Real-Time 3D Imaging and Inhibition Analysis of Various Amyloid Aggregations Using Quantum Dots
Abstract
:1. Introduction
2. Results
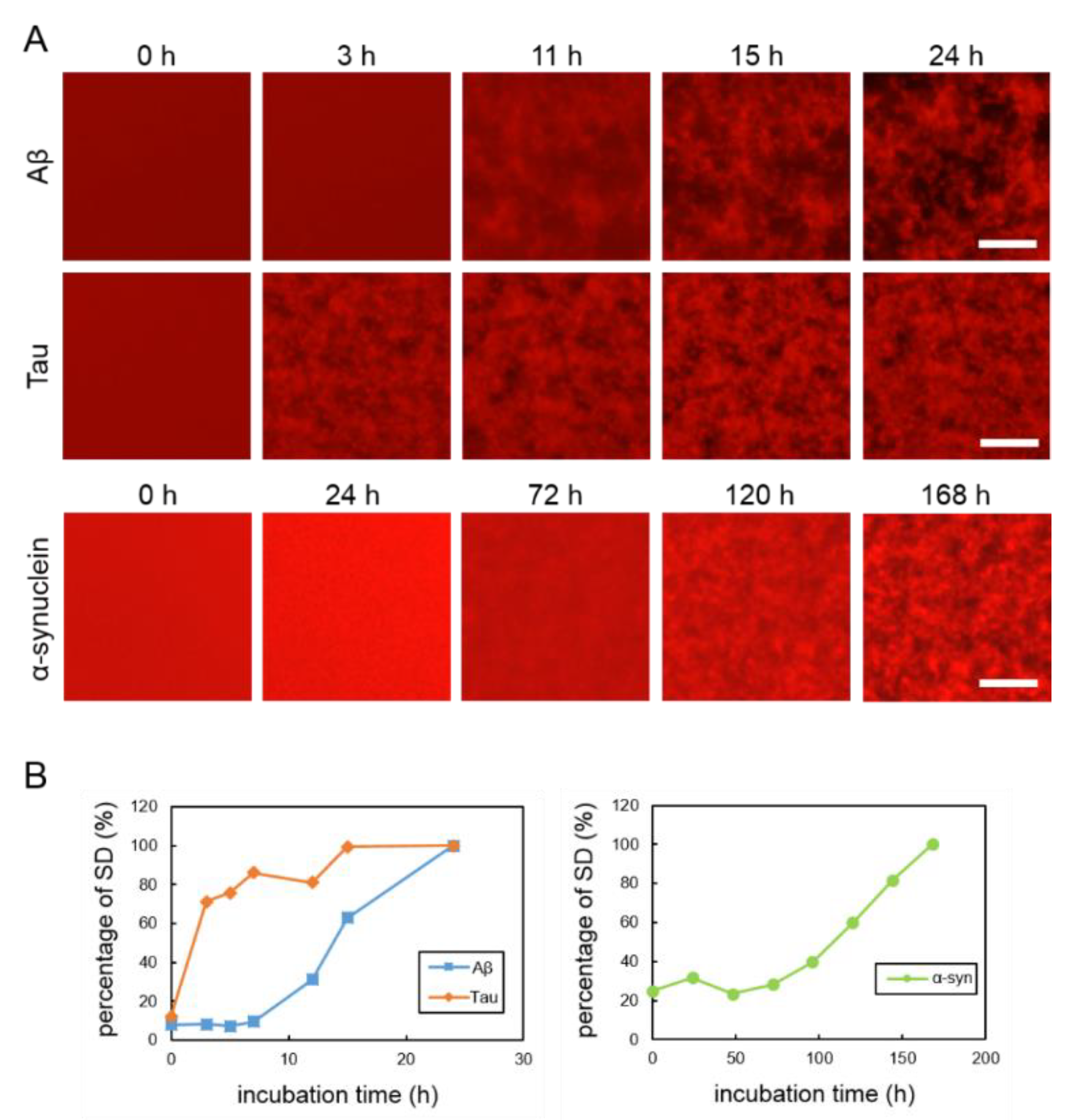
2.1. Real-Time Imaging of Aggregation Processes of Various Amyloid Proteins
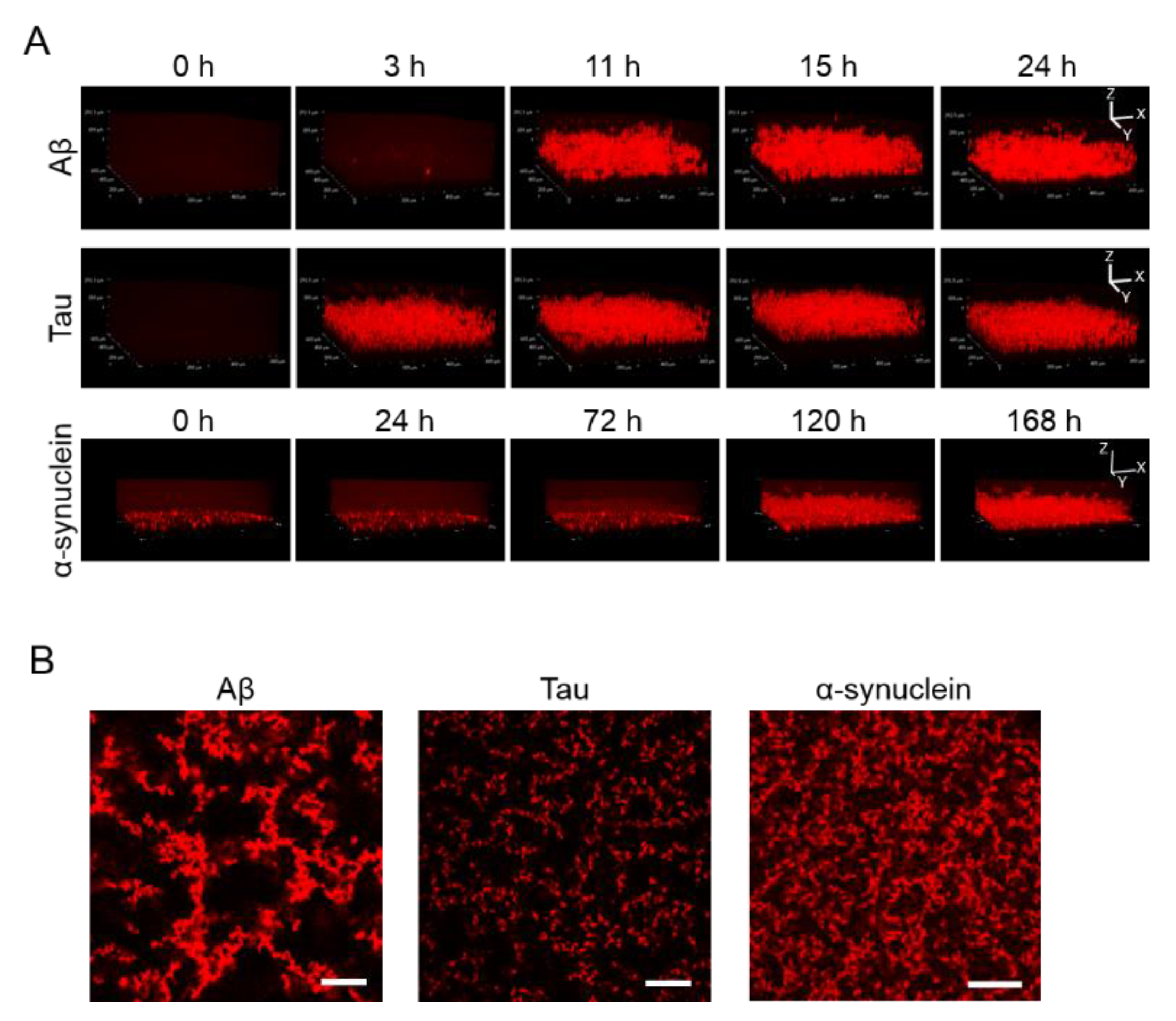
2.2. 3D Observation of Aggregation of Various Amyloid Protein
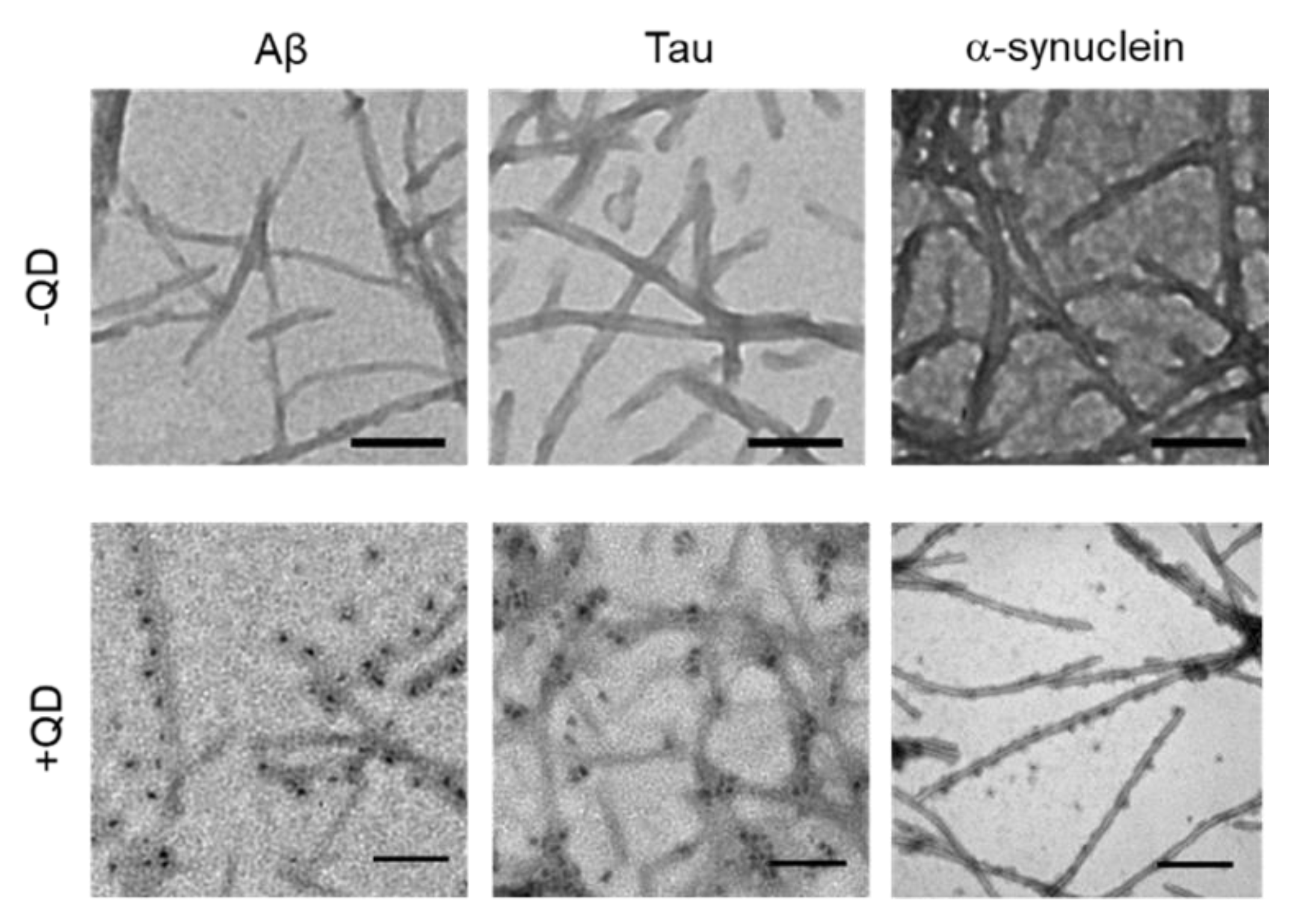
2.3. Transmission Electron Microscopy Observation of Various Amyloid Fibrils
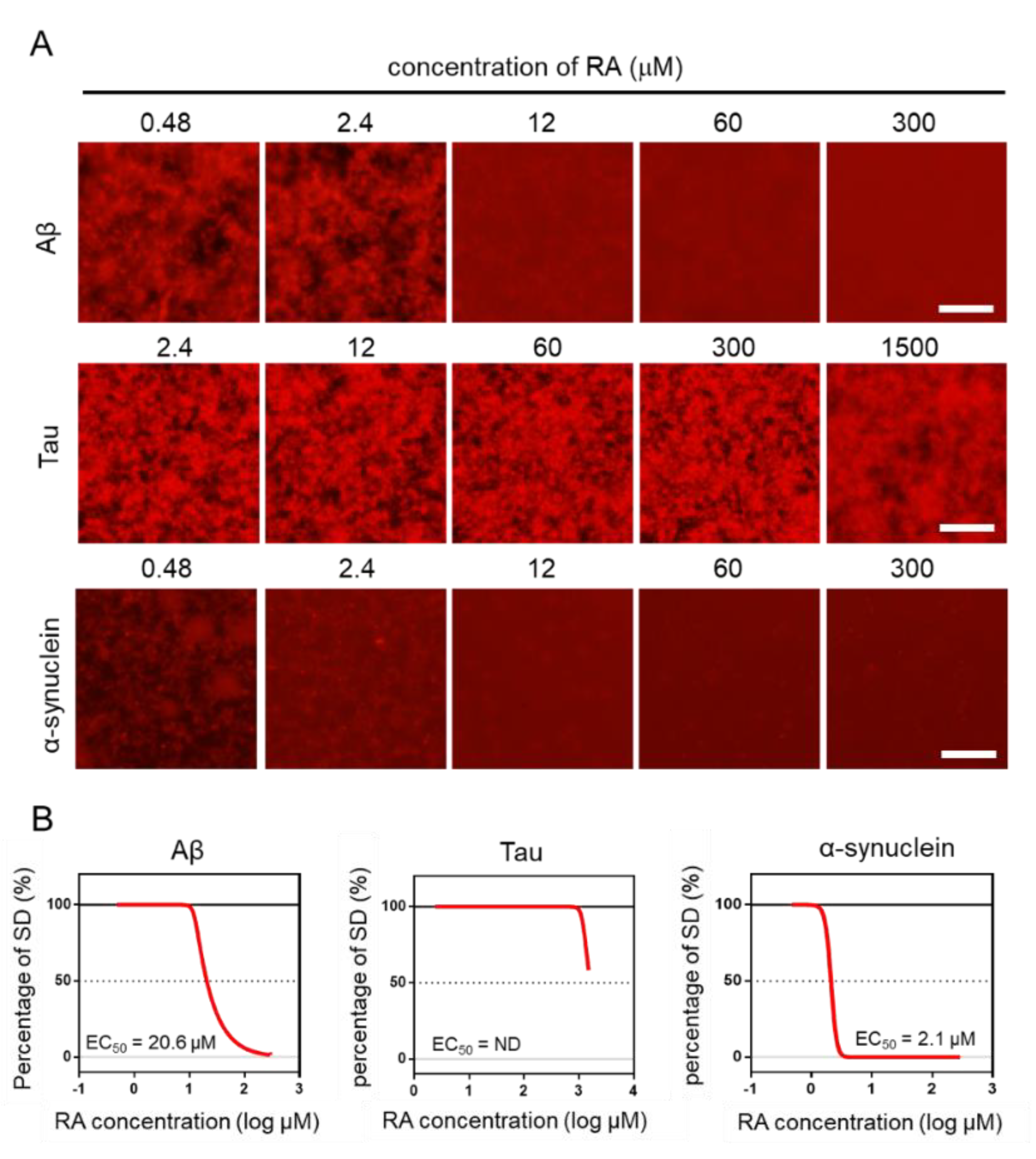
2.4. Effect of RA on the Aggregation of Three Amyloid Proteins
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Preparation of tau and α-synuclein
4.3. Microliter-Scale High-Throughput Screening System
4.4. Aggregation of Three Amyloid Proteins
4.5. Transmission Electron Microscopy Observation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Sipe, J.D.; Benson, M.D.; Buxbaum, J.N.; Ikeda, S.I.; Merlini, G.; Saraiva, M.J.M.; Westermark, P. Amyloid fibril proteins and amyloidosis: Chemical identification and clinical classification International Society of Amyloidosis 2016 Nomenclature Guidelines. Amyloid 2016, 23, 209–213. [Google Scholar] [CrossRef]
- Sipe, J.D.; Benson, M.D.; Buxbaum, J.N.; Ikeda, S.I.; Merlini, G.; Saraiva, M.J.M.; Westermark, P. Amyloid fibril protein nomenclature: 2012 recommendations from the Nomenclature Committee of the International Society of Amyloidosis. Amyloid 2012, 19, 167–170. [Google Scholar] [CrossRef]
- Watanabe, K.; Uchida, K.; Chambers, J.K.; Tei, M.; Shoji, A.; Ushio, N.; Nakayama, H. Experimental Transmission of AA Amyloidosis by Injecting the AA Amyloid Protein Into Interleukin-1 Receptor Antagonist Knockout (IL-1raKO) Mice. Vet. Pathol. 2015, 52, 505–512. [Google Scholar] [CrossRef] [Green Version]
