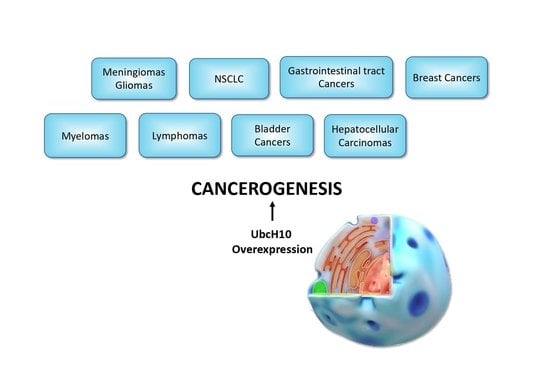
UbcH10 a Major Actor in Cancerogenesis and a Potential Tool for Diagnosis and Therapy
Abstract
1. Introduction
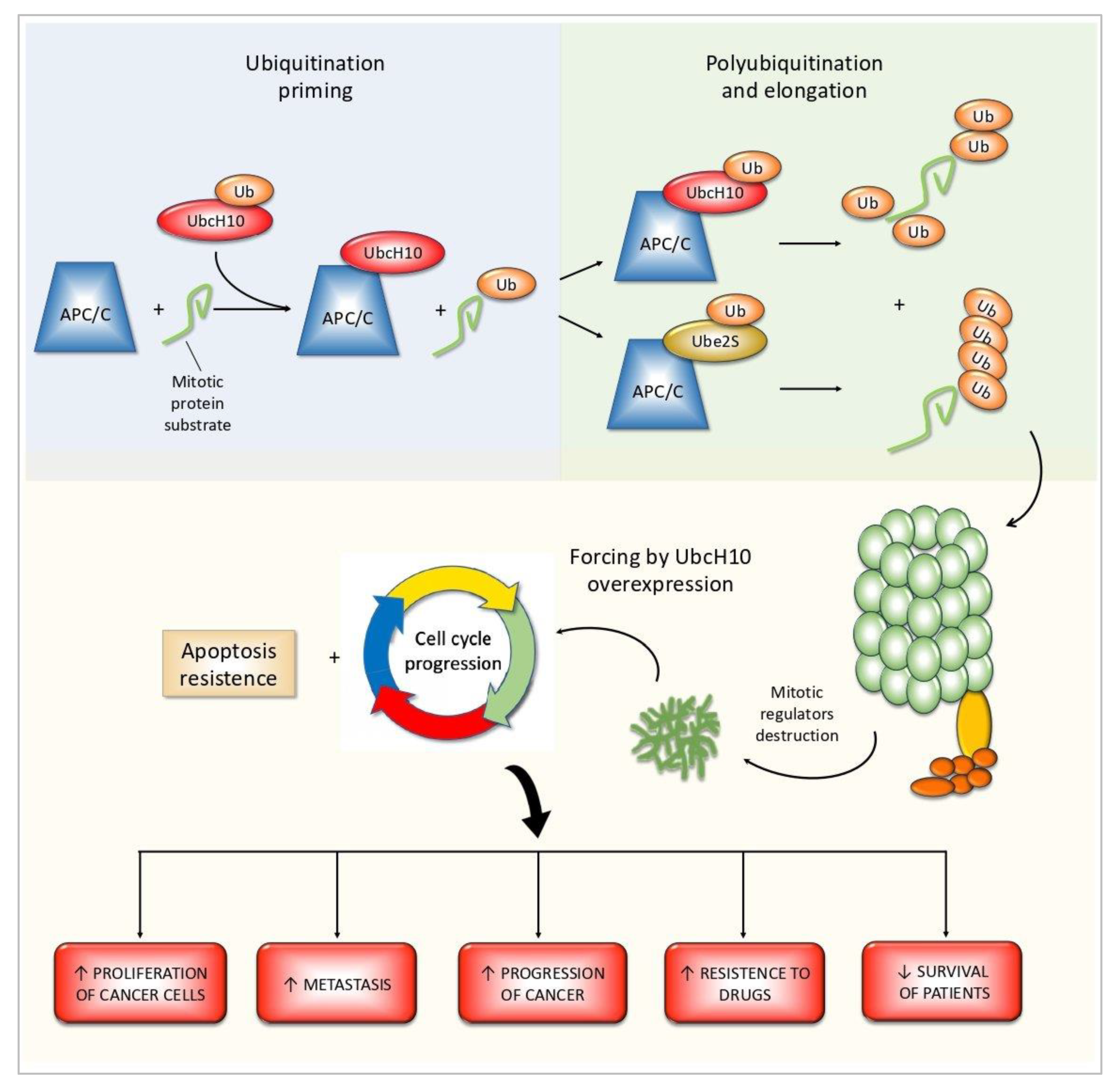
2. The Role in Cell Cycle Regulation
3. UbcH10 Studies in Various Types of Tumors
3.1. Brain Tumors
3.2. Lung Cancer
3.3. Breast Cancer
3.4. Thyroid Cancer
3.5. Gastroenteric Tumors
3.6. Gynecologic Tumors
3.7. Pancreas and Liver
3.8. Other Neoplasms
4. Discussion
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ABCB1 | ATP binding cassette subfamily B member 1 |
| Akt/PKB | AKT Serine/Threonine Kinase 1 |
| ALLN | N-acetyl-Leu-Leu-Norleu-al |
| APC/C | Anaphase-promoting complex/cyclosome |
| ATC | Anaplastic thyroid carcinoma |
| BTZ | Bortezomib |
| Cdh1 | APC coactivator Cdh1 |
| CdK1 | Cyclin-dependent kinase 1 |
| CdK2 | Cyclin dependent kinase 2 |
| CDKs | Cyclin-depended kinases |
| c-Myc | Avian myelocytomatosis viral oncogene homolog |
| CTCs | Circulating tumor cells |
| E1s | Ubiquitin-activating enzymes |
| E2s | Ubiquitin-conjugating enzymes |
| E3s | Ubiquitin ligases |
| EGFR | Epidermal growth factor receptor |
| ErbB2 | Erb-b2 receptor tyrosine kinase 2 |
| ERK | Extracellular regulated MAP kinase |
| E2~Ub | E2 enzyme and ubiquitin conjugated by thioester bond |
| FNA | Fine-needle aspiration |
| HCC | Hepatocellular carcinoma |
| HL | Hodgkin’s lymphoma |
| hsa-miR-631 | microRNA 631 |
| hsa-miR661-3p | microRNA 661 |
| NF-κB | Nuclear factor kappa B subunit |
| NHL | non-Hodgkin’s lymphoma |
| NSCLC | non-small cell lung carcinomas |
| p38 | Mitogen-activated protein kinase p38 |
| p53 | Tumor suppressor p53 |
| RIZ1 | Retinoblastoma protein-interacting zinc finger protein1 |
| RNAi | RNA interference |
| qRT-PCR | Quantitative real time polymerase chain reaction |
| siRNA | Small interfering RNA |
| SKY | Spectral karyotyping |
| UPS | Ubiquitin proteasome system |
References
- Ye, Y.; Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 2009, 10, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Scheffner, M.; Nuber, U.; Huibregtse, J.M. Protein ubiquitination involving an E1–E2–E3 enzyme ubiquitin thioester cascade. Nature 1995, 373, 81–83. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.G.; VanderLinden, R.; Watson, E.R.; Weissmann, F.; Ordureau, A.; Wu, K.-P.; Zhang, W.; Yu, S.; Mercredi, P.Y.; Harrison, J.S.; et al. Dual RING E3 Architectures Regulate Multiubiquitination and Ubiquitin Chain Elongation by APC/C. Cell 2016, 165, 1440–1453. [Google Scholar] [CrossRef] [PubMed]
- Deshaies, R.J.; Joazeiro, C.A.P. RING Domain E3 Ubiquitin Ligases. Annu. Rev. Biochem. 2009, 78, 399–434. [Google Scholar] [CrossRef] [PubMed]
- Glickman, M.H.; Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: Destruction for the sake of construction. Physiol. Rev. 2002, 82, 373–428. [Google Scholar] [CrossRef] [PubMed]
- Leestemaker, Y.; Ovaa, H. Tools to Investigate the Ubiquitin Proteasome System. Available online: https://pubmed.ncbi.nlm.nih.gov/?term=Tools+to+investigate+the+ubiquitin+proteasome+system (accessed on 19 February 2020).
- Primorac, I.; Musacchio, A. Panta rhei: The APC/C at steady state. J. Cell Biol. 2013, 201, 177–189. [Google Scholar] [CrossRef]
- Okamoto, Y.; Ozaki, T.; Miyazaki, K.; Aoyama, M.; Miyazaki, M.; Nakagawara, A. UbcH10 is the cancer-related E2 ubiquitin-conjugating enzyme. Cancer Res. 2003, 63, 4167–4173. [Google Scholar]
- Poon, R.Y.C. Cell cycle control: A system of interlinking oscillators. Methods Mol. Biol. 2016, 1342, 3–19. [Google Scholar]
- Lukas, C.; Sørensen, C.S.; Kramer, E.; Santoni-Ruglu, E.; Lindeneg, C.; Peters, J.M.; Bartek, J.; Lukas, J. Accumulation of cyclin B1 requires E2F and cyclin-A-dependent rearrangement of the anaphase-promoting complex. Nature 1999, 401, 815–818. [Google Scholar] [CrossRef]
- Pagano, M.; Pepperkok, R.; Verde, F.; Ansorge, W.; Draetta, G. Cyclin A is required at two points in the human cell cycle. EMBO J. 1992, 11, 961–971. [Google Scholar] [CrossRef]
