Identification and Characterization of the First Virulent Phages, Including a Novel Jumbo Virus, Infecting Ochrobactrum spp.
Abstract
1. Introduction
2. Results and Discussion
2.1. Identification and Characterization of the Plaque and Virion Morphologies of Ochrobactrum-Specific Phages
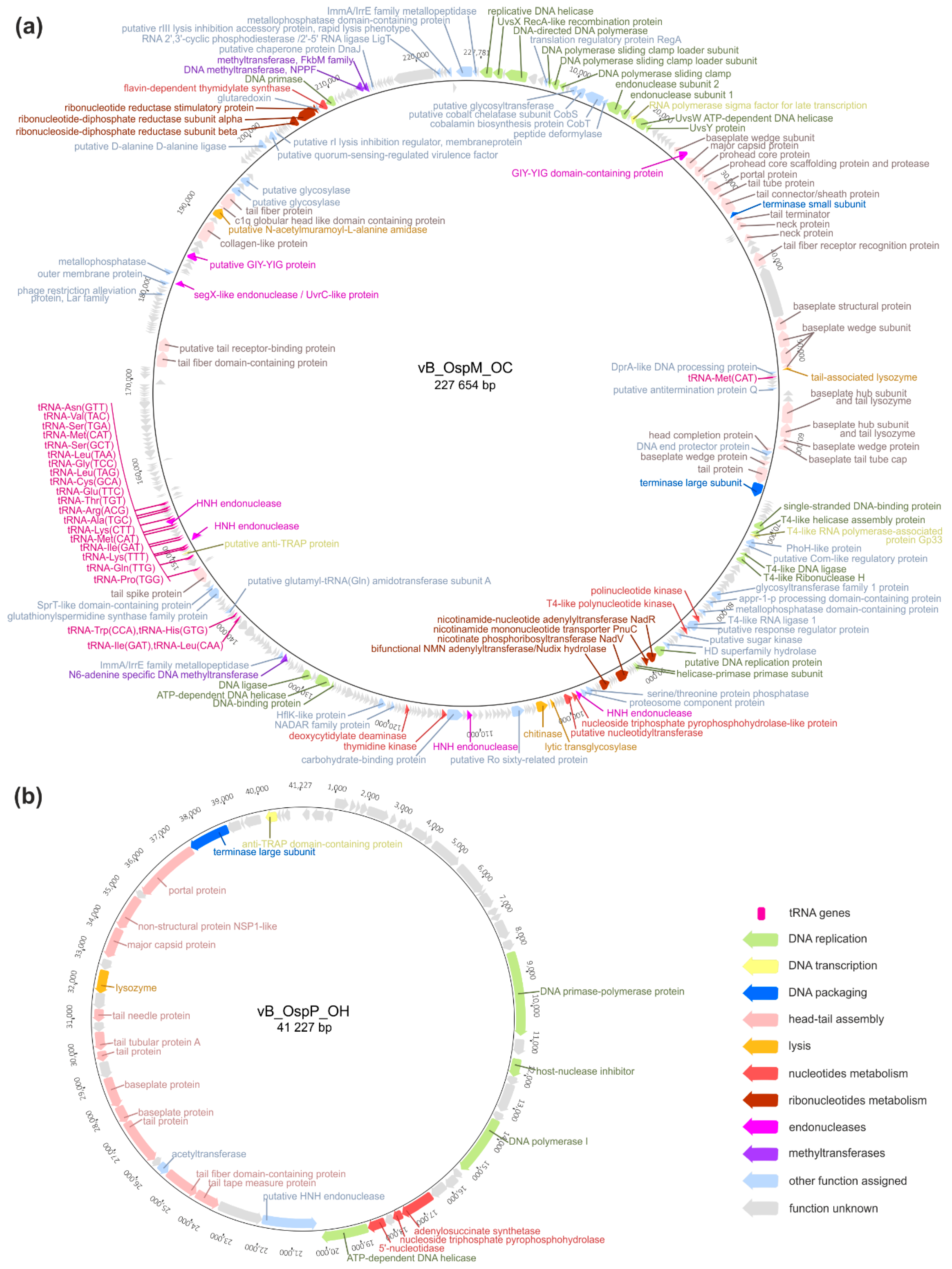
2.2. Genomic Analysis of the vB_OspM_OC and vB_OspP_OH Phages
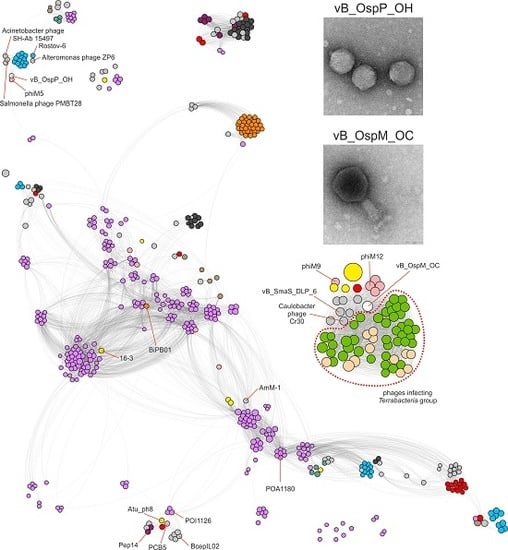
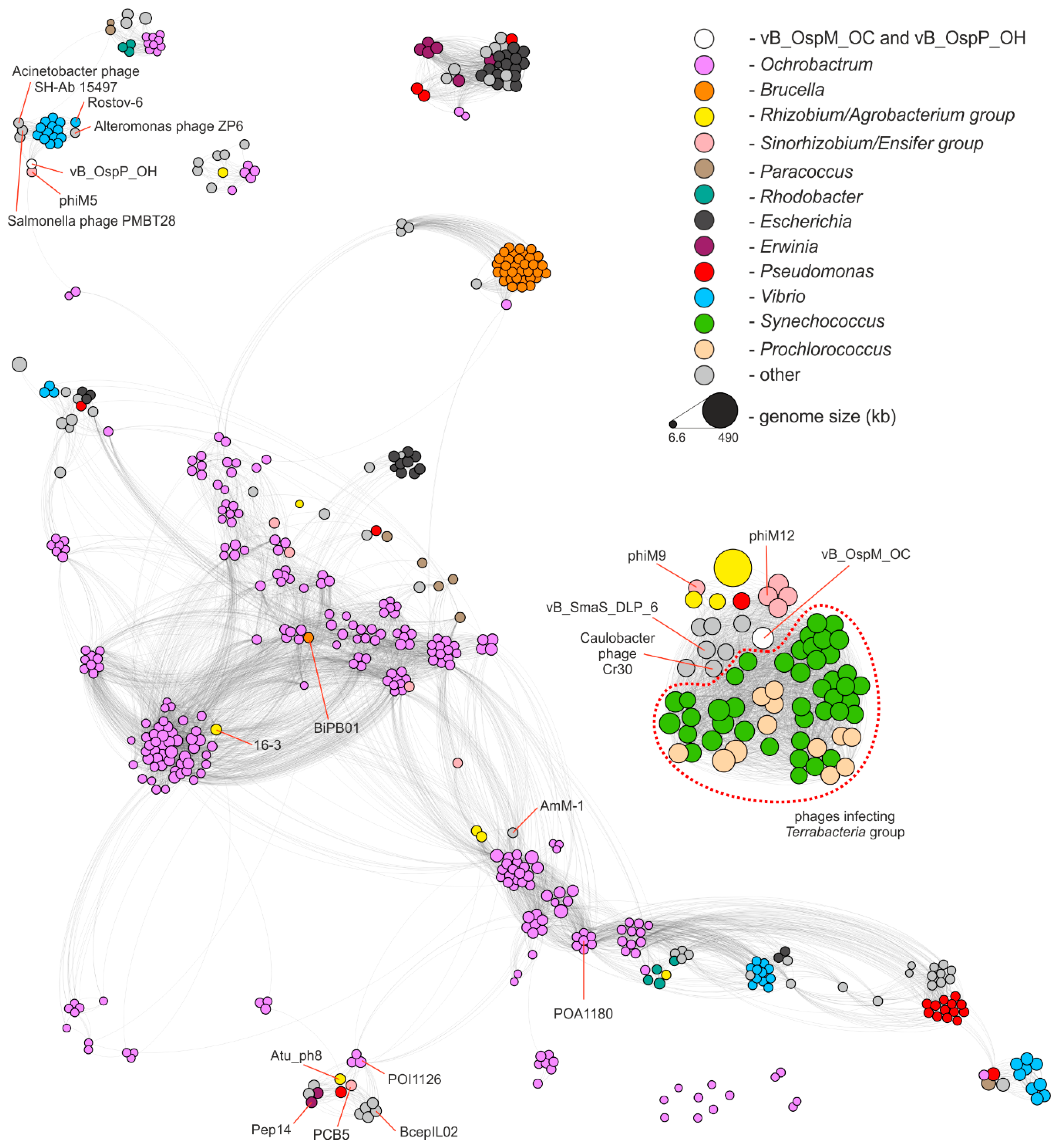
2.3. Comparative Genomics of Ochrobactrum Phages
2.4. Functional Characterization of vB_OspM_OC and vB_OspP_OH
2.4.1. Host Range Analysis
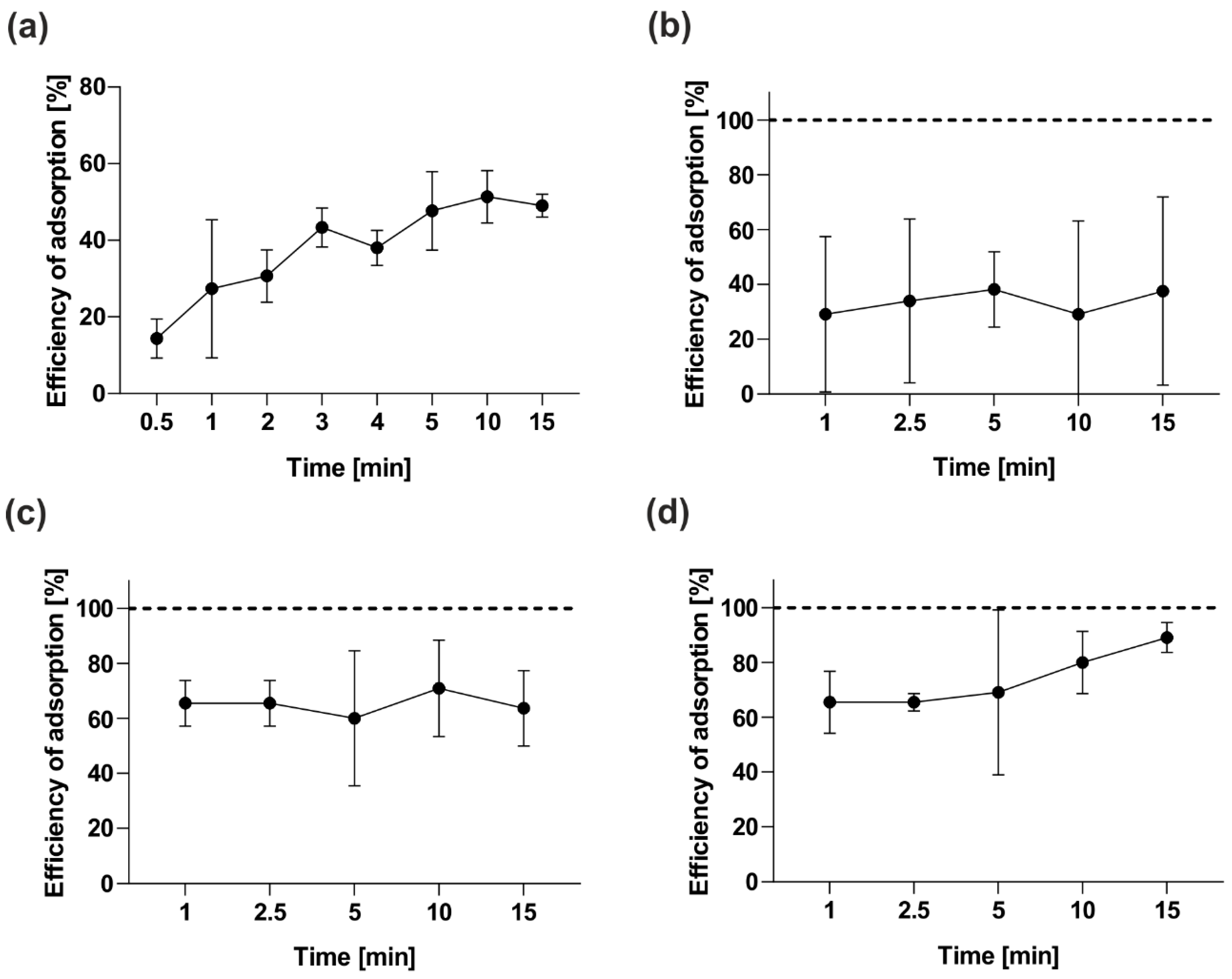
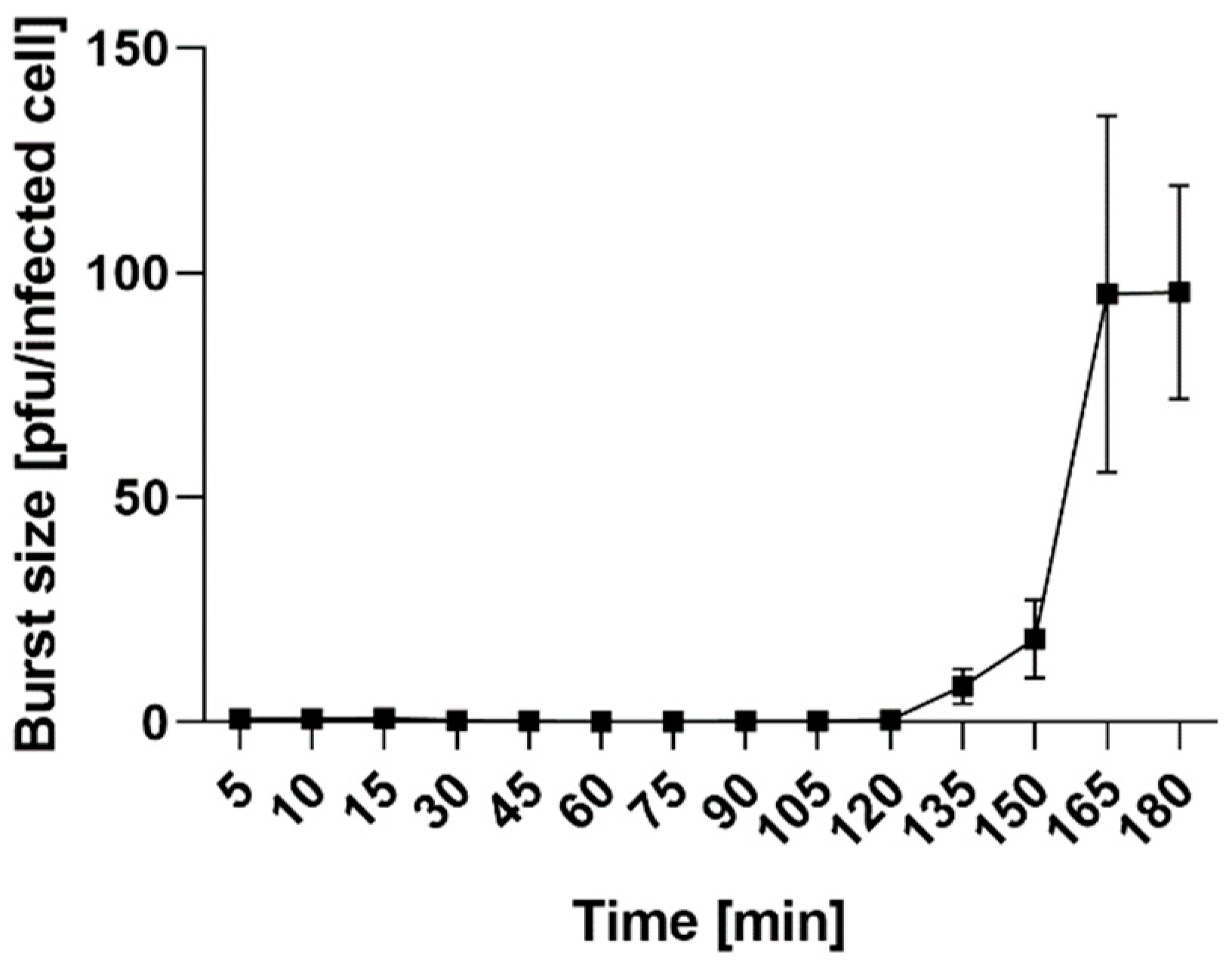
2.4.2. Adsorption Assay and One-Step Growth Curve
2.4.3. Killing Assay
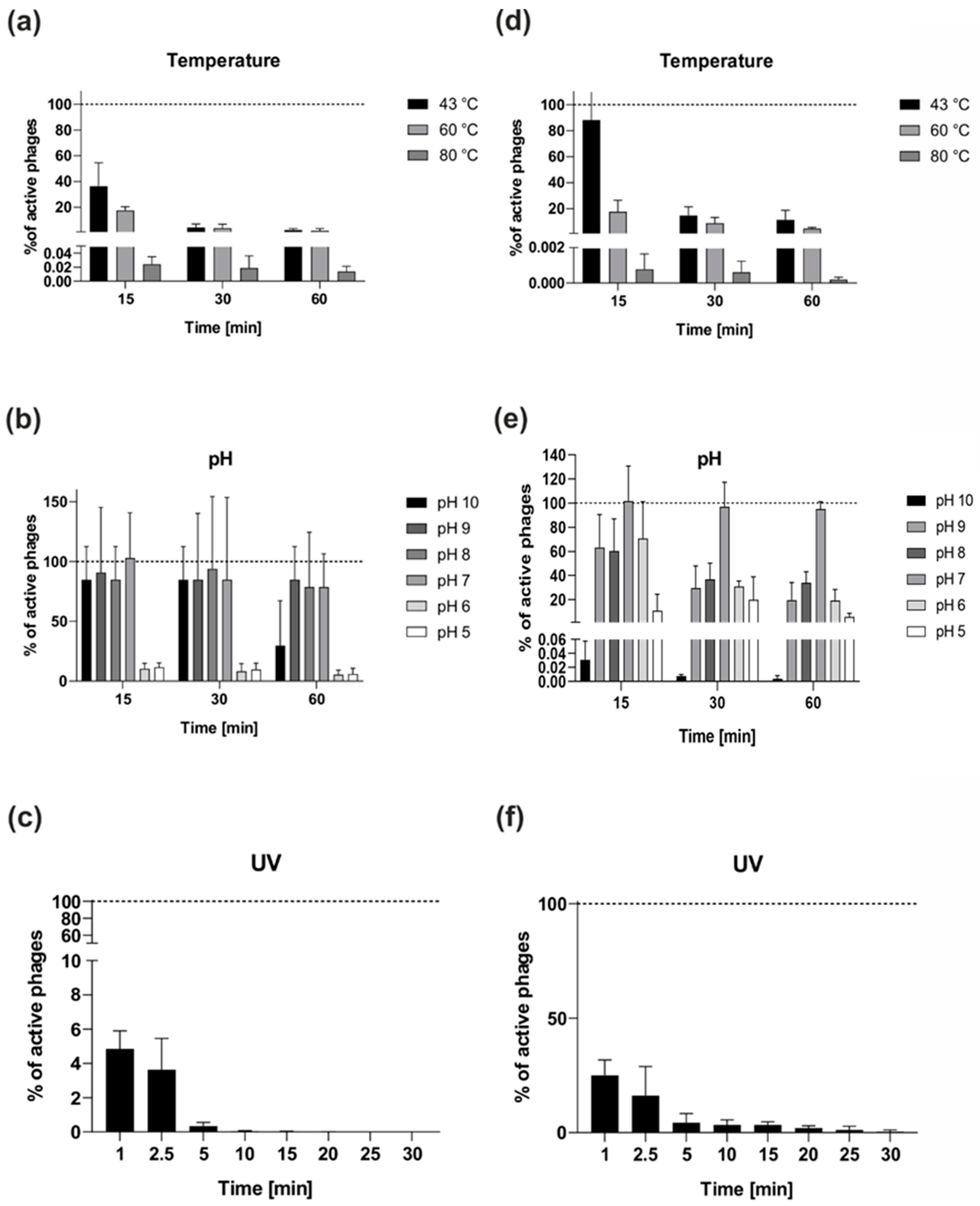
2.4.4. Stability of the vB_OspM_OC and vB_OspP_OH Phages
3. Materials and Methods
3.1. Bacterial Strains, Plasmids and Culture Conditions
3.2. DNA Manipulations and Introduction of Plasmid DNA into Bacterial Cells
3.3. Cloning, Overexpression, Purification, and Testing of Putative DNA MTase Activities
3.4. Bacteriophages Isolation
3.5. Transmission Electron Microscopy (TEM)
3.6. Host Range Analysis
3.7. Adsorption Kinetics Assay of vB_OspM_OC and vB_OspP_OH
3.8. Killing Assay
3.9. One-Step Growth Curve
3.10. Bacteriophage Stability at Various Values of pH and Temperature
3.11. Bacteriophage Resistance to UV Light
3.12. DNA Sequencing
3.13. Detection of Prophages in Bacterial Genomes and Classification of Phages
3.14. Genome Annotation
3.15. Comparative Genomics
3.16. Nucleotide Sequence Accession Numbers
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Thomas, R.; Janardhanan, A.; Varghese, R.T.; Soniya, E.V.; Mathew, J.; Radhakrishnan, E.K. Antibacterial properties of silver nanoparticles synthesized by marine Ochrobactrum sp. Braz. J. Microbiol. 2014, 45, 1221–1227. [Google Scholar] [CrossRef] [PubMed]
- Mishra, S.K.; Khan, M.H.; Misra, S.; Dixit, V.K.; Khare, P.; Srivastava, S.; Chauhan, P.S. Characterisation of Pseudomonas spp. and Ochrobactrum sp. isolated from volcanic soil. Antonie Leeuwenhoek Int. J. Gen. Mol. Microbiol. 2017, 110, 253–270. [Google Scholar] [CrossRef] [PubMed]
- Sipahutar, M.K.; Vangnai, A.S. Role of plant growth-promoting Ochrobactrum sp. MC22 on triclocarban degradation and toxicity mitigation to legume plants. J. Hazard. Mater. 2017, 329, 38–48. [Google Scholar] [CrossRef] [PubMed]
- Kampfer, P.; Huber, B.; Busse, H.J.; Scholz, H.C.; Tomaso, H.; Hotzel, H.; Melzer, F. Ochrobactrum pecoris sp. nov., isolated from farm animals. Int. J. Syst. Evol. Microbiol. 2011, 61, 2278–2283. [Google Scholar] [CrossRef] [PubMed]