- Sipe, J.D.; Cohen, A.S. Review: History of the amyloid fibril. J. Struct. Biol. 2000, 130, 88–98. [Google Scholar] [CrossRef] [Green Version]
- Howe, H.S. Rheumatoid arthritis—A review. Ann. Acad. Med. Singap. 1998, 27, 83–88. [Google Scholar]
- Ruipérez, V.; Darios, F.; Davletov, B. Alpha-synuclein, lipids and Parkinson’s disease. Prog. Lipid Res. 2010, 49, 420–428. [Google Scholar] [CrossRef]
- Selkoe, D.J.; Hardy, J.; Sciences, B.; Hu, N.-W.; Nicoll, A.J.; Zhang, D.; Mably, A.J.; O’Malley, T.; Purro, S.A.; Terry, C.; et al. Amyloid β-Protein Dimers Isolated Directly from Alzheimer Brains Impair Synaptic Plasticity and Memory. Nat. Med. 2016, 7, 3374. [Google Scholar]
- Koo, E.H.; Lansbury, P.T.; Kelly, J.W. Amyloid diseases: Abnormal protein aggregation in neurodegeneration. Proc. Natl. Acad. Sci. USA 1999, 96, 9989–9990. [Google Scholar] [CrossRef] [Green Version]
- Ising, C.; Venegas, C.; Zhang, S.; Scheiblich, H.; Schmidt, S.V.; Vieira-Saecker, A.; Schwartz, S.; Albasset, S.; McManus, R.M.; Tejera, D.; et al. NLRP3 inflammasome activation drives tau pathology. Nature 2019, 575, 669–673. [Google Scholar] [CrossRef]
- Pfefferkorn, C.M.; Jiang, Z.; Lee, J.C. Biophysics of α-synuclein membrane interactions. Biochim. Biophys. Acta Biomembr. 2012, 1818, 162–171. [Google Scholar] [CrossRef] [Green Version]
- Baglioni, S.; Casamenti, F.; Bucciantini, M.; Luheshi, L.M.; Taddei, N.; Chiti, F.; Dobson, C.M.; Stefani, M. Prefibrillar amyloid aggregates could be generic toxins in higher organisms. J. Neurosci. 2006, 26, 8160–8167. [Google Scholar] [CrossRef]
- Bucciantini, M. Inherent cytotoxicity of aggregates implies a common origin for protein misfolding diseases. Nature 2002, 416, 507–511. [Google Scholar] [CrossRef]