- Geley, S.; Kramer, E.; Gieffers, C.; Gannon, J.; Peters, J.M.; Hunt, T. Anaphase-promoting complex/cyclosome-dependent proteolysis of human cyclin A starts at the beginning of mitosis and is not subject to the spindle assembly checkpoint. J. Cell Biol. 2001, 153, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Elzen Den, N.; Pines, J. Cyclin A is destroyed in prometaphase and can delay chromosome alignment and anaphase. J. Cell Biol. 2001, 153, 121–135. [Google Scholar] [CrossRef] [PubMed]
- Peters, J.M. The anaphase-promoting complex: Proteolysis in mitosis and beyond. Mol. Cell 2002, 9, 931–943. [Google Scholar] [CrossRef]
- Townsley, F.M.; Aristarkhov, A.; Beck, S.; Hershko, A.; Ruderman, J.V. Dominant-negative cyclin-selective ubiquitin carrier protein E2-C/UbcH10 blocks cells in metaphase. Proc. Natl. Acad. Sci. USA 1997, 94, 2362–2367. [Google Scholar] [CrossRef] [PubMed]
- Yamanaka, A.; Hatakeyama, S.; Kominami, K.I.; Kitagawa, M.; Matsumoto, M.; Nakayama, K.I. Cell cycle-dependent expression of mammalian E2-C regulated by the anaphase-promoting complex/cyclosome. Mol. Biol. Cell 2000, 11, 2821–2831. [Google Scholar] [CrossRef][Green Version]
- Rape, M.; Kirschner, M.W. Autonomous regulation of the anaphase-promoting complex couples mitosis to S-phase entry. Nature 2004, 432, 588–595. [Google Scholar] [CrossRef]
- Cai, Z.; Zou, Y.; Hu, H.; Lu, C.; Sun, W.; Jiang, L.; Hu, G. RIZ1 negatively regulates ubiquitin-conjugating enzyme E2C/UbcH10 via targeting c-Myc in meningioma. Am. J. Transl. Res. 2017, 9, 2645–2655. [Google Scholar]
- Jiang, L.; Wang, T.; Bao, Y.; Qian, J.; Wu, X.J.; Hu, G.H.; Lu, Y.C. A study of UbcH10 expression and its association with recurrence of meningiomas. J. Surg. Oncol. 2012, 106, 327–331. [Google Scholar] [CrossRef]
- Jiang, L.; Huang, C.G.; Lu, Y.C.; Luo, C.; Hu, G.H.; Liu, H.M.; Chen, J.X.; Han, H.X. Expression of ubiquitin-conjugating enzyme E2C/UbcH10 in astrocytic tumors. Brain Res. 2008, 1201, 161–166. [Google Scholar] [CrossRef]
- Donato, G.; Iofrida, G.; Lavano, A.; Volpentesta, G.; Signorelli, F.; Pallante, P.L.; Berlingieri, M.T.; Pierantoni, M.G.; Palmieri, D.; Conforti, F.; et al. Analysis of UbcH10 expression represents a useful tool for the diagnosis and therapy of astrocytic tumors. Clin. Neuropathol. 2008, 27, 219–223. [Google Scholar] [CrossRef]
- Jiang, L.; Bao, Y.; Luo, C.; Hu, G.; Huang, C.; Ding, X.; Sun, K.; Lu, Y. Knockdown of ubiquitin-conjugating enzyme E2C/UbcH10 expression by RNA interference inhibits glioma cell proliferation and enhances cell apoptosis in vitro. J. Cancer Res. Clin. Oncol. 2010, 136, 211–217. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Gu, X.; Liu, F.; Rui, Z.; Liu, M.; Zhao, L. Antitumor effects of hsa-miR661-3p on non-small cell lung cancer in vivo and in vitro. Oncol. Rep. 2019, 41, 2987–2996. [Google Scholar] [CrossRef] [PubMed]
- Perrotta, I.; Bruno, L.; Maltese, L.; Russo, E.; Donato, A.; Donato, G. Immunohistochemical Analysis of the Ubiquitin-conjugating Enzyme UbcH10 in Lung Cancer: A Useful Tool for Diagnosis and Therapy. J. Histochem. Cytochem. 2012, 60, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Pallante, P.; Malapelle, U.; Berlingieri, M.T.; Bellevicine, C.; Sepe, R.; Federico, A.; Rocco, D.; Galgani, M.; Chiariotti, L.; Sanchez-Cespedes, M.; et al. UbcH10 overexpression in human lung carcinomas and its correlation with EGFR and p53 mutational status. Eur. J. Cancer 2013, 49, 1117–1126. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Jiang, L.; Wang, L.; He, J.; Yu, H.; Sun, G.; Chen, J.; Xiu, Q.; Li, B. UbcH10 expression provides a useful tool for the prognosis and treatment of non-small cell lung cancer. J. Cancer Res. Clin. Oncol. 2012, 138, 1951–1961. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.C.; Chang, T.W.; Chen, F.M.; Hou, M.F.; Hung, S.Y.; Chong, I.W.; Lee, S.C.; Zhou, T.H.; Lin, S.R. Combination of multiple mRNA markers (PTTG1, survivin, UbcH10 and TK1) in the diagnosis of Taiwanese patients with breast cancer by membrane array. Oncology 2007, 70, 438–446. [Google Scholar] [CrossRef]
