabidopsis by the nuclear protein ETHYLENE-INSENSITIVE3 and related proteins. Cell 1997, 89, 1133–1144. [Google Scholar] [CrossRef]
- Solano, R.; Stepanova, A.N.; Chao, Q.; Ecker, J.R. Nuclear events in ethylene signaling: A transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genome Res. 1998, 12, 3703–3714. [Google Scholar] [CrossRef] [PubMed]
- Mao, C.; Wang, S.; Jia, Q.; Wu, P. OsEIL1, a Rice Homolog of the Arabidopsis EIN3 Regulates the Ethylene Response as a Positive Component. Plant Mol. Biol. 2006, 61, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, A.; Zhang, Z.; Huang, Z.; Lu, P.; Zhang, D.; Liu, X.; Zhang, Z.-F.; Huang, R. Ethylene Response Factor TERF1, Regulated by ETHYLENE-INSENSITIVE3-like Factors, Functions in Reactive Oxygen Species (ROS) Scavenging in Tobacco (Nicotiana tabacum L.). Sci. Rep. 2016, 6, 29948. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Tang, L.; Yang, Y.; Zouine, M.; Bouzayen, M. A conserved phosphorylation site regulates the transcriptional function of ETHYLENE-INSENSITIVE3-like1 in tomato. J. Exp. Bot. 2011, 63, 427–439. [Google Scholar] [CrossRef][Green Version]
- Quan, R.; Wang, J.; Yang, D.; Zhang, H.; Zhang, Z.; Huang, R. EIN3 and SOS2 synergistically modulate plant salt tolerance. Sci. Rep. 2017, 7, 44637. [Google Scholar] [CrossRef]
- Tieman, D.M.; Ciardi, J.A.; Taylor, M.G.; Klee, H.J. Members of the tomato LeEIL (EIN3-like) gene family are functionally redundant and regulate ethylene responses throughout plant development. Plant J. 2001, 26, 47–58. [Google Scholar] [CrossRef]
- Yokotani, N.; Tamura, S.; Nakano, R.; Inaba, A.; Kubo, Y. Characterization of a novel tomato EIN3-like gene (LeEIL4). J. Exp. Bot. 2003, 54, 2775–2776. [Google Scholar] [CrossRef]
- Yin, X.-R.; Allan, A.C.; Chen, K.-S.; Ferguson, I.B. Kiwifruit EIL and ERF Genes Involved in Regulating Fruit Ripening1[W]. Plant Physiol. 2010, 153, 1280–1292. [Google Scholar] [CrossRef]
- Tacken, E.; Ireland, H.; Wang, Y.-Y.; Putterill, J.; Schaffer, R. Apple EIN3 BINDING F-box 1 inhibits the activity of three apple EIN3-like transcription factors. AoB Plants 2012, 2012, 034. [Google Scholar] [CrossRef]
- Huang, S.; Sawaki, T.; Takahashi, A.; Mizuno, S.; Takezawa, K.; Matsumura, A.; Yokotsuka, M.; Hirasawa, Y.; Sonoda, M.; Nakagawa, H.; et al. Melon EIN3-like transcription factors (CmEIL1 and CmEIL2) are positive regulators of an ethylene- and ripening-induced 1-aminocyclopropane-1-carboxylic acid oxidase gene (CM-ACO1). Plant Sci. 2010, 178, 251–257. [Google Scholar] [CrossRef]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jin, Q.; Lin, Y.; Cai, Y. Genome-wide analysis suggests high level of microsynteny and purifying selection affect the evolution of EIN3/EIL family in Rosaceae. PeerJ 2017, 5, e3400. [Google Scholar] [CrossRef] [PubMed]