- Winner, B.; Jappelli, R.; Maji, S.K.; Desplats, P.A.; Boyer, L.; Aigner, S.; Hetzer, C.; Loher, T.; Vilar, M.; Campioni, S.; et al. In vivo demonstration that α-synuclein oligomers are toxic. Proc. Natl. Acad. Sci. USA 2011, 108, 4194–4199. [Google Scholar] [CrossRef] [Green Version]
- Radamaker, L.; Lin, Y.H.; Annamalai, K.; Huhn, S.; Hegenbart, U.; Schönland, S.O.; Fritz, G.; Schmidt, M.; Fändrich, M. Cryo-EM structure of a light chain-derived amyloid fibril from a patient with systemic AL amyloidosis. Nat. Commun. 2019, 10, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Liberta, F.; Loerch, S.; Rennegarbe, M.; Schierhorn, A.; Westermark, P.; Westermark, G.T.; Hazenberg, B.P.C.; Grigorieff, N.; Fändrich, M.; Schmidt, M. Cryo-EM fibril structures from systemic AA amyloidosis reveal the species complementarity of pathological amyloids. Nat. Commun. 2019, 10, 1–12. [Google Scholar]
- Iadanza, M.G.; Jackson, M.P.; Hewitt, E.W.; Ranson, N.A.; Radford, S.E. A new era for understanding amyloid structures and disease. Nat. Rev. Mol. Cell Biol. 2018, 19, 755–773. [Google Scholar] [CrossRef]
- Sevigny, J.; Chiao, P.; Bussière, T.; Weinreb, P.H.; Williams, L.; Maier, M.; Dunstan, R.; Salloway, S.; Chen, T.; Ling, Y.; et al. The antibody aducanumab reduces Aβ plaques in Alzheimer’s disease. Nature 2016, 537, 50–56. [Google Scholar] [CrossRef]
- Nakamura, A.; Kaneko, N.; Villemagne, V.L.; Kato, T.; Doecke, J.; Doré, V.; Fowler, C.; Li, Q.X.; Martins, R.; Rowe, C.; et al. High performance plasma amyloid-β biomarkers for Alzheimer’s disease. Nature 2018, 554, 249–254. [Google Scholar] [CrossRef]
- Ishihara, S.; Tada, Y.; Kawashima, K.; Kataoka, M.; Sonoyama, H.; Yamashita, N.; Oka, A.; Kusunoki, R.; Fukuba, N.; Mishima, Y.; et al. Serum amyloid A level correlated with endoscopic findings in patients with Crohn’s disease—Possible biomarker for evaluating mucosal healing. Dig. Liver Dis. 2018, 50, 553–558. [Google Scholar] [CrossRef]