- Berlingieri, M.T.; Pallante, P.; Sboner, A.; Barbareschi, M.; Bianco, M.; Ferraro, A.; Mansueto, G.; Borbone, E.; Guerriero, E.; Troncone, G.; et al. UbcH10 is overexpressed in malignant breast carcinomas. Eur. J. Cancer 2007, 43, 2729–2735. [Google Scholar] [CrossRef]
- Wang, C.; Pan, Y.H.; Shan, M.; Xu, M.; Bao, J.L.; Zhao, L.M. Knockdown of UbcH10 enhances the chemosensitivity of dual drug resistant breast cancer cells to epirubicin and docetaxel. Int. J. Mol. Sci. 2015, 16, 4698–4712. [Google Scholar] [CrossRef]
- Pallante, P.; Berlingieri, M.T.; Troncone, G.; Kruhoffer, M.; Orntoft, T.F.; Viglietto, G.; Caleo, A.; Migliaccio, I.; Decaussin-Petrucci, M.; Santoro, M.; et al. UbcH10 overexpression may represent a marker of anaplastic thyroid carcinomas. Br. J. Cancer 2005, 93, 464–471. [Google Scholar] [CrossRef]
- Lee, J.J.; Foukakis, T.; Hashemi, J.; Grimelius, L.; Heldin, N.E.; Wallin, G.; Rudduck, C.; Lui, W.O.; Höög, A.; Larsson, C. Molecular cytogenetic profiles of novel and established human anaplastic thyroid carcinoma models. Thyroid 2007, 17, 289–301. [Google Scholar] [CrossRef]
- Guerriero, E.; Ferraro, A.; Desiderio, D.; Pallante, P.; Berlingieri, M.T.; Iaccarino, A.; Palmieri, E.; Palombini, L.; Fusco, A.; Troncone, G. UbcH10 expression on thyroid fine-needle aspirates. Cancer Cytopathol. 2010, 118, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Fujita, T.; Ikeda, H.; Taira, N.; Hatoh, S.; Naito, M.; Doihara, H. Overexpression of UbcH10 alternates the cell cycle profile and accelerate the tumor proliferation in colon cancer. BMC Cancer 2009, 9, 87. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.-M.; Jiang, C.-Y.; Wu, J.-Y.; Liu, B.; Chen, Y.-J.; Hu, C.-J.; Liu, X.-X. Rna Interference-Mediated Silencing of Ubch10 Gene Inhibits Colorectal Cancer Cell Growth in Vitro and in Vivo. Clin. Exp. Pharmacol. Physiol. 2009, 37. [Google Scholar]
- Li, S.Z.; Song, Y.; Zhang, H.H.; Jin, B.X.; Liu, Y.; Liu, W.B.; Zhang, X.D.; Du, R.L. UbcH10 overexpression increases carcinogenesis and blocks ALLN susceptibility in colorectal cancer. Sci. Rep. 2014, 4. [Google Scholar] [CrossRef] [PubMed]
- Cacciola, N.A.; Calabrese, C.; Malapelle, U.; Pellino, G.; De Stefano, A.; Sepe, R.; Sgariglia, R.; Quintavalle, C.; Federico, A.; Bianco, A.; et al. UbcH10 expression can predict prognosis and sensitivity to the antineoplastic treatment for colorectal cancer patients. Mol. Carcinog. 2016, 55, 793–807. [Google Scholar] [CrossRef]
- Pellino, G.; Pallante, P.; Malapelle, U.; Ferraro, A.; Bellevicine, C.; Milone, M.; Troncone, G.; Fusco, A.; Selvaggi, F. UbcH10 overexpression is less pronounced in older colorectal cancer patients. Int. J. Colorectal Dis. 2016, 31, 1367–1368. [Google Scholar] [CrossRef]
- Berlingieri, M.T.; Pallante, P.; Guida, M.; Nappi, C.; Masciullo, V.; Scambia, G.; Ferraro, A.; Leone, V.; Sboner, A.; Barbareschi, M.; et al. UbcH10 expression may be a useful tool in the prognosis of ovarian carcinomas. Oncogene 2007, 26, 2136–2140. [Google Scholar] [CrossRef][Green Version]
- Zhao, Z.K.; Wu, W.G.; Chen, L.; Dong, P.; Gu, J.; Mu, J.S.; Yang, J.H.; Liu, Y. Bin Expression of UbcH10 in pancreatic ductal adenocarcinoma and its correlation with prognosis. Tumor Biol. 2013, 34, 1473–1477. [Google Scholar] [CrossRef]
- Matsumoto, A.; Ishibashi, Y.; Urashima, M.; Omura, N.; Nakada, K.; Nishikawa, K.; Shida, A.; Takada, K.; Kashiwagi, H.; Yanaga, K. High UBCH10 protein expression as a marker of poor prognosis in esophageal squamous cell carcinoma. Anticancer Res. 2014, 34, 955–962. [Google Scholar]
- Yang, M.; Qu, Y.; Shi, R.; Wu, X.; Su, C.; Hu, Z.; Chang, Q.; Liu, S.; Pan, G.; Lei, M.; et al. Ubiquitin-conjugating enzyme UbcH10 promotes gastric cancer growth and is a potential biomarker for gastric cancer. Oncol. Rep. 2016, 36, 779–786. [Google Scholar] [CrossRef]