- Nakano, T.; Suzuki, K.; Fujimura, T.; Shinshi, H. Genome-Wide Analysis of the ERF Gene Family in Arabidopsis and Rice. Plant Physiol. 2006, 140, 411–432. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Gomes, B.L.; Mila, I.; Purgatto, E.; Peres, L.E.; Frasse, P.; Maza, E.; Zouine, M.; Roustan, J.-P.; Bouzayen, M.; et al. Comprehensive Profiling of Ethylene Response Factor Expression Identifies Ripening-Associated ERF Genes and Their Link to Key Regulators of Fruit Ripening in Tomato. Plant Physiol. 2016, 170, 1732–1744. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.-C.; Ma, B.; Collinge, D.P.; Pogson, B.J.; He, S.-J.; Xiong, Q.; Duan, K.-X.; Chen, H.; Yang, C.; Lu, X.; et al. Ethylene responses in rice roots and coleoptiles are differentially regulated by a carotenoid isomerase-mediated abscisic acid pathway. Plant Cell 2015, 27, 1061–1081. [Google Scholar] [CrossRef] [PubMed]
- Harkey, A.; Watkins, J.; Olex, A.L.; DiNapoli, K.; Lewis, D.R.; Fetrow, J.; Binder, B.; Muday, G.K. Identification of Transcriptional and Receptor Networks That Control Root Responses to Ethylene. Plant Physiol. 2017, 176, 2095–2118. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Joung, J.-G.; McQuinn, R.P.; Chung, M.-Y.; Fei, Z.; Tieman, D.; Klee, H.; Giovannoni, J.J. Combined transcriptome, genetic diversity and metabolite profiling in tomato fruit reveals that the ethylene response factor SlERF6 plays an important role in ripening and carotenoid accumulation. Plant J. 2012, 70, 191–204. [Google Scholar] [CrossRef]
- Yin, X.-R.; Xie, X.; Xia, X.; Yu, J.; Ferguson, I.B.; Giovannoni, J.J.; Chen, K.-S. Involvement of an ethylene response factor in chlorophyll degradation during citrus fruit degreening. Plant J. 2016, 86, 403–412. [Google Scholar] [CrossRef]
- Li, X.; Xu, Y.; Shen, S.; Yin, X.; Klee, H.; Zhang, B.; Chen, K. Transcription factor CitERF71 activates the terpene synthase gene CitTPS16 involved in the synthesis of E-geraniol in sweet orange fruit. J. Exp. Bot. 2017, 68, 4929–4938. [Google Scholar] [CrossRef]
- Robles, L.; Stepanova, A.N.; Alonso, J. Molecular Mechanisms of Ethylene–Auxin Interaction. Mol. Plant 2013, 6, 1734–1737. [Google Scholar] [CrossRef]
- Tadiello, A.; Longhi, S.; Moretto, M.; Ferrarini, A.; Tononi, P.; Farneti, B.; Busatto, N.; Vrhovsek, U.; Molin, A.D.; Avanzato, C.; et al. Interference with ethylene perception at receptor level sheds light on auxin and transcriptional circuits associated with the climacteric ripening of apple fruit (Malus x domestica Borkh.). Plant J. 2016, 88, 963–975. [Google Scholar] [CrossRef]
- Thagun, C.; Imanishi, S.; Kudo, T.; Nakabayashi, R.; Ohyama, K.; Mori, T.; Kawamoto, K.; Nakamura, Y.; Katayama, M.; Nonaka, S.; et al. Jasmonate-responsive erf transcription factors regulate steroidal glycoalkaloid biosynthesis in tomato. Plant Cell Physiol. 2016, 57, 961–975. [Google Scholar] [CrossRef] [PubMed]
- Bakshi, A.; Piya, S.; Fernandez, J.C.; Chervin, C.; Hewezi, T.; Binder, B. Ethylene Receptors Signal via a Noncanonical Pathway to Regulate Abscisic Acid Responses. Plant Physiol. 2017, 176, 910–929. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Huang, Z.; Chen, Q.; Zhang, Z.; Zhang, H.; Wu, Y.; Huang, D.; Huang, R. Ectopic overexpression of tomato JERF3 in tobacco activates downstream gene expression and enhances salt tolerance. Plant Mol. Biol. 2004, 55, 183–192. [Google Scholar] [CrossRef] [PubMed]