- Bathe, S.; Achouak, W.; Hartmann, A.; Heulin, T.; Schloter, M.; Lebuhn, M. Genetic and phenotypic microdiversity of Ochrobactrum spp. FEMS Microbiol. Ecol. 2006, 56, 272–280. [Google Scholar] [CrossRef]
- Alonso, C.A.; Kwabugge, Y.A.; Anyanwu, M.U.; Torres, C.; Chah, K.F. Diversity of Ochrobactrum species in food animals, antibiotic resistance phenotypes and polymorphisms in the blaOCH gene. FEMS Microbiol. Lett. 2017, 364. [Google Scholar] [CrossRef]
- Chudasama, K.S.; Thaker, V.S. Genome sequence of Ochrobactrum anthropi strain SUBG007, a plant pathogen and potential xenobiotic compounds degradation bacterium. Genom. Data 2017, 11, 116–117. [Google Scholar] [CrossRef]
- Bhattacharya, M.; Biswas, D.; Sana, S.; Datta, S. Biodegradation of waste lubricants by a newly isolated Ochrobactrum sp. C1. 3 Biotech 2015, 5, 807–817. [Google Scholar] [CrossRef]
- Katsivela, E.; Moore, E.R.B.; Kalogerakis, N. Biodegradation of aliphatic and aromatic hydrocarbons: Specificity among bacteria isolated from refinery waste sludge. Water Air Soil Pollut. 2003, 3, 103–115. [Google Scholar] [CrossRef]
- Arulazhagan, P.; Vasudevan, N. Biodegradation of polycyclic aromatic hydrocarbons by a halotolerant bacterial strain Ochrobactrum sp. VA1. Mar. Pollut. Bull. 2011, 62, 388–394. [Google Scholar] [CrossRef]
- Poszytek, K.; Karczewska-Golec, J.; Ciok, A.; Decewicz, P.; Dziurzynski, M.; Gorecki, A.; Jakusz, G.; Krucon, T.; Lomza, P.; Romaniuk, K.; et al. Genome-guided characterization of Ochrobactrum sp. POC9 enhancing sewage sludge utilization - biotechnological potential and biosafety considerations. Int. J. Environ. Res. Public Health 2018, 15, 1501. [Google Scholar] [CrossRef] [PubMed]
- Pujar, N.K.; Laad, S.; Premakshi, H.G.; Pattar, S.V.; Mirjankar, M.; Kamanavalli, C.M. Biodegradation of phenmedipham by novel Ochrobactrum anthropi NC-1. 3 Biotech 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.; Zhang, N.; Xing, Y.; Lian, L.; Chen, Y.; Zhang, D.; Li, G.; Sun, G.; Song, Y. Microbial degradation of organophosphorus pesticides: Novel degraders, kinetics, functional genes, and genotoxicity assessment. Environ. Sci. Pollut. Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Liu, S.J.; Du, W. Chemotactic screening of imidazolinone-degrading bacteria by microfluidic SlipChip. J. Hazard. Mater. 2019, 512–519. [Google Scholar] [CrossRef] [PubMed]