- Kefeli, M.; Yildiz, L.; Celik, H.; Tosun, M.; Karagoz, F. UbcH10 expression in benign, hyperplastic, and malignant endometrial curetted materials: A tissue microarray study. Int. J. Surg. Pathol. 2012, 20, 360–366. [Google Scholar] [CrossRef] [PubMed]
- Morikawa, T.; Kawai, T.; Abe, H.; Kume, H.; Homma, Y.; Fukayama, M. UBE2C is a marker of unfavorable prognosis in bladder cancer after radical cystectomy. Int. J. Clin. Exp. Pathol. 2013, 6, 1367–1374. [Google Scholar] [PubMed]
- Wang, S.T.; Li, D.Z.; Li, J.M.; Fang, J.; Li, H.Z.; Tong, P.J.; Liu, F.C. Lentivirus-mediated RNA interference targeting UbcH10 reduces cell growth and invasion of human osteosarcoma cells via inhibition of ki-67 and matrix metalloproteinases. Oncol. Lett. 2015, 9, 2171–2176. [Google Scholar] [CrossRef] [PubMed]
- Troncone, G.; Guerriero, E.; Pallante, P.; Berlingieri, M.T.; Ferraro, A.; Del Vecchio, L.; Gorrese, M.; Mariotti, E.; Iaccarino, A.; Palmieri, E.A.; et al. UbcH10 expression in human lymphomas. Histopathology 2009, 54, 731–740. [Google Scholar] [CrossRef]
- Xi, H.; Li, L.; Du, J.; An, R.; Fan, R.; Lu, J.; Wu, Y.X.; Wu, S.X.; Hou, J.; Zhao, L.M. Hsa-miR-631 resensitizes bortezomib-resistant multiple myeloma cell lines by inhibiting UbcH10. Oncol. Rep. 2017, 37, 961–968. [Google Scholar] [CrossRef]
- Han, S.S.; Liu, Q.G.; Zhou, Z.Y.; Yao, Y.M.; Song, T.; Zan, X.F.; Sun, H. FAK expression and its clinicopathological significance in hepatocellular carcinoma. J. Xi’an Jiaotong Univ. Medical Sci. 2013, 34, 280–284. [Google Scholar]
- Wang, G.; Benasutti, H.; Jones, J.F.; Shi, G.; Benchimol, M.; Pingle, S.; Kesari, S.; Yeh, Y.; Hsieh, L.E.; Liu, Y.T.; et al. Isolation of Breast cancer CTCs with multitargeted buoyant immunomicrobubbles. Colloids Surfaces B Biointerfaces 2018, 161, 200–209. [Google Scholar] [CrossRef]
- Hall, C.; Valad, L.; Lucci, A. Circulating tumor cells in breast cancer patients. Crit. Rev. Oncog. 2016, 21, 125–141. [Google Scholar] [CrossRef]
- Tabach, Y.; Sakin, I.K.; Buganim, Y.; Solomon, H.; Goldfinger, N.; Hovland, R.; Ke, X.S.; Oyan, A.M.; Kalland, K.H.; Rotter, V.; et al. Amplification of the 20q chromosomal arm occurs early in tumorigenic transformation and may initiate cancer. PLoS ONE 2011, 6, e14632. [Google Scholar] [CrossRef]
- Wang, Y.; Lu, P.; Liu, M. Expression of eIF3h in hepatocellular carcinoma and its clinicopathological significance. J. Pract. Oncol. 2017, 32, 345–348. [Google Scholar]
- Van Ree, J.H.; Jeganathan, K.B.; Malureanu, L.; Van Deursen, J.M. Overexpression of the E2 ubiquitin-conjugating enzyme UbcH10 causes chromosome missegregation and tumor formation. J. Cell Biol. 2010, 188, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liu, X.; Yu, G.; Liu, L.; Wang, J.; Chen, X.; Bian, Y.; Ji, Y.; Zhou, X.; Chen, Y.; et al. UBE2C is a potential biomarker of intestinal-type gastric cancer with chromosomal instability. Front. Pharmacol. 2018, 9, 847. [Google Scholar] [CrossRef] [PubMed]
- Sheffer, M.; Bacolod, M.D.; Zuk, O.; Giardina, S.F.; Pincas, H.; Barany, F.; Paty, P.B.; Gerald, W.L.; Notterman, D.A.; Domany, E. Association of survival and disease progression with chromosomal instability: A genomic exploration of colorectal cancer. Proc. Natl. Acad. Sci. USA 2009, 106, 7131–7136. [Google Scholar] [CrossRef] [PubMed]
- Rizvi, S.; Fischbach, S.R.; Bronk, S.F.; Hirsova, P.; Krishnan, A.; Dhanasekaran, R.; Smadbeck, J.B.; Smoot, R.L.; Vasmatzis, G.; Gores, G.J. YAP-associated chromosomal instability and cholangiocarcinoma in mice. Oncotarget 2018, 9, 5892–5905. [Google Scholar] [CrossRef]
- Adams, J. Proteasome inhibition in cancer: Development of PS-341. Semin. Oncol. 2001, 28, 613–619. [Google Scholar] [CrossRef]
- Palombella, V.J.; Rando, O.J.; Goldberg, A.L.; Maniatis, T. The ubiquitinproteasome pathway is required for processing the NF-κB1 precursor protein and the activation of NF-κB. Cell 1994, 78, 773–785. [Google Scholar] [CrossRef]
- Hyer, M.L.; Milhollen, M.A.; Ciavarri, J.; Fleming, P.; Traore, T.; Sappal, D.; Huck, J.; Shi, J.; Gavin, J.; Brownell, J.; et al. A small-molecule inhibitor of the ubiquitin activating enzyme for cancer treatment. Nat. Med. 2018, 24, 186–193. [Google Scholar] [CrossRef]