- McGrath, K.; Dombrecht, B.; Manners, J.M.; Schenk, P.; Edgar, C.I.; MacLean, D.J.; Scheible, W.-R.; Udvardi, M.K.; Kazan, K. Repressor- and Activator-Type Ethylene Response Factors Functioning in Jasmonate Signaling and Disease Resistance Identified via a Genome-Wide Screen of Arabidopsis Transcription Factor Gene Expression[w]. Plant Physiol. 2005, 139, 949–959. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, J.; Quan, R.; Pan, X.; Wan, L.; Huang, R. EAR motif mutation of rice OsERF3 alters the regulation of ethylene biosynthesis and drought tolerance. Planta 2013, 237, 1443–1451. [Google Scholar] [CrossRef]
- Liu, M.; Diretto, G.; Pirrello, J.; Roustan, J.-P.; Li, Z.; Giuliano, G.; Regad, F.; Bouzayen, M. The chimeric repressor version of an Ethylene Response Factor ( ERF ) family member, Sl-ERF.B3, shows contrasting effects on tomato fruit ripening. New Phytol. 2014, 203, 206–218. [Google Scholar] [CrossRef]
- Xie, X.-L.; Yin, X.-R.; Chen, K.-S. Roles of APETALA2/Ethylene-Response Factors in Regulation of Fruit Quality. Crit. Rev. Plant Sci. 2016, 35, 120–130. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, H.; Quan, R.; Wang, X.-C.; Huang, R. Transcriptional Regulation of the Ethylene Response Factor LeERF2 in the Expression of Ethylene Biosynthesis Genes Controls Ethylene Production in Tomato and Tobacco1[W][OA]. Plant Physiol. 2009, 150, 365–377. [Google Scholar] [CrossRef]
- Li, T.; Jiang, Z.; Zhang, L.; Tan, D.; Wei, Y.; Yuan, H.; Li, T.; Wang, A. Apple (Malus domestica) MdERF2 negatively affects ethylene biosynthesis during fruit ripening by suppressing MdACS1 transcription. Plant J. 2016, 88, 735–748. [Google Scholar] [CrossRef]
- 4Hao, P.-P.; Wang, G.-M.; Cheng, H.-Y.; Ke, Y.-Q.; Qi, K.-J.; Zhang, S.; Zhang, S. Transcriptome analysis unravels an ethylene response factor involved in regulating fruit ripening in pear. Physiol. Plant 2018, 163, 124–135. [Google Scholar] [CrossRef]
- Liu, L.; White, M.J.; Macrae, T.H. Transcription factors and their genes in higher plants functional domains, evolution and regulation. J. Biol. Inorg. Chem. 1999, 262, 247–257. [Google Scholar] [CrossRef] [PubMed]
- Ohta, M.; Matsui, K.; Hiratsu, K.; Shinshi, H.; Ohme-Takagi, M. Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 2001, 13, 1959–1968. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Hu, Z.; Grierson, D. Differential regulation of tomato ethylene responsive factor LeERF3b, a putative repressor, and the activator Pti4 in ripening mutants and in response to environmental stresses. J. Plant Physiol. 2008, 165, 662–670. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Barbero, G.F.; Geerinck, J.; Tilleman, S.; Grunewald, W.; Perez, A.C.; Chico, J.M.; Bossche, R.V.; Sewell, J.; Gil, E. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 2010, 464, 788–791. [Google Scholar] [CrossRef]
- Kagale, S.; Rozwadowski, K. EAR motif-mediated transcriptional repression in plants: An underlying mechanism for epigenetic regulation of gene expression. Epigenetics 2011, 6, 141–146. [Google Scholar] [CrossRef]
- Mathooko, F.M.; Tsunashima, Y.; Owino, W.Z.; Kubo, Y.; Inaba, A. Regulation of genes encoding ethylene biosynthetic enzymes in peach (Prunus persica L.) fruit by carbon dioxide and 1-methylcyclopropene. Postharvest. Biol. Techol. 2001, 21, 265–281. [Google Scholar] [CrossRef]