- Veeranagouda, Y.; Emmanuel Paul, P.V.; Gorla, P.; Siddavattam, D.; Karegoudar, T.B. Complete mineralisation of dimethylformamide by Ochrobactrum sp. DGVK1 isolated from the soil samples collected from the coalmine leftovers. Appl. Microbiol. Biotechnol. 2006, 71, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Moller, L.V.; Arends, J.P.; Harmsen, H.J.; Talens, A.; Terpstra, P.; Slooff, M.J. Ochrobactrum intermedium infection after liver transplantation. J. Clin. Microbiol. 1999, 37, 241–244. [Google Scholar] [CrossRef]
- Menezes, F.G.; Abreu, M.G.; Kawagoe, J.Y.; Warth, A.N.; Deutsch, A.D.; Dornaus, M.F.; Martino, M.D.; Correa, L. Ochrobactrum anthropi bacteremia in a preterm infant with cystic fibrosis. Braz. J. Microbiol. 2014, 45, 559–561. [Google Scholar] [CrossRef]
- Galanakis, E.; Bitsori, M.; Samonis, G.; Christidou, A.; Georgiladakis, A.; Sbyrakis, S.; Tselentis, Y. Ochrobactrum anthropi bacteraemia in immunocompetent children. Scand. J. Infect. Dis. 2002, 34, 800–803. [Google Scholar] [CrossRef]
- Mahmood, M.S.; Sarwari, A.R.; Khan, M.A.; Sophie, Z.; Khan, E.; Sami, S. Infective endocarditis and septic embolization with Ochrobactrum anthropi: Case report and review of literature. J. Infect. 2000, 40, 287–290. [Google Scholar] [CrossRef]
- Jelveh, N.; Cunha, B.A. Ochrobactrum anthropi bacteremia. Hear. Lung J. Acute Crit. Care 1999, 28, 145–146. [Google Scholar] [CrossRef]
- Jäckel, C.; Hertwig, S.; Scholz, H.C.; Nockler, K.; Reetz, J.; Hammerl, J.A. Prevalence, host range, and comparative genomic analysis of temperate Ochrobactrum phages. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Scholz, H.C.; Mühldorfer, K.; Shilton, C.; Benedict, S.; Whatmore, A.M.; Blom, J.; Eisenberg, T. The change of a medically important genus: Worldwide occurrence of genetically diverse novel Brucella species in exotic frogs. PLoS ONE 2016, 11, e0168872. [Google Scholar] [CrossRef] [PubMed]
- Poszytek, K.; Karczewska-Golec, J.; Dziurzynski, M.; Stepkowska-Kowalska, O.; Gorecki, A.; Decewicz, P.; Dziewit, L.; Drewniak, L. Genome-wide and functional view of proteolytic and lipolytic bacteria for efficient biogas production through enhanced sewage sludge hydrolysis. Molecules 2019, 24, 2624. [Google Scholar] [CrossRef] [PubMed]
- Van Twest, R.; Kropinski, A.M. Bacteriophage enrichment from water and soil. Methods Mol. Biol. 2009, 501, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Adriaenssens, E.M.; Rodney Brister, J. How to name and classify your phage: An informal guide. Viruses 2017, 9, 70. [Google Scholar] [CrossRef]
- Miller, E.S.; Kutter, E.; Mosig, G.; Arisaka, F.; Kunisawa, T.; Ruger, W. Bacteriophage T4 Genome. Microbiol. Mol. Biol. Rev. 2003, 67, 86–156. [Google Scholar] [CrossRef]