- Harper, J.W.; King, R.W. Stuck in the middle: Drugging the ubiquitin system at the E2 step. Cell 2011, 145, 1007–1009. [Google Scholar] [CrossRef][Green Version]
- Kothayer, H.; Spencer, S.M.; Tripathi, K.; Westwell, A.D.; Palle, K. Synthesis and in vitro anticancer evaluation of some 4,6-diamino-1,3,5-triazine-2-carbohydrazides as Rad6 ubiquitin conjugating enzyme inhibitors. Bioorganic Med. Chem. Lett. 2016, 26, 2030–2034. [Google Scholar] [CrossRef]
- Orlicky, S.; Tang, X.; Neduva, V.; Elowe, N.; Brown, E.D.; Sicheri, F.; Tyers, M. An allosteric inhibitor of substrate recognition by the SCF Cdc4 ubiquitin ligase. Nat. Biotechnol. 2010, 28, 733–737. [Google Scholar] [CrossRef]
- Pooja Naik, L.C. Diabetes Mellitus and Blood-Brain Barrier Dysfunction: An Overview. J. Pharmacovigil. 2014, 2, 125. [Google Scholar] [CrossRef] [PubMed]
- Chan, C.H.; Morrow, J.K.; Li, C.F.; Gao, Y.; Jin, G.; Moten, A.; Stagg, L.J.; Ladbury, J.E.; Cai, Z.; Xu, D.; et al. Pharmacological inactivation of Skp2 SCF ubiquitin ligase restricts cancer stem cell traits and cancer progression. Cell 2013, 154, 556–568. [Google Scholar] [CrossRef] [PubMed]
- Sabitha, K.; Rajkumar, T. Identification of small molecule inhibitors against UBE2C by using docking studies. Bioinformation 2012, 8, 1047–1058. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Haynes, B.; Zhang, Y.; Liu, F.; Li, J.; Petit, S.; Kothayer, H.; Bao, X.; Westwell, A.D.; Mao, G.; Shekhar, M.P.V. Gold nanoparticle conjugated Rad6 inhibitor induces cell death in triple negative breast cancer cells by inducing mitochondrial dysfunction and PARP-1 hyperactivation: Synthesis and characterization. Nanomed. Nanotechnol. Biol. Med. 2016, 12, 745–757. [Google Scholar] [CrossRef]
- Malara, N.; Guzzi, G.; Mignogna, C.; Trunzo, V.; Camastra, C.; Della Torre, A.; Di Vito, A.; Lavecchia, A.M.; Gliozzi, M.; Ceccotti, C.; et al. Non-invasive real-time biopsy of intracranial lesions using short time expanded circulating tumor cells on glass slide: Report of two cases. BMC Neurol. 2016, 16, 127. [Google Scholar] [CrossRef] [PubMed]
- Simone, G.; Malara, N.; Trunzo, V.; Perozziello, G.; Neuzil, P.; Francardi, M.; Roveda, L.; Renne, M.; Prati, U.; Mollace, V.; et al. Protein-carbohydrate complex reveals circulating metastatic cells in a microfluidic assay. Small 2013, 9, 2152–2161. [Google Scholar] [CrossRef] [PubMed]
- Malara, N.; Trunzo, V.; Foresta, U.; Amodio, N.; De Vitis, S.; Roveda, L.; Fava, M.; Coluccio, M.L.; Macrì, R.; Di Vito, A.; et al. Ex-vivo characterization of circulating colon cancer cells distinguished in stem and differentiated subset provides useful biomarker for personalized metastatic risk assessment. J. Transl. Med. 2016, 14, 133. [Google Scholar] [CrossRef]
- Guadagno, E.; Presta, I.; Maisano, D.; Donato, A.; Pirrone, C.K.; Cardillo, G.; Corrado, S.D.; Mignogna, C.; Mancuso, T.; Donato, G.; et al. Role of macrophages in brain tumor growth and progression. Int. J. Mol. Sci. 2018, 19, 1005. [Google Scholar] [CrossRef]
- Kim, W.T.; Jeong, P.; Yan, C.; Kim, Y.H.; Lee, I.S.; Kang, H.W.; Kim, Y.J.; Lee, S.C.; Kim, S.J.; Kim, Y.T.; et al. UBE2C cell-free RNA in urine can discriminate between bladder cancer and hematuria. Oncotarget 2016, 7, 58193–58202. [Google Scholar] [CrossRef]
| Tumor/Organ | Methods | Syntesis Results | References |
|---|---|---|---|
| Intracranial meningioma | Four primary meningioma cells cultures and Immunohistochemistry on tissue samples. | UbcH10 played an important role in the proliferation, apoptosis, and progression of human meningioma cells. RIZ1 regulated UbcH10 in a c-Myc dependent manner. | [18] |
| Immunohistochemistry. | Nuclear and cytoplasm immunopositivity is observed in advanced stages; low immunoreactivity in meningiomas with low histological grade. UbcH10 immunoreactivity significantly overexpressed in meningioma patients with recurrence (p < 0.001). | [19] | |