- Strømland, Ø.; Kakubec, M.; Halskau, Ø. Detection of mis-folded protein aggregates from a clinical perspective. J. Clin. Transl. Res. 2016, 1, 11–26. [Google Scholar]
- Sasaki, R.; Tainaka, R.; Ando, Y.; Hashi, Y.; Deepak, H.V.; Suga, Y.; Murai, Y.; Anetai, M.; Monde, K.; Ohta, K.; et al. An automated microliter-scale high-throughput screening system (MSHTS) for real-time monitoring of protein aggregation using quantum-dot nanoprobes. Sci. Rep. 2019, 9, 1–9. [Google Scholar]
- Chan, W.C.W.; Nie, S. Quantum dot bioconjugates for ultrasensitive nonisotopic detection. Science 1998, 281, 2016–2018. [Google Scholar] [CrossRef] [Green Version]
- Bruchez, M.; Moronne, M.; Gin, P.; Weiss, S.; Alivisatos, A.P. Semiconductor nanocrystals as fluorescent biological labels. Science 1998, 281, 2013–2016. [Google Scholar] [CrossRef] [Green Version]
- Ishigaki, Y.; Tanaka, H.; Akama, H.; Ogara, T.; Uwai, K.; Tokuraku, K. A Microliter-Scale High-throughput Screening System with Quantum-Dot Nanoprobes for Amyloid-β Aggregation Inhibitors. PLoS ONE 2013, 8, e72992. [Google Scholar] [CrossRef] [Green Version]
- Tokuraku, K.; Marquardt, M.; Ikezu, T. Real-time imaging and quantification of amyloid-β peptide aggregates by novel quantum-dot nanoprobes. PLoS ONE 2009, 4, e8492. [Google Scholar] [CrossRef]
- Taguchi, R.; Hatayama, K.; Takahashi, T.; Hayashi, T.; Sato, Y.; Sato, D.; Ohta, K.; Nakano, H.; Seki, C.; Endo, Y.; et al. Structure–activity relations of rosmarinic acid derivatives for the amyloid β aggregation inhibition and antioxidant properties. Eur. J. Med. Chem. 2017, 138, 1066–1075. [Google Scholar] [CrossRef] [Green Version]
- Lu, J.; Yu, Y.; Zhu, I.; Cheng, Y.; Sun, P.D. Structural mechanism of serum amyloid A-mediated inflammatory amyloidosis. Proc. Natl. Acad. Sci. USA 2014, 111, 5189–5194. [Google Scholar] [CrossRef] [Green Version]