- Yuan, Y.; Gao, M. Jumbo bacteriophages: An overview. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef]
- Hendrix, R.W. Jumbo bacteriophages. Curr. Top. Microbiol. Immunol. 2009, 328, 229–240. [Google Scholar] [CrossRef]
- Kreuzer, K.N.; Brister, J.R. Initiation of bacteriophage T4 DNA replication and replication fork dynamics: A review in the Virology Journal series on bacteriophage T4 and its relatives. Virol. J. 2010, 7. [Google Scholar] [CrossRef]
- Lee, J.Y.; Li, Z.; Miller, E.S. Vibrio phage KVP40 encodes a functional NAD+ salvage pathway. J. Bacteriol. 2017, 199. [Google Scholar] [CrossRef]
- Wood, E.J. Fundamentals of biochemistry: Life at the molecular level (Third Edition) by Voet, D., Voet, J. and Pratt. C.W. Biochem. Mol. Biol. Educ. 2008, 36, 319–320. [Google Scholar] [CrossRef]
- Miller, E.S.; Heidelberg, J.F.; Eisen, J.A.; Nelson, W.C.; Durkin, A.S.; Ciecko, A.; Feldblyum, T.V.; White, O.; Paulsen, I.T.; Nierman, W.C.; et al. Complete genome sequence of the broad-host-range vibriophage KVP40: Comparative genomics of a T4-related bacteriophage. J. Bacteriol. 2003, 185, 5220–5233. [Google Scholar] [CrossRef] [PubMed]
- Decewicz, P.; Radlinska, M.; Dziewit, L. Characterization of Sinorhizobium sp. LM21 prophages and virus-encoded DNA methyltransferases in the light of comparative genomic analyses of the sinorhizobial virome. Viruses 2017, 9, 161. [Google Scholar] [CrossRef] [PubMed]
- Decewicz, P.; Dziewit, L.; Golec, P.; Kozlowska, P.; Bartosik, D.; Radlinska, M. Characterization of the virome of Paracoccus spp. (Alphaproteobacteria) by combined in silico and in vivo approaches. Sci. Rep. 2019, 9, 7899. [Google Scholar] [CrossRef]
- Dziewit, L.; Oscik, K.; Bartosik, D.; Radlinska, M. Molecular characterization of a novel temperate Sinorhizobium bacteriophage, ΦLM21, encoding DNA methyltransferase with CcrM like specificity. J. Virol. 2014, 88, 13111–13124. [Google Scholar] [CrossRef]
- Coulby, J.N.; Sternberg, N.L. Characterization of the phage P1 dam gene. Gene 1988, 74, 191. [Google Scholar] [CrossRef]
- Radlinska, M.; Bujnicki, J.M. Cloning of enterohemorrhagic Escherichia coli phage VT-2 dam methyltransferase. Acta Microbiol. Pol. 2001, 50, 161–167. [Google Scholar]
- Scherzer, E.; Auer, B.; Schweiger, M. Identification, purification, and characterization of Escherichia coli virus T1 DNA methyltransferase. J. Biol. Chem. 1987, 262, 15225–15231. [Google Scholar]