- Tatsuki, M.; Haji, T.; Yamaguchi, M. The involvement of 1-aminocyclopropane-1-carboxylic acid synthase isogene, Pp-ACS1, in peach fruit softening. J. Exp. Bot. 2006, 57, 1281–1289. [Google Scholar] [CrossRef]
- Prinsi, B.; Negri, A.S.; Fedeli, C.; Morgutti, S.; Negrini, N.; Cocucci, M.; Espen, L. Peach fruit ripening: A proteomic comparative analysis of the mesocarp of two cultivars with different flesh firmness at two ripening stages. Phytochemistry 2011, 72, 1251–1262. [Google Scholar] [CrossRef]
- Gong, D.; Cao, S.; Sheng, T.; Shao, J.; Song, C.; Wo, F.; Chen, W.; Yang, Z. Effect of blue light on ethylene biosynthesis, signalling and fruit ripening in postharvest peaches. Sci. Hortic. 2015, 197, 657–664. [Google Scholar] [CrossRef]
- Zhang, C.H.; Shangguan, L.F.; Ma, R.J.; Sun, X.; Tao, R.; Guo, L.; Korir, N.K.; Yu, M.L. Genome-wide analysis of the AP2/ERF superfamily in peach (Prunus persica). Genet. Mol. Res. 2012, 11, 4789–4809. [Google Scholar] [CrossRef]
- Wang, X.; Ding, Y.; Wang, Y.; Pan, L.; Niu, L.; Lu, Z.; Cui, G.; Zeng, W.; Wang, Z. Genes involved in ethylene signal transduction in peach ( Prunus persica ) and their expression profiles during fruit maturation. Sci. Hortic. 2017, 224, 306–316. [Google Scholar] [CrossRef]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhao, S.; Gu, C.; Zhou, Y.; Zhou, H.; Ma, J.; Cheng, J.; Han, Y. Deep RNA-Seq uncovers the peach transcriptome landscape. Plant Mol. Biol. 2013, 83, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Busatto, N.; Rahim, A.; Trainotti, L. Peach ripening transcriptomics unveils new and unexpected targets for the improvement of drupe quality. Plant Genom. Plant Biotechnol. 2013, 165–182. [Google Scholar] [CrossRef]
- Lü, P.; Yu, S.; Zhu, N.; Chen, Y.-R.; Zhou, B.; Pan, Y.; Tzeng, D.; Fabi, J.P.; Argyris, J.; Garcia-Mas, J.; et al. Genome encode analyses reveal the basis of convergent evolution of fleshy fruit ripening. Nat. Plants 2018, 4, 784–791. [Google Scholar]
- Martel, C.; Vrebalov, J.; Tafelmeyer, P.; Giovannoni, J.J. The tomato MADS-box transcription factor RIPENING INHIBITOR interacts with promoters involved in numerous ripening processes in a COLORLESS NONRIPENING-dependent manner. Plant Physiol. 2011, 157, 1568–1579. [Google Scholar] [CrossRef] [PubMed]
- Yanagisawa, S.; Yoo, S.-D.; Sheen, J. Differential regulation of EIN3 stability by glucose and ethylene signalling in plants. Nature 2003, 425, 521–525. [Google Scholar] [CrossRef]
- Potuschak, T.; Lechner, E.; Parmentier, Y.; Yanagisawa, S.; Grava, S.; Koncz, C.; Genschik, P. EIN3-Dependent Regulation of Plant Ethylene Hormone Signaling by Two Arabidopsis F Box Proteins. Cell 2003, 115, 679–689. [Google Scholar] [CrossRef]
- Guo, H.; Ecker, J.R. Plant responses to ethylene gas are mediated by SCFEBF1/EBF2-dependent proteolysis of EIN3 transcription factor. Cell 2003, 115, 667–677. [Google Scholar] [CrossRef]
- Liu, M.; Pirrello, J.; Kesari, R.; Mila, I.; Roustan, J.-P.; Li, Z.; Latché, A.; Pech, J.; Bouzayen, M.; Regad, F. A dominant repressor version of the tomato Sl-ERF. B3 gene confers ethylene hypersensitivity via feedback regulation of ethylene signaling and response components. Plant J. 2013, 76, 406–419. [Google Scholar] [CrossRef]