| Glioma/Brain | Cell culture of astrocytic tumors cells. Immunohistochemistry on tissue samples. | Elevated expression levels of UbcH10 messenger RNA (mRNA) in high- versus low-grade astrocytomas or normal controls. Immunohistochemistry shows increased UbcH10 in high-grade astrocytomas versus low-grade tumors or normal controls. | [20] |
| Immunohistochemistry on tissue samples. | No UbcH10 detection in normal and gliotic brain. Correlation between UbcH10 expression and histological grade in astrocytic tumors. | [21] | |
| Cell culture of U251 human glioblastoma cells and small interfering RNA (siRNA) transfection. | RNA interference targeting UbcH10 induces growth inhibition, apoptosis, and cell cycle arrest of U251 cells. | [22] | |
| Carcinoma/Lung | NSCLC tumors and cell lines A549 and SK-MES-1. | Impairment of NSCLC cells growth through the modulation of UbcH10 expression after hsa-miR661-3p restoration. | [23] |
| Immunohistochemistry | UbcH10 positivity in lung adenocarcinoma, squamous cell carcinoma, large cell, and small cell carcinoma. Progressive increase of UbcH10 levels associated to decrease of tumor differentiation. Statistically significant difference in UbcH10 positivity between grade I/III of lung adenocarcinoma and squamous cell carcinoma. | [24] | |
| Quantitative RT-PCR and tissue microarray immunohistochemistry. | UbcH10 expression significantly higher in squamous cell and large cell carcinomas compared to adenocarcinomas. Opposite correlations with mutational status of p53 and EGFR. | [25] | |
| Immunohistochemistry Cell culture and transfection of siRNA. | UbcH10 overexpression in poorly differentiated NSCLC compared to well-differentiated ones. Higher levels in squamous cell carcinoma than in adenocarcinoma. Higher UbcH10 expression associated with shorter postoperative survival in NSCLC. Inhibition of cell proliferation and Increase of chemosensitivity after UbcH10 suppression in SK-MES-1 cells. Concomitant decrease of MDR1 gene expression. | [26] | |
| Carcinoma/Breast | Array of mRNA markers expressed by circulating tumor cells (CTCs). | mRNA multimarker panel detected in CTCs from breast cancer patients included UbcH10. | [27] |
| Tissue micro-arrays Immunohistochemistry MB231; MDA468; MDA436; MCF7; T47D and ZR 75-1 human breast carcinoma cell lines. | Association between high UbcH10 levels and ductal histotype. ErbB2 positivity and high Ki-67 staining. siRNA mediated UbcH10 downregulation inhibits breast carcinoma cell growth. | [28] | |
| Immunohistochemistry on cancer tissues. Molecular analysis in breast cancer cell lines. | Increased expression levels of UbcH10 in cancer, in comparison with adjacent tissues. UbcH10 mRNA and protein levels analysis in breast cancer cell lines are significantly higher compared to normal mammary epithelial cells. The knockdown of UbcH10 inhibits proliferation of MCF-7 and MCF-7/EPB/TXT cells increasing sensitivity to chemotherapy drugs. | [29] | |
| Carcinoma/Thyroid | Immunohistochemistry on thyroid cancer tissues and normal thyroid tissues. RT-PCR and western blot analyses in human thyroid carcinoma cell lines. | Normal thyroid, nodular goiter, and HT almost always negative for UbcH10 expression. Weak staining in follicular adenomas; higher levels in papillary, follicular and poorly differentiated thyroid carcinomas. Abundancy of UbcH10 protein in diverse thyroid carcinoma cell lines, but barely detectable in normal thyroid cells. Western blot analysis, confirmed the immunohistochemical and RT-PCR data. | [30] |
| Spectral karyotyping (SKY) and G-banding in two novel ATC lines and six frequently used ATC lines. Comparative genomic hybridization. | Frequent gain of 20q, including the UBE2C locus in 20q13.12. | [31] | |
| Immunohistochemistry and RT-PCR on thyroid FNA samples. | Quantitative RT-PCR is more useful than immunohistochemistry to evaluate UbcH10as marker of malignancy in thyroid FNAs. | [32] | |