- Hattman, S.; Wilkinson, J.; Swinton, D.; Schlagman, S.; Macdonald, P.M.; Mosig, G. Common evolutionary origin of the phage T4 dam and host Escherichia coli dam DNA-adenine methyltransferase genes. J. Bacteriol. 1985, 164, 932–937. [Google Scholar] [CrossRef]
- Mizuno, C.M.; Rodriguez-Valera, F.; Kimes, N.E.; Ghai, R. Expanding the marine virosphere using metagenomics. PLoS Genet. 2013, 9. [Google Scholar] [CrossRef]
- Stojković, E.A.; Rothman-Denes, L.B. Coliphage N4 N-acetylmuramidase defines a new family of murein hydrolases. J. Mol. Biol. 2007, 366, 406–419. [Google Scholar] [CrossRef] [PubMed]
- Edwards, R.; Katelyn, D.P.; Daniel, S. Linsalrob/PhiSpy: Version 3.4; Zenodo: Geneva, Switzerland, 2019. [Google Scholar] [CrossRef]
- Akhter, S.; Aziz, R.K.; Edwards, R.A. PhiSpy: A novel algorithm for finding prophages in bacterial genomes that combines similarity- and composition-based strategies. Nucleic Acids Res. 2012, 40, e126. [Google Scholar] [CrossRef] [PubMed]
- Bin Jang, H.; Bolduc, B.; Zablocki, O.; Kuhn, J.H.; Roux, S.; Adriaenssens, E.M.; Brister, J.R.; Kropinski, A.M.; Krupovic, M.; Lavigne, R.; et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 2019, 37, 632–639. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.C.; Sena-Velez, M.; Washburn, B.K.; Platt, G.N.; Lu, S.; Brewer, T.E.; Lynn, J.S.; Stroupe, M.E.; Jones, K.M. Structure, proteome and genome of Sinorhizobium meliloti phage ΦM5: A virus with LUZ24-like morphology and a highly mosaic genome. J. Struct. Biol. 2017, 200, 343–359. [Google Scholar] [CrossRef]
- Ely, B.; Gibbs, W.; Diez, S.; Ash, K. The Caulobacter crescentus transducing phage Cr30 is a unique member of the T4-Like family of myophages. Curr. Microbiol. 2015, 70, 854–858. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Solís-Sánchez, A.; Hernández-Chiñas, U.; Navarro-Ocaña, A.; De La Mora, J.; Xicohtencatl-Cortes, J.; Eslava-Campos, C. Genetic characterization of ØvC8 lytic phage for Vibrio cholerae O1. Virol. J. 2016, 13. [Google Scholar] [CrossRef]
- Koberg, S.; Brinks, E.; Albrecht, V.; Neve, H.; Franz, C.M.A.P. Complete genome sequence of the novel virulent phage PMBT28 with lytic activity against thermotolerant Salmonella enterica subsp. enterica serovar Senftenberg ATCC 43845. Genome Announc. 2018, 6. [Google Scholar] [CrossRef]
- Golec, P.; Karczewska-Golec, J.; Voigt, B.; Albrecht, D.; Schweder, T.; Hecker, M.; Wȩgrzyn, G.; Łoś, M. Proteomic profiles and kinetics of development of bacteriophage T4 and its ri and riii mutants in slowly growing Escherichia coli. J. Gen. Virol. 2013, 94, 896–905. [Google Scholar] [CrossRef]