- Bhat, A.; Selmi, C.; Naguwa, S.M.; Cheema, G.S.; Gershwin, M.E. Currents concepts on the immunopathology of amyloidosis. Clin. Rev. Allergy Immunol. 2010, 38, 97–106. [Google Scholar] [CrossRef]
- Jayaraman, S.; Gantz, D.L.; Haupt, C.; Gursky, O. Serum amyloid A forms stable oligomers that disrupt vesicles at lysosomal pH and contribute to the pathogenesis of reactive amyloidosis. Proc. Natl. Acad. Sci. USA 2017, 114, E6507–E6515. [Google Scholar] [CrossRef] [Green Version]
- Rodriguez, J.A.; Ivanova, M.I.; Sawaya, M.R.; Cascio, D.; Reyes, F.E.; Shi, D.; Sangwan, S.; Guenther, E.L.; Johnson, L.M.; Zhang, M.; et al. Structure of the toxic core of α-synuclein from invisible crystals. Nature 2015, 525, 486–490. [Google Scholar] [CrossRef] [Green Version]
- Fitzpatrick, A.W.P.; Falcon, B.; He, S.; Murzin, A.G.; Murshudov, G.; Garringer, H.J.; Crowther, R.A.; Ghetti, B.; Goedert, M.; Scheres, S.H.W. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 2017, 547, 185–190. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Merlini, G.; Bellotti, V. Molecular mechanisms of amyloidosis. N. Engl. J. Med. 2003, 349, 583–596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van der Kant, R.; Goldstein, L.S.B.; Ossenkoppele, R. Amyloid-β-independent regulators of tau pathology in Alzheimer disease. Nat. Rev. Neurosci. 2019, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Long, J.M.; Holtzman, D.M. Alzheimer Disease: An Update on Pathobiology and Treatment Strategies. Cell 2019, 179, 312–339. [Google Scholar] [CrossRef] [PubMed]