| Carcinoma/Colon | Cell-line based assay and tissue array analyses. | UbcH10 overexpression led to a significant acceleration of cellular growth in colon cancer cells. Knockdown of UbcH10 largely reduced cellular proliferation. Immunohistochemical analysis indicates that UbcH10 was significantly higher in colon cancer tissue compared to normal colon epithelia. | [33] |
| Silencing UBE2C gene by RNAi in colorectal cancer cell growth in vitro and in vivo in a nude mouse xenografts model. | Suppression of colorectal cancer cells growth, with arrest in the G2-M phase, upon UBE2C gene silencing. The downregulation of UBE2C gene in vivo inhibited tumor growth in a nude mouse. | [34] | |
| DDL1 colorectal cancer cells line Xenograft into nude mice. Colony formation assay, growth curve, soft agar and xenograft assays. Blocking of proteasome pathway by ALLN. | High expression of UbcH10 drives resistance to ALLN-induced cell death, while cells deficient in UbcH10 are susceptible. The depletion of UbcH10 hindered tumorigenesis both in vitro and in vivo. | [35] | |
| UbcH10 mRNA and protein evaluation in colon carcinoma and normal samples from CRC patients. Growth assays and pharmacological treatments on Caco-2 and DLD-1 cell lines. | Overexpression of UbcH10 mRNA and protein observed in the vast majority of tumoral samples analyzed. UbcH10 suppression decreases Colorectal cancer cells growth rate and sensitizes them to pharmacological treatments with irinotecan, SN-38 and cetuximab. | [36] | |
| Analysis of protein expression in CRC samples and normal colonic tissue. | Levels of UbcH10 overexpression related to the age of patient at surgery and to the lymph node spread. | [37] | |
| Carcinoma/Ovary | Immunohistochemistry on tumor tissues. Analysis on ovary carcinomas cell lines SKOV-3 also with siRNA targeting the UBE2C mRNA. | Cell growth inhibition after efficient knock down of the UbcH10 protein. Immunohistochemistry demonstrated that UbcH10 expression significantly correlates with the tumor grade. Clear correlation between UbcH10 overexpression and a reduced survival in ovarian carcinoma patients. | [38] |
| Ductal adenocarcinoma/pancreas | Real-time qRT-PCR Immunohistochemistry. | High levels of UbcH10 significantly associated with poor overall survival in PDA patients, with clinical stage, degree of histological differentiation, and lymph node metastasis. | [39] |
| Esophagus/Squamous Cell Carcinoma | Immunohistochemistry. | Detection of UbcH10 protein in cancerous lesions and some dysplastic lesions surrounding cancerous tissue, but not in normal tissue. association between high levels of UbcH10 protein expression in esophageal tissues and poor prognosis in patients with esophageal squamous cell carcinoma. | [40] |
| Stomach/carcinoma | Immunohistochemistry on cancer tissues. Gene silencing in human gastric carcinoma cell lines SGC-7901, AGS, NCI-N87, HS 746T, MKN-45, KATO III, NCI-SNU-1, SNU-5 and SNU-16. | Immunohistochemistry showed high levels of UbcH10 protein in most gastric cancer tissues but is unable to detect it in adjacent normal tissues. siRNA Knockdown of UbcH10 in gastric cancer cell lines (high expression of UbcH10) resulted in reduced proliferation, increased cisplatin-induced apoptosis, and reduced serum induced ERK, Akt, and p38 phosphorylation. | [41] |
| Uterus/Carcinoma | Immunohistochemistry on endometrial curettage biopsies of proliferative endometrium, disordered proliferative endometrium, complex atypical hyperplasia, nonatypical hyperplasia endometrioid adenocarcinoma. | A statistically significant difference was found only between the carcinoma group and the other groups, except the complex atypical hyperplasia group. | [42] |
| Bladder/Cancer | Immunohistochemistry on cancer tissues. Gene silencing in UM-UC-3 bladder cancer cell line. | SiRNA mediated suppression of UbcH10 in UM-UC-3 cells inhibited cell proliferation in vitro. Immunohistochemical UbcH10 positivity observed in 62% bladder urothelial carcinoma. Instead, UbcH10 negative in all non-neoplastic urothelium examined. UbcH10 positivity significantly associated with higher tumor stage and lymphovascular invasion. UbcH10 positivity significantly associated with shorter cancer-specific survival after cystectomy. | [43] |