- Łoś, M.; Wegrzyn, G. Pseudolysogeny. In Advances in Virus Research; Academic Press Inc.: Cambridge, MA, USA, 2012; Volume 82, pp. 339–349. [Google Scholar] [CrossRef]
- Hampton, H.G.; Watson, B.N.J.; Fineran, P.C. The arms race between bacteria and their phage foes. Nature 2020, 577, 327–336. [Google Scholar] [CrossRef]
- Rostøl, J.T.; Marraffini, L. (Ph)ighting phages: How bacteria resist their parasites. Cell Host Microbe 2019, 25, 184–194. [Google Scholar] [CrossRef]
- Dziewit, L.; Pyzik, A.; Szuplewska, M.; Matlakowska, R.; Mielnicki, S.; Wibberg, D.; Schlüter, A.; Pühler, A.; Bartosik, D. Diversity and role of plasmids in adaptation of bacteria inhabiting the Lubin copper mine in Poland, an environment rich in heavy metals. Front. Microbiol. 2015, 6, 152. [Google Scholar] [CrossRef] [PubMed]
- Hooykaas, P.J.; den Dulk-Ras, H.; Schilperoort, R.A. Molecular mechanism of Ti plasmid mobilization by R plasmids: Isolation of Ti plasmids with transposon-insertions in Agrobacterium tumefaciens. Plasmid 1980, 4, 64–75. [Google Scholar] [CrossRef]
- DiCenzo, G.C.; Debiec, K.; Krzysztoforski, J.; Uhrynowski, W.; Mengoni, A.; Fagorzi, C.; Gorecki, A.; Dziewit, L.; Bajda, T.; Rzepa, G.; et al. Genomic and biotechnological characterization of the heavy-metal resistant, arsenic-oxidizing bacterium Ensifer sp. M14. Genes 2018, 9, 379. [Google Scholar] [CrossRef]
- Hanahan, D. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 1983, 166, 557–580. [Google Scholar] [CrossRef]
- Urakami, T.; Tamaoka, J.; Suzuki, K.; Komagata, K. Paracoccus alcaliphilus sp. nov., an alkaliphilic and facultatively methylotrophic bacterium. Int. J. Syst. Bacteriol. 1989, 39, 116–121. [Google Scholar] [CrossRef]
- Urakami, T.; Araki, H.; Oyanagi, H.; Suzuki, K.; Komagata, K. Paracoccus aminophilus sp. nov. and Paracoccus aminovorans sp. nov., which utilize N,N-dimethylformamide. Int. J. Syst. Bacteriol. 1990, 40, 287–291. [Google Scholar] [CrossRef] [PubMed]
- Bartosik, A.A.; Glabski, K.; Jecz, P.; Mikulska, S.; Fogtman, A.; Koblowska, M.; Jagura-Burdzy, G. Transcriptional profiling of ParA and ParB mutants in actively dividing cells of an opportunistic human pathogen Pseudomonas aeruginosa. PLoS ONE 2014, 9, e87276. [Google Scholar] [CrossRef] [PubMed]
- Matlakowska, R.; Sklodowska, A. The culturable bacteria isolated from organic-rich black shale potentially useful in biometallurgical procedures. J. Appl. Microbiol. 2009, 107, 858–866. [Google Scholar] [CrossRef]