- Pirrello, J.; Jaimes-Miranda, F.; Sanchez-Ballesta, M.T.; Tournier, B.; Khalil-Ahmad, Q.; Regad, F.; Latché, A.; Pech, J.; Bouzayen, M. Sl-ERF2, a Tomato Ethylene Response Factor Involved in Ethylene Response and Seed Germination. Plant Cell Physiol. 2006, 47, 1195–1205. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.-C.; Kuang, J.-F.; Chen, J.-Y.; Liu, X.; Xiao, Y.-Y.; Fu, C.-C.; Wang, J.-N.; Wu, K.-Q.; Lu, W.-J. Banana Transcription Factor MaERF11 Recruits Histone Deacetylase MaHDA1 and Represses the Expression of MaACO1 and Expansins during Fruit Ripening. Plant Physiol. 2016, 171, 1070–1084. [Google Scholar] [CrossRef] [PubMed]
- Howe, E.A.; Sinha, R.; Schlauch, D.; Quackenbush, J. RNA-Seq analysis in MeV. Bioinformatics 2011, 27, 3209–3210. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Gao, Z.; Wang, F.; Jun, Z.; Zhang, Z. Selection of reliable reference genes for gene expression studies in peach using real-time PCR. BMC Mol. Biol. 2009, 10, 71. [Google Scholar] [CrossRef] [PubMed]
- Hellens, R.; Edwards, E.A.; Leyland, N.; Bean, S.; Mullineaux, P. pGreen: A versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol. Biol. 2000, 42, 819–832. [Google Scholar] [CrossRef]
- Espley, R.; Hellens, R.P.; Putterill, J.; Stevenson, D.E.; Kutty-Amma, S.; Allan, A.C. Red colouration in apple fruit is due to the activity of the MYB transcription factor, MdMYB10. Plant J. 2007, 49, 414–427. [Google Scholar] [CrossRef]



| Gene | Accession No. | Flower | Leaf | Fruit | |||||
|---|---|---|---|---|---|---|---|---|---|
| WF | RF | GL | RL | YS3 | RS3 | YS4 | RS4 | ||
| PpEIL1 | Prupe.6G018200 | 13.2 | 13.0 | 13.5 | 15.6 | 12.5 | 15.6 | 15.3 | 21.6 |
| PpEIL2 | Prupe.2G058400 | 78.8 | 77.7 | 130.8 | 146.3 | 198.6 | 171.7 | 167.9 | 225.2 |
| PpEIL3 | Prupe.2G058500 | 42.9 | 35.8 | 68.9 | 68.9 | 61.5 | 44.0 | 46.8 | 65.5 |
| PpEIL4 | Prupe.6G181600 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| PpEIL5 | Prupe.2G070300 | 14.8 | 13.4 | 19.2 | 23.1 | 22.2 | 22.6 | 8.1 | 30.6 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhou, H.; Zhao, L.; Yang, Q.; Amar, M.H.; Ogutu, C.; Peng, Q.; Liao, L.; Zhang, J.; Han, Y. Identification of EIL and ERF Genes Related to Fruit Ripening in Peach. Int. J. Mol. Sci. 2020, 21, 2846. https://doi.org/10.3390/ijms21082846
Zhou H, Zhao L, Yang Q, Amar MH, Ogutu C, Peng Q, Liao L, Zhang J, Han Y. Identification of EIL and ERF Genes Related to Fruit Ripening in Peach. International Journal of Molecular Sciences. 2020; 21(8):2846. https://doi.org/10.3390/ijms21082846
Chicago/Turabian StyleZhou, Hui, Lei Zhao, Qiurui Yang, Mohamed Hamdy Amar, Collins Ogutu, Qian Peng, Liao Liao, Jinyun Zhang, and Yuepeng Han. 2020. "Identification of EIL and ERF Genes Related to Fruit Ripening in Peach" International Journal of Molecular Sciences 21, no. 8: 2846. https://doi.org/10.3390/ijms21082846
APA StyleZhou, H., Zhao, L., Yang, Q., Amar, M. H., Ogutu, C., Peng, Q., Liao, L., Zhang, J., & Han, Y. (2020). Identification of EIL and ERF Genes Related to Fruit Ripening in Peach. International Journal of Molecular Sciences, 21(8), 2846. https://doi.org/10.3390/ijms21082846