| Bone/Osteosarcoma | UbcH10 knockdown in osteosarcoma U2OS and SaOS2 cell lines using lentivirus-mediated RNA interference. | UbcH10 knockdown in osteosarcoma cells exhibited impaired invasion and migration capacities. The downregulation of UbcH10 suppressed osteosarcoma cell proliferation and colony formation ability decreasing Ki-67 expression. Furthermore, UbcH10 knockdown led to decreased levels of matrix metalloproteases MMP-3 and MMP-9. | [44] |
| Lymphoid cells/Lymphoma | RT-PCR and tissue microarray immunohistochemistry to screening cell lines and tissue samples from HL and NHL for UbcH10 expression at transcriptional and translational levels. | Low UbcH10 expression in indolent tumors and higher expression in a variety of HL and NHL cell lines and in aggressive lymphomas. Highest expression in Burkitt’s lymphoma. UbcH10 plays a relevant role in lymphoid cell proliferation, since blocking of its synthesis by RNA interference inhibited cell growth. | [45] |
| Multiple myeloma/Plasma cells | Bortezomib (BTZ) -resistant myeloma cell lines U-266/BTZ, NCI-H929/BTZ and RPMI-8226/BTZ. Assessment of specific microRNAs (miRNAs) in both resistant and their parental cells. Luciferase reporter assay. | The development of BTZ resistance in U-266 cell lines is associated with decrease of hsa-miR-631 levels and increased expression of UbcH10. Positive correlation between MDR1 and UbcH10 due to reduced ubiquitination of MDR1. Overexpression of miR-631 enhanced both BTZ sensitivity and BTZ-induced apoptosis in resistant cells. Re-sensitization by miR-631 overexpression is blocked by exogenous UbcH10 not regulated by intracellular miR-631. Verification that hsa-miR-631 may inhibit translation by binding UbcH10-3’UTR. | [46] |
| Hepatocellular carcinoma/Liver | Immunohistochemistry in hepatocellular carcinoma tissue, the adjacent tissue and normal liver tissue. Study of UbcH10 mRNA expression using RT-PCR in normal liver cell line LO2, cancer cell lines BEL-7402, Hep3B, HepG2 and SMMC-7721. | Stronger UbcH10 expression in hepatocellular carcinoma tissues compared to adjacent tissues and normal liver tissues. Normal liver cell line showed relative lower UbcH10 mRNA levels compared to cancer cell lines. UbcH10 mRNA expression is significantly higher in hepatocellular carcinoma tissues compared to the corresponding non-tumor tissues. Clinicopathological evaluation suggests that UbcH10 expression is associated with tumor invasion of the portal vein, tumor size, TNM staging, and tumor differentiation. | [47] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Presta, I.; Novellino, F.; Donato, A.; La Torre, D.; Palleria, C.; Russo, E.; Malara, N.; Donato, G. UbcH10 a Major Actor in Cancerogenesis and a Potential Tool for Diagnosis and Therapy. Int. J. Mol. Sci. 2020, 21, 2041. https://doi.org/10.3390/ijms21062041
Presta I, Novellino F, Donato A, La Torre D, Palleria C, Russo E, Malara N, Donato G. UbcH10 a Major Actor in Cancerogenesis and a Potential Tool for Diagnosis and Therapy. International Journal of Molecular Sciences. 2020; 21(6):2041. https://doi.org/10.3390/ijms21062041
Chicago/Turabian StylePresta, Ivan, Fabiana Novellino, Annalidia Donato, Domenico La Torre, Caterina Palleria, Emilio Russo, Natalia Malara, and Giuseppe Donato. 2020. "UbcH10 a Major Actor in Cancerogenesis and a Potential Tool for Diagnosis and Therapy" International Journal of Molecular Sciences 21, no. 6: 2041. https://doi.org/10.3390/ijms21062041
APA StylePresta, I., Novellino, F., Donato, A., La Torre, D., Palleria, C., Russo, E., Malara, N., & Donato, G. (2020). UbcH10 a Major Actor in Cancerogenesis and a Potential Tool for Diagnosis and Therapy. International Journal of Molecular Sciences, 21(6), 2041. https://doi.org/10.3390/ijms21062041