- Dziewit, L.; Cegielski, A.; Romaniuk, K.; Uhrynowski, W.; Szych, A.; Niesiobedzki, P.; Zmuda-Baranowska, M.J.; Zdanowski, M.K.; Bartosik, D. Plasmid diversity in arctic strains of Psychrobacter spp. Extremophiles 2013, 17, 433–444. [Google Scholar] [CrossRef][Green Version]
- Jamieson, W.D.; Pehl, M.J.; Gregory, G.A.; Orwin, P.M. Coordinated surface activities in Variovorax paradoxus EPS. BMC Microbiol. 2009, 9, 124. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2001. [Google Scholar]
- Kushner, S.R. An improved method for transformation of E. coli with ColE1 derived plasmids. In Genetic Engineering; Boyer, H.B., Nicosia, S., Eds.; Elsevier/North-Holland: Amsterdam, The Netherlands, 1978; pp. 17–23. [Google Scholar]
- Drozdz, M.; Piekarowicz, A.; Bujnicki, J.M.; Radlinska, M. Novel non-specific DNA adenine methyltransferases. Nucleic Acids Res. 2012, 40, 2119–2130. [Google Scholar] [CrossRef] [PubMed]
- Golec, P.; Karczewska-Golec, J.; Loś, M.; Wegrzyn, G. Bacteriophage T4 can produce progeny virions in extremely slowly growing Escherichia coli host: Comparison of a mathematical model with the experimental data. FEMS Microbiol. Lett. 2014, 351, 156–161. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Li, H. Aligning Sequence Reads, Clone Sequences and Assembly Contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Robinson, J.T.; Thorvaldsdóttir, H.; Winckler, W.; Guttman, M.; Lander, E.S.; Getz, G.; Mesirov, J.P. Integrative genomics viewer. Nat. Biotechnol. 2011, 29, 24–26. [Google Scholar] [CrossRef]
- Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 2011, 7. [Google Scholar] [CrossRef]
- Grazziotin, A.L.; Koonin, E.V.; Kristensen, D.M. Prokaryotic Virus Orthologous Groups (pVOGs): A resource for comparative genomics and protein family annotation. Nucleic Acids Res. 2017, 45, D491–D498. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2014, 12, 59–60. [Google Scholar] [CrossRef]
- Nepusz, T.; Yu, H.; Paccanaro, A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat. Methods 2012, 9, 471–472. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.; Harris, S.R.; Berriman, M.; Parkhill, J.; McQuillan, J.A. Artemis: An integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 2012, 28, 464–469. [Google Scholar] [CrossRef] [PubMed]
- McNair, K.; Zhou, C.; Dinsdale, E.A.; Souza, B.; Edwards, R.A. Phanotate: A novel approach to gene identification in phage genomes. Bioinformatics 2019. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R.; et al. CDD: A Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef]
- Claudel-Renard, C.; Chevalet, C.; Faraut, T.; Kahn, D. Enzyme-specific profiles for genome annotation: PRIAM. Nucleic Acids Res. 2003, 31, 6633–6639. [Google Scholar] [CrossRef]