- Bernini, F.; Malferrari, D.; Pignataro, M.; Bortolotti, C.A.; Di Rocco, G.; Lancellotti, L.; Brigatti, M.F.; Kayed, R.; Borsari, M.; Del Monte, F.; et al. Pre-amyloid oligomers budding: A metastatic mechanism of proteotoxicity. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Tijms, B.M.; Vermunt, L.; Zwan, M.D.; van Harten, A.C.; van der Flier, W.M.; Teunissen, C.E.; Scheltens, P.; Visser, P.J. Pre-amyloid stage of Alzheimer’s disease in cognitively normal individuals. Ann. Clin. Transl. Neurol. 2018, 5, 1037–1047. [Google Scholar] [CrossRef] [Green Version]
- Reixach, N.; Deechongkit, S.; Jiang, X.; Kelly, J.W.; Buxbaum, J.N. Tissue damage in the amyloidoses: Transthyretin monomers and nonnative oligomers are the major cytotoxic species in tissue culture. Proc. Natl. Acad. Sci. USA 2004, 101, 2817–2822. [Google Scholar] [CrossRef] [Green Version]
- Levine, H. Soluble multimeric Alzheimer β(1-40) pre-amyloid complexes in dilute solution. Neurobiol. Aging 1995, 16, 755–764. [Google Scholar] [CrossRef]
- Lundmark, K.; Westermark, G.T.; Nyström, S.; Murphy, C.L.; Solomon, A.; Westermark, P. Transmissibility of systemic amyloidosis by a prion-like mechanism. Proc. Natl. Acad. Sci. USA 2002, 99, 6979–6984. [Google Scholar] [CrossRef] [Green Version]
- Glabe, C.G. Common mechanisms of amyloid oligomer pathogenesis in degenerative disease. Neurobiol. Aging 2006, 27, 570–575. [Google Scholar] [CrossRef]
- Qiang, W.; Yau, W.; Lu, J.; Collinge, J.; Tycko, R. Structural variation in amyloid-β fibrils from Alzheimer’s disease clinical subtypes. Nature 2017, 541, 217–221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jameson, L.P.; Smith, N.W.; Dzyuba, S.V. Dye-binding assays for evaluation of the effects of small molecule inhibitors on amyloid (Aβ) self-assembly. ACS Chem. Neurosci. 2012, 3, 807–819. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Giau, V.; Bagyinszky, E.; Youn, Y.C.; An, S.S.A.; Kim, S.Y. APP, PSEN1, and PSEN2 mutations in asian patients with early-onset alzheimer disease. Int. J. Mol. Sci. 2019, 20, E4757. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogawa, S.; Murakami, T.; Inoshima, Y.; Ishiguro, N. Effect of heating on the stability of amyloid A (AA) fibrils and the intra- and cross-species transmission of AA amyloidosis. Amyloid 2015, 22, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Blesl, A.; Rainer, F.; Pollheimer, M.; Kump, P.; Durchschein, F.; Wenzl, H.; Eller, K.; Krisper, P.; Petritsch, W.; Högenauer, C. Successful Pregnancies After Regression of AA Amyloidosis by Anti-inflammatory Therapy in Chronic Active Crohn’s Disease. Dig. Dis. Sci. 2019, 20, 4757. [Google Scholar] [CrossRef] [PubMed]
- Sperry, B.W.; Tang, W.H.W. Amyloid heart disease: Genetics translated into disease-modifying therapy. Heart 2017, 103, 812–817. [Google Scholar] [CrossRef]
- Garber, K. Alnylam’s RNAi therapy targets amyloid disease. Nat. Biotechnol. 2015, 33, 577. [Google Scholar] [CrossRef]
- Moreira, G.G.; Cristóvão, J.S.; Torres, V.M.; Carapeto, A.P.; Rodrigues, M.S.; Landrieu, I.; Cordeiro, C.; Gomes, C.M. Zinc Binding to Tau Influences Aggregation Kinetics and Oligomer Distribution. Int. J. Mol. Sci. 2019, 20, 5979. [Google Scholar] [CrossRef] [Green Version]
- Kollmer, M.; Close, W.; Funk, L.; Rasmussen, J.; Bsoul, A.; Schierhorn, A.; Schmidt, M.; Sigurdson, C.J.; Jucker, M.; Fändrich, M. Cryo-EM structure and polymorphism of Aβ amyloid fibrils purified from Alzheimer’s brain tissue. Nat. Commun. 2019, 10, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Sidhu, A.; Segers-Nolten, I.; Subramaniam, V. Solution conditions define morphological homogeneity of α-synuclein fibrils. Biochim. Biophys. Acta Proteins Proteom. 2014, 1844, 2127–2134. [Google Scholar] [CrossRef]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [Google Scholar] [PubMed]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Hoyer, W.; Antony, T.; Cherny, D.; Heim, G.; Jovin, T.M.; Subramaniam, V. Dependence of α-synuclein aggregate morphology on solution conditions. J. Mol. Biol. 2002, 322, 383–393. [Google Scholar] [CrossRef] [Green Version]