- Soding, J.; Biegert, A.; Lupas, A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 2005, 33, W244–W248. [Google Scholar] [CrossRef]
- Lowe, T.M.; Chan, P.P. tRNAscan-SE On-line: Integrating search and context for analysis of transfer RNA genes. Nucleic. Acids Res. 2016, 44, W54–W57. [Google Scholar] [CrossRef]
- Laslett, D.; Canback, B. Aragorn, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 2004, 32, 11–16. [Google Scholar] [CrossRef]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An open source software for exploring and manipulating networks. In Proceedings of the Third International Conference on Weblogs and Social Media (ICWSM 2009), San Jose, CA, USA, 17–20 May 2009; pp. 361–362. [Google Scholar]
- Jacomy, M.; Venturini, T.; Heymann, S.; Bastian, M. ForceAtlas2, a continuous graph layout algorithm for handy network visualization designed for the Gephi software. PLoS ONE 2014, 9, e98679. [Google Scholar] [CrossRef]
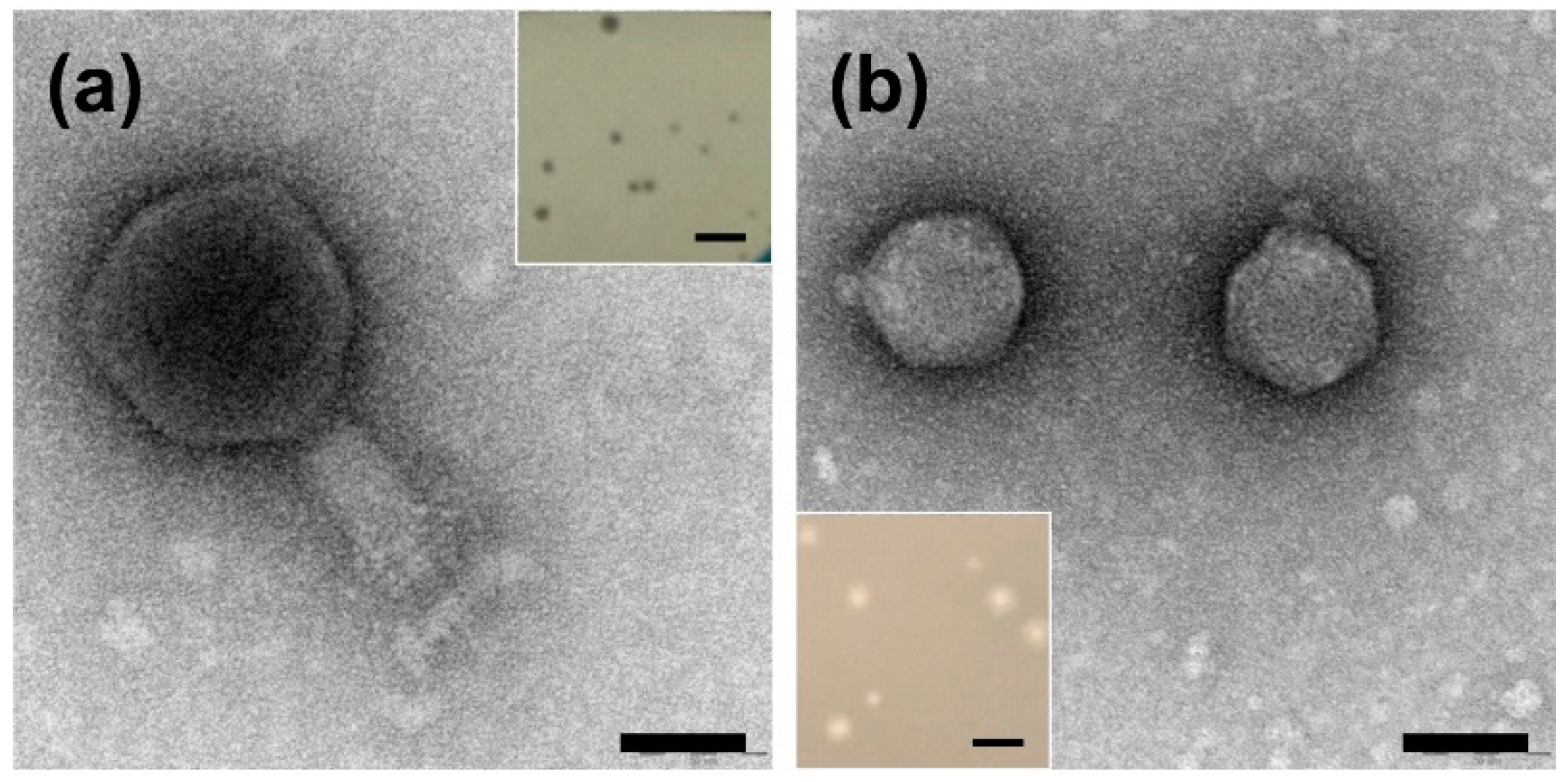
| Phage | Head Width (nm) * | Head Length (nm) * | Tail Width (nm) * | Tail Length (nm) * |
|---|---|---|---|---|
| vB_OspM_OC | 97.6 ± 7.8 | 102.9 ± 8.0 | 28.9 ± 2.9 | 68.3 ± 11.4 |
| vB_OspP_OH | 55.1 ± 1.6 | 55.8 ± 1.9 | 11.5 ± 1.6 | 7.6 ± 0.9 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Decewicz, P.; Golec, P.; Szymczak, M.; Radlinska, M.; Dziewit, L. Identification and Characterization of the First Virulent Phages, Including a Novel Jumbo Virus, Infecting Ochrobactrum spp. Int. J. Mol. Sci. 2020, 21, 2096. https://doi.org/10.3390/ijms21062096
Decewicz P, Golec P, Szymczak M, Radlinska M, Dziewit L. Identification and Characterization of the First Virulent Phages, Including a Novel Jumbo Virus, Infecting Ochrobactrum spp. International Journal of Molecular Sciences. 2020; 21(6):2096. https://doi.org/10.3390/ijms21062096
Chicago/Turabian StyleDecewicz, Przemyslaw, Piotr Golec, Mateusz Szymczak, Monika Radlinska, and Lukasz Dziewit. 2020. "Identification and Characterization of the First Virulent Phages, Including a Novel Jumbo Virus, Infecting Ochrobactrum spp." International Journal of Molecular Sciences 21, no. 6: 2096. https://doi.org/10.3390/ijms21062096
APA StyleDecewicz, P., Golec, P., Szymczak, M., Radlinska, M., & Dziewit, L. (2020). Identification and Characterization of the First Virulent Phages, Including a Novel Jumbo Virus, Infecting Ochrobactrum spp. International Journal of Molecular Sciences, 21(6), 2096. https://doi.org/10.3390/ijms21062096