- Buell, A.K.; Galvagnion, C.; Gaspar, R.; Sparr, E.; Vendruscolo, M.; Knowles, T.P.J.; Linse, S.; Dobson, C.M. Solution conditions determine the relative importance of nucleation and growth processes in α-synuclein aggregation. Proc. Natl. Acad. Sci. USA 2014, 111, 7671–7676. [Google Scholar] [CrossRef] [Green Version]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, X.; Galaqin, N.; Tainaka, R.; Shimamori, K.; Kuragano, M.; Noguchi, T.Q.P.; Tokuraku, K. Real-Time 3D Imaging and Inhibition Analysis of Various Amyloid Aggregations Using Quantum Dots. Int. J. Mol. Sci. 2020, 21, 1978. https://doi.org/10.3390/ijms21061978
Lin X, Galaqin N, Tainaka R, Shimamori K, Kuragano M, Noguchi TQP, Tokuraku K. Real-Time 3D Imaging and Inhibition Analysis of Various Amyloid Aggregations Using Quantum Dots. International Journal of Molecular Sciences. 2020; 21(6):1978. https://doi.org/10.3390/ijms21061978
Chicago/Turabian StyleLin, Xuguang, Nuomin Galaqin, Reina Tainaka, Keiya Shimamori, Masahiro Kuragano, Taro Q. P. Noguchi, and Kiyotaka Tokuraku. 2020. "Real-Time 3D Imaging and Inhibition Analysis of Various Amyloid Aggregations Using Quantum Dots" International Journal of Molecular Sciences 21, no. 6: 1978. https://doi.org/10.3390/ijms21061978
APA StyleLin, X., Galaqin, N., Tainaka, R., Shimamori, K., Kuragano, M., Noguchi, T. Q. P., & Tokuraku, K. (2020). Real-Time 3D Imaging and Inhibition Analysis of Various Amyloid Aggregations Using Quantum Dots. International Journal of Molecular Sciences, 21(6), 1978. https://doi.org/10.3390/ijms21061